Abstract
Malaysian Mahseer (
Keywords
Introduction
Malaysian Mahseer (
In some species, the presence of bacteria in sperm microbiota affects the sperm during spermatogenesis or sperm maturation. In insects, development of sperm in alpha-proteobacterium
Materials and Methods
Collection of Malaysian Mahseer sperm
Sperm samples of Malaysian Mahseer were randomly collected from 3 locations: Hatchery Kelah at Universiti Malaysia Terengganu (UMT), Kuala Terengganu (5°24′11.39″ N, 103°05′9.60″ E), Ajil, Terengganu (4°53′15.173″ N, 103°2′24.529″ E), and Kuantan, Pahang (3°47′39.811″ N, 103°16′0.174″ E). Sperm was collected from 3 male broodstock (N = 3) at each location and anesthetized using 0.04% of clove oil and sterilized by spraying of 95% ethanol at fish abdominal area before sperm collection. The body weights (BWs) of the selected brooders were 2.34 ± 0.16, 0.47 ± 0.13, and 1.13 ± 0.26 kg for the UMT, Ajil, and Pahang samples, respectively. The total lengths (TLs) of the selected brooders were 59 ± 1.0, 35 ± 3.1, and 46 ± 2.6 cm for the UMT, Ajil, and Pahang samples, respectively.
Sperm sample was then collected from males by gentle abdominal pressure 14 and put into a sterile 1.5-mL microcentrifuge tube. Then, the sperm samples were immediately transferred to the laboratory and further experiments were performed.
Sperm quality evaluation
One microliter of sperm was transferred into the upper part/wall of a 1.5-mL centrifuge tube that contained 999 µL of freshwater. The sperm was then mixed into freshwater and, immediately, 8 µL of the mixture was transferred into slide glass for observation under microscope at 20× magnification. The sperm motility was recorded using video camera (Dino eye) using Dino capture 2.0 software (version 1.4.3) starting from the time sperm was mixed with the activation solution (freshwater). The sperm motility videos were then analyzed manually to determine percent motility. At least n = 50 sperm were observed for each video recording during each sperm motility evaluation.
Sperm concentration (cells/mL) was estimated by diluting sperm with the extender solution at a final milt-to-extender dilution ratio of 1:9999 µL and counting the number of sperm in a hemocytometer chamber (Thoma Neu; chamber volume 0.00025 mm3) under a phase contrast microscope. In each instance, 5 chambers were counted. 15
Bacterial genomic DNA extraction and polymerase chain reaction amplification of 16S rDNA
Each sperm sample was suspended with sterile phosphate buffered saline (PBS; pH 7.4, Invitrogen, San Diego, USA) and sonicated using an ultrasound sonicator for 5 minutes to detach sperm-attached bacteria, and then sperm-related bacteria were collected using centrifugation at 14 000×
Denaturing gradient gel electrophoresis
Denaturing gradient gel electrophoresis (DGGE) fingerprinting analysis was performed in 6% (v/w) polyacrylamide gel with 35% to 55% gradient of urea and formamide in 1× Tris-acetate-EDTA (TAE) buffer. A total of 20 µL of each PCR sample was loaded into each well and electrophoresis was initiated by pre-running for 20 minutes at 60 V and subsequently run at a constant voltage of 150 V for 7 hours at 60°C using the C.B.S Scientifix Model DGGE-2401 (C.B.S. Scientifix, Del Mar, CA, USA). Thereafter, the gels were stained for 1 hour in SYBR Safe DNA gel stain (Invitrogen, Thermo Fisher Scientific, Waltham, MA, USA), washed for 10 minutes, and photographed using Omega Lum G Imaging System (Aplegen, San Francisco, CA, USA). The DNA from unique band was retrieved by elution of the excised band into 20 µL of sterile double-distilled water (ddw) at 4°C overnight. Then 1 µL of eluted DNA was amplified using the same primers without GC clamp. A total of 30 µL of PCR reaction mixture contained the following: sterile ddw, master mix (MyTaq Mix, Bioline, London, UK), primers (each 0.2 mM), and 1 µL of eluted DNA. After a 4-minute initial denaturation at 95°C, the reaction mixture was run for 30 cycles based on the following condition: 95°C for 0.5 minutes, 56°C for 0.5 minutes, and 72°C for 1.5 minutes followed by 72°C for 7 minutes. Thereafter, all PCR products were analyzed on 1.5% agarose gel for 30 minutes at 100 V.
