Abstract
Per- and polyfluoroalkylated substances (PFAS) are persistent anthropogenic chemicals widely distributed in the environment that are known to have toxic effects in animals and humans following exposure. Some PFAS have been shown to activate peroxisome proliferator-activated receptor α (PPARα), a transcription factor involved in lipid metabolism, leading to dyslipidemia or liver toxicity. PFAS comprise a wide range of compounds, and variations in their structural characteristics could reveal important details regarding the level of PPARα activation. In this work, using a Chemically Activated LUciferase eXpression (CALUX) assay, we experimentally tested the PPARα activation efficiency of several PFAS compounds of varying chain lengths and functional groups. Activation and potency were compared across and within PFAS class based on chemical differences. When compounds with the same number of carbons or perfluorinated carbons were compared across class, the rank from high to low activator class remained the same. Perfluorocarboxylated ether was found to be the strongest class, while polyfluorotelomer was the weakest, suggesting the importance of structural features in PPARa activation. Perfluorocarboxylates were consistently better PPARα activators than perfluorosulfonates. Comparing within these 2 classes, the number of perfluorinated carbon atoms better predicted activation than the number of carbon atoms. In the perfluorocarboxylated ether, perfluorocarboxylate, and perfluorosulfonate classes, a direct correlation existed between potency and the percentage of PPARα activation (R2 = 0.702), a novel observation. These findings provide new insights regarding distinct chemical characteristics of PFAS compounds which may be predictive of PPARα activation level.
Introduction
More than 14,000 distinct PFAS have been identified to date, and many are ubiquitous due to prolonged biological and environmental half-lives and widespread usage in many industrial products. 1 PFAS have been reported to have toxic effects in animals,2–4 and their epidemiologic association with human health endpoints5,6 is a significant public health concern resulting in restrictions and the phased-out usage of legacy PFAS by industrial groups.7,8
PFAS toxicity may involve the activation of PPARα,9–12 a nuclear receptor isoform expressed in metabolically active cells which coordinates the regulation of a broad array of genes. 13 PFAS were reported to activate mouse and human PPARα in reporter gene assays.14,15 Activation by PFAS occurs in human primary hepatocytes,16,17 and in mouse models the alteration of hepatic gene expression is regulated primarily by PPARα. 18 PPARα plays a critical role in lipid homeostasis, inducing the expression of numerous genes involved in lipid metabolic pathways.19,20
Multiple studies have linked adverse PFAS effects to PPARα activation. Hari et al. showed that the up-regulation of PPARα genes involved in lipid metabolism by PFAS leads to liver steatosis in rats. 21 PPARα activation by PFAS in human hepatocytes led to triglyceride accumulation and altered the expression of genes related to cholesterol metabolism. 11 Attema et al. demonstrated that long-term exposure of mice to PFAS disrupts hepatic lipid metabolism with the effects mediated entirely by PPARα. 22 Systemic toxicity was induced by the topical application of PFAS in a murine model with alterations in PPAR genes. 23
Here, we use a luciferase cell-based assay to compare the differences between classes of PFAS in their capacity to activate human PPARα. PFAS comprises a wide range of compounds, and variations in their chemical characteristics could reveal new details regarding the extent of PPARα activation. This study seeks to probe the linkage of PFAS structural features with PPARα activation efficiency which may further enable identification and prioritization of PFAS chemicals for risk assessment.
Materials and Methods
Chemicals
PFAS were purchased at the highest available purity from SynQuest Labs (Alachua, FL): perfluoropentane sulfonic acid (PFPeS), perfluorohexane sulfonic acid (PFHxS), perfluoroheptane sulfonic acid (PFHpS), 1H,1H,2H,2H-perfluoro-1-hexanol (4:2 FTOH), 1H,1H,2H,2H-perfluoro-1-octanol (6:2 FTOH), and 1H,1H,2H,2H-perfluoro-1-decanol (8:2 FTOH); and Sigma Aldrich (St. Louis, MO): perfluorobutanoic acid (PFBA), perfluoropentanoic acid (PFPeA), perfluorohexanoic acid (PFHxA), perfluoroheptanoic acid (PFHpA), perfluorooctanoic acid (PFOA), perfluorononanoic acid (PFNA), perfluorodecanoic acid (PFDA), perfluorobutane sulfonic acid (PFBS), perfluorooctane sulfonic acid (PFOS), and hexafluoropropylene oxide dimer acid (HFPO-DA). These PFAS can be divided into 4 classes based on specific chemical features. Perfluoroether carboxylic acids (PFECA) are ether compounds with perfluorinated carbons and a carboxylic acid functional group. Perfluorocarboxylic acids (PFCA) and perfluorosulfonic acids (PFSA) have perfluorinated carbons and either a carboxylic acid or sulfonic acid functional group, respectively. Polyfluorotelomer alcohols (FTOH) have polyfluorinated carbons and an alcoholic functional group.
