Abstract
DNA damage is an established initiating event in the mutagenicity and carcinogenicity of genotoxic chemicals. Accordingly, assessment of this endpoint is critical for chemicals which are being developed for use in humans. To assess the ability of the Chicken Egg Genotoxicity Assay (CEGA) to detect genotoxic pharmaceuticals, a set of 23 compounds with different pharmacological and reported genotoxic effects was tested for the potential to produce nuclear DNA adducts and strand breaks in the embryo-fetal livers using the 32P-nucleotide postlabeling (NPL) and comet assays, respectively. Due to high toxicity, two aneugens, colchicine and vinblastine, and an autophagy inhibitor, hydroxychloroquine, could not be evaluated. Out of the 20 remaining pharmaceuticals, 10 including estrogen modulators, diethylstilbestrol and tamoxifen, antineoplastics cyclophosphamide, etoposide, and mitomycin C, antifungal griseofulvin, local anesthetics lidocaine and prilocaine, and antihistamines diphenhydramine and doxylamine, yielded clear positive outcomes in at least one of the assays. The antihypertensive vasodilator hydralazine and antineoplastics streptozotocin and teniposide, produced only DNA strand breaks, which were not dose-dependent, and thus, the results with these 3 pharmaceuticals were considered equivocal. No DNA damage was detected for 7 compounds, including the purine antagonist 6-thioguanine, antipyretic analgesics acetaminophen and phenacetin, antibiotic ciprofloxacin, antilipidemic clofibrate, anti-inflammatory ibuprofen, and sedative phenobarbital. However, low solubility of these compounds limited dosages tested in CEGA. Overall, results in CEGA were largely in concordance with the outcomes in other systems in vitro and in vivo, indicating that CEGA provides reliable detection of DNA damaging activity of genotoxic compounds. Further evaluations with a broader set of compounds would support this conclusion.
Keywords
Introduction
Chemicals being developed as potential human medications for treatment of non-life threatening, sometimes chronic, conditions are typically required by regulatory agencies to undergo testing for carcinogenicity in rodent models.1-3 Such testing takes years to complete and results are usually not available when clinical trials are first undertaken. To provide a level of assurance of safety, short term assays for mutagenicity are required as surrogates. 4 These generally assess bacterial and mammalian cell mutagenicity, as well as structural and numerical chromosomal aberrations, ie, clastogenicity and aneugenicity.5,6
A common underlying molecular mechanism of the mutagenicity and carcinogenicity of some chemicals is a reaction with nuclear DNA, referred to as genotoxicity.7,8 For example, formation of pro-mutagenic DNA lesions is considered to be an initiating event in the adverse outcome pathway for induction of hepatocellular carcinoma by aflatoxin B1. 9 Thus, comet assay and DNA covalent binding assays are listed as other in vivo genotoxicity assays 3 and the comet assay is frequently used in Option 2 of International Conference on Harmonisation of Technical Requirements for Registration of Pharmaceuticals for Human Use (ICH) S2 (R1). 4 However, none of the currently required genotoxicity and mutagenicity batteries, includes an assay for DNA reactivity. Also, although the liver is the organ with the broadest xenobiotic biotransformation capability of any tissue, 10 none of the recommended assays utilizes the liver as the target organ. While the in vivo transgenic mutation assay includes the liver, this is a mutation assay and not one for DNA reactivity.
An assay which incorporates both of these features (DNA reactivity and liver bioactivation) is the hepatocyte DNA damage assay, with the endpoints of either DNA repair 11 or DNA adduct formation. 12 It utilizes intact hepatocytes with intrinsic metabolic capability, which is more comprehensive than that of subcellular preparations13,14 used in other in vitro assays. Results in the DNA damage assay were highly concordant with demonstrated genotoxicity or the presence of chemical structures underlying genotoxicity for a wide variety of genotoxic agents, 15 confirming the versatility of liver cells in bioactivating carcinogens, including ones that do not have the liver as their target organ. The assay also has the important feature that human cells can be used.12,16
