Abstract
Objectives
Autosomal dominant polycystic kidney disease (ADPKD), the most frequently diagnosed hereditary disease affecting Persian cats, is caused by a cytosine-to-adenine transversion (10063C>A) in PKD1, the gene that codes for polycystin-1. The objective of this study was to provide a preliminary estimate of the frequency of the pathogenic 10063C>A single nucleotide polymorphism (SNP) of PKD1 in Persian and Persian-related cat breeds in western Mexico.
Methods
Blood samples were collected from 104 cats (89 Persian, seven Persian crossbreed, five Siamese and three Himalayan cats). Genotyping was performed with our proposed PCR restriction fragment length polymorphism (RFLP) assay, as well as a previously established PCR-RFLP method for validation. The genotypes of control cats were corroborated by a commercial veterinary genetics laboratory.
Results
Our proposed PCR-RFLP assay and the validated PCR-RFLP methodology indicated that 24/104 (23.1%) cats in this study were heterozygous carriers of the 10063C>A SNP, including 23/89 Persian cats (25.8%) and 1/7 Persian crossbreed cats (14.3%). No Siamese or Himalayan cats were carriers. There were no discrepancies between the results obtained with our proposed assay and those obtained with the validation method or with commercial laboratory results.
Conclusions and relevance
The carrier frequency of the PKD1 10063C>A SNP in Persian and Persian-related cat breeds in western Mexico was found to be 23.1%. ADPKD frequencies among cat populations in Mexico have not been published previously. Genotyping assays can be used to facilitate the selection of breeding stocks by local breeders and veterinarians to avoid propagation of ADPKD.
Keywords
Introduction
Autosomal dominant polycystic kidney disease (ADPKD) is a common hereditary disorder in Persian and Persian-related cat breeds, with reported estimates of ADPKD prevalence in these breeds often exceeding 40%.1–5 Affected cats develop cysts in the renal parenchyma, 6 and exhibit clinical signs, including vomiting, anorexia, fatigue, polyuria and polydipsia. 4 The average age at onset of chronic kidney disease in affected cats is 7 years. 7 ADPKD pathogenesis has been linked to a cytosine-to-adenine transversion in position 10,063 of exon 29 of PKD1, a gene that codes for polycystin-1 and is located within chromosome E3. This single nucleotide polymorphism (SNP) generates a termination codon at amino acid 3284, resulting in the production of a truncated and non-functional protein. 8
Prior to characterization of the 10063C>A SNP in PKD1 and its association with ADPKD, the disease was diagnosed by ultrasound examination. However, cysts must grow to a volume large enough to be detected before a diagnosis can be made. 9 This limitation can delay diagnosis, and such cysts are not detected reliably before a cat has reached 10 months of age. 1
Earlier diagnosis of ADPKD is possible with molecular biology techniques, including PCR sequencing, PCR restriction fragment length polymorphism (RFLP) and real-time quantitative PCR.8,10,11 However, false negatives have been reported when the primers originally designed for detecting the 10063C>A SNP are used with other molecular techniques.6,11 To address these false negatives, multiple primer pairs have been designed to corroborate and optimize molecular testing results. 12 In the PCR-RFLP technique, a restriction enzyme is used to introduce a double-stranded break in a specific nucleotide sequence of amplified DNA segments, and the sizes of the fragments produced differ depending on genotype. Thus, separation of the fragment products by size via gel electrophoresis and subsequent observation of the gel under ultraviolet light can reveal the PKD1 genotype. Notably, because PCR-RFLP does not require costly reaction equipment, it is accessible to laboratories worldwide.
The objective of the present study was to provide a preliminary estimate of the frequency of the pathogenic 10063C>A PKD1 SNP using a PCR-RFLP assay in Persian and Persian-related cat breeds in western Mexico.
Materials and methods
Study population and sampling
The study population consisted of 104 cats (51 females, 53 males), including 89 Persian, seven Persian crossbreed, five Siamese and three Himalayan cats. Authorization and verbal consent were obtained from the cats’ owners prior to sample collection. Blood samples were taken from the jugular vein and placed directly into EDTA-coated Vacutainer tubes. The study was approved by the Internal Bioethics Regulation of the Biological and Agricultural Sciences Center of the University of Guadalajara, Mexico (Centro Universitario de Ciencias Biológicas y Agropecuarias, Universidad de Guadalajara, México; approval number CC/NN11-12/00/2012).
