Abstract
Major advances in scientific discovery and insights that stem from the development and use of new techniques and models can bring remarkable progress to conventional toxicology. Although animal testing is still considered as the "gold standard" in traditional toxicity testing, there is a necessity for shift from animal testing to alternative methods regarding the drug safety testing owing to the emerging state-of-art techniques and the proposal of 3Rs (replace, reduce, and refine) towards animal welfare. This review describes some recent research methods in drug discovery toxicology, including in vitro cell and organ-on-a-chip, imaging systems, model organisms (
Keywords
Introduction
Drug discovery and development toxicology is an evolving area of science within biopharmaceutical industry and academic research. Routine in-vitro safety screening, as well as GLP (Good Laboratory Practice) and non-GLP toxicity testing, follows regulatory framework from USFDA (Food and Drug Administration), ICH (International Council for Harmonization), and EMA (European Medicines Agency) guidelines among others. However, in recent years, industry is moving towards increasing safety screening in lead optimization stages of the drug discovery. Owing to this shift, the role of "exploratory" or "investigative" toxicologist holds key in the progress of the toxicological sciences.
In the routine safety screening for small molecule drug candidates, mutagenicity and cardiovascular toxicity are commonly assessed with in vitro assays. Larger panel-based hit testing for G-protein coupled receptors, ion channels, neurotransmitter channels, nuclear receptors, kinases, and certain non-kinase enzymes are tested with a battery of in vitro assays. The goal of these assays is to identify early risks associated with discovery candidates. The utility of these assays is universal across the biopharmaceutical industry, as they provide minimum necessary safety information. Once the compounds show positive results in safety tests, dedicated specific assays will be performed. However, the compounds move forward unless the potency of the identified hit is beyond certain value. For example, for cardiovascular in vitro screening, if a compound shows the potential to inhibit hERG (human ether-a-go-go related gene) channel which might be an indicator of cardiovascular risk, a follow-up study will be performed to calculate the IC50 of this compound to inhibit hERG channel. USFDA guideline provides a specific criterion to calculate safety window for in vitro hERG channel inhibition. Also, these criteria drive the go–no-go decisions for the drug candidates. Such specific definition of risk identification is not available for all the screening level assays in toxicology. Moreover, the mechanism of the safety hit observed, or chemical matter involved is not generally understood in these assays. Investigative toxicology addresses these questions based on in vitro, in vivo and in silico analyses. Some of the common approaches within investigative toxicology include gene expression signature, two and three-dimensional tissue culture models, miniature organ-on-chip (also called micro physiological systems), and proteomic or metabolomic signatures. Additionally, high resolution microscopy-based techniques, as well as in-silico evaluations are increasingly being used in industry. To mimic human physiology as a whole, zebrafish, nematode, or fruit fly models have also earned significant place in the toxicology screening.
This review discusses the advances in the research for these investigative toxicology tools and approaches. The focus of the chapter is the latest technological advances in toxicology for early drug screening within the biopharmaceutical industry.
In Vitro Models
Drug discovery and development is an expensive and slow process. The least expensive part of this process is the early discovery, that includes lead identification and lead optimization. Once the candidates move past "development" nomination, the expensive and intense GLP toxicology program is initiated. By this point, in the life cycle of the drug molecule, it’s critical to have basic safety information during early screening stage, including genotoxicity potential, cardiovascular risk and off-target effects including drug-induced organ injury. Early in vitro screening assays are vital to assess the risks that inform the programs teams of the expected outcome in the preclinical GLP toxicology.
In vitro models provide an affordable alternative to animal testing. Cell-based 2D (two-dimensional) and 3D (three-dimensional) models recapitulate organ system physiology to some extent. For studying specific mechanisms within organ injury, cell-based models are preferred. For instance, bile acid induced liver injury is one of the major mechanisms of DILI (drug-induced liver injury). Perturbation of regular bile acid homeostasis, accumulation in the hepatocytes or excessive levels in the systemic circulation could lead to potential toxic effects. Primary hepatocyte culture in a Matrigel-based sandwich arrangement creates a cellular system with bile ducts, and polarized cells with basolateral and canalicular distinction. 1 This special culture mimics the bile circulation physiology of the liver quite accurately, providing a useful tool for DILI risk assessment of small molecules. More advanced 3D liver models include sphere or bioprinted liver tissues. One of the benefits of these models is the longevity compared to regular 2D culture, and hence the scope to perform multiday exposure experiments. 3D models generally are more sensitive to drug-induced injury. Micro-patterned co-culture of hepatocytes with fibroblasts is another example of physiologically relevant model that supports longer duration studies. 2
The advent of micro physiological systems (MPS) has the potential to revolutionize the toxicity testing paradigm.