Liver spheroids, including 3D HepaRG liver models, 3D reconstructed primary human hepatocyte model, liver-chip, can be used to assess metabolism-dependent drug-induced liver injury (DILI) as a part of screening for drug toxicity.17,18 These models are capable of intrinsic metabolic activation and can identify drug-specific metabolites that are formed in humans. However, it should be taken into consideration that the activity of certain metabolic enzymes in vitro declines with time.
To further explore the capability of the liver in detecting DNA-reactive chemicals, the Chicken Egg Genotoxicity Assay (CEGA), and related Turkey Egg Genotoxicity Assay (TEGA) were developed.19-24 These models use intact embryo-fetal livers of fertilized avian eggs, which provide bioactivation in a self-supporting but non-animal organism. The effects of chemicals injected into the eggs on liver nuclear DNA (genotoxicity) is assessed by 32P-nucleotide postlabelling (NPL) and comet assays. These endpoints detect the main types of chemical-induced DNA damage, namely, nuclear DNA adduct formation measured by thin-layer chromatography of hydrolyzed DNA or DNA breaks measured by electrophoretic migration of nuclear DNA.25,26
A wide range ot chemicals has been tested in CEGA and TEGA (almost 70 have already been reported, including alkenyl benzenes, flavor and fragrance materials, aromatic amines, dialkylnitrosamines, polycyclic aromatic hydrocarbons, aflatoxins). The results yielded 100% specificity with a high degree of sensitivity.19,20,22-24,27 In support of the validity of the NPL assay, the chemical structures of DNA adducts produced by alkenylbenznes detected by the NPL method have been confirmed by mass spectrometry. 28
Here, we report testing in CEGA of 23 pharmaceuticals for which published results on mutagenicity and/or carcinogenicity are available. The 23 pharmaceutical agents were selected to comprise mutagenic and non-mutagenic compounds. Investigated pharmaceuticals were of the following types: antineoplastic agents for hormonal therapy (tamoxifen and diethylstilbestrol) and chemotherapeutics (cyclophosphamide, mitomycin C, streptozotocin, etoposide, teniposide, 6-thioguanine, and vinblastine), an antifungal (griseofulvin), local anesthetics (lidocaine and prilocaine), antihistamines (diphenhydramine and doxylamine), an antihypertensive vasodilator (hydralazine), a sedative barbiturate (phenobarbital), a nonsteroidal anti-inflammatory drug (ibuprofen), an anti-inflammatory tubulin blocker for prevention of gout attacks (colchicine), an antimalarial (hydroxychloroquine), antipyretic analgesics (phenacetin and its metabolite acetaminophen), an antibiotic (ciprofloxacin), and an antilipidemic fibrate (clofibrate) (Figure 1). Chemical structures of tested pharmaceuticals. PC, positive comparator.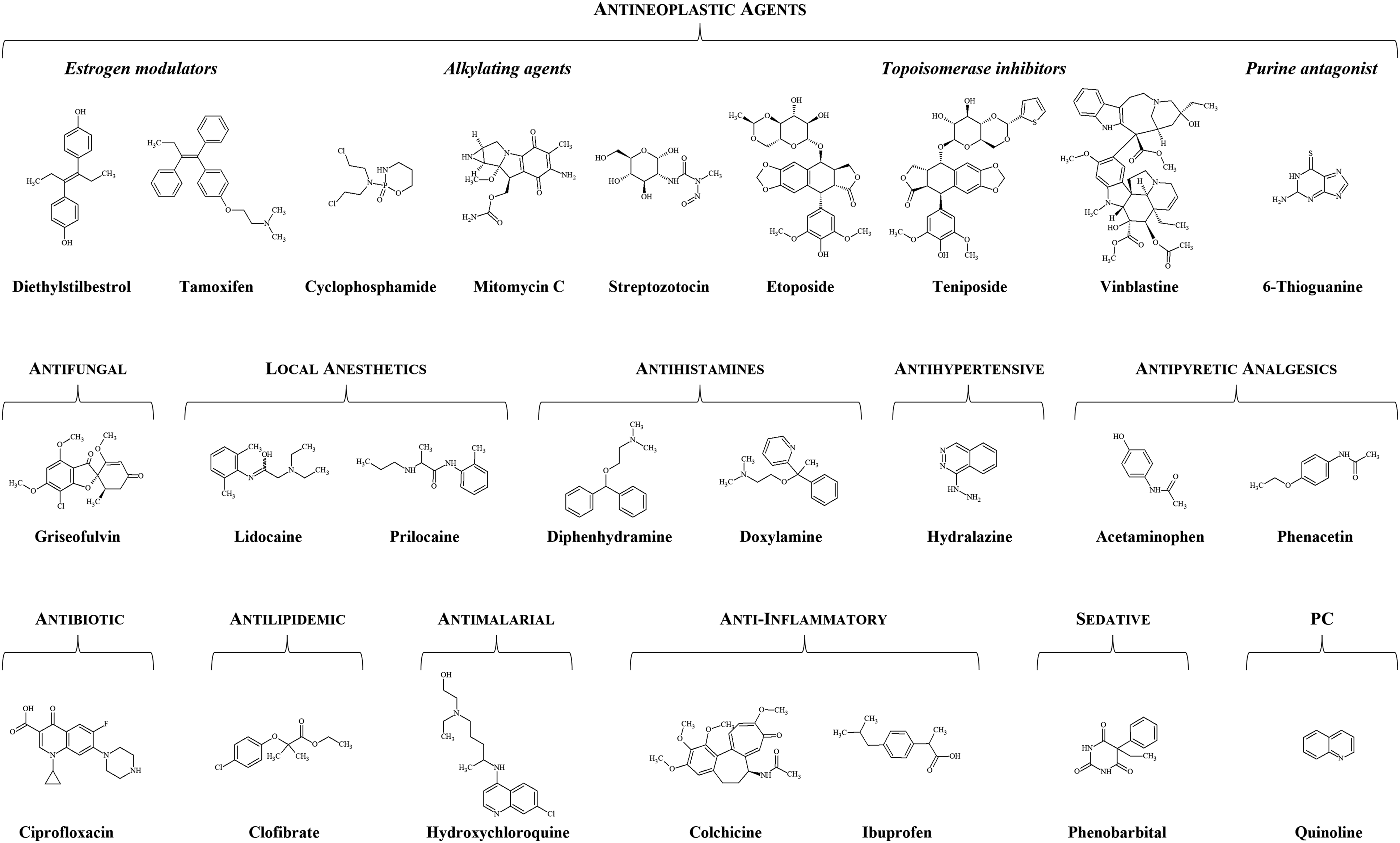
Findings in this study confirm that DNA reactivity detected in CEGA is a reliable indicator of genotoxicity.
Materials and Methods
Chemicals
Results of Testing of Pharmaceuticals in the CEGA.
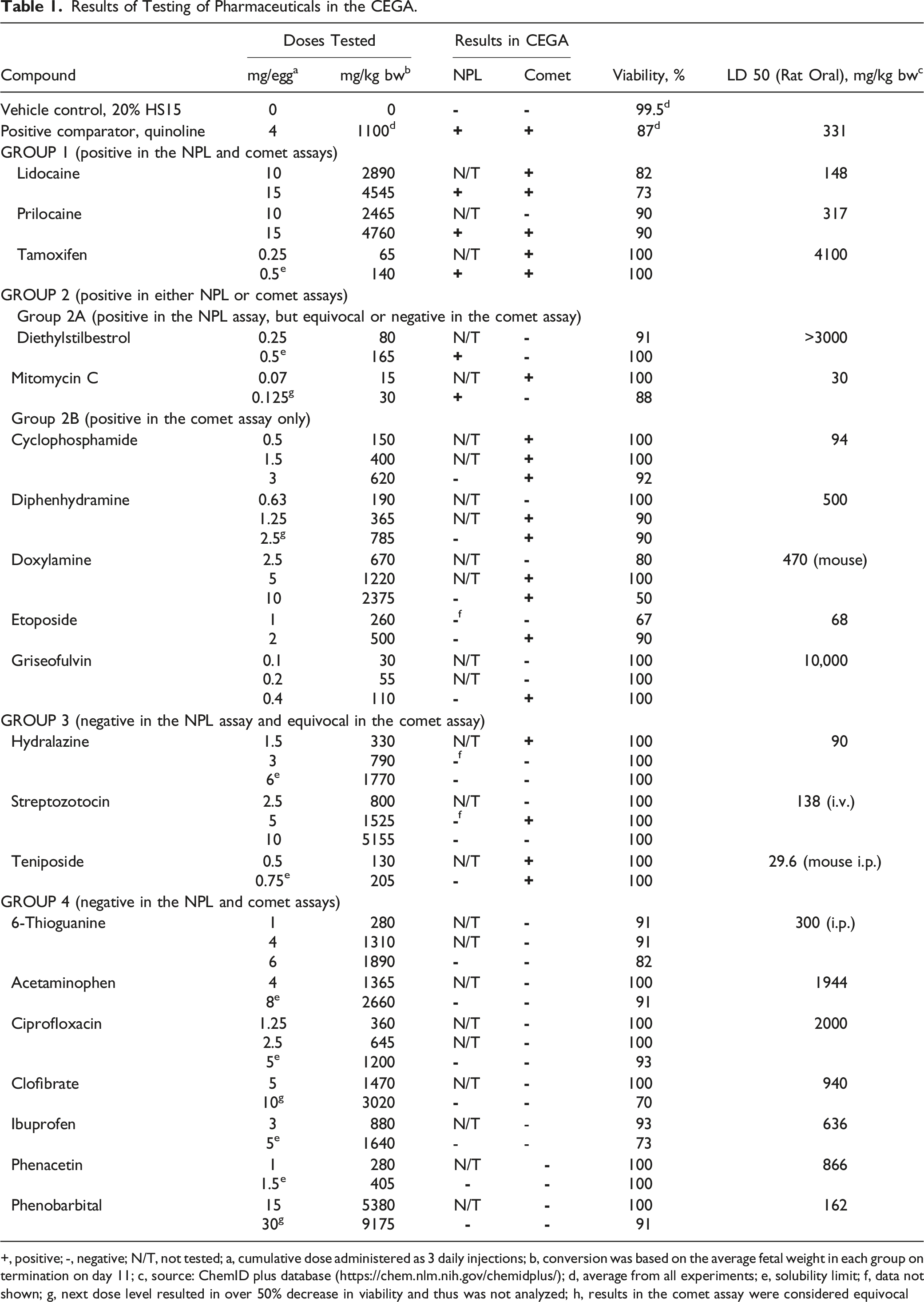
+, positive; -, negative; N/T, not tested; a, cumulative dose administered as 3 daily injections; b, conversion was based on the average fetal weight in each group on termination on day 11; c, source: ChemID plus database (https://chem.nlm.nih.gov/chemidplus/); d, average from all experiments; e, solubility limit; f, data not shown; g, next dose level resulted in over 50% decrease in viability and thus was not analyzed; h, results in the comet assay were considered equivocal
All tested compounds were dissolved in Kolliphor HS15 (CAS: 70,142-34-6) obtained from Sigma-Aldrich (St Louis, MO, USA) and prepared as a 20% aqueous solution (20% HS15). This vehicle was selected on the basis of its ability to dissolve both, water soluble and lipophilic compounds. It was demonstrated to be nontoxic to the developing avian embryo-fetuses in the previous studies. Also, it demonstrated no genotoxicity in the comet and NPL assays. Quinoline (CAS: 91-22-5; 98%) was purchased from Fisher Scientific (Waltham, MA, USA) and served as a positive comparator (PC).