Sequencing and biodiversity analysis
PCR amplicon products from re-amplification of eluted DGGE gel bands were purified and sequenced in an external laboratory (FIRST BASE LABORATORIES SDN BHD, Selangor, Malaysia). The obtained chromatograms of DNA sequences were analyzed using Chromas 2.6.2 software (Technelysium, Brisbane, QLD, Australia). Then, all sequences were checked for chimeric sequences using a chimeric sequence detection software Decipher. 17 Homology searches were performed using sequences of −550 nucleotides, and close relatives were assigned by GenBank database through National Center for Biotechnology Information (NCBI) using BLAST program (https://blast.ncbi.nlm.nih.gov/Blast.cgi). Multiple alignments were conducted using MEGA version 7. 18 A phylogenetic tree was constructed using the Neighbor-joining analysis function of MEGA version 7 with 1000 replicates in bootstrap analysis. Distances were estimated with Jukes-Cantor correction. Thereafter, 3 alpha diversity indices (Margalef species richness, Simpson diversity index, and Shannon-Wiener index) and cluster analysis to compare the bacterial communities based on Bray-Curtis dissimilarity index were analyzed using PAST program version 3.15. 19
Statistical analysis
All data were presented as mean ± standard error in the table. Data analysis was performed using one-way analysis of variance (ANOVA) to determine the significance between the sperm quality and sperm microbiota from different locations using SPSS (version 18). Correlation was analyzed based on Pearson correlations using R version 3.5.1 with R Studio v. 1.1.45 (https://www.r-project.org/; https://www.rstudio.com/).
Nucleotide sequence accession numbers
The 16S rDNA sequences of the DGGE bands obtained in this study have been deposited to DDBJ/EMBL/GenBank under Accession No. MK779776-MK779795.
Results
Sperm quality and biodiversity of sperm microbiota
The results of sperm quality (sperm motility and concentration) and biodiversity index of the sperm microbiota in samples from different locations based on DGGE fingerprinting result (Figure 1) are presented in Table 1. The Pahang sample showed the highest sperm quality (sperm motility and concentration) and the UMT sample shows the lowest sperm quality, although there were no significant differences (

Denaturing gradient gel electrophoresis (DGGE) profile of PCR products of the V3-V5 region of 16S rDNA from Malaysian Mahseer sperm samples from different locations. Triplicates of samples were prepared for each location sample. PCR indicates polymerase chain reaction; UMT, Universiti Malaysia Terengganu.
Bacterial diversity and sperm quality of Malaysian Mahseer from different locations.
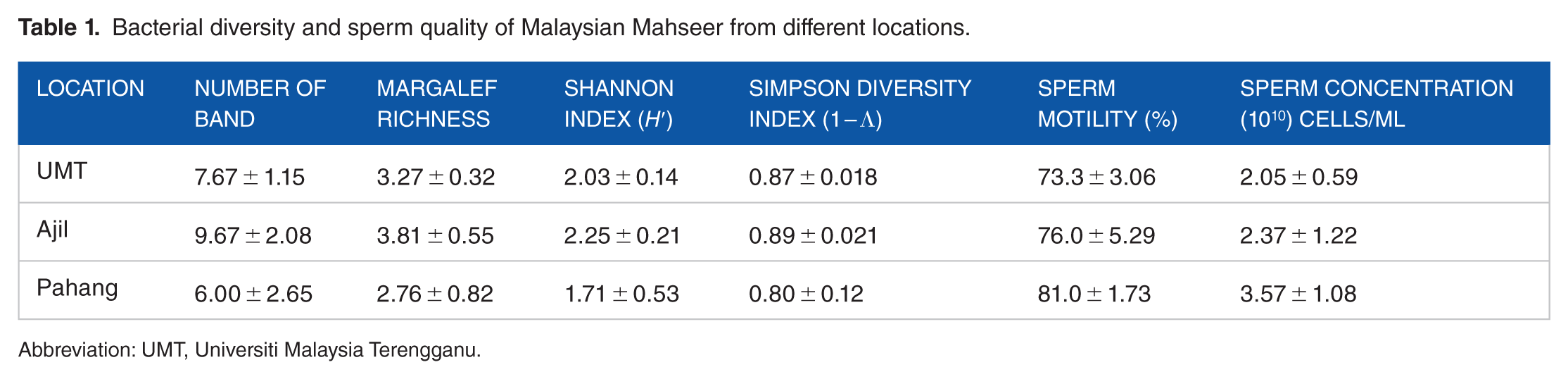
Abbreviation: UMT, Universiti Malaysia Terengganu.