Microphysiological Systems
MPS consists of cellular constructs that are interconnected with fluid network. Cellular constructs are designed from primary or immortalized cells, or stem cells from embryonic origin or pluripotent stem cells. These miniature cellular constructs represent a specific organ system based on the origin of the cells used, and multiple of these organ systems are connected with a network of fluidics on a small chip. Figure 1 displays representative MPS for neuronal system. MPS for neuronal system and resources needed for its assembly.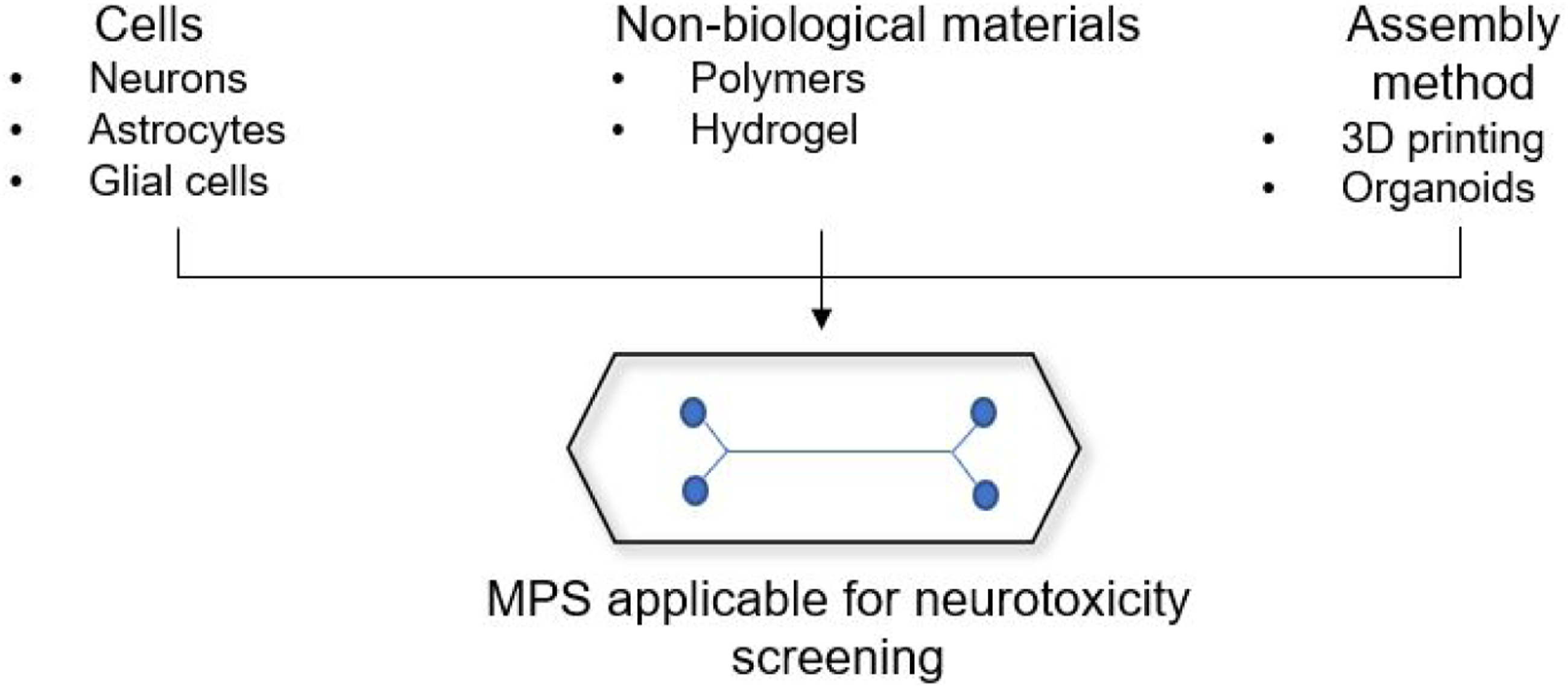
Viravaidya et al. 3 first demonstrated the importance of connecting multiple single organ MPS to detect naphthalene toxicity. In the study, a two-cell system which contains lung epithelial cells and hepatocytes was developed to study naphthalene toxicity. Their multi single organ MPS successfully demonstrated the process that naphthalene was first metabolized to reactive metabolites (i.e., 1,2-naphthalenediol and 1,2-naphthoquinone) in the liver cells, which then circulate to the “lung,” depleting glutathione (GSH) in lung cells. The inter-organ mechanistic connections are key to understanding toxicity for multiple chemicals. This discovery paved the way for the development of multi-organ MPS. Multiple cellular constructs representing an individual organ are connected to mimic the physiological interplay as a response to xenobiotic exposure of the body. The multiple organ constructs are connected with fluidics and electronic miniature pumps or passive gravity are used to drive the circulation. For instance, an MPS system comprised of human intestinal model, liver spheroids, and circulation provide appropriate tool for accurately measuring oral bioavailability of compounds. A model pairing kidney and liver MPS was described to measure hepatic first pass and bioavailability. 4 Interpretation of the data from multi-organelle MPS requires the use of computational analysis. Scaling of the MPS to overall physiology, flow rates and time, and geometry of the system need to be accurately considered. Physiological-based pharmacokinetic modeling (PBPK), PK-PD (pharmacokinetic-pharmacodynamic) models for exposure-response, and quantitative systems pharmacology (QSP) models are of great utility in data extrapolation from MPS.
MPS systems have been developed for multiple organ systems including blood–brain barrier,5,6 female reproductive track, 7 heart,8,9 and so on. Specifically, from the toxicology point of view, several MPS have been described with experiments that involve exposure to xenobiotics and environmental chemicals.
Mathur et al. described human iPSC derived cardiac MPS which measured the beat frequency as a measure of cardiotoxicity. 10 The data obtained for known agents including isoproterenol were within acceptable range of the EC50 values from traditional cardiac muscle model. Cerivastatin, which was removed from the market owing to rhabdomyolysis, 11 was tested in skeletal muscle MPS and displayed toxicity at nanomolar levels. These data from MPS clearly showed the sensitivity of the system and clinical applications. In the same study, Lovastatin was tested, and its toxicity levels were far above the pharmacological concentrations.12,13 Brain organoid derived MPS were tested for effects of nicotine, and authors concluded that brain cell differentiation was affected commensurate with fetal effects of nicotine in pregnant smokers. 14
To date, MPS is primarily adopted for preclinical safety in early drug discovery (e.g., drug toxicology and metabolism).15,16 Several MPS tools have been developed and applied by the pharmaceutical industry. TissUse, which were developed by Technische Universität Berlin, has been developed to evaluate hepatotoxicity during drug development using human hematopoietic stem cells. 17 Later, two-organ-chip models were developed based on TissUse to investigate the crosstalk between different organs. For example, Kühnlenz et al. co-cultured liver cells and thyroid follicles using TissUse to assess hepatic and thyroidal associated dysregulations of thyroid metabolism separately or in combination. 18 Bauer et al. co-cultured pancreatic islet microtissues and liver spheroids on TissUse to investigate pancreatic islet–liver crosstalk. 19 Emulate was developed by the Wyss Institute for Biologically Inspired Engineering and has been applied to preclinical safety testing. For instance, Jang et al. developed a liver-chip using Emulate consisting of rat, dog, and human hepatocytes, endothelial cells, Kupffer cells, and stellate cells to assess cross-species toxicities of known hepatotoxic drugs and experimental compounds. 20 Mimetas OrganoPlate® developed from Leiden University is another MPS system that has been used in for investigating cardiovascular, 21 renal, 22 and blood vessel 23 toxicities. Since its rapidly developing novel area of science, MPS has several challenges ahead, for instance, identifying a universal medium that caters to the nutritional and physiological needs of individual organ constructs, volume limitations, and developing statistical simulations to accurately interpret the data. Regenerating cellular structural arrangements for specialized tissues is also a challenge. Using MPS for high throughput screening which simultaneously assess a large number of test samples of various safety endpoints is also an area under investigation.