Tamoxifen citrate (CAS: 54,965-24-1; ≥98% pure as reported by supplier) was obtained from Fisher Scientific (Waltham, MA, USA). Diethylstilbestrol (CAS: 56-53-1; ≥99%), cyclophosphamide monohydrate (CAS: 6055-19-2), mitomycin C (CAS: 50-07-7; ≥98%), etoposide (CAS: 33 419-42-0; 98%) teniposide (CAS: 29 767-20-2; ≥97%), griseofulvin (CAS: 126-07-8; ≥97%), hydralazine hydrochloride (CAS: 304-20-1), phenobarbital sodium (CAS: 57-30-7), ibuprofen (CAS: 15,687-27-1; ≥98%), hydroxychloroquine sulphate (CAS: 747-36-4; ≥98%), ciprofloxacin (CAS: 85,721-33-1; ≥98%), and clofibrate (CAS: 637-07-0; >99%) were purchased from Sigma-Aldrich (St Louis, MO, USA). Streptozotocin (CAS: 18,883-66-4; ≥97%), 6-thioguanine (CAS: 154-42-7; 98%), diphenhydramine hydrochloride (CAS: 147-24-0; 99%), and colchicine (CAS: 64-86-8; 98%) were purchased from Alfa Aesar (Tewksbury, MA, USA). Vinblastine sulphate (CAS: 18,883-66-4; >99%), lidocaine hydrochloride (CAS: 6108-05-0; >99%), prilocaine hydrochloride (CAS: 1786-81-8), and doxylamine succinate (CAS: 562-10-7; >99%) were purchased from MP Biomedicals Inc. Phenacetin (CAS: 62-44-2; ≥98%) was obtained from Aldrich Chemical Company, Inc. (Milwaukee, WI, USA). Acetaminophen (paracetamol) (CAS: 103-90-2) was obtained from ICN Biomedicals, Inc. (Aurora, OH, USA). (Solon, OH, USA).
Dosages were selected based on available toxicity data (oral LD50 in rodents), assuming the average mass of the chicken egg is 60 g. As indicated in Table 1, the highest tested doses of tamoxifen, diethylstilbestrol, teniposide, hydralazine, ibuprofen, phenacetin, acetaminophen, and ciprofloxacin were close to the solubility limit determined by serial dilution. Selection of the highest dose for testing of mitomycin C, vinblastine, diphenhydramine, phenobarbital, colchicine, hydroxychloroquine, and clofibrate was limited by cytotoxicity.
Experimental Design
Specific pathogen-free (SPF), fertilized white leghorn chicken (Gallus gallus) eggs of undetermined sex were obtained from Charles River Laboratories (North Franklin, CT, USA). The experimental protocol is described in detail in Williams et al. 24 Eggs were incubated in 2GIF Styrofoam incubators with automatic egg turners (Murray McMurray Hatchery, Webster City, IA, USA) at 37 ± 0.5°C and 60 ± 5% humidity. The repeated dose regimen consisted of 3 daily injections with vehicle or test substances administered into the air sac on days 9 through 11 of incubation. The application volume for vehicle and tested substances was 50 μl/egg/day (0.15 ml/egg total volume). Eggs were terminated three hours after the last injection on day 11. Upon termination, body and liver weights, as well as viability, were recorded. Three liver samples per group were processed immediately for the comet assay, and the remaining liver samples were frozen at −80°C for subsequent NPL assay. Groups chosen for analysis had viability levels higher than 50% (at least half of the fetuses in the group are viable upon termination) to avoid false-positive results due to cytotoxicity. In each assay 3 liver samples per group were analyzed, owing to the limitation in the number of samples which can be analyzed simultaneously.
Detection of Nuclear DNA Adducts
The NPL assay was conducted on DNA extracted from frozen liver samples (n = 3 samples (eggs) per group) using genomic QIAGEN G100 columns (Valencia, CA, USA) following the manufacturer’s protocol as previously described. 24 Nuclear DNA (10 μg per sample) was processed for the assay, with nuclease P1 (NP1) digestion. 29 Where adducts were not detected, hydrophilic-lipophilic balance (HLB) column enrichment was used. 30 Labelled modified bases were resolved using two- or three-directional thin-layer chromatography (TLC) systems. The radioactivity on the TLC plates was detected using a Molecular Dynamics Storm 860 system with exposure times of 2 h to overnight (GE Health Care Life Sciences, Edison, NJ, USA). Only groups which received the highest tested doses of pharmaceuticals were analysed in the NPL assay, since they are most likely to reveal the presence of adducts.
Detection of Nuclear DNA Strand Breaks
For alkaline single cell gel electrophoresis (comet) assay, 200 mg of each liver sample (n = 3 samples (eggs) per group) were minced into fine pieces in 1 ml of ice-cold Hank’s balanced salt solution (HBSS, Gibco, Grand Island, NY, USA) containing 20 mM EDTA and 10% dimethylsulfoxide (DMSO). Aliquots of the cell suspensions (about 5 × 105 cells in 10 µl of suspension) were then embedded in 1.0% low melting point agarose on a slide and processed in the standard comet assay according to OECD guidelines, 31 as previously reported. 24 Electrophoresis was applied and nuclear DNA migration was analyzed by fluorescence microscopy using a Nikon OPTIPHOT microscope. The percentage of DNA-in-tail in samples was determined using the Comet Score software v 1.5 (TriTek Corp, Sumerduck, Virginia), counting >150 cells per sample. Median % tail DNA for each slide was determined and the mean was calculated for each embryo-fetus.
The evaluation criteria for determining positive comet results are described in detail in Kobets et al. 20 The historical vehicle (20% HS15) control range (95% confidence interval (CI) established in the laboratory based on over 40 experiments is 7.3-8.2 with a margin of error of 0.45).
Statistical Analyses
Results are presented as mean ± SD. Statistical analyses were performed using GraphPad Prism software version 7 (GraphPad Software Inc, La Jolla, CA). Data were analyzed by the one-way analysis of variance (ANOVA) with the pairwise multiple comparison being made by Dunnett’s method. Linear regression analysis was used to determine dose-related trends. P-values < 0.05 were considered significant.
Results
Findings in CEGA for tested pharmaceuticals with a variety of pharmacologic activities and chemical structures are summarized in Table 1. Low solubility limits of acetaminophen, ciprofloxacin, diethylstilbestrol, hydralazine, ibuprofen, phenacetin, tamoxifen, and teniposide (Table 1), limited the highest dose tested in CEGA, which could have affected the outcomes in the assay.
The average viability of control groups in all experiments was 99.5%, while in groups that received the positive comparator, quinolone, average viability was 87% (Table 1; Figures 3-5). Three pharmaceuticals, two of which, colchicine and vinblastine, cause mitotic spindle dysfunction, and one, hydroxychloroquine, inhibits lysosomes and autophagy, exhibited high lethality in the system. Specifically, no embryo-fetuses were alive in the groups dosed with colchicine at 0.005-3 mg/egg, vinblastine at total dosages of 0.31-1.25 mg/egg, and hydroxychloroquine at 5-20 mg/egg. Thus, these compounds could not be evaluated in the NPL or comet assays. In other dosed groups viability was at least 50% which is still acceptable for analyses (Table 1).