Cell Culture
PPARα Chemically Activated LUciferase eXpression (CALUX) assay was obtained from BioDetection Systems (Amsterdam, Netherlands). The method uses human osteosarcoma cells (U2OS) transfected with the firefly luciferase reporter gene to produce the enzyme luciferase in response to the activation of PPARα. The cells were subcultured and passaged twice in Dulbecco’s Modified Eagle Medium/Nutrient Mixture F12 (DMEM/F12), supplemented with 7.5% fetal calf serum (FCS), 10 U/mL penicillin, and 10 μg/mL streptomycin (Gibco-Fisher, Waltham, MA). U2OS (1.5 × 104 cells/well) were plated in 96-well plates at 100 μl/well and maintained under standardized conditions, 37°C, 5% CO2, and 95% humidity.
XTT Viability Assay
To determine U2OS viability after PFAS exposure, the XTT assay (CyQUANT™ XTT Cell Viability Assay, Thermo Fisher Scientific) was used. The tetrazolium salt, XTT, is reduced to highly colored formazan dye by dehydrogenase enzymes in metabolically active cells. U2OS was added to 96=well plates (1.5 × 104 cells/well in 200 μL culture media). The cells were exposed to PFAS compounds (0.001–1000 μM) and incubated for 20 h under standardized conditions. Activated XTT reagent (1:50 v:v) was added, and the cells were incubated for 4 hours. The product generated was proportional to the number of living cells and quantified using an Agilent Cytation 5 plate reader to measure absorbance at wavelength 475 nm. The background signal at 660 nm was subtracted to obtain normalized absorbance values.
CALUX Assay
Culture medium was removed after 24 h, and exposure medium (DMEM/F12 without phenol red, 5% charcoal-stripped FCS, 10 U/mL penicillin, and 10 μg/mL streptomycin) was added. Following this, 0.1 % dimethyl sulfoxide (DMSO) vehicle or PFAS compounds serially diluted to final concentrations of 3, 30, 60, and 100 μM dissolved in the DMSO vehicle were added to selected wells to a final volume of 200 μL. Plates had 0.01 to 100 nM of the reference control, 2-[4-[2-[4-cyclohexylbutyl(cyclohexylcarbamoyl)amino]ethyl]phenyl]sulfanyl-2-methylpropanoic acid (GW7647), a full PPARα agonist, for an entire response curve (Figure S1). After incubation for 20 hours, the exposure medium was aspirated off, and the cells were lysed with 30 μL/well of Triton-lysis buffer. Luminescence reagent (BioDetection Systems) (100 μL/well) was added and measured for luciferase activity for 4 s with a multimode plate reader (Agilent Cytation 5, Santa Clara, CA). Activity per well was measured as relative light units (RLUs).
Data Analysis
Each experimental point was performed in triplicate. Each 96-well plate contained its own reference compound series and vehicle solvent. All test samples, vehicle solvent, and reference compound were analyzed with the acceptable maximum coefficient of variation defined as <15%. For normalization and minimization of intra-/inter-experimental variation, the background luminescent signal in wells with vehicle was subtracted from that in experimental wells. PPARα activation percentage was determined by dividing the corrected signal by the positive control signal of the maximally efficacious response of GW7647, set at 100%. Data for each PFAS chemical were represented as mean luminescent signal from 3 to 5 independent experiments. Values for the chemicals at each concentration tested were scaled accordingly. Differences between PFAS chemicals were determined via a two-sample t-test assuming unequal variances where P-values < 0.05 were considered statistically significant.