DGGE fingerprinting, cluster analysis, and identification of sperm microbiota
To compare the structure of sperm microbiota from different locations, PCR-DGGE fingerprinting was conducted. In total, 21 unique bands (SPM1-20) were successfully excised and retrieved from DGGE gel (Figure 1). DGGE banding patterns and cluster analysis differed substantially between UMT and both Ajil and Pahang samples (Figures 1 and 2). Four bands (SPM 4, 10, 11, and 19) were found in samples from all locations. SPM 2, 3, 5, 6, 7, 16, 18, and 20 were found in either the Ajil and Pahang samples, the UMT and Ajil samples, or the UMT and Pahang samples. Eight bands (SPM 1, 8, 9, 12-15, and 17) were unique in each location sample (Supplemental Table 1). Partial 16S rDNA sequences (–550 bp) were assigned to their closest relatives based on comparison with the deposited sequences in GenBank database (Supplemental Table 1). The sequences of DGGE bands were diversely affiliated with α-Proteobacteria, β-Proteobacteria, γ-Proteobacteria, Bacilli, Bacteroidia, Cytophagia, Planctomycetia, and Actinobacteria (Figure 3). Bands SPM 3, 9, and 11 were closely related to the genus

Cluster dendrogram for comparison of sperm microbiota composition in samples from different locations based on PCR-DGGE fingerprinting profile using the Ward method. PCR-DGGE indicates polymerase chain reaction denaturing gradient gel electrophoresis; UMT, Universiti Malaysia Terengganu.
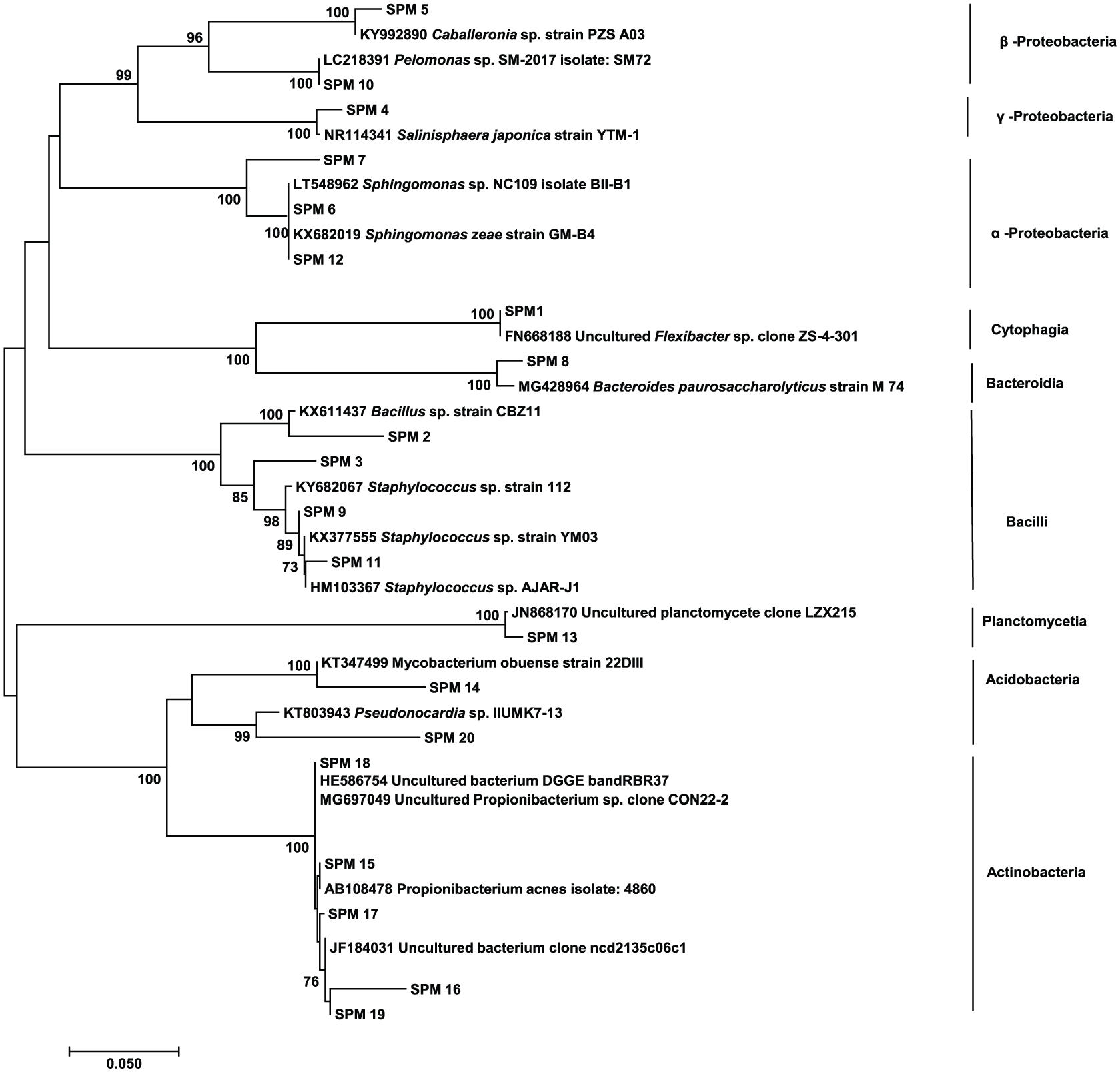
Neighbor-joining tree of partial 16S rDNA nucleotide sequence (ca. 550 bp) retrieved from DGGE bands shown in Figure 1. Bootstrap resampling was performed 1000 times. Bootstrap values above 70 are shown. DGGE indicates denaturing gradient gel electrophoresis.