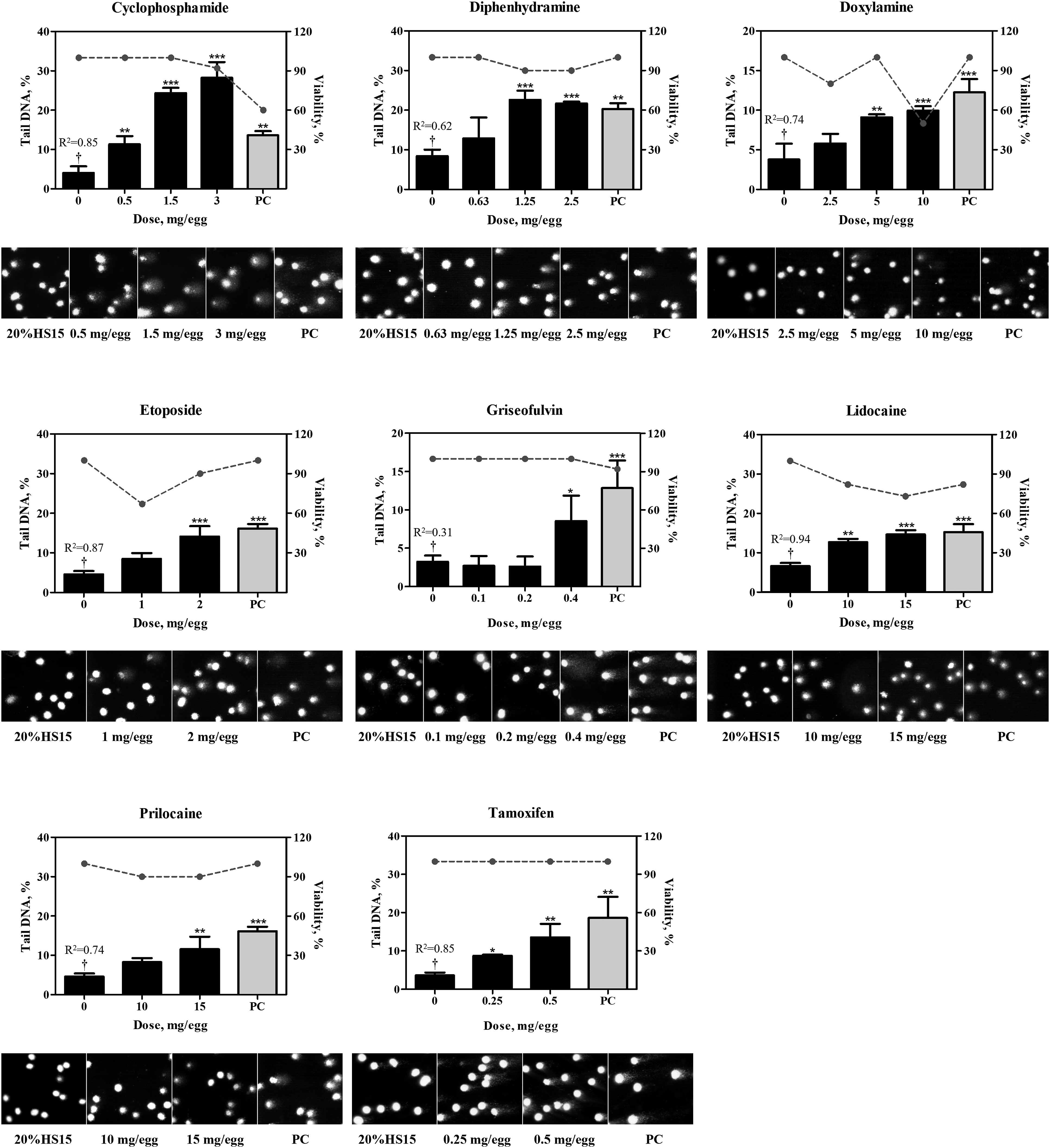
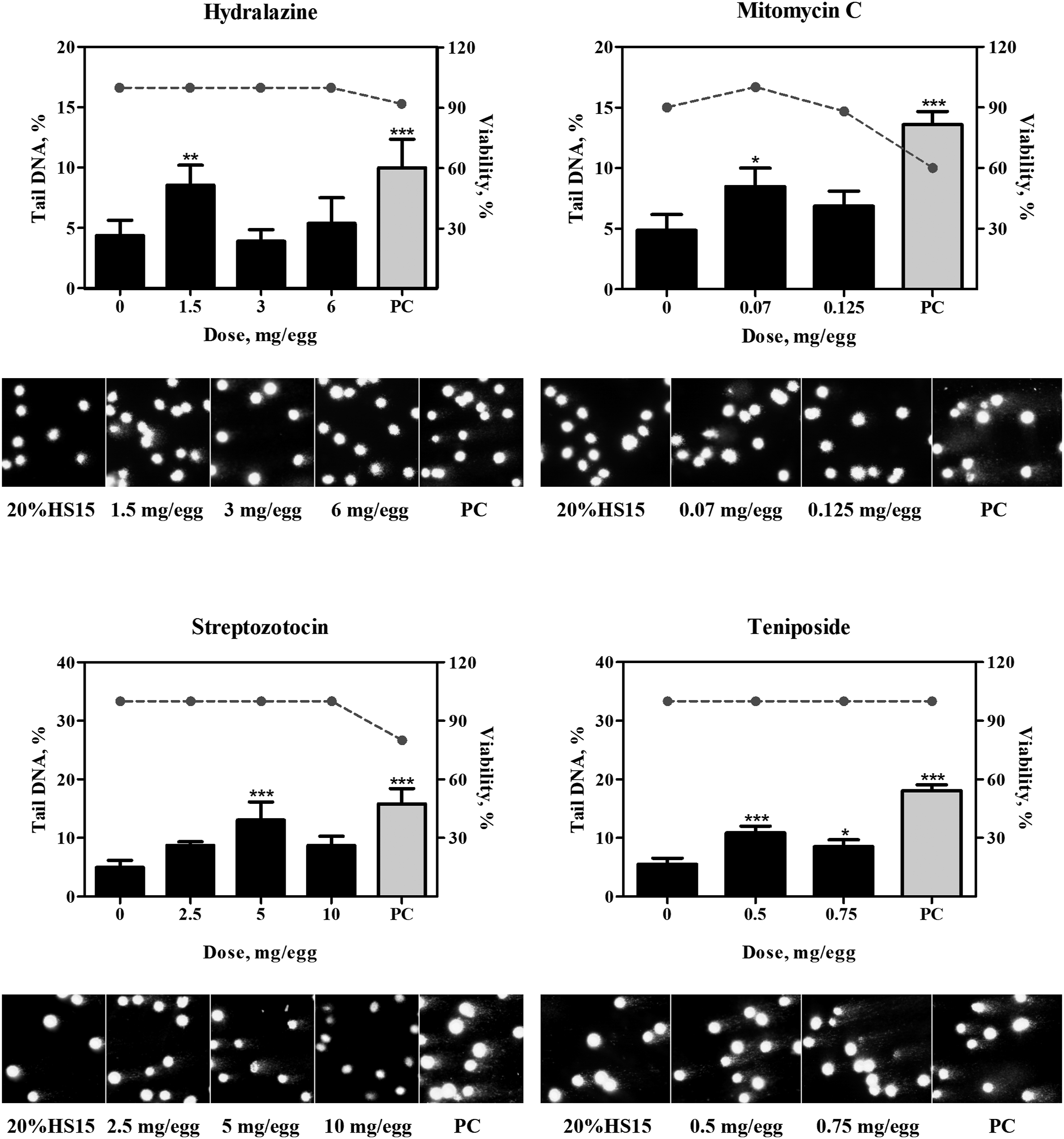
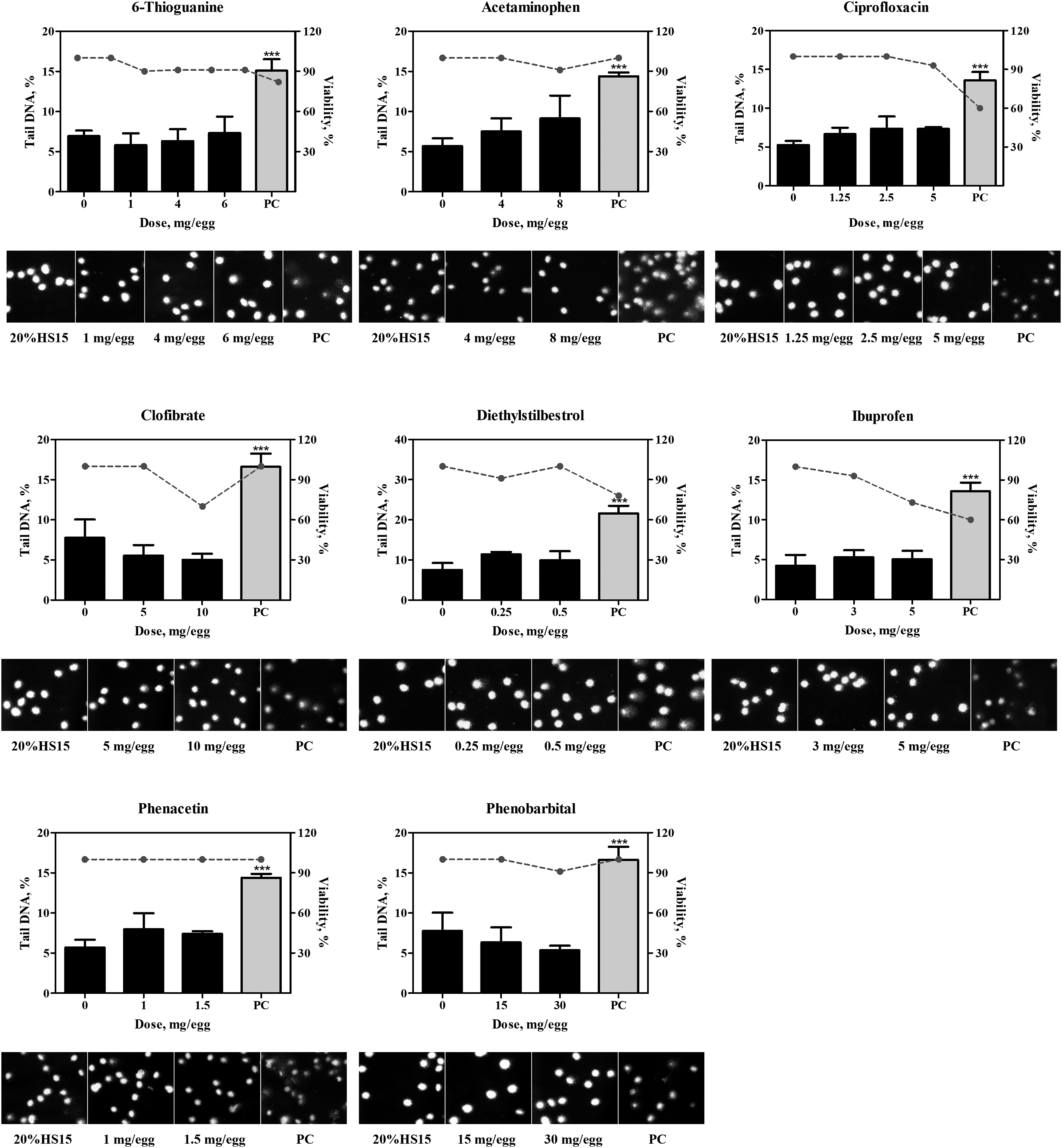
In the control groups that received vehicle, ie, 20% HS 15, no increases above the historical control range in DNA adducts or DNA migration were detected. The positive comparator, quinoline, at the total dose of 4 mg/egg consistently produced positive results in both, the NPL and comet assays (Table 1; Figures 2-5). 32P-nucleotide postlabeling assay (NPL). Chromatograms for compounds which yielded positive (A.) or negative (B.) responses. DNA adducts were resolved in the second and third directions of chromatography and are indicated by arrows. C. Comparison of DNA-binding potencies of pharmaceuticals with positive response in the NPL assay in the embryo-fetal chicken livers. Conversion to adducts per µmol was conducted using the molar mass of the chemicals (Suppl. Table 1). PC, positive comparator, quinoline 4 mg/egg. Pharmaceuticals which produced positive results in the comet assay. Bar graphs present comet values (percentage of DNA in tail, Suppl. Table 2) as mean ± SD (n = at least 3 samples per group), dotted line represent percent viability in the corresponding group. Representative photomicrographs of comet assay for each group are provided below each chart. *, represents significant (P < 0.05) difference from corresponding control group; **, represents significant (P < 0.01) difference from corresponding control group; ***, represents significant (P < 0.001) difference from corresponding control group. †, represents significant (P < 0.05) dose-related trend. PC, positive comparator, quinoline 4 mg/egg. Pharmaceuticals which produced equivocal results in the comet assay. Bar graphs present comet values (percentage of DNA in tail, Suppl. Table 2) as mean ± SD (n = at least 3 samples per group), dotted line represent percent viability in the corresponding group. Representative photomicrographs of comet assay for each group are provided below each chart. *, represents significant (P < 0.05) difference from corresponding control group; **, represents significant (P < 0.01) difference from corresponding control group; ***, represents significant (P < 0.001) difference from corresponding control group. PC, positive comparator, quinoline 4 mg/egg. Pharmaceuticals which produced negative results in the comet assay. Bar graphs present comet values (percentage of DNA in tail, Suppl. Table 2) as mean ± SD (n = at least 3 samples per group), dotted line represent percent viability in the corresponding group. Representative photomicrographs of comet assay for each group are provided below each chart. *, represents significant (P < 0.05) difference from corresponding control group. PC, positive comparator, quinoline 4 mg/egg.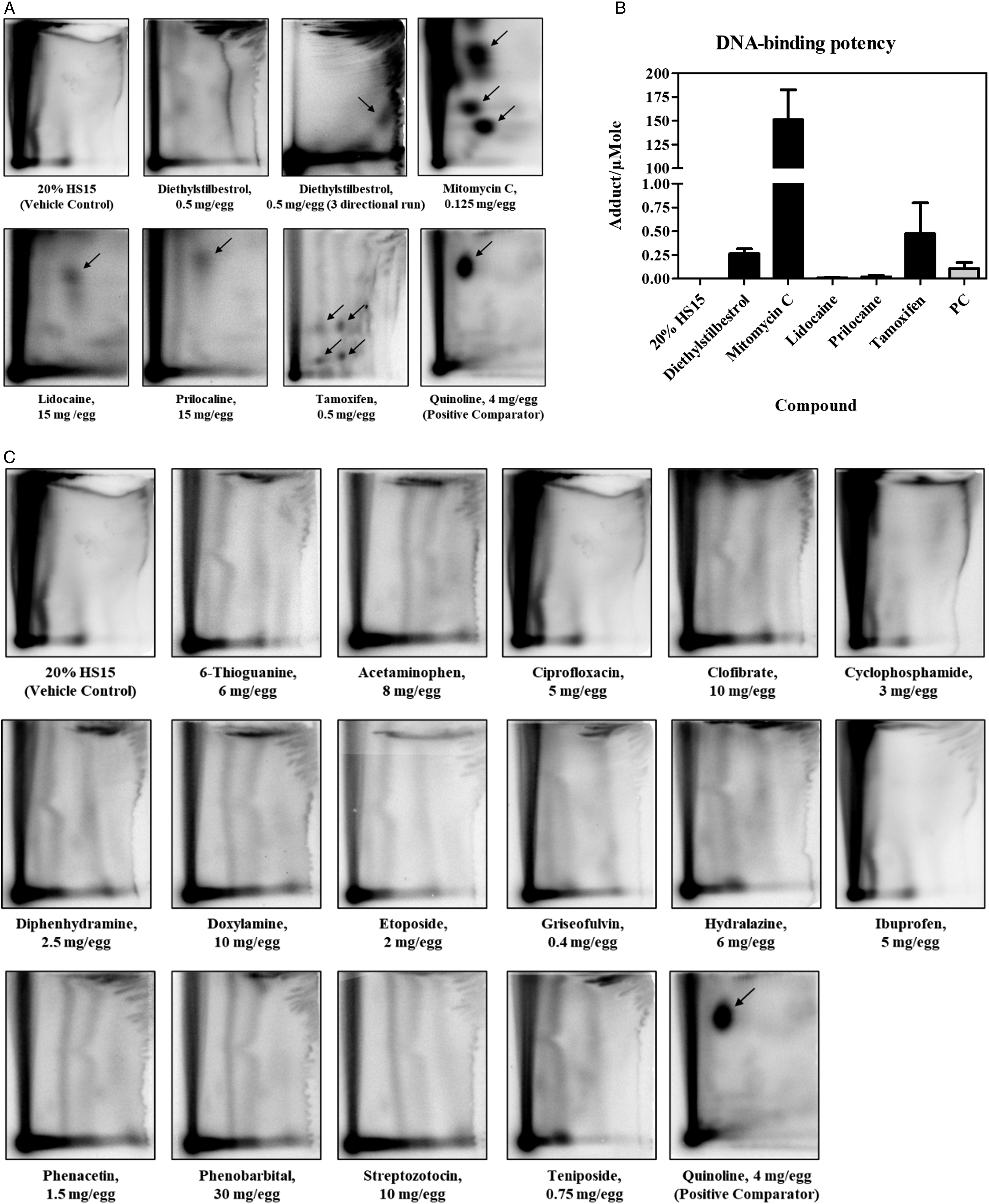
