Abstract
Objectives
Domestic cat hepadnavirus (DCHBV), belonging to the Orthohepadnavirus genus and closely related to the human hepatitis B virus (HBV), is detected in domestic cats; however, its disease progression and pathological impact remain unclear. This study investigates the longitudinal dynamics of DCHBV infection in naturally infected cats over a period of up to 310 days after detection, focusing on blood parameters and viral load fluctuations, liver pathology and extrahepatic dissemination.
Methods
Among 87 screened cats, four (4.6%) tested positive for DCHBV using quantitative PCR (qPCR). These cats were monitored longitudinally through repeated health checks, including viral load monitoring. After natural death, necropsy, histopathology, in situ hybridisation (ISH) and immunohistochemistry (IHC) were conducted to assess lesion distribution and viral localisation. Whole-genome sequencing and phylogenetic analysis were also performed on DCHBV-positive cases.
Results
Three of four DCHBV-positive cats exhibited persistent high viremia (>7 log10 genomic copies/ml) for over 6 months. Despite sustained viremia, liver enzyme levels showed variable trends, with some cases maintaining normal alanine aminotransferase and alkaline phosphatase levels. Histopathological analysis revealed various degrees of interface hepatitis, consistent with immune-mediated liver injury. DCHBV DNA was most abundant in the liver and confirmed through qPCR, ISH and IHC. Viral DNA was also detected in extrahepatic tissue, including the spleen, lung and salivary glands. Complete genome sequencing confirmed clustering within genotype A, with low genetic variability. Coinfection with feline leukaemia virus (FeLV) was noted in two cats, which may influence host immune responses.
Conclusions and relevance
Persistent viremia and liver inflammation suggest DCHBV may contribute to hepatic pathology, likely influenced by host immune responses and coinfections. However, because of the limited number of cases and the presence of FeLV coinfection in some cats, firm conclusions cannot be drawn. These findings provide a foundational basis that warrants confirmation and expansion in larger cohorts.
Keywords
Introduction
Domestic cat hepadnavirus (DCHBV) is a DNA virus classified within the Orthohepadnavirus genus, which also includes hepatitis B virus (HBV) in humans and woodchuck hepatitis B virus (WHV) in animals.1,2 Infections with HBV and WHV in their respective hosts are strongly associated with severe liver diseases, including hepatitis, cirrhosis and hepatocellular carcinoma (HCC). Although the clinical significance of DCHBV infection in domestic cats remains uncertain, emerging evidence indicates its potential association with various hepatic and extrahepatic manifestations.3,4 These include elevated liver enzyme levels – such as alanine transaminase (ALT), aspartate aminotransferase (AST) and alkaline phosphatase (ALKP) – as well as varying degrees of hepatopathy, hepatitis, HCC and extrahepatic conditions, such as lymphoma.4 –8
Although DCHBV shares virological and pathological characteristics with other Orthohepadnavirus members, its pathogenesis remains largely undefined. In humans, HBV infection can lead to acute hepatitis or progress to chronic hepatitis B (CHB), the latter being characterised by the persistent presence of hepatitis B surface antigen (HBsAg) for more than 6 months.9 –13 The progression of CHB is influenced by a complex and dynamic interplay between viral and host factors. Therefore, multiple clinical parameters – including ALT levels, hepatitis B e antigen (HBeAg) serostatus, HBV DNA levels, liver histology, HBV genotype, viral mutations and hepatitis B core-related antigen (HBcrAg) – are continuously monitored to assess disease progression and clinical outcomes.9,11
Given the similarities between HBV and DCHBV, it is important to explore whether DCHBV follows a comparable disease progression. Persistent DCHBV infection has been reported in domestic cats; however, evidence supporting chronic DCHBV infection remains inconclusive, as this observation is based on a single report with only one documented case of chronic viremia. 14 This finding suggests that DCHBV infection may follow distinct clinical phases and outcomes, akin to those observed in CHB. 13 However, because of the scarcity of longitudinal studies, the natural course of DCHBV infection remains poorly understood and warrants further investigation.
Chronic HBV infection in humans is also found to be associated with a range of systemic and extrahepatic complications, including vasculitis, non-rheumatoid and rheumatoid arthritis, neuropathies, glomerulopathies and non-Hodgkin’s lymphoma. 15 These manifestations are thought to arise from immune-mediated injury and direct viral invasion of non-hepatic tissues. 16 Like HBV, DCHBV has also been detected in extrahepatic tissues, including the lungs, kidneys, spleen, urinary bladder and lymph nodes of infected cats.4,17 In addition, DCHBV DNA has been identified in body fluid specimens, including those collected from auricular, nasal, oral and rectal swabs, and pleural effusion, suggesting possible haematogenous dissemination of the virus to extrahepatic sites.3,18
Despite attempts to understand DCHBV infection, most studies have been limited to single time-point assessments, and longitudinal studies remain scarce. To address this knowledge gap, the present study investigates the natural course of prolonged DCHBV infection in domestic cats by analysing viral genotypes, viral load and liver-associated pathogenesis. In addition, we examine viral localisation across multiple tissues in chronically infected cats to better characterise the extent of extrahepatic involvement. This study includes a longitudinal observation with multiple time-point determinations of viral loads and blood profiles until the infected cats succumbed to underlying diseases of unknown association with DCHBV infection. After their deaths, post-mortem examinations, histopathological assessments, and analyses of viral distribution and tissue localisation were conducted to further elucidate the pathological significance of DCHBV. These findings aim to provide deeper insights into the pathogenesis of DCHBV and its potential parallels with HBV infection in humans.