DNA extraction and genotyping
DNA was extracted from blood with Quick-DNA Universal Kits (Zymo Research). To identify the PKD1 10063C>A SNP, specific forward (5′-TTTGACAAGCATCTCTGGCTCTC-3′) and reverse primers (5′-AGAAACAGAATGACCAGGTAGACG-3′) were designed from the E3 chromosome sequence in GenBank (CM001393.3). The amplicon produced with this primer pair has a length of 346 base pairs (bp). PCRs were performed with DreamTaq polymerase (Thermo-Fisher Scientific), each in a reaction volume of 20 µl containing blood DNA lysate (~100 ng), 2 µl 1× PCR buffer containing 20 mM MgCl2, 1 µl 10 mM dNTP mix, 0.5 µl DNA polymerase, 5 pmol of each primer and double distilled water for the remaining volume. The PKD1 fragments obtained were amplified with a Techne TC-5000 Thermal Cycler with the following program: 5 mins at 95°C, thirty-five 20 s cycles at 94°C, 30 s at 62°C, 30 s at 72°C and a final 5 mins extension step at 72°C. After amplification, the samples were digested with the restriction enzyme MLY1 according to the manufacturer’s protocol (New England Biolabs), separated in a 4% agarose gel and then stained with GelRed (Biotium) intercalating dye to enable visualization of bands under ultraviolet light. Homozygosity for the wildtype PDK1 allele was identified by the presence of an undigested 346 bp band. PDK1 heterozygosity for the 10063C>A SNP was recognized by the presence of 89 bp and 257 bp fragments of the SNP allele, together with the aforementioned 346 bp band.
For corroboration, all samples were also genotyped by the PCR methodology described by Lyons et al (primers: 5′-CAGGTAGACGGGATAGACGA-3′ and 5′TTCTTCCTGGTCAACGACTG-3′). 8 The amplification conditions were 94°C for 4 mins, thirty 30 s cycles at 94°C, a 30 s annealing step at 58°C, 30 s at 72°C and a final 10 mins extension step at 72°C. The products were digested by the restriction enzyme MLY1, as described previously. 4 Genotype assignments were further corroborated by the Veterinary Genetics Laboratory at University of California, Davis.
Results
Genotyping for the 10063C>A SNP of PKD1 was completed for 104 cats with our proposed PCR-RFLP assay (Figure 1). The wild-type PKD1 allele was readily detected as an undigested 346 bp band. The genotypes, allele frequencies and percentages of cats affected overall and by breed are reported in Table 1. About 1/4 Persian cats, about 1/6 Persian crossbreed cats and no Siamese or Himalayan cats were found to be 10063C>A SNP allele carriers. All genotypes obtained using our PCR-RFLP assay were confirmed with the established validation methodology. The results obtained by a commercial veterinary genetics laboratory confirmed our results.
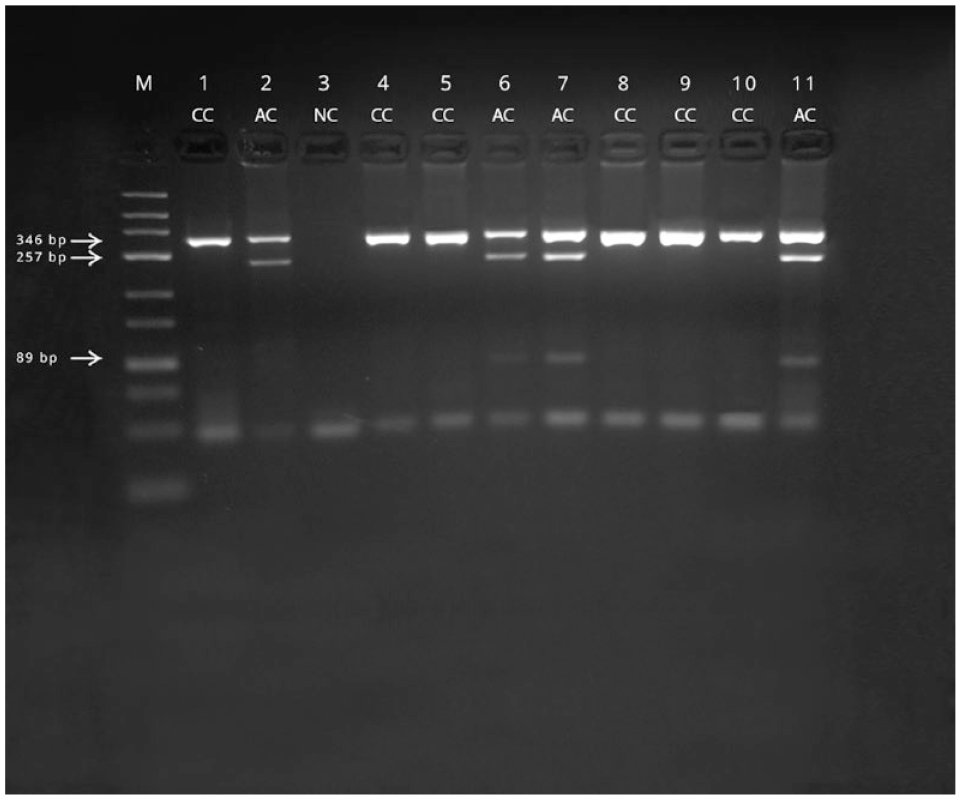
Agarose gel electrophoresis (4%) of PKD1 fragments produced by restriction enzyme MLY1 digestion. Lanes 1 and 2: controls for each genotype (AC and CC). Lane 3: negative control (NC). Lanes 6, 7 and 11: cats carrying 10063C>A SNP. Lanes 4, 5, 8, 9 and 10: cats homozygous for wildtype allele. Arrows indicate the sizes of the separated restriction fragments.