In the NPL assay, four compounds, mitomycin C, lidocaine, prilocaine and tamoxifen, under the conditions of exposure produced definitive DNA adducts (Figure 2A). Another pharmaceutical, diethylstilbestrol produced weak spots on the chromatogram after two-directional TLC run, with adducts close to the detection limit (below 1 per 109 normal nucleotides). However, a three-directional TLC run revealed an adduct spot which was measurable (Figure 2A). Based on the observation of patterns of DNA adduct spots on chromatograms, it was concluded that mitomycin and tamoxifen produced 3 and 4 different adducts, respectively, while diethylstilbestrol, lidocaine and prilocaine generated one adduct (Figure 2A).
To compare DNA-binding potencies of pharmaceuticals with positive responses in the NPL assay, the total levels of DNA adducts per 108 nucleotides were normalized to dosages of the compound in µmol (calculated based on the molecular weight) (Figure 2B; Suppl. Table 1). This approach was used since the NPL assay was conducted using only one dose level for each tested material. Based on these calculations, mitomycin C was the most potent genotoxic pharmaceutical with DNA binding potency of approximately 150 adducts/µmol. It was followed by tamoxifen and diethylstilbestrol (∼0.5 and 0.25 adducts/µmol, respectively), and about 10-fold lower DNA binding potencies were exhibited by lidocaine and prilocaine (∼0.02 and 0.01 adducts/µmol, respectively). DNA binding potency of the positive comparator quinoline was calculated to be ∼0.1 adducts/µmol. (Figure 2B; Suppl. Table 1). The other fifteen pharmaceuticals, including 6-thioguanine, acetaminophen, ciprofloxacin, clofibrate, cyclophosphamide, diphenhydramine, doxylamine, etoposide, hydralazine, ibuprofen, griseofulvin, phenacetin, phenobarbital, streptozotocin, and teniposide, did not exhibit DNA binding at the dosages tested as evident from chromatograms in Figure 2C.
In the comet assay, eight pharmaceuticals produced positive results as evident by dose-dependent, statistically significant increases in the percentage of DNA-in-tail in the dosed groups (Figure 3; Suppl. Table 2). These included three compounds, lidocaine, prilocaine, and tamoxifen, which also produced DNA adducts, as described above, as well as cyclophosphamide, diphenhydramine, doxylamine, etoposide, and