Results
CALUX Assay Functionality
A concentration response curve for reference compound GW7647 was plotted using 9 serial dilutions ranging from 0.01 - 100 nM. The curve was fit using a sigmoidal 4-parameter logistic equation. The sigmoidal response curve verified CALUX PPARα assay functionality (Figure S1). To determine PFAS dose responsiveness, the U2OS were exposed to increasing concentrations of each chemical. XTT assay assessment demonstrated that the cells were not viable when exposed to some PFAS chemicals over 100 μM concentration. Due to this, there was no access to the upper plateau of concentration response curves, precluding comparisons at higher PFAS concentrations (data not shown). Other studies also reported cytotoxicity in HepG2 hepatocytes at concentrations over 100 μM for the same compounds. 24
PFAS Activation of PPARα
All PFAS demonstrated concentration-responsive (3 to 100 μM) activation of PPARα (Figure 1). None of the compounds reached the maximum efficacy level of the full agonist GW7647 (P < 0.05 to P < 0.001 by two-sample t-test assuming unequal variances; Table S1). The mean activation percentage varied from a low of 0.5% (3 μM, 8:2 FTOH) to a high of 57% (100 μM, PFOA) (Figure 1). Mean PPARα activation percentage by PFAS at 3 μM, 30 μM, 60 μM, and 100 μM. The classes perfluoroether carboxylic acid (PFECA), perfluorocarboxylic acid (PFCA), perfluorosulfonic acid (PFSA), and polyfluorotelomer (FTOH) are depicted in green, blue, red, or orange, respectively. The X-axis corresponds to increasing concentration of each PFAS displayed in increasingly darker color. The Y-axis corresponds to the mean PPARα activation percentage +/−SEM, where activation of the reference, GW7647, was set at 100%. C# = number of carbons; CF# = number of perfluorinated carbons. Data are expressed as mean +/− SEM; n = 3–5.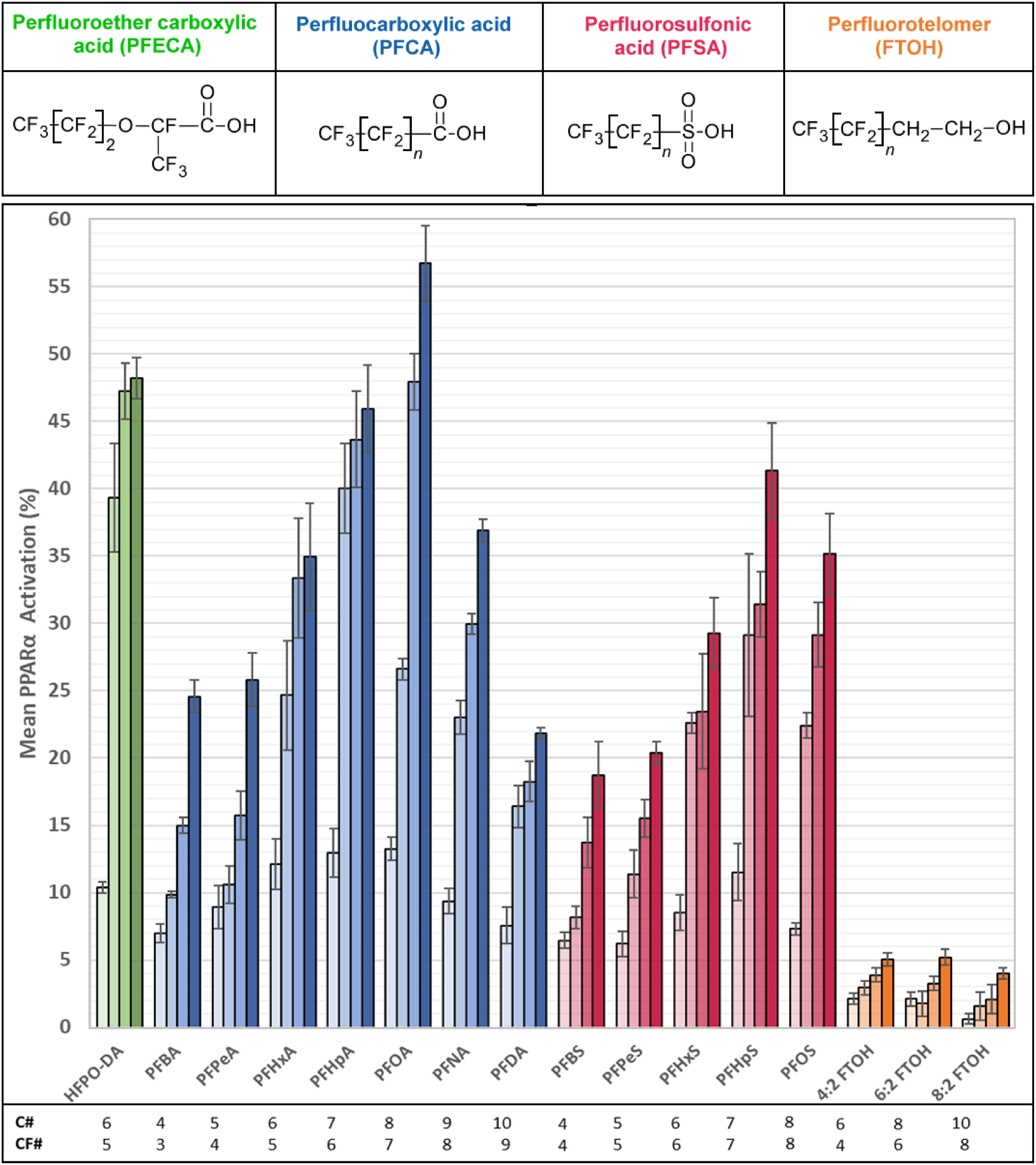
PPARα activation was compared across the PFECA, PFCA, PFSA, and FTOH classes by comparing compounds with the same number of carbon atoms. The ranking from high to low activator class remained the same regardless of the number. PFAS with 6 carbons were HFPO-DA, PFHxA, PFHxS, and 4:2 FTOH, and the ranked order was PFECA > PFCA > PFSA > FTOH (Figure 1). Comparisons were made in PFAS with 4C (PFBA > PFBS), 5C (PFPeA > PFPeS), 7C (PFHpA > PFHpS), 8C (PFOA > PFOS > 6:2 FTOH), and 10C (PFDA > 8:2 FTOH). In each comparison, the ranked order remained PFCA > PFSA > FTOH (Figure 1). The comparisons were made again across the classes, here on compounds with the same perfluorinated carbon number. These included PFAS with 4CF (PFPeA > PFBS > 4:2 FTOH), 5CF (HFPO-DA > PFHxA > PFPeS), 6CF (PFHpA > PFHxS > 6:2 FTOH), 7CF (PFOA > PFHpS), and 8CF (PFNA > PFOS > 8:2 FTOH). The ranking from high to low remained the same: PFECA > PFCA > PFSA > FTOH (Figure 1).