Center for Drug Evaluation and Research (CDER)/USFDA promote use of MPS as a part of new approach methodologies to replace, reduce and refine the use of animals in safety testing. ICH M3(R2) nonclinical safety guidelines encourage using new in vitro alternative methods for safety testing prior to clinical trials if the alternative methods are validated. ICH S5(R3) guidance on reproductive toxicity lists the qualification criteria for alternative test systems for regulatory acceptance in Annex 2. The initiative from regulatory agencies to work alongside academic and industry researchers demonstrates the immense potential MPS hold in speeding up drug discovery process and reducing animal use.
Model Organisms
In the drug discovery process, cell culture-based testing is one of the most recognized in vitro models to identify therapeutic compounds. However, it has been challenged that the impact of chemical microenvironments cannot be investigated in the cell model and the drug-toxicity profile in cell models cannot reflect tissue-specific toxic response and is poorly correlated with in vivo results. 24
Starting from the proposal of 3Rs (reduce, replace, and refine) by Bill Russel and Rex Burch in 1959, 25 followed by European Union (EU) legislation clearly set out the full replacement of animal methods as the ultimate policy goal in 2010. 26 This was echoed by the National Institutes of Health (NIH) and the U.S. Environmental Protection Agency (EPA) to move away from in vivo studies in mammalian models.27,28 Regulatory agencies are seeking New Approach Methods (NAMs) as alternative methods to test chemicals for health effects.
Among the many non-mammalian model systems that can contribute to fundamental understanding in toxicology,
Caenorhabditis Elegans
The adult hermaphrodites have 959 somatic cells while the adult male has 1031, and these cells were found to remarkably consistent from animal to animal.31-33 The nervous system of
Toxicity testing aims to obtain information acquired in a particular model that can be applied to other biological systems, such as toxicity data obtained from
The use of
Toxicity comparison study conducted in
The application of
To summarize, the great diversity of endpoints and possibilities of
However,
Danio Rerio (Zebrafish)
The zebrafish is a member of the minnow family of fish. Zebrafish as a model organism can provide a significant amount of information for vertebrates that enables biochemical, genetic, and cellular toxicity mechanism investigation and testing. 58 Phylogenetic analysis indicates a similar morphology and physiology of the nervous, cardiovascular, and digestive systems between fish and human. 59
Zebrafish have a short life cycle and a large capacity to generate offspring. The heart, liver, brain, pancreas, and other organs are developed by 5 days post fertilization (dpf). 60 A pair of adult fish can produce 200-300 transparent embryos. 58 The fast growth rate, transparent embryo, small size, higher fecundity, and ease of handling, compared to mammalians, have made zebrafish amenable to high-throughput toxicity screening. 61
To investigate environmental and developmental toxicology, zebrafish embryos can be exposed to chemicals which can be simply added to the medium. Lethality and developmental abnormalities from fertilization through larval stages can be observed and scored under microscope on the transparent embryos. Although juvenile and adult zebrafish are not transparent, the generation of unpigmented zebrafish mutant strain can be crossed to fluorescent reporters or the addition of pigment formation inhibitors can be used to aid observation and imaging.62,63 For example, Torregroze et al. and Das et al. screened a set of retinoic acid derivatives and compounds designed to modulate transforming growth factor beta (TGFβ) in zebrafish embryos and observed cardiovascular defects in early embryogenesis that were similarly seen in mammalian models.64,65 Another study exposed zebrafish embryos to 31 compounds respectively at 4-6 hours post fertilization (hpf) and cultured for 5 days. 66 Several dysmorphologies on indicators of embryotoxicity, such as tail, fin, brain, and jaw morphology defects, were observed. The observation was 87% consistent with in vivo mammalian data for all 31 compounds. The no observed adverse effect level (NOAEL) and LC25 were determined based on these results. Later, the ratio between LC25 and NOAEL was used to separate teratogens from non-teratogens, as defined by the Interagency Coordinating Committee on the Validation of Alternative Methods.67,68
The zebrafish genome has been fully sequenced and annotated,
69
making it a fast model for genetic and
It has been demonstrated that the expression of genes involved in controlling neurotransmitters in zebrafish are similar to those in mammals. 73 Zebrafish and mammalian brains share may features, including well-conserved neurotransmitter structures and systems, such as GABA, glutamate, histamine, acetylcholine, noradrenaline, serotonin, and dopamine. 74 Moreover, zebrafish also share broad anatomical features, including the presence of the cerebellum, telencephalon, diencephalon, spinal cord, and enteric-autonomic nervous systems with mammalian brains. 73 In addition to the similarities in nervous systems, zebrafish also develop a blood brain barrier at 3 dpf, which is reported to be similar to those found in mammals as well, 75 making it an ideal model to investigate neurotoxicity. However, zebrafish do not have a neocortex, which means it is not a proper model for cognitive studies. 76
Zebrafish express a full range of cytochrome P450 (cyp) genes and a total of 86 cyp genes were identified with many of the metabolic characteristics of the related human enzymes. 77 However, the information on specific mechanisms of zebrafish cyp activity remains unclear. But mutant zebrafish line with abnormally large or small livers is documented. 63
Zebrafish and human eye have conserved gene expression, cellular makeup, tissue architecture, and anatomical similarities (e.g., cornea, lens, choroid, retina, etc.), making it an excellent model for studying ocular toxicity. 78 Several studies have demonstrated consistent results between zebrafish vision assays and human ocular reactions to drugs, such as chlorpromazine, cisplatin, quinine, vardenafil, and so on. 79 Recently, retinal toxicity was tested in zebrafish larvae in pharmaceutical pipeline discovery compounds, replacing studies in mammalian models and indicating values of zebrafish in drug discovery toxicology. 80
In general, zebrafish are a useful and promising tool for pharmaceutical toxicity testing. However, several obvious limitations with this platform includes physical barriers (chemical solubility), kinetic factor (absorption, distribution, metabolism, and excretion), and dynamic factors (susceptibility and responsiveness).