griseofulvin, which did not form adducts. DNA strand breaks produced by hydralazine, mitomycin C, streptozotocin, and teniposide, were not dose-dependent (Figure 4) and did not satisfy all of the criteria for a positive response in the comet assay (detailed in the Materials and Methods section). As such, findings in the comet assay for these four compounds were judged equivocal. No increases in DNA migration were observed in the groups that received 6-thioguanine, acetaminophen, ciprofloxacin clofibrate, ibuprofen, phenacetin, and phenobarbital, in concordance with negative NPL outcomes. In addition to these seven compounds, diethylstilbestrol was also negative in the comet assay (Figure 5).
Based on the weight of evidence for genotoxicity from results of testing of 20 pharmaceuticals in the NPL and comet assays, compounds were grouped into 4 groups (Table 1): • Group 1 – compounds that yielded clear positive outcomes in both, NPL and comet assays • Group 2 – compounds that yielded a clear positive outcome in either NPL or comet assay o Group 2A –compounds the yielded clear positive outcomes in the NPL assay, but either equivocal or negative results in the comet assay o Group 2 B - compounds that yielded clear positive outcomes in the comet assay, but negative in the NPL assay • Group 3 – compounds that produced equivocal outcomes in the comet assay • Group 4 – compounds that produced clear negative results in both, NPL and comet assays
Compounds belonging to groups 1 (lidocaine, prilocaine, and tamoxifen) and 2A (diethylstilbestrol and mitomycin C) were considered genotoxic in CEGA, since ability to produce nuclear DNA adducts indicates chemical-specific DNA reactivity. No definitive conclusion regarding direct DNA reactivity can be made for the compounds in group 2B (cyclophosphamide, diphenhydramine, doxylamine, etoposide, and griseofulvin), since they were positive only for nuclear DNA strand breaks which can result from either direct or indirect DNA damage. Compounds in group 3 (hydralazine, streptozotocin, and teniposide) were considered equivocal, since they did not produce DNA adducts and did not meet all of the requirements for either a clear positive or negative response in the comet assay. Compounds in group 4 (6-thioguanine, acetaminophen, ciprofloxacin, clofibrate, ibuprofen, phenacetin, and phenobarbital) were deemed clearly negative.
Discussion
Results of Testing of Pharmaceuticals in the CEGA and Comparison of Genotoxic and Carcinogenic Potentials of Tested Pharmaceuticals in ovo, in vitro and in vivo.