Materials and methods
Sample collection
EDTA- and heparinised-anticoagulated blood samples were initially collected from 87 shelter-housed cats in Bangkok, Thailand, with prior consent from their owners. These samples were first screened for DCHBV infection using molecular detection methods to determine the infection status.7,17,18 Haematological and blood chemistry analyses were also performed on the original samples. For DCHBV-positive cats, additional serial blood samples were subsequently collected at multiple time points, from day 0 (initial detection) to day 310, to monitor the progression of infection. Sampling was conducted by licensed veterinarians using a convenience-based approach. After the initial DCHBV diagnosis, the infected cats were individually isolated in designated sentinel areas within the same shelter facility. They were monitored under routine care by separate personnel, and were physically separated from both healthy and other infected animals to reduce potential cross-contamination. To further assess the impact of prolonged DCHBV infection, necropsy examinations were performed on DCHBV-positive cats that died during this observation at the Department of Pathology, Faculty of Veterinary Science, Chulalongkorn University. Fresh tissues, including the brain, heart, intestine, kidney, liver, lung, lymph nodes, salivary glands, spleen and urinary bladder, were collected and individually stored for molecular detection and pathological analysis. All procedures were reviewed and approved by the Institutional Biosafety Committee of Chulalongkorn University (IBC number 2331046) and the Chulalongkorn Institutional Animal Care and Use Committee (CU-IACUC number 2331075).
Viral nucleic acid extraction and selective molecular screening
Viral nucleic acids were extracted from 200 µl of blood or 5 g of collected tissue using the IndiSpin Pathogen Kit (QIAGEN), following the manufacturer’s instructions. The quality and quantity of extracted nucleic acids were assessed using a Nano UV/Vis Nano Spectrophotometer (Nabi; MicroDigital) at a 260:280 absorbance ratio. Extracted nucleic acids were stored at −80°C until further use. To investigate common feline viral infections that may directly or indirectly influence liver activities through persistent infection or systemic illness, 19 DCHBV-positive blood samples were further screened using diagnostic PCR assays for feline coronavirus (FCoV), feline herpesvirus (FHV), feline calicivirus (FCV) and for proviral DNA of feline leukaemia virus (FeLV) and feline immunodeficiency virus (FIV), as previously described.20 –22
DCHBV detection and quantification
DCHBV detection and quantification were performed using quantitative PCR (qPCR), following previously established protocols.7,17,18 The qPCR master mix was prepared using the KAPA SYBR FAST qPCR Master Mix (Kapa Biosystems), with 1 µl of DNA template and primers (DCHBVq-F and DCHBVq-R) targeting the overlapping open reading frames (ORFs) of the DCHBV polymerase and surface proteins. The qPCR thermocycling conditions included an initial denaturation at 95°C for 10 mins, followed by 35 cycles of denaturation at 95°C for 10 s and annealing/extension at 60°C for 20 s. For further analysis, DCHBV DNA load was categorised based on previous studies as low viral load (<5 log10 genomic copies [log10GC]/ml), high viral load (5–7 log10GC/ml) or very high viral load (>7 log10GC/ml). 18
DCHBV complete genome sequencing and phylogenetic analysis
Complete genome sequencing was performed on DCHBV-positive blood samples following established protocols.17,18 A 25 µl PCR master mix was prepared using GoTaq Green Master Mix (Promega), along with three different sets of primers (see Table S1 in the supplementary material). Conventional PCR (cPCR) was conducted using a thermocycler, with optimised annealing temperatures for each primer set. The obtained complete sequences were aligned and compared with available DCHBV sequences in GenBank (see Table S2 in the supplementary material). A colour-coded pairwise identity matrix was generated using the freely available Sequence Demarcation Tool (SDTv1.2). Phylogenetic analysis was performed using Molecular Evolutionary Genetic Analysis (MEGA) 11 software. A maximum likelihood phylogenetic tree was constructed with 1000 bootstrap replicates, employing the Tamura-Nei (TN93) model with gamma distribution and invariable sites (G+I). The best-fit model was selected based on the lowest Bayesian Information Criterion (BIC) score among candidate substitution models.
Haematological and blood chemistry analyses
For complete blood count (CBC) analysis, EDTA-anticoagulated blood specimens were processed using the Procyte Dx Hematology Analyzer (IDEXX Laboratories). For blood chemistry tests, heparinised-anticoagulated blood specimens were centrifuged at 3500 g for 5 mins to obtain plasma. The collected plasma was then analysed using the Catalyst One Chemistry Analyzer and Chem 10 CLIP (IDEXX Laboratories) to measure glucose, blood urea nitrogen (BUN), creatinine, ALT, ALKP, total protein (TP), albumin and globulin.