63
Since gills are not functioning and embryos are not yet free feeding during the first 5 dpf, the main exposure route is dermal, making dosing difficult. There is also debate on whether zebrafish are able to metabolize all xenobiotics which results in difficulty in studying toxicities of chemicals that require metabolic activation. Furthermore, zebrafish do require special equipment for culture and extra efforts for maintenance, compared to
Drosophila Melanogaster (Fruit Flies)
Moreover,
Another promising application of
Short life cycle, ease of culture, propagation, and maintenance, cost effectiveness, ease of genetic modification, and fully mapped neurons in the brain make
All three model organisms mentioned above can provide significant information on genetic and cellular process in a rapid, cost-effective, and good correlation to rodents’ results. The selection of which model to use can be considered based on the nature of disease being studied, targeted dosing route, the questions being asked, etc.
Imaging-Based Approaches
Along with serum biomarkers of organ injury, classic toxicological evaluations heavily rely on histopathological findings from the tissues. However, novel in vivo imaging techniques could potentially revolutionize the translational aspect of drug safety screening. Non-invasive or minimally invasive imaging techniques, including positron emission tomography (PET), magnetic resonance imaging (MRI), computed tomography (CT) and ultrasound are increasingly gaining more attention in clinical toxicology.
Along with clinical studies, imaging has wide applicability in preclinical safety testing as well. Often, information that could be obtained from imaging technique cannot be obtained with another method of assessment. For instance, internal tissue disorders such as fibrosis of the liver are difficult to accurately diagnose for its severity and scoring through biomarkers. Histology from liver biopsy was considered as a gold standard until imaging techniques were utilized. CT scans can reveal changes in the tissue density accurately and reproducibly, and without having to subject animals or patients to painful biopsy. In fact, growing fibrotic phenotype in the liver can be monitored over longer duration of experiments in same subjects using imaging.92,93
MRI techniques allow assessment of blood flow, perfusion, cellular density as well as detection of necrosis and inflammation. Stoica et al. described MRI-based diagnosis of femoral head necrosis, which leads to musculoskeletal disorder. 94 For hepatic injury, alanine transaminase and aspartate transaminase are commonly used serum biomarkers. However, MRI-based tools provide much advanced data from liver including iron toxicity levels, and its correlation with cardiac iron levels. 95 Lipid accumulation in liver, also termed steatosis, is common drug safety observation. However, steatosis severity does not always correlate with increase in serum biomarker analysis. Imaging-based techniques can avoid the necessity for biopsy. Vigabatrin is a marketed anticonvulsant drug. During its preclinical evaluation, rats and dogs demonstrated intramyelinic edema in distinct regions of the brain.96,97 MRI has been shown as a sensitive tool to diagnose structural changes in brain regions in preclinical species. 98
Imaging mass spectrometry has gained widespread popularity in drug distribution studies in recent years. Biological samples are mapped by pixel arrays, and analytes are localized by sequential mass spectra acquisition. 99 MALDI (matrix assisted laser desorption ionization) further enhanced the MS-based imaging by allowing small molecules as well as large peptides and proteins mapping without special tagging or labeling. 100 MALDI utilizes laser induced ionization of the molecules, which are then routed to time-of-flight (TOF) mass-based analysis. Moreover, MALDI allows simultaneous detection of parent molecule and metabolites. To avoid background non-specific signal from matrix clusters, stable isotope labeling of the matrix can be employed. 101 Kadar et al. described brain distribution of the MPTP (1-methyl-4-phenyl-1,2,3,6-tetrahydropyridine) metabolite MPP+ (1-methyl-4-phenylpyridinium) after oral administration in mice. 102 This is an example of the direct utilization of the MALDI imaging in toxicology. For environmental chemical accumulation in tissues, MALDI-based imaging has been applied with success in preclinical studies.103,104
For the treatment of Parkinson’s disease (PD), L-DOPA chronic administration is commonly employed. About 80% of the patients experience L-DOPA induced dyskinesia (LID) several years into the treatment. 105 Rat models of 6-OHDA induced brain lesions mimic PD to some extent. MALDI imaging of rat brains to discern spatial dynorphin opioid peptides has been described.106,107 Interestingly, in this PD model, rodents develop pathology on one side of the brain, which means other half can be utilized as internal control, addressing animal to animal variability issues. 108
Toxicogenomics
High throughput screening approaches are essential in discovery toxicology to identify safety signals early in the process and terminate candidates or chemical series without investing too much time or resources. Systems biology approach has been recently used to garner power of genomics, proteomics and metabolomics under one umbrella of toxicogenomics.
109
Stierum et al. demonstrated rodent hepatotoxicity of butylated hydroxytoluene and propyl galate using gene expression profiling coupled with conventional biochemistry.
110
Toxicogenomics studies the interaction between genes and xenobiotics/toxicants. Based on central dogma of life, DNA encodes mRNA that carries vital information necessary for protein synthesis. The proteins then carry out the cellular functions including enzymatic reactions, barriers, and channel formation, synthesize or regulate intracellular metabolites/endogenous chemical homeostasis. Transcript, protein, metabolite profiling can be combined with the computational tools of bioinformatics to aid conventional toxicology on investigating how xenobiotics/toxicants affect gene expression (transcriptomics), protein production (proteomics), and small molecule metabolism and function (metabolomics) (Figure 2).