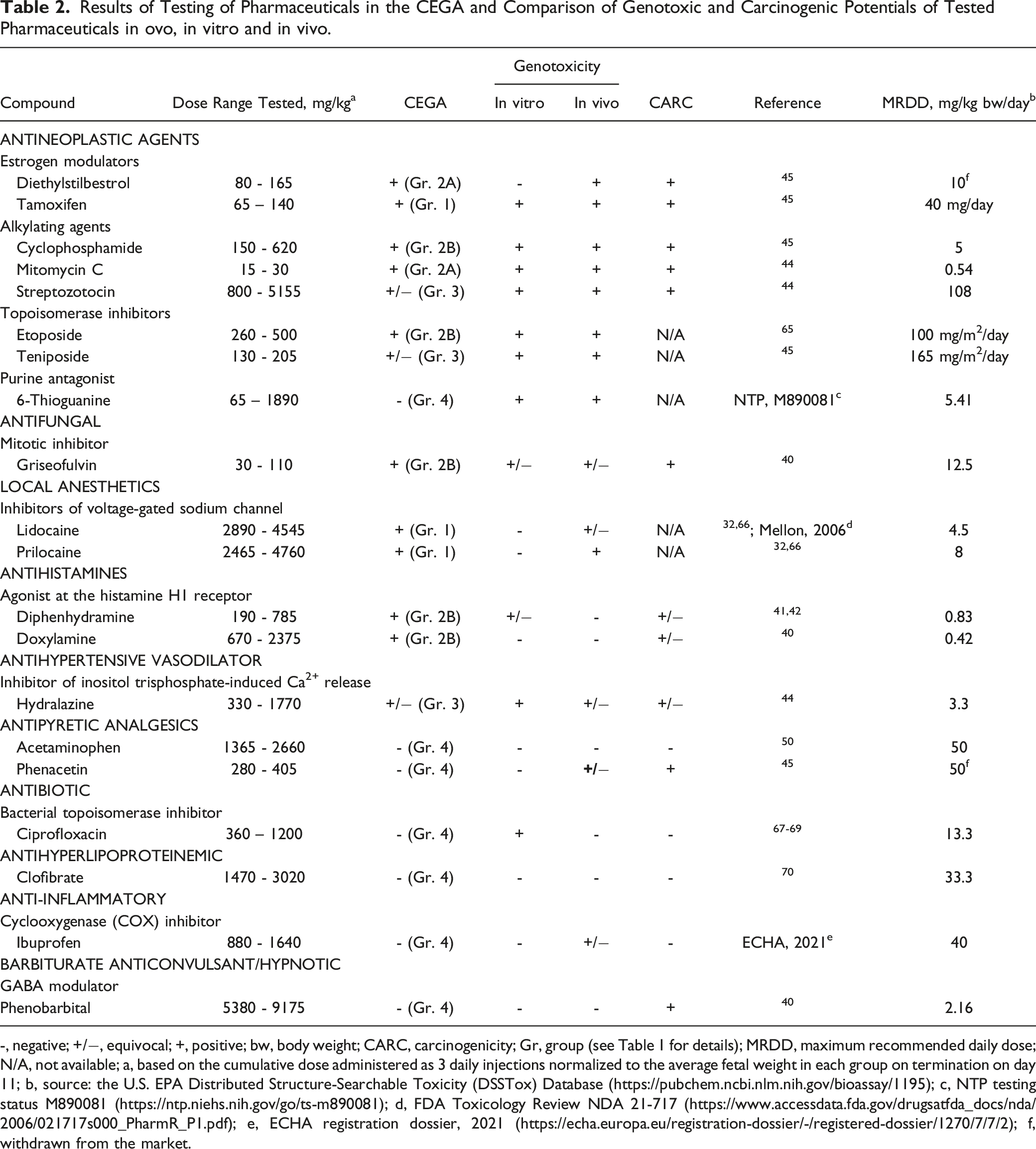
-, negative; +/−, equivocal; +, positive; bw, body weight; CARC, carcinogenicity; Gr, group (see Table 1 for details); MRDD, maximum recommended daily dose; N/A, not available; a, based on the cumulative dose administered as 3 daily injections normalized to the average fetal weight in each group on termination on day 11; b, source: the U.S. EPA Distributed Structure-Searchable Toxicity (DSSTox) Database (https://pubchem.ncbi.nlm.nih.gov/bioassay/1195); c, NTP testing status M890081 (https://ntp.niehs.nih.gov/go/ts-m890081); d, FDA Toxicology Review NDA 21-717 (https://www.accessdata.fda.gov/drugsatfda_docs/nda/2006/021717s000_PharmR_P1.pdf); e, ECHA registration dossier, 2021 (https://echa.europa.eu/registration-dossier/-/registered-dossier/1270/7/7/2); f, withdrawn from the market.
The 10 clearly positive compounds (Groups 1 and 2) included the estrogen modulator diethylstilbestrol and estrogen receptor ligand tamoxifen, alkylating agents cyclophosphamide and mitomycin C, topoisomerase inhibitor, etoposide, antifungal griseofulvin, local anesthetics lidocaine and prilocaine, antihistamines diphenhydramine and doxylamine. Three of these, lidocaine, prilocaine, and tamoxifen, were positive in both, comet and NPL assays (Group 1), indicating that these compounds directly react with DNA. The majority of these compounds have produced genotoxic effects in other test systems (Table 2). For example, patterns of DNA adducts produced by tamoxifen, diethylstilbestrol, lidocaine and prilocaine in rat livers32-35 were chromatographically similar to those detected in CEGA (Figure 6). The difference in relative adduct labelling between the rodent studies and CEGA, indicates that caution should be given to quantitative readouts when assessing DNA binding potency of genotoxic compounds. Nevertheless, the findings in CEGA demonstrate that the embryo-fetal chicken liver is capable of the two-step activation processes required for genotoxicity of these carcinogens.36,37 Schematic patterns of DNA adduct spots obtained using 32P-postlabeling assay from liver DNA of chicken embryo-fetuses or rodents after dosing with tamoxifen, diethylstilbestrol, mitomycin C, lidocaine and prilocaine (see Figure 2 for reference). Schematic patterns of DNA adducts in rodents were created based on chromatograms in the published literature (references provided below each chromatogram in parenthesis).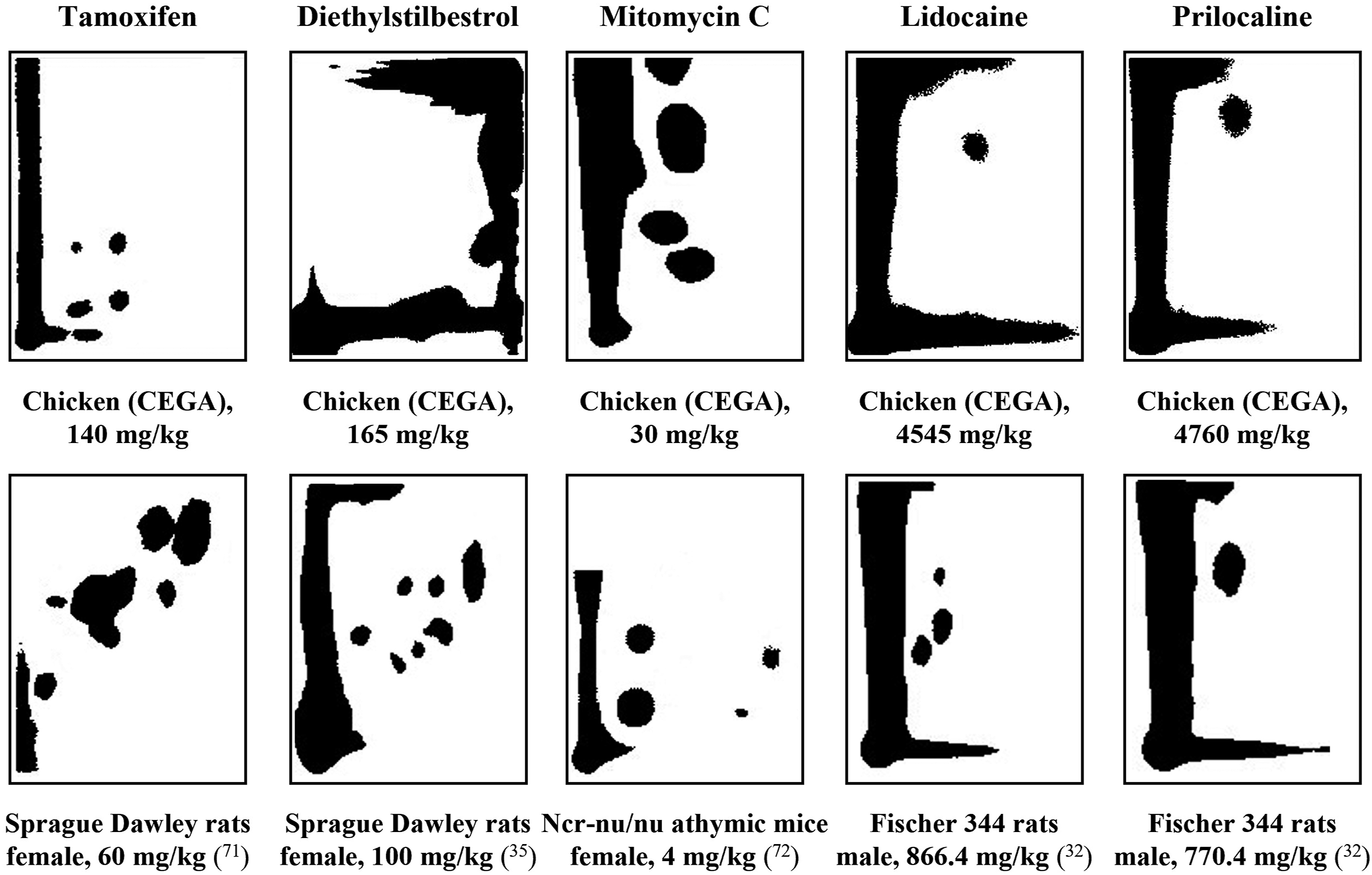
Chemotherapeutics that produced DNA damage in the model act as cross-linkers and alkylating agents and hence have established genotoxic potentials. The structures of all these compounds contain recognized DNA-reactive moieties. 38 The positive results indicate that measurement of DNA damage is a reliable mean of assessment of potential genotoxicity, as also demonstrated in the hepatocyte DNA damage assay. 15 These findings support the role of DNA damage in mutagenicity. 39
Positive findings in the comet assay in CEGA for two antihistamines, diphenhydramine and doxylamine (Figure 3), indicate that these compounds can exhibit genotoxicity. However, no definitive conclusion can be made regarding the mechanisms producing DNA strand breaks. DNA damage detected in the comet assay may stem from either direct DNA reactivity, or from indirect mechanisms, such as oxidative DNA damage. Diphenhydramine and doxylamine were mainly negative in a number of mutagenicity assays, and thus, were not considered to be genotoxic.40,41 Nevertheless, diphenhydramine induced formation of micronuclei and chromosomal aberrations in the Chinese hamster lung cell line, produced weak evidence of DNA strand breaks in primary rat hepatocytes and chromosomal aberrations in human cells in vitro, and doxylamine induced DNA damage in primary rat hepatocytes and chromosomal aberrations in mice treated transplacental.42,43 In addition, both pharmaceuticals produced equivocal evidence of carcinogenicity in rodent bioassays.40,41 Specifically, diphenhydramine hydrochloride produced marginal increases in alveolarhronchiolar neoplasms and in astrocytomas or gliomas in the brain of male F344/N rats, as well as pituitary gland adenomas in female rats, while no carcinogenic activity was reported in male or female B6C3F1 mice. 41 Doxylamine succinate administration increased incidences of hepatocellular and thyroid follicular-cell adenomas in male and female mice and marginally increased incidences of hepatocellular adenomas and carcinomas combined in male rats. 40 The mechanisms underlying these findings in carcinogenicity bioassays remain not well understood.