Histopathological examination and DCHBV localisation in tissues
To evaluate tissue morphology, histopathological examination was performed on formalin-fixed paraffin-embedded (FFPE) tissues from DCHBV-positive succumbed cats with haematoxylin and eosin (H&E) staining. Microscopic changes were assessed by an American-boarded veterinary pathologist (TK). In addition, DCHBV localisation was further assessed using two different assays, immunohistochemistry (IHC) and in situ hybridisation (ISH). Slides were independently analysed by board-certified veterinary pathologists (TK and ST). For ISH assay, a 230 bp non-radioactive digoxigenin (DIG)-labelled nucleotide probe was constructed using specific primers for DCHBV, based on previous studies.4,7 Hybridisation was performed on FFPE tissues after overnight incubation at 37°C. Positive signals were detected using an ALKP-labelled anti-DIG antibody (anti-DIG-AP Fab fragments; Roche Diagnostics) and a chromogenic substrate (PermaRed AP chromogen; Diagnostics BioSystems). A non-specific probe incubation using feline bocavirus-3 probe was used as a negative control to confirm specificity. 4
IHC assay for DCHBV localisation was conducted following a previously published protocol. 17 FFPE tissue sections were deparaffinised and treated with heat-induced antigen retrieval using citrate buffer (pH 6.0, 20 mins at 95°C). In the absence of a commercially available DCHBV antibody, the sections were incubated overnight at 4°C with a primary antibody against hepatitis B virus core antigen (HBcAg) (rabbit polyclonal anti-HBcAg B0586, 1:4,000; DAKO). The cross-reactivity of DCHBV core antigen and HBcAg has been confirmed and described in previous studies.17,23,24 After primary incubation, the slides were incubated with an anti-rabbit secondary antibody (DAKO REAL EnVision Detection System) for 1 h at room temperature. The chromogen 3,3’-diaminobenzidine (DAB) was applied, and positive signals were visualised under a light microscope. An isotype-matched control antibody (normal rabbit IgG antibody NI01; Sigma-Aldrich) was used to assess non-specific binding.
Results
DCHBV viral load and haematological profile during prolonged infection
Of the 87 screened cats, four (4.6%) tested positive for DCHBV using qPCR on day 0 and were designated as cases 1–4. All four cats were neutered and lived in the same shelter housing over 200 cats. Clinical signs observed before death in DCHBV-positive cats were non-specific and included mild to moderate anorexia and depression. Case 1 had a low DCHBV DNA load (4.33 log10GC/ml), whereas cases 2–4 exhibited very high viral loads in the range of 8.73–9.40 log10GC/ml (Table 1). Molecular screening for coinfections with other feline viruses revealed positive detection for FeLV provirus in cases 2 and 3, while cases 1 and 4 tested negative for all screened proviral retroviruses. No additional feline viruses were detected in any of the DCHBV-positive cats. Demographic characteristics, retroviral coinfection status and tentative cause of death based on necropsy findings for the four DCHBV-positive cats are summarised in Table 2.
Haematological and clinical chemistry parameters of cats during domestic cat hepadnavirus (DCHBV) infection

Numbers in bold represent abnormal values above or below the reference interval (RI)
Below the RI
Above the RI
ALKP = alkaline phosphatase; ALT = alanine transaminase; HBG = haemoglobin; HCT = haematocrit; N/A = not available; PCT = plateletcrit; PLT = platelets; RBCs = red blood cells; WBCs = white blood cells; – = negative result
Demographic characteristics, retroviral infection status and tentative cause of death in domestic cat hepadnavirus-positive cats

Diagnosis based on necropsy findings
DSH = domestic shorthair cat; FeLV = feline leukaemia virus
Longitudinal monitoring of viral load revealed persistent DCHBV viremia in cases 2–4, while case 1 tested positive only at the initial screening. Throughout the study period, case 2 was monitored at eight time points (days 0–280), exhibiting consistently high viremia with a mean genomic copy of 8.91 ± 0.52 log10GC/ml (range 8.25–9.90). Despite this, ALT levels remained largely within the normal range, except for a transient elevation on day 60. This cat showed persistent anaemia, thrombocytopenia and leukocytosis. ALKP levels remained persistently low throughout the observation period (Table 1).
Case 3 was monitored at six time points (days 0–120) before succumbing to illness. Viral loads remained very high, at a mean of 9.17 ± 0.32 log10GC/ml (range 8.85–9.77). Haematological abnormalities included persistent anaemia and thrombocytopenia, while the blood chemistry profile revealed elevated ALT levels during the first half of the study, followed by a return to normal values in the latter half. ALKP levels showed no significant variation (Table 1).