There was a positive correlation between sperm motility and concentration in all location samples, albeit in various ranges (Figure 4). There was a positive correlation between bacterial biodiversity and sperm motility in the Ajil (0.98 as very strong) and Pahang (0.50 as moderate) samples, whereas the UMT sample had a moderately negative correlation (>–0.50 as moderate). On the other hand, in the case of bacterial diversity and sperm concentration, there was a strong negative correlation in the UMT and Pahang samples (>–0.70), whereas there was a positive moderate correlation in the Ajil sample.

Pearson correlation results among bacterial diversity, sperm motility, and sperm concentration in samples from different locations. No_B represents the number of DGGE band; Simp represents Simpson diversity index; Shan represents Shannon index; Marg represents Margalef richness; SM represents sperm motility; and SC represents sperm concentration. DGGE indicates denaturing gradient gel electrophoresis; UMT, Universiti Malaysia Terengganu.
Discussion
There are numerous studies of bacterial populations and their role in fish skin, gills, and the digestive tract, as well as internal organs such as the kidney, liver, and spleen.
20
To date, there are only a few reports on sperm-related bacteria, mainly from the perspective of biosafety, associated with the risk of pathogenic contamination during the semen preservation process.21–23 Loch and Faisal
24
suggested the possibility that fish sperm may be responsible for the transfer of fish pathogen, such as flavobacterial species, to fish egg during the process of fertilization. Similarly, in vitro insemination of cryopreserved sperm showed that bacteria present in cryopreserved sperm affected the fertilization capacity of sperm and was responsible for the bacterial transmission to embryos.
25
These results suggest the importance of revealing sperm microbiota composition and its relationship with sperm quality, as there is still a paucity of investigations on sperm microbiota. Boonthai et al
21
indicated that appropriate sperm collection methods could reduce sperm-related bacterial abundance; however, there was no significant difference in sperm quality. In this study, there was a correlation between bacterial diversity and sperm quality, and no known pathogenic bacteria, such as
There is only speculation on the role of intracellular bacteria on fish sperm, such as the novel theory that the bacteria Verrucomicrobia is responsible for the origin of the flagellum, as it contains tubulins, which are a prerequisite for flagellum formation.
27
Although spermatogenesis in fish is similar to that of other non-mammalian organisms, the control mechanisms are still unclear. In this study, sperm microbiota showed phylogenetic diversity, including Proteobacteria, Firmicutes, and Actinobacteria. Some bacteria, such as
In our study, the sequence result of unique DGGE band showed the presence of 1 bacterial clone (
This is the first report on Malaysian Mahseer (
Supplemental Material
supplemental_table_1_revision – Supplemental material for Molecular Characterization of Microbiota Associated With Sperm of Malaysian Mahseer Tor tambroides
Supplemental material, supplemental_table_1_revision for Molecular Characterization of Microbiota Associated With Sperm of Malaysian Mahseer Tor tambroides by Ivan Chong Chu Koh, Bin Hassan Badrul Nizam, Yazid Muhammad Abduh, Ambok Bolong Abol Munafi and Shumpei Iehata in Evolutionary Bioinformatics
Footnotes
Acknowledgements
We would like to thank Institute of Marine Biotechnology for allowing usage of the Decipher Denaturing Gradient Gel Electrophoresis machine.
Funding:
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research was supported financially by the Research Acculturation Grant Scheme (RAGS) vot 57120, Ministry of Higher Education, Malaysia.
Declaration of conflicting interests:
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Author Contributions
ICCK and SI designed the study, and SI wrote manuscript. BHBN conducted culture experiment and data analysis. ABAM and YMA provide and assisted in fish rearing. All authors read and approved the final manuscript.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