These results between classes indicate that the chemical features of PFECAs can be predictive of increased PPARα activation. PFECA, the highest activation class, has structural features absent in FTOH, the lowest class. As exemplified by HFPO-DA, PFECAs are PFAS ethers with perfluorinated carbons and a carboxylic acid functional group. In contrast, FTOHs are polyfluorinated PFAS with an alcoholic functional group. The results also show that PFCAs were consistently better PPARα activators than PFSAs, whether compared by carbon number or perfluorinated carbon number. This further supports the hypothesis that the carboxylic acid functional group is a chemical feature predictive of higher PPARα activation.
PPARα activation was also studied within classes. PFAS here were compared by the number of carbons and perfluorocarbons making up the chain length. If comparing the PFCAs by carbon number, the 8 carbon chain perfluorooctanoic acid (PFOA) had the highest activation percentage: 4C PFBA < 5C PFPeA < 6C PFHxA < 7C PFHpA < 8C PFOA > 9C PFNA > 10 C PFDA. But among PFSAs, the 7C carbon chain perfluoroheptane sulfonic acid (PFHpS) had the highest percentage: 4C PFBS < 5C PFPeS < 6C PFHxS < 7C PFHpS > 8C PFOS. However, if compared by the number of perfluorocarbons in their linear chain, PFOA and PFHpS have the same number, 7, suggesting that perfluorocarbon chain number better predicts a higher PPARα activation than simply carbon number (Figure 1, C# and CF#, Table S2).
C20 Potency
Potency, the concentration of PFAS (μM) activating PPARα to a given level, was determined for PFECA, PFCAs, and PFSAs to further compare activation in these classes. Activation outcomes were adjusted to the same scale using a percentage of maximal response achieved by the reference. The C20 potency was thus defined as that concentration that achieved 20% of the maximal response obtained by GW7647 on the Y-axis of the linear regression curve. The horizontal line intercepting the regression line was used to determine the concentration of each PFAS on the X-axis. Potency was reported as EC50 in another study, 10 but as previously noted, here EC50 was precluded due to U2OS cytotoxicity at higher PFAS concentrations.