111
Overview of "omics" screening platforms in toxicology.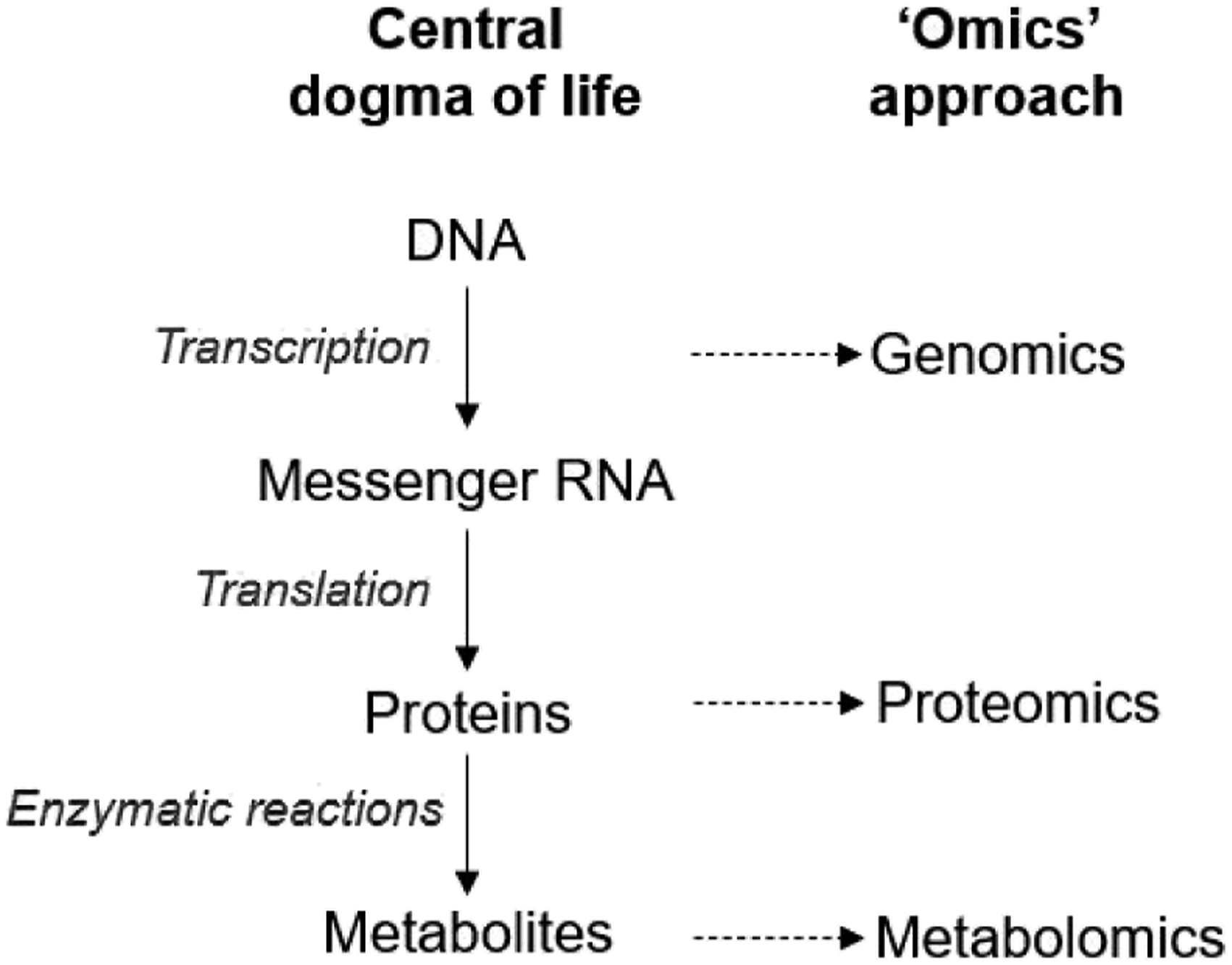
Genomics
Traditional toxicology relied on phenotypic and pathological changes in biological systems because of chemical exposure. With a move towards mechanistic understanding of the toxicology mechanisms, genomics can serve as a critical tool. 112 Genomic bioassays can also assist in early detection of genotoxicity or carcinogenicity potential of new chemical entities, as opposed to traditional two-year rodent studies. The advent of microarray technologies along with statistical data processing tools enables genomic analysis in toxicology. Moreover, genomic screening can also provide a translational tool to better identify relevance of the preclinical species to the clinical predictions. Genomic sequence information is available for preclinical species, and detailed human genome has been mapped as well. Genome comparisons can enable identification of conserved sequences of the genomes across the species, making it easier to choose the relevant species for specific effect.
Changes in expression of genes coding the family of enzymes can relate to a specific effect of chemical exposure. For instance, stress response genomic arrays were described in multiple investigations relevant to xenobiotic exposure/treatment.113,114 Pharmacological benefits of curcumin in cardio-protection were revealed through the use of microarray. 115 The data from microarray were further bolstered with RT-PCR-based individual gene expression studies in curcumin treated rat models of myocardial infarction.