Case 4, which was monitored at six time points (days 0–310), exhibited persistently high viremia (mean 9.14 ± 0.41 log10GC/ml; range 8.73–9.88) before succumbing shortly after the final monitoring. Haematological analysis revealed persistent anaemia, leukocytosis and thrombocytopenia. The blood chemistry profile demonstrated dynamic fluctuations in ALT levels, closely paralleling DCHBV DNA load. Between days 0 and 60, both ALT levels and viral load increased, although ALT remained within the normal range. Between days 60 and 180, both parameters exhibited a slight decline, followed by a significant drop on day 220. By day 310, ALT levels increased sharply, exceeding the normal reference interval, alongside an increase in DCHBV DNA load. In contrast, ALKP levels remained consistently low in all monitored cats (Table 1).
DCHBV detection and localisation in various tissues
DCHBV DNA was detected in liver tissue of all four cases. In case 1, DCHBV DNA was found exclusively in the liver tissue with DNA level of 2.90 log10GC/g, with all other organs testing negative (Table 3). In case 2, DCHBV DNA was detected in the heart, intestine, kidney, liver, lung, lymph nodes and spleen, while brain tissue tested negative. The highest viral load was detected in the liver (6.86 log10GC/g), whereas the lowest was found in the intestine (3.92 log10GC/g). Cases 3 and 4 exhibited widespread tissue positivity, including the brain. The liver again harboured the highest viral load (8.35 log10GC/g in case 3, 8.48 log10GC/g in case 4), while the brain had the lowest but detectable viral loads (6.24 log10GC/g in case 3, 6.79 log10GC/g in case 4) (Table 3).
Detection and quantification of domestic cat hepadnavirus (DCHBV) DNA in various tissues

N/A = not available; – = negative result
Histopathological investigation revealed multifocal lymphoplasmacytic portal-to-interface hepatitis with centrilobular hepatocellular degeneration as the prominent pathological change in liver sections, with varying degrees of severity observed across all investigated cases (Figure 1a). Within the liver sections of cases 2 and 4, several portal tracts were surrounded by variably dense infiltrates of predominantly small lymphocytes and fewer plasma cells, which often extended beyond the limiting plate and dissected around the surrounding hepatocytes, along with rare individualised, hypereosinophilic hepatocytes consistent with acidophils (Figure 1b). Centrilobular hepatocytes were multifocally swollen with variable cytoplasmic vacuolation and scant accumulations of granular to globular yellow-brown bile or lipofuscin pigment. Scant aggregates of homogeneous yellow bile were noted within occasional canaliculi in some areas. In addition, emperipolesis of mononuclear cells was observed in several hepatocytes, predominantly near the portal areas. Similar lesions were also present in case 1, albeit with lesser severity.

Domestic cat hepadnavirus (DCHBV) infection. Liver. (a) Multifocal lymphoplasmacytic inflammation (arrows) extending from the portal regions to the surrounding liver tissue, accompanied by centrilobular hepatocellular degeneration. Haematoxylin and eosin (H&E). (b) Interface hepatitis, characterised by lymphocyte and plasma cell infiltration surrounding a portal tract, breaching the limiting plate and extending into adjacent hepatocytes. An acidophil body is also observed (inset). H&E. (c) Immunohistochemistry (IHC) revealing DCHBV immunolabelling (brown) localised in the nuclei of hepatocytes. (d) Cytoplasmic expression of DCHBV antigens is also detected (inset). IHC. (e) In situ hybridisation (ISH) demonstrating rare DCHBV DNA hybridisation (red) within the nucleus of a hepatocyte. (f) DCHBV DNA hybridisation detected in the nucleus of infiltrating lymphocytes within a region of portal hepatitis. ISH. Scale bars: 50 µm
Within case 3, hepatocytes were diffusely disorganised and dissociated, resulting in diffuse individualisation. Mild to moderate infiltrates of lymphocytes, plasma cells and variable numbers of neutrophils were observed surrounding bile ducts within the portal regions, occasionally extending to and dissecting around adjacent hepatocytes. The lumens of some bile ducts contained variable aggregates of neutrophils and karyorrhectic debris or scant accumulations of eosinophilic proteinaceous to slightly yellow-brown homogeneous material. In addition, small random foci of hepatocellular loss were observed, accompanied by aggregates of karyorrhectic debris, neutrophils and fewer lymphocytes. In the centrilobular regions, variable numbers of hepatocytes contained scant amounts of yellow-brown granular to globular pigment, consistent with bile or lipofuscin. Several Kupffer cells were reactive and laden with the same yellow-brown pigment. Throughout all liver sections from case 4, there were high numbers of circulating mononuclear cells, suspected to be small lymphocytes or extramedullary haematopoietic precursors. Variable numbers of central veins were thickened by thin to moderately thick bands of fibrosis. Additional post-mortem findings and tentative causes of death for the DCHBV-positive cases – namely hypertrophic cardiomyopathy with pneumonia, chronic renal disease and renal lymphoma in cases 1 to 3, respectively – are summarised in Table 2.