The potency of PFAS compounds which achieved a C20 value is shown in Figure 2. This included the PFAS in all classes except FTOH, the weakest class of PPAR α activators. A lower C20 indicates higher potency. There was a direct correlation between C20 potency and the mean percent PPARα activation after treatment with PFAS (30 μM, 60 μM, and 100 μM; R2 > 0.702). The PFAS compounds fell into 2 groups categorized by C20 as higher or lower PPARα potency activators. Higher potency activators included the PFECA, 5CF to 8CF PFCAs, and 6CF to 8CF PFSA and were ranked as follows: PFHpA > HFPO-DA > PFOS > PFNA > PFOA > PFHxS = PFHxA > PFHpS. At 100 μM PFAS, the PPARα percent activation of the higher potency PFAS was over 28%. Lower potency PFAS included 3CF, 4CF PFCAs and 4CF, 5CF PFSAs, and the 9CF PFCA and were ranked as follows: PFDA > PFPeA > PFBA > PFPeS > PFBS. The PPARα percent activation of the lower potency PFAS was less than 27% (Figure 2). C20 Potency (μM) by Mean PPARα Activation (%). Perfluoroether carboxylic acid (PFECA), perfluorocarboxylic acid (PFCA), and perfluorosulfonic acid (PFSA) results are again depicted in green, blue, or red, respectively. The X-axis corresponds to the mean PPARα activation percentage after exposure to each chemical at 100 μM. GW7647 was tested several logs lower than 100 μM. The Y-axis corresponds to C20, the concentration of each chemical (μM) needed to activate PPARα at 20% of the maximal response reached by GW7647.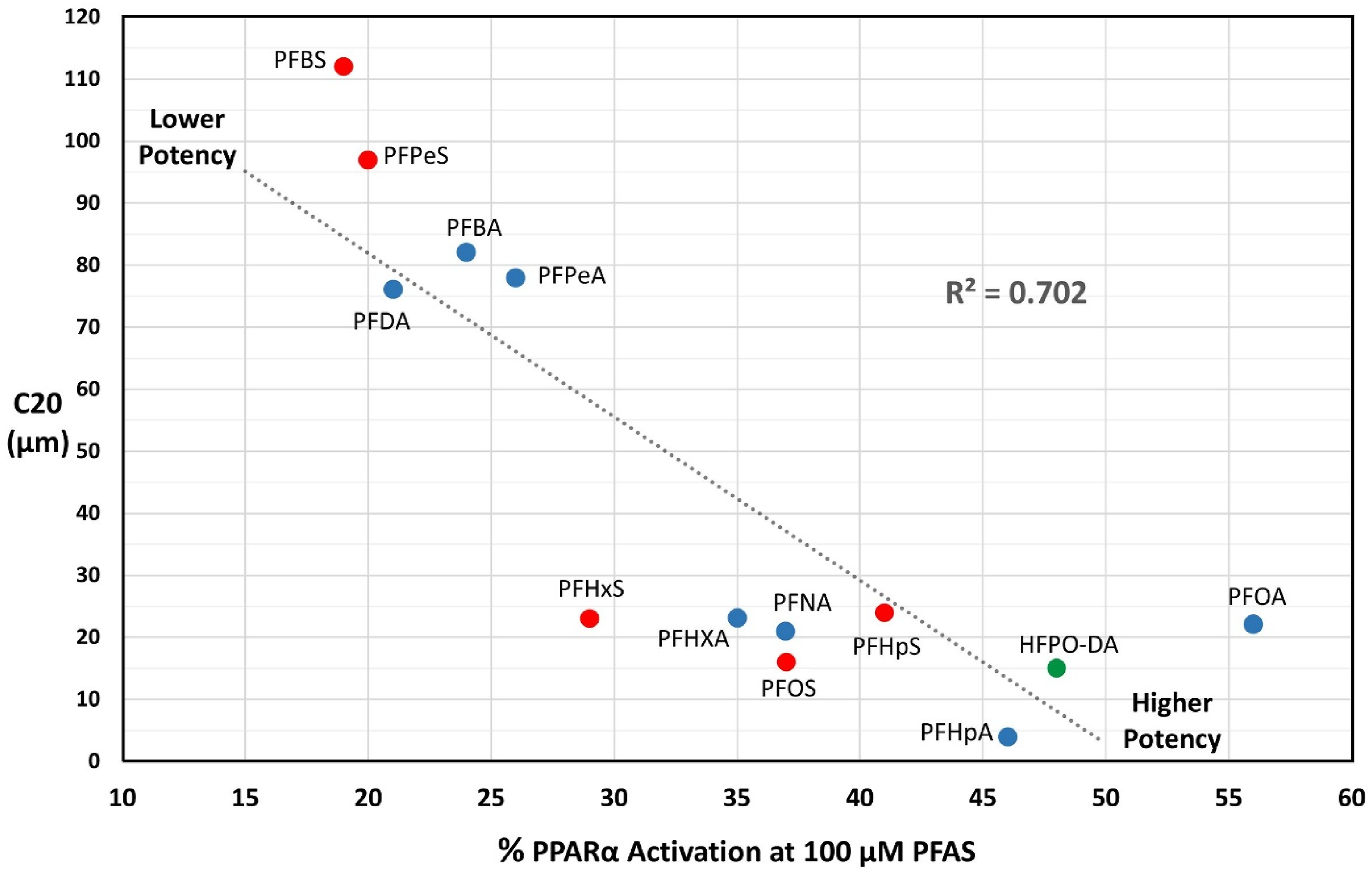
Higher potency activators included the PFECA, 5CF to 8CF PFCAs, and 6CF to 8CF PFSA and were ranked as follows: PFHpA > HFPO-DA > PFOS > PFNA > PFOA > PFHxS = PFHxA > PFHpS. At 100 μM PFAS, the PPARα percent activation of the higher potency PFAS was over 28%. Lower potency PFAS included 3CF, 4CF PFCAs and 4CF, 5CF PFSAs, and the 9CF PFCA and were ranked as follows: PFDA > PFPeA > PFBA > PFPeS > PFBS. The PPARα percent activation of the lower potency PFAS was less than 27% (Figure 2).
Discussion