Currently, there are three genomic techniques in use, genome-wide association (GWAS), exome sequencing, and whole genome sequencing. 116 GWAS uses high-density genotyping of common variants, which account for 1-5% of allele frequency in the population, to provide information on the potential mechanisms that underlie human diseases and drug responses. 117 The identification of genetic variants, such as CYP2C19, interleukin-28B (IFNL3), HLA-B*5701, and HLA-B*1502, guide the choice of dosage for clinicians. 117 However, there are fewer GWAS studies focused on drug toxicities. According to European Bioinformatics Institute GWAS Catalog list (a total of 6401 publications), only 15 publications used GWAS to investigate drug-induced toxicity. 118 Exome sequencing that captures coding sequence mutations achieves good quality coverage of ∼1.5% of the human genome and primarily is used for drug development at clinical stage. For example, Gréen et al. exome sequenced whole blood from lung cancer patients and identified two new genetic markers with the potential to predict myelosuppression induced by gemcitabine/carboplatin chemotherapy. 119 But efforts to bring exome sequencing to drug discovery continue. The UK Biobank (UKB) and eight biopharmaceutical companies established the UK Biobank Exome Sequencing Consortium (UKB-ESC) that aims at sequencing exomes for all ∼500,000 UKB participants. 120 The data yield will provide rich information on rare coding variations that may contribute to drug discovery. Whole genome sequencing that detects high-frequency variants can cover ∼85% of the genome. 121 This technique was initially used to clarify genetic factors of disease and now it is helping to understand the genetic basis of variable drug responses and adverse drug events. 122 Whole genome sequencing has also been used in preclinical drug safety evaluation. For example, it can be used to generate hypotheses for finding differentially expressed genes due to drug treatment which may be further linked to the toxicity, for mechanistic investigations 123 and/or for identifying genes previously unknown to be involved in toxic responses. 124
Genomic approaches inherently have certain limitations like any other technology. Subtle changes in genetic sequence can result in massive functional outcomes in an organism. The human genome has 35000 genes.125,126 Chemical exposure can lead to multiple changes in various regions of the genome, and sophisticated statistical programs enable identification of changes that could be functional or not. Despite the availability of these tools, the data interpretation process is difficult and time consuming. Moreover, genomic data have predictive significance, but cannot yield definitive toxicology endpoints. 123
Proteomics
Proteomics encompasses high throughput separation and identification, and even semi-quantitation of proteins in the biological matrices. In conventional toxicology assessments, biomarkers from blood and other matrices play a key role. Most of the biomarkers utilized in toxicology are small molecule metabolites or proteins that are perturbed because of chemical exposure. Along with providing definitive biomarker data, proteomics can also assist in mechanistic understanding of an adverse reaction in
Han et al. combined conventional toxicology assessment methods with proteomic analysis to evaluate iron oxide nanoparticle effects in rats. 128 Iron oxide nanoparticles were intravenously administered to rats as a part of conventional toxicology and toxicokinetic study. Circulating biomarkers and histopathology revealed spleen as a target organ, and macrophage lysosomes were the organelle for toxicity. Proteomic data revealed upregulation of 197, and downregulation of 75 spleen relevant proteins, agreeing mechanistically with biomarker findings. Charlwood et al. studied gentamicin nephrotoxicity using proteomics. 129 They determined that gentamicin perturbed twenty proteins that were involved in energy production and mitochondrial function, leading to cellular toxicity in rat kidneys. Moller et al. used 2D gel electrophoresis followed by proteomic analysis to investigate daunorubicin effects on pancreatic carcinoma cells. 130 With daunorubicin exposure, they demonstrated dose-dependent protein upregulation, which was undetected in RT-PCR-based gene expression studies. This study supported the notion that protein level changes signify functional outcome better than mRNA level changes.
Fluorescence-based 2D gel electrophoresis was utilized to study the biochemical basis of acetaminophen (APAP) hepatotoxicity in mice. 131 Mouse liver homogenates from treated and control mice were run on dye-based 2D gel, and differences in the protein expressions were quantified. A total of twenty different proteins were identified, and nine of them have been matched to known APAP cytotoxicity pathway proteins. However, eleven of these identified proteins (apolipoprotein Al, fatty acid binding protein, malate dehydrogenase, heat shock cognate protein, hemoglobin beta chain, antioxidant protein 2, arginase 1, UV excision repair homolog, alpha enolase, glucose regulated protein, and apolipoprotein A4) have not been previously linked to APAP-toxicity. Authors postulate that proteomic analysis points to further details into the mechanism of APAP hepatotoxicity. As LC-MS technology makes advances towards more sensitive and sophisticated instruments, proteomic analysis benefits from it. Additionally, proteomic analysis in combination with pathology can be a powerful predictive tool in drug discovery. The scope of proteomic analysis is limited with available platforms. Also, the sample processing and identification of relevant protein changes remains a challenge. Transcriptomics studies prior to proteomic analysis can be beneficial in guiding specific proteome studies. Given the practical considerations and limits of the technology, global proteomic analysis remains largely uncovered.
Metabolomics
Metabolomics is defined as comprehensive and detailed endogenous metabolite evaluation to provide biochemical status of the system under the influence of xenobiotics. 132 Cellular metabolism routinely produces small molecules including carbohydrates, amino acids, nucleotides, lipids, steroids, fatty acids, and so on. Metabolomics can be applied to rodent body fluids, tissues or whole organisms, and also to the in vitro systems. The experimental approaches in metabolomics can be divided according to various criteria, for instance, in vitro vs in vivo, or targeted vs non-targeted.