Both IHC and ISH assays were performed on liver tissues from cases 2 and 4. Despite the high DCHBV DNA load, immunostaining levels were consistently low across all investigated cats. DCHBV localisation was observed in the cytoplasm and nuclei of scattered hepatocytes (Figure 1c,d). This was confirmed by ISH, which detected low levels of viral signals in both the nuclei and cytoplasm of hepatocytes in both cases (Figure 1e,f). In the spleen, both ISH and IHC assays demonstrated widespread DCHBV localisation, predominantly within the nuclei of cells in the red pulp of case 2 (Figure 2a,b). In the lungs, although ISH did not detect viral presence, IHC revealed strong DCHBV immunoreactivity in the bronchial and bronchiolar epithelial cells of case 2 (Figure 2c). In addition, focal IHC staining was observed in the endothelial lining of blood vessels within the salivary glands of case 4 (Figure 2d). No positive signals were detected in other examined organs by either ISH or IHC. As expected, neither DNA hybridisation nor immunoreaction was observed in the negative control section (see Figure S1 in the supplementary material).
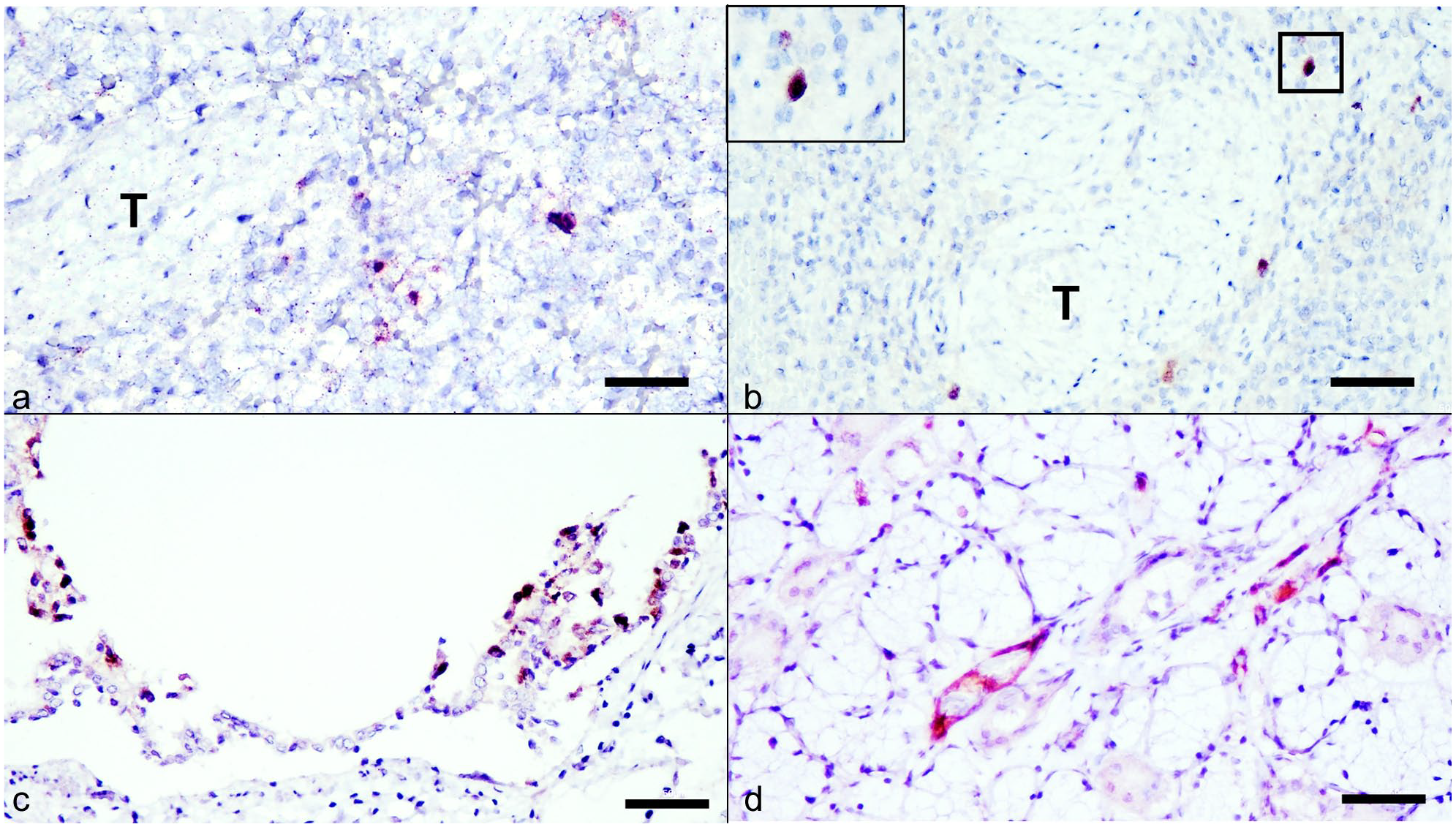
Domestic cat hepadnavirus (DCHBV) infection. Extrahepatic tissues. (a) Nuclear hybridisation (red) indicating DCHBV DNA within inflammatory mononuclear cells in the spleen. In situ hybridisation (ISH). (b) DCHBV antigen (brown) detected in the nucleus of mononuclear inflammatory cells located at a splenic trabecula. Immunohistochemistry (IHC). (c) DCHBV antigens (brown) observed in the cytoplasm of bronchial epithelial cells in the lung. IHC. (d) DCHBV localisation detected in a spindle-shaped cell, suspected to be an endothelial cell of a small capillary, within the parotid salivary gland. IHC. Scale bars: 50 µm. T = trabecula
Complete genome and phylogenetic analyses
Complete 3184-bp genome sequences were successfully obtained from cases 2–4 and designated as DCHBV CV-3/2023, DCHBV CV-5/2023 and DCHBV SH-48/2023 (GenBank accession numbers PV392816–PV392818). Pairwise identity analysis using SDT software confirmed a high genetic similarity (>99%) among these sequences. They also shared a 95–98% identity with previously reported DCHBV genomes in GenBank, except for DCHBV Rara/2022 (Accession number LC685967.1), which showed a notably lower identity (<90%), suggesting greater genetic divergence (Figure 3). Phylogenetic analysis confirmed that all three sequences clustered within genotype A, consistent with previous classifications. In contrast, DCHBV Rara/2022 (Accession number LC685967.1) from Japan formed a separate branch within genotype B, indicating significant genetic divergence from other known DCHBV strains. The phylogenetic tree further showed that the sequences retrieved in this study clustered within the A1 clade, closely related to previously identified DCHBV isolates from Thailand (Accession numbers OQ362114, OQ362110, MT506041, MT506046, MT506039M and OQ362106), as well as isolates from Hong Kong, Italy, Australia, Turkey and the USA. In contrast, other Thai isolates formed a distinct sister clade (A2), suggesting potential intra-genotypic diversity within genotype A (Figure 4). The combined results from pairwise identity analysis and phylogenetic reconstruction confirm that DCHBV sequences retrieved from cats with prolonged infection are genetically consistent with previously reported genotype A isolates. This reinforces their classification within this lineage and suggests a high degree of conservation among circulating strains.