This study used human osteosarcoma cells transfected with a luciferase reporter construct to investigate the chemical characteristics of PFAS that may predict the level of PPARα activation. Previous studies demonstrate that some PFAS compounds can activate PPARα.9,14–16 Because the adverse effects of PFAS have been linked to this activation,21–23 the chemical features of PFAS which could affect activation efficacy and potency are of significance.
When comparing PPARα activation across PFAS class, the order from high to low rank was PFECA > PFCA > PFSA > FTOH. PFECA, the strongest class, has chemical characteristics that are missing in FTOH, the weakest class. This suggests that those features, an ether linkage, perfluorinated carbons, and a carboxylic acid functional group, may be indicators for higher activation. HFPO-DA, the PFECA in this study, has been reported in other investigations as an effective activator.9,10 Using a full length human reporter construct in COS7 cells, Nielsen demonstrated that HFPO-DA had the highest efficacy and potency of all PFAS tested. 10 Referred to as Gen X, this compound is used in the manufacturing industry as a replacement PFAS for PFOA, PFOS, and other so-called legacy PFAS. While still unclear if PPARα activation is the initiating molecular event leading to hepatic toxicity downstream, the results of this and prior studies suggest that HFPO-DA warrants additional testing to determine its usage as a safe PFAS alternate.
Further comparison across classes showed the findings were in agreement with others showing PFCAs to be better PPARα activators than PFSAs.9,12,15,25 The PFCA, which differed from the PFSA only in having a carboxylic acid end group rather than a sulfonic acid end group, were consistently stronger. When comparing within each class, the highest activators of PFCA and PFSA compounds, PFOA and PFHpS, respectively, have an equal chain number of perfluorinated carbons, 7CF, but a different number of carbons. Previous studies reported that PFAS carbon number plays a role in the degree of PPARα activation,9,24,26 but the results here indicate that perfluorinated carbon chain number better predicts activation efficacy.