LC-MS-based techniques are leading instrumentation used in metabolomic analysis. NMR is the second most used instrumentation besides MS. Initial metabolic footprints of the xenobiotic exposure can be further characterized by targeted analysis of specific set of metabolites using LC-MS. This is a commonly used approach to obtain confirmatory quantification data. Overview of typical metabolomic assay is displayed in Figure 3. Samples that can be used include plasma, urine, serum, or tissue extracts. The data obtained from NMR or MS-based assay is then subjected to multivariate analysis through hierarchical clusters, or principal component analysis among other methods. These statistical methods facilitate the classification of the data and point out which changes in metabolites could be significant. After data are sorted through statistical programs, the results are sorted against existing database, including human metabolome database, reactome database, Ingenuity pathway database, and Cytoscape among others. These are commonly used databases in metabolic analysis. Overview of the metabolomic assay.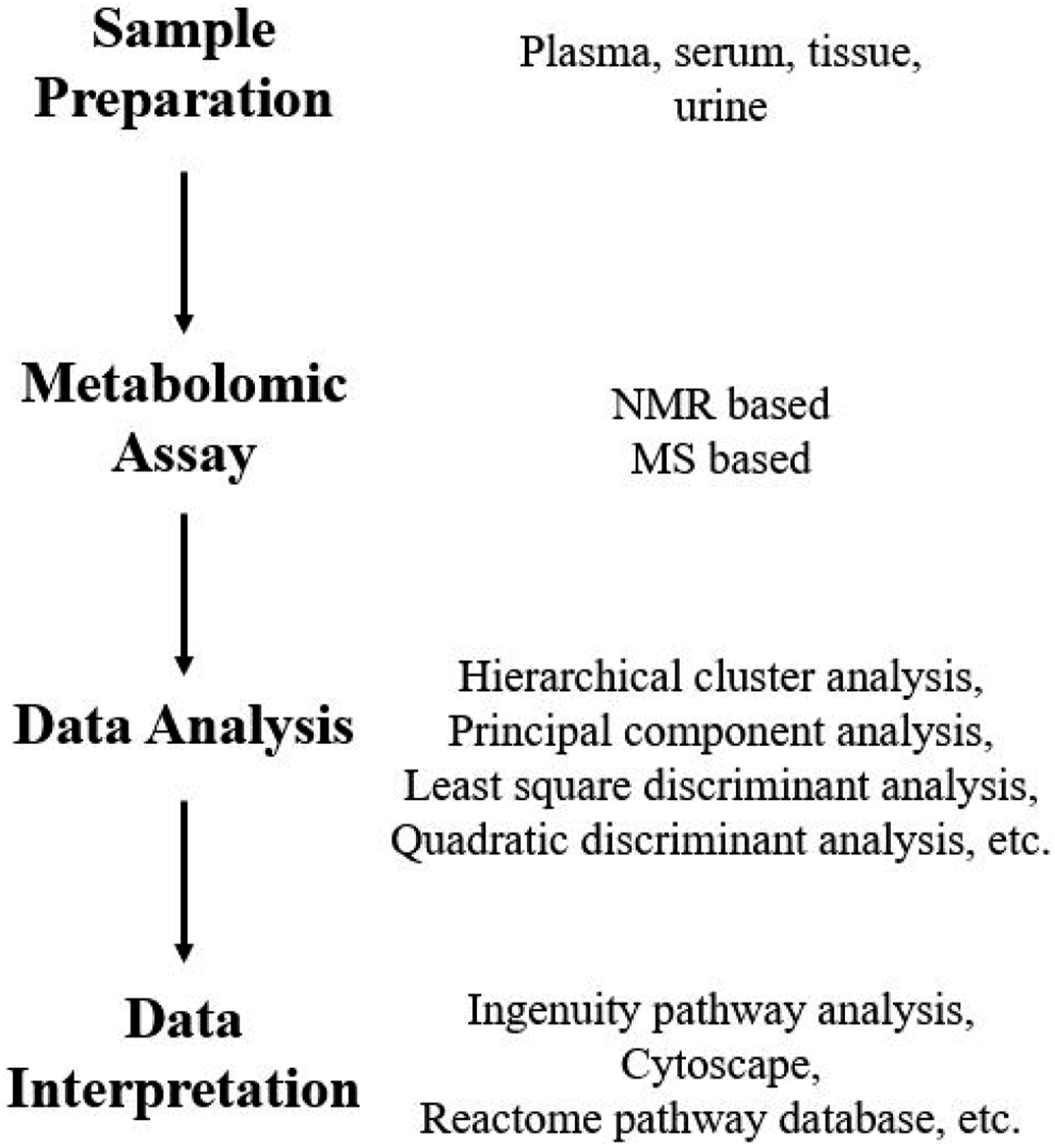
Increasing need to move towards relevant non-animal models, metabolomics analysis in
Targeted metabolomics focuses on a specific molecular pathway, organ system, or pathological mechanism. The metabolites to be identified are pre-decided. Primary bile acids are synthesized in the liver and released into the intestinal lumen, wherein microflora synthesizes secondary bile acids. Most bile acids are returned to the liver through enterohepatic recirculation. 135 Increase in the bile acid levels in systemic circulation or urine could potentially indicate hepatic elimination disturbances, transporter expression changes, hepatotoxicity or intestinal inflammation. Want et al. described an ultra-performance LC-MS method to measure bile acids in biofluids. 136 Bile acids were extracted from serum and quantified on Q-TOF mass spectrometer. In rats exposed to galactosamine, taurine-conjugated bile acid levels were elevated, indicating hepatotoxicity. These data were supported by histopathological scoring of the liver tissue.
Nontargeted metabolomics generates a comprehensive list of metabolites from the biological system. This information, when sorted according to specific pathways, can provide mechanistic information about xenobiotic exposure. Nontargeted metabolic signature of the known toxicants can serve as a guideline for understanding the unknown compound effects.
Bristol Myers Squibb published a transcriptomic-metabolomic combination study that identified ascorbate and gluonic acid as preclinical markers of microsomal CYP induction. 137 This was the first attempt at identifying the microsomal inducers so early in the screening stages. This provided non-invasive method to identify CYP induction, and the data were confirmed with conventional primary hepatocyte cultures. Cyclosporin’s clinical use as an immunosuppressant is limited because of its nephrotoxicity. 138 University of Colorado conducted an open label single oral dose study with cyclosporin in 13 healthy volunteers. 139 NMR and LC-MS-based metabolomic analysis of urine samples revealed that oxidative stress biomarker isoprostane levels were elevated compared to the placebo group. Additionally, citrate, hippurate, lactate, creatinine, and phenylalanine levels also showed significant change after single dose cyclosporin administration. These changes clearly indicated cyclosporin-induced adverse effects on renal function. Duan et al. reviewed utility of metabolomics in hepatotoxicity, nephrotoxicity and cardiotoxicity for traditional Chinese medicine. 140
With increasing utility of metabolomics, early discovery predictions relevant to drug safety are possible. This is increasingly becoming common practice across the biopharmaceutical industry.