The Sequence Demarcation Tool colour-coded pairwise identity matrix of domestic cat hepadnavirus (DCHBV) sequences. Sequences obtained in this study (highlighted within the red box) are compared with all complete DCHBV genome sequences available in the GenBank database. Each coloured cell represents the pairwise identity between sequences, with the identity score key displayed in the top-right corner of the figure

Phylogenetic tree of domestic cat hepadnavirus (DCHBV) based on the complete genome. The maximum likelihood tree is constructed using the Tawamura–Nei model, with a gamma distribution and invariable sites and analysed through bootstrap analysis with 1000 replications. Sequences retrieved in this study are indicated by red triangles ( ), while other isolates from Thailand are labelled with blue triangles (
), while other isolates from Thailand are labelled with blue triangles ( ) and rectangles (
) and rectangles ( ). The scale bar indicates the rate of nucleotide substitution
). The scale bar indicates the rate of nucleotide substitution
Discussion
DCHBV has been detected worldwide; however, understanding of its pathology remains a limitation.5,6,17,25 –28 Previous studies have reported a range of clinical outcomes from elevated liver enzymes and hepatopathies to asymptomatic infections.7,14,25,26 Similarly, in humans, HBV infection presents diverse manifestations influenced by viral genotype, viral load and host immune responses.11,29 –32 However, information on DCHBV genotypes, viral load dynamics and liver-associated effects during prolonged infection remains poorly understood. This study addresses these gaps by conducting a longitudinal assessment of viral load fluctuations, blood parameters and tissue localisation in DCHBV-infected cats.
Among 87 tested cats, four were DCHBV-positive, with one case (case 1) exhibiting only transient viremia and three cases (cases 2–4) displaying persistent infections. High initial DCHBV DNA loads were associated with prolonged viremia, whereas the cat with a low initial viral load tested negative in subsequent screenings. These findings suggest that a high initial DCHBV viral load may serve as a predictor for persistent infection, mirroring HBV infection, where viral load at onset influences disease progression.9,33,34 Although persistently high DCHBV DNA levels were observed in prolonged infections, these high DNA loads were not consistently coupled with increased ALT or ALKP levels, indicating that DCHBV infection alone may not always induce liver enzyme abnormalities. Notably, the presence of normal ALT levels despite high viral loads, particularly in case 2, echoes the immune-tolerant phase of chronic HBV, characterised by active viral replication with minimal liver damage.35,36 However, transient ALT elevations in case 3, particularly in an FeLV-positive cat, suggest that coinfections or immune activation may contribute more to liver enzyme alterations than viral load alone. Given that FeLV suppresses immune function, its presence may influence DCHBV infection dynamics.