After PFCA and PFSA compounds achieved maximal activation at 7CF perfluorinated carbons, a decrease was observed at higher number. Increasing compound size could decrease its membrane permeability making PFAS with chain length over 7CF less effective crossing the U2OS cell membrane. Wolf postulated that the lack of a stronger effect by PFDA compared to shorter chain compounds could be its larger size made it sterically impractical to bind to the ligand-binding domain site. 26 Differences in PFAS efficacy of transportation to the nucleus may also play a role. Zhang showed that liver fatty binding protein transported long chain PFCAs to the nucleus to a lesser extent than shorter chain PFCAs. 27 Organic anion transporters (OATS), polypeptides differentially expressed in multiple cell types, facilitate membrane transport of PFAS,6,28,29 and Liedauer reported that 2 OATS transporters, OATP3A1 and OATP4A1s, are expressed in human osteosarcoma cells, the cell type in this study. 30 These explanations require additional investigation to determine their validity.
While the findings here are generally consistent with other studies, the relative activation of PPARα by PFOA versus HFPO-DA was inconsistent. Although we observed PFOA achieved a higher activation, Behr reported that HFPO-DA was more effective than PFOA in stimulating PPARα in HEK293T cells under the same conditions. 9 Nielsen’s work with COS7 cells supported that result. 10 Variations in ligand-binding efficacies across species may explain some differences in PFAS-induced hepatotoxicity between humans and rodents, 31 and species variation could also explain the divergence in results between U2OS, a human cell line, from COS7, a monkey fibroblast cell line. HEK293T is a human cell line, but the differences here could be based on variance in cellular coregulators required for nuclear receptor activity. 32 For example, the expression of OATS transporter in U2OS cells, but not HEK293T or COS7, might affect the efficacy of transportation to the nucleus. Further investigations would determine if these possibilities provide a valid explanation.
In this study, the potency and efficacy of PPARα activation were evaluated to assess the risk of PFAS exposure. A novel result was the observation of a direct correlation between PPARα activation level at the higher concentrations, 30, 60, and 100 μM, and the C20 potency of PFAS. PFECA, PFCA, and PFSA compound broke into 2 types: higher potency-activation compounds or lower potency-activation compounds. The first group consisted of PFECA, 5CF to 8CF PFCAs, and 6CF to 8CF PFSAs. Thus, both metrics generally gave similar results with regard to the efficacy of activating the nuclear receptor. Compounds with high potency but low efficacy may not reach the threshold of activation necessary to initiate toxicity downstream. On the other hand, a compound with high efficacy but a low potency can still pose a risk if enough exposure occurs. Whether a specific activation threshold is needed to trigger toxicity is unresolved, and further studies should address this question.
Conclusions
In this investigation, the chemical features of a large number of PFAS compounds which activate PPARα to varying extent were studied. PFECA, the strongest activator class, has specific characteristics, an ether linkage and a carboxylic acid functional group, that are absent in FTOH, the weakest class. PFCAs were stronger activators than PFSAs, further supporting carboxylic acid functional group as an important feature. By comparing chemicals within the PFCA and PFSA classes, perfluorinated carbon rather than carbon number better predicted highest activation. Assessing the role of polypeptide transporters and other coregulators could explain the divergent results observed between studies. A direct correlation was noted between PPARα activation percentage and C20 potency. If PPARα activation is the triggering event for subsequent intracellular processes leading to a systemic toxicological outcome, this study has significance by revealing those chemical characteristics of PFAS which enhance that activation.
Supplemental Material
Supplemental Material - Differential Activation of Peroxisome Proliferator-Activated Receptor Alpha by Per- and Polyfluoroalkyl Substances
Supplemental Material for Differential Activation of Peroxisome Proliferator-Activated Receptor Alpha by Per- and Polyfluoroalkyl Substances by Mary Helen Kanitz and Cynthia Striley in International Journal of Toxicology
Footnotes
Acknowledgments
Research herein was performed with support from National Institute for Occupational Safety and Health (NIOSH) internal research funding.
Author Contributions
Kanitz, M.: contributed to conception and design, contributed to acquisition, analysis, and interpretation, drafted manuscript; critically revised manuscript; gave final approval; agrees to be accountable for all aspects of work ensuring itegrity and accuracy.
Striley, C.: contributed to design; contributed to analysis and interpretation; critically revised manuscript; gave final approval; agrees to be accountable for all aspects of work ensuring itegrity and accuracy.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Disclaimer
The findings and conclusions in this report are those of the author(s) and do not necessarily represent the official position of the National Institute for Occupational Safety and Health (NIOSH) and Centers for Disease Control and Prevention (CDC). Mention of any company or product does not constitute endorsement by the NIOSH/CDC.
Data Availability Statement
Data will be made available on request.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