Epigenetics: Micro-RNA
Toxicants may act on areas "above or in addition" to genes. 141 Epigenetics establishes tissue-specific and heritable patterns of gene expression without changing the DNA sequence. Several epigenetic mechanisms, including DNA methylation, histone modification, and microRNA expression have been demonstrated to play a significant role in modern toxicology. DNA methylation and histone modification have been well studied, so these topics will not be discussed here. 142
Micro-RNAs (miRNA) are a class of small non-coding single-stranded RNAs with lengths of 19 to 25 nucleotides that are involved both in the nuclear and the posttranscriptional regulation of gene expression.
143
Since the first miRNA, lin-4, was discovered more than 20 years ago by Victor Ambros and his colleagues in
Extracellular miRs are also called circulation miRNAs. It was found that miRNAs are stable and resistant to RNase activity and extreme pH and temperatures in different body fluids, including serum, plasma, urine, saliva, at different concentrations.150,151 For cancer diagnosis, using circulating miRNAs is invasive as traditional diagnostic method commonly involves a biopsy of the cancer tissue. 152 For instance, miRNA levels in the liver have been suggested to be associated with serum levels for some of miRNAs. 153 MiRNAs can be released through passive outflow due to cellular damage, such as inflammation and necrosis, or through extracellular vesicles such as apoptotic bodies, exosomes and micro vesicles, or through macromolecular complexes with Argonaute proteins (AGO) and high-density lipoprotein (HDL) to form RNA-binding proteins.154,155 Chim et al. were the first to report using circulating miRNAs as diagnostic tools to detect placental miRNAs in maternal plasma. 156 Then another study reported that miR-155, miR-210, and miR-21 were significantly up-regulated in the serum of patients with diffuse large B-cell lymphoma. 157 Serum miRNAs are usually found in exosomes. 158 The miRNAs in some exosomes have been shown to be quite different from the miRNAs in secreting cells and it might be regarded as hormones delivering epigenetic information. However, some studies have suggested that most exosomes lack miRNA. 159 Unfortunately, these are only hypotheses now and the molecular mechanism for this is still unclear. But at least, some of these extracellular miRNAs may act as death signal in cancer cells.
Apart from being promising as a toxicity biomarker, miRNAs also play an important role in predicting xenobiotic toxicological effects. Cytochrome P450s (CYPs) are enzymes that metabolize xenobiotics and the regulation of CYPs and nuclear receptors by miRNAs has been well investigated. 160 For example, miR-142-3p and miR-200a have been found to regulate CYP1A1 and 1A2, miR-134a regulates CYP2C19, miR-10a and let-7g regulates CYP2E1, and let-7b is reported to regulate CYP2D6. 161 Nuclear receptors, including aryl hydrocarbon receptor (AhR, targeted by miR-375), pregnane X receptor (PXR, targeted by miR-148a), and constitutive androstane receptor (CAR, targeted by miR-122), are targeted by several miRNA and represent CYP regulation.162-164
Conclusions
Drug safety and toxicology are essential disciplines within drug discovery and development. Before new chemical entities are ready to be administered in the healthy volunteer studies, its thorough safety characterization is requested from regulatory agencies. The late stage GLP safety studies are time-consuming, long-term and expensive, hence the number of compounds that can be selected for GLP toxicology studies have to be limited. This makes the discovery safety screening studies significant and imperative. Given the larger number of compounds at the early discovery stage, high throughput screening assays are most appropriate for safety screening.
Here, we described recent advances in the discovery safety screening technologies. Given the increasing need to accommodate non-animal screening methods in drug discovery research, we focused mainly on in vitro models and multicellular model organisms. In addition, biomarkers are described in greater detail as "omics" platforms are exploding in technological space, and their applications in toxicology are endless.
In the last decade, MPS technology has improved a lot. Combination of multiple organoid scaffolds connected with common circulating medium, and its operation monitored with mechanical pumps and other engineering tools has made the MPS more feasible, relevant, and applicable biological systems. For further exploration of MPS into toxicology, sophisticated in vitro to in vivo extrapolation methods are needed. Also, common media conducive of supporting diverse organ scaffolds limit the combinations possible. Imaging methods provide support for ex vivo and in vitro biomarker and pathology exploration. With sensitive and specific mass spectrometry-based methods, imaging technologies have a certain future in the discovery safety screening. Importantly, localization of the tissue characteristic and specific region-based data are prime advantages of the imaging techniques in toxicology.
Model organisms including nematodes and zebrafish have emerged as whole biological system alternatives to the in vitro organoids or tissue culture. These systems represent independent functioning biological systems that mimic the human body in a unique way. For heavy metal toxicity screening, for mechanistic studies, and for screening level assays, model organisms are a priority in academic toxicology. Industrial acceptance of model organisms for toxicology screening remains an evolving area of science. Specifically, neurotoxicology screening, model organisms hold promise to replace other in vitro techniques.
For toxicology biomarkers, omics platforms are gaining more traction within industry environment. Toxicogenomic approaches involve combination of genomic, transcriptomic, proteomic and metabolomic technologies. Industry-wide acceptance of toxicogenomics has grown over the last two decades. Early signals obtained from omics assays allow drug discovery teams to structurally modify the risky chemical scaffolds and select the least troubling chemotypes. The translation rate of omics predictions to in vivo observations is quite high, which makes these technologies reliable. Along with genomic evaluations, epigenetic markers including microRNA have a place in diagnostic and biomarker research. Circulating microRNAs have become popular in organ toxicity screening. The ease of sample collection and analysis makes these markers unique.
With the advent of sophisticated in vitro and imaging technologies, toxicology screening is gaining newer tools. With regulatory push for reduced use of animals, and increasing translational ability of in vitro tools, the discipline of toxicology is evolving into a modern branch of science that is vital to the drug discovery process.
Footnotes
Author Contributions
Tang, B. and More, V. contributed to conception and design, drafted manuscript, critically revised manuscript, gave final approval and agree to be accountable for all aspects of work ensuring integrity and accuracy.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