In case 3, multifocal hepatitis was identified as the principal lesion at necropsy. The presence of DCHBV DNA within affected hepatocytes, confirmed by ISH, supports the virus’s direct involvement in the lesion. Nonetheless, FeLV coinfection – known to cause immunosuppression and potentially contribute to hepatic changes – may have acted synergistically or independently. Although a definitive cause of death could not be established, these findings highlight the potential for DCHBV to contribute to liver pathology, even in the context of coinfection. However, without direct measures of immune function, such interactions remain speculative. Furthermore, haematological abnormalities, such as anaemia, thrombocytopenia and leukocytosis, were observed in some DCHBV-positive cats, particularly those coinfected with FeLV. These findings are non-specific and may reflect a range of underlying processes, including retroviral-induced myelosuppression, immune activation or chronic inflammation. 37
Although a direct causal role of DCHBV in these changes cannot be established, their presence underscores the complexity of interpreting clinical parameters in coinfected animals. Future studies in FeLV/FIV-negative cats are necessary to better define DCHBV-specific haematological effects. Further studies are needed to determine whether immunosuppression affects viral persistence or liver pathology, as seen in HBV–HIV coinfections in humans.38,39 In HBV infection, viral persistence is influenced not only by viral load but also by host immune responses. 11 Interestingly, although case 4 generally exhibited normal ALT levels throughout the observation period, a consistent parallel trend between DCHBV DNA load and fluctuations in ALT was observed. It is also worth noting that in case 4, no coinfection with other feline viruses, including FIV and FeLV, was detected, making the observed parallelism between ALT dynamics and DCHBV DNA load particularly intriguing. Although an intriguing temporal pattern between ALT levels and viral load was noted in case 4, the findings are limited to a single animal and cannot support any statistically valid conclusions. Furthermore, in case 1, despite the absence of sustained DCHBV viraemia, mild hepatic changes were observed at necropsy. Elevated leukocyte counts before death may be attributable to pneumonia or stress-associated inflammatory responses, as confirmed by post-mortem findings.
The absence of liver enzyme abnormalities in some persistently infected cats suggests a possible immune-tolerant state, whereas ALT fluctuations in other cases may indicate immune-mediated liver injury. Given the critical role of host immunity in HBV-associated liver disease, future studies assessing immune parameters in DCHBV-infected cats could provide valuable insights into the mechanisms of liver injury and disease progression. In addition, the transient viraemia observed in case 1 raises the question of whether DCHBV can establish occult infection, similar to HBV, where viral DNA persists in hepatocytes despite undetectable viremia.11,40 The potential persistence of DCHBV in tissue reservoirs, even when blood viral loads become undetectable, remains an open question. Future studies incorporating liver tissue analysis could help clarify this possibility.
The lesions observed in DCHBV-positive cases in this study, particularly interface hepatitis, align with findings from previous studies on DCHBV infection in domestic cats.7,25 Similar hepatic changes, characterised by inflammatory infiltration at the interface between hepatocytes and portal tracts, have been reported in naturally infected cats, suggesting a potential role of DCHBV in liver pathology.7,24,25 Notably, interface hepatitis is a well-documented histopathological feature in HBV-infected humans, where it is considered a marker of immune-mediated liver injury and is associated with progression of chronic hepatitis.41,42 The presence of similar lesions in DCHBV-infected cats highlights the need for further investigation into the immunopathogenesis of DCHBV and its long-term hepatic consequences.
DCHBV DNA was most consistently detected in the liver, reinforcing hepatic tropism. However, extrahepatic distribution – particularly in the spleen, lungs and salivary glands – was also observed, consistent with previous reports.3,17 These findings, supported by qPCR, IHC and ISH, suggest systemic dissemination. Notably, the discrepancy between high DNA loads and low IHC/ISH signals may reflect differences in the molecular targets detected by each assay. The primers used in the qPCR assay amplify a region overlapping the polymerase and surface ORFs and thus may detect surface protein-encoding DNA that persists from earlier replication events. In hepadnaviruses, viral surface proteins can be released independently as incomplete viral particles, and their expression often remains stable within tissues. 43 In contrast, the IHC assay utilised an antibody targeting the DCHBV core antigen, which is more indicative of active viral replication. Therefore, the discordance between qPCR and IHC/ISH findings may reflect prior viral activity followed by transcriptional dormancy, particularly in extrahepatic tissues, similar to what has been observed in HBV infection of non-hepatic reservoirs such as the kidney and lymphoid tissue. 44
The highest viral burden was consistently observed in the liver, reinforcing its role as the primary site of viral replication. However, the detection of DCHBV DNA in multiple extrahepatic tissues suggests a potential role for systemic dissemination.3,4,17 The presence of DCHBV in the endothelial cells of the salivary glands, despite the absence of pathological lesions, raises questions about whether the virus found in saliva originates from infected salivary glands, and if infectious saliva could facilitate viral transmission, similar to HBV in infected patients. 45 Alternatively, the virus may simply disseminate to the salivary glands without contributing to transmission. Although tissue damage was not prominent in extrahepatic organs, the presence of viral nucleic acids and antigens raises questions about possible long-term consequences or transmission potential via body fluids. Despite minor intra-genotypic differences, no clear correlation was found between sequence variation and clinical outcomes. This suggests that host factors may play a larger role than genotype in determining disease course, contrasting with HBV,29,33,34 where genotypes often influence progression and treatment response.