Genotype and allele frequencies for the PKD1 10063C>A single nucleotide polymorphism (SNP)

Heterozygous 10063C>A SNP carriers were identified by the additional presence of the enzymatic digestion-yielded 89 bp and 257 bp fragments of the SNP allele.
Discussion
Using molecular PKD1 genotyping, this study indicated that 23.1% of the sampled cat population in western Mexico were 10063C>A SNP carriers, including 25.8% of Persian cats and 14.3% of Persian crossbreed cats. This study provides the first report of the frequency of this mutation in cats from this region. These frequencies are higher than those reported recently in Turkey (Persian cats: 17.5%; Persian-related cats [Scottish Fold, Exotic Shorthair and British Shorthair]: 2.7%) 4 but lower than frequencies reported in Iran (46.8%), 3 Australia (43%), 1 France (41.8%), 2 Japan (40%), 5 Italy (37.1%) 6 and the UK (27.5%). 10 Although the frequency reported in the Japanese study was likely inflated owing to a large number of ill cats being sampled, the values reported for Iran, Italy and the UK did not have preliminary diagnoses, suggesting that the PKD1 10063C>A SNP is still being propagated in these countries, despite their having access to molecular tests. The high frequencies reported in the Australian and French studies occurred before genetic testing was available and hence the prevalence and incidence of the disease should be lower. Allele frequencies may be declining in Brazil. A study carried out in Brazil in 2020 yielded frequencies of 5.03% and 1.62% for Persian and Persian-related cat breeds, respectively (4.54% overall), 13 representing sharp decreases compared with a 2014 Brazilian study, 14 which yielded frequencies of 33% and 7%, respectively. This decrease may reflect the breed stock selection effect of molecular assays. In Taiwan, molecular assays have been used in addition to ultrasonography diagnostics for more than a decade to pre-emptively identify carrier kittens. 11 Hence, there remains a need to increase awareness of the importance of PKD1 genotyping among cat owners and breeders to exclude carrier cats from breeding populations.
A number of molecular diagnostic methodologies have been employed for identification of the PKD1 10063C>A SNP allele. Lyons et al have demonstrated consistent identification results with their PCR-RFLP methodology.5,8,13,14 Compared with real-time quantitative PCR, PCR-RFLP is less expensive and easier to implement, which should facilitate greater use. Although diagnostic ultrasonography offers high sensitivity after cyst formation in adult cats, it is not reliable in cats younger than 10 months old. 1 Molecular testing can identify affected kittens as early as the first day of life, thus enabling breeders to better control and eradicate ADPKD from cat populations.
Conclusions
The present data provide a preliminary carrier frequency estimate of 23.1% for the PKD1 10063C>A SNP in Persian and Persian-related cat breeds in western Mexico. To the best of our knowledge, this is the first ADPKD frequency estimate to be reported for cats in Mexico.
Footnotes
Conflict of interest
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This work was supported by the University of Guadalajara through the Research Development project of the Department of Animal Production (grant number 263719).
Ethical approval
The work described in this manuscript involved the use of non-experimental (owned or unowned) animals. Established internationally recognized high standards (‘best practice’) of veterinary clinical care for the individual patient were always followed and/or this work involved the use of cadavers. Ethical approval from a committee was therefore not specifically required for publication in JFMS. Although not required, where ethical approval was still obtained, it is stated in the manuscript.
Informed consent
Informed consent (verbal or written) was obtained from the owner or legal custodian of all animal(s) described in this work (experimental or non-experimental animals, including cadavers) for all procedure(s) undertaken (prospective or retrospective studies). No animals or people are identifiable within this publication, and therefore additional informed consent for publication was not required.