Although this study provides insights into DCHBV persistence, viral load dynamics and tissue distribution, interpretation of the infection phase remains limited because of the current lack of DCHBV-specific serological markers – tools that are well established for staging HBV infection but not yet available for DCHBV.34,46,47 In addition, although this study followed a small cohort of cats longitudinally, a larger sample size is needed to fully characterise disease progression. Future research should focus on developing serological assays, investigating immune response dynamics and assessing long-term liver pathology to further clarify the clinical implications of DCHBV infection. It is important to acknowledge that 2/4 DCHBV-positive cats were coinfected with FeLV, complicating the attribution of pathological findings exclusively to DCHBV. FeLV is known to induce immunosuppression, haematological abnormalities and immunopathological reactions in various organs, potentially influencing the course of concurrent infections. 37 However, this study was designed as an observational analysis of naturally occurring infections in a shelter population, where retroviral coinfection is common. Although recent studies report FeLV prevalence rates in Thailand to be in the range of 4.2–16.5% depending on the cat population and health status,48,49 conducting longitudinal studies on DCHBV monoinfected cats remains challenging. Extended follow-up requires repeated sampling, consistent care and controlled housing, which are difficult to maintain in typical shelter settings. In addition, naturally infected cats may develop unrelated diseases over time, complicating the interpretation of DCHBV-specific effects. Furthermore, several studies have reported DCHBV detection in cats also infected with FeLV or FIV,4,7,26 indicating that coinfection is not uncommon and may reflect overlapping risk factors or host susceptibilities. Nonetheless, the detection of persistent DCHBV viremia and liver pathology across multiple animals – including at least one without retroviral infection – supports a potential role for DCHBV in chronic liver changes. Future controlled studies in FeLV/FIV-negative cats will be essential to confirm these findings and fully clarify DCHBV-specific effects. Furthermore, it is also important to note that coinfection screening in this study was limited to blood samples, which may have reduced sensitivity for detecting certain viruses such as FCoV and FCV, particularly in non-viremic animals.
Given that some investigated cats were coinfected with FeLV, the potential contributory role of DCHBV as a coinfection should be considered when interpreting the findings. In addition, as observed in previous studies,5,17,26 coinfection with retroviruses in DCHBV-infected cats parallels the interaction between HBV and HIV in humans.38,39 This suggests that DCHBV may share similarities with retroviral infection dynamics in cats, highlighting the need for further investigation into the potential impact of coinfection and its clinical significance.
Conclusions
This study demonstrates that DCHBV infection in domestic cats can result in persistent viremia and interface hepatitis, suggesting a potential role in chronic liver pathology. Although high viral loads were observed in several cases, liver enzyme levels varied, indicating that viral replication alone may not directly correlate with hepatic dysfunction. DCHBV DNA was consistently detected in the liver and occasionally in extrahepatic tissues, supporting systemic dissemination, although active replication appeared largely confined to hepatic tissue. Notably, FeLV coinfection was present in 2/4 DCHBV-positive cats, limiting the ability to attribute clinical and pathological findings solely to DCHBV. Although this study highlights associations between persistent DCHBV infection and hepatic changes, the small number of cases and presence of comorbidities in most cats preclude definitive conclusions about DCHBV pathogenicity. Future studies specifically designed to investigate DCHBV monoinfected cats, particularly those free of FeLV/FIV and other severe conditions, will be essential to clarify the independent pathogenic potential of DCHBV. Despite this limitation, our findings provide foundational insights into DCHBV infection dynamics in naturally infected cats and suggest its relevance as a feline hepatotropic virus with possible parallels to chronic HBV infection in humans.
Supplemental Material
Table S1
Primer sets used to obtain the complete genome of domestic cat hepadnavirus.
Supplemental Material
Table S2
Domestic cat hepadnavirus sequences from this study and reference sequences used to construct phylogenetic tree.
Supplemental Material
Figure S1
Domestic cat hepadnavirus (DCHBV) infections of DCHBV immunohistochemistry (IHC) and in situ hybridisation (ISH).
Footnotes
Acknowledgements
SWW and CP are supported by the Second Century Fund (C2F), Chulalongkorn University.
Supplementary material
The following files are available as supplementary material:
Conflict of interest
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This research project is supported by the Thailand Science Research and Innovation Fund at Chulalongkorn University (HEA_FF_68_051_3100_009 to ST).
Ethical approval
The work described in this manuscript involved the use of non-experimental (owned or unowned) animals. Established internationally recognised high standards (‘best practice’) of veterinary clinical care for the individual patient were always followed and/or this work involved the use of cadavers. Ethical approval from a committee was therefore not specifically required for publication in JFMS. Although not required, where ethical approval was still obtained, it is stated in the manuscript.
Informed consent
Informed consent (verbal or written) was obtained from the owner or legal custodian of all animal(s) described in this work (experimental or non-experimental animals, including cadavers, tissues and samples) for all procedure(s) undertaken (prospective or retrospective studies). No animals or people are identifiable within this publication, and therefore additional informed consent for publication was not required.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