Three compounds, the antihypertensive vasodilator hydralazine, the alkylating agent streptozotocin, and topoisomerase inhibitor teniposide, produced some evidence of DNA damage, although responses were not dose dependent. Thus, their genotoxic potential was deemed equivocal in CEGA. All of these compounds were shown to produce DNA damage in vivo.44,45 In the case of hydralazine, slow acetylation was shown to enhance DNA reactivity. 46 The activity of chicken acetyltransferases has not been compared to that in mouse or rat, while activity of arylamine N-acetyltransferase isolated from chicken liver, has been shown to be over 4-times greater compared to that in rabbit. 47
Seven compounds that tested negative in CEGA included purine antagonist 6-thioguanine, antipyretic analgesics acetaminophen and phenacetin, antibiotic ciprofloxacin, antihyperliporoteinemic clofibrate, anti-inflammatory cyclooxygenase inhibitor ibuprofen, as well as barbiturate phenobarbital. The negative results with clofibrate and phenobarbital confirm those previously reported in CEGA. 19 Four of the negative compounds also have been reported as negative in other systems (Table 2). For example, acetaminophen and phenobarbital did not form DNA adducts in mouse 48 or rat livers. 49 Acetaminophen (p-hydroxyacetanilide) at cytotoxic doses increased the incidences of liver tumors in mice. 50 Interestingly, when administered together with a genotoxic hepatocarcinogen, acetaminophen inhibited the induction of rat liver tumors.51,52 Phenobarbital is a weak rodent hepatocarcinogen which produces tumors through non-genotoxic tumor-promoting mechanisms.40,53,54
No DNA damage was observed with the purine analog 6-thioguanine in the CEGA, in contrast to reports of DNA incorporation of 6-thioguanine in mammalian cells, leading to formation of DNA strand breaks and mutagenicity.55,56 Once incorporated into DNA 6-thioguanine undergoes methylation and is processed by DNA mismatch repair, which can result in DNA damage and cytotoxicity. 57 Mechanisms other than DNA incorporation also can be involved in toxicity produced by 6-thioguanine, since it was also shown to be cytotoxic to non-proliferating primary hepatocytes. 58 Nevertheless, the activity of DNA mismatch repair system correlated with the formation of single strand DNA breaks. 59 It can be speculated that lack of genotoxicity of 6-thioguanine in ovo could indicate a difference in activity of DNA mismatch repair system in the developing avian embryo compared to mammals, however, this hypothesis requires further investigation.
Two alkaloid pharmaceuticals with aneugenic activity, colchicine and vinblastine, were extremely lethal in CEGA. This indicates that the actively proliferating cells of the developing avian organism are highly sensitive to disruption of the mitotic spindle. An autophagy inhibitor used for treatment of malaria, hydroxychloroquine, also exhibited high toxicity in the system. While the mechanisms underlying this effect in the chicken embryo-fetuses are unknown, in mammalian cells choloroquine had an inhibitory effect on the repair of DNA damage. 60 In addition, hydroxychloroquine was reported to suppress metabolic activity and cell proliferation while inducing cell death via autophagy, particularly in human dermal fibroblasts, through inhibition of extracellular signal-regulated kinase ERK1/2 activation. 61 It is interesting that the chicken embryo-fetuses were more sensitive to chemotherapeutic drugs which induce cytotoxicity through mechanisms involving microtubule dynamics and spindle assembly, than those acting through other mechanisms, eg, alkylation.
To conclude, the CEGA demonstrated high reliability in the assessment of the genotoxicities of pharmaceuticals, as it has with a variety of other types of chemicals, of course, further evaluations with a broader set of compounds would be needed to support this conclusion.19,21-24 Moreover, CEGA has other valuable features as an assay for genotoxicity of pharmaceuticals. It allows facile delivery of the test substance, both lipo- and hydrophilic compounds, into the air sac of an egg, providing ready bioavailability through absorption into the embryo-fetus by the chorioallantoic membrane. The reliable dosing allows testing of complex mixtures (including oils) in this model. With the target being the intact liver of a self-supporting, but non-animal, organism, the test material is subjected to metabolism by a wide range of enzymes including some that are not functional in subcellular preparations, such as phase II conjugation enzymes.21,62 It is noteworthy that tamoxifen which is bioactivated by α-hydroxylation and subsequent sulfate conjugation63,64 was positive in CEGA indicating function of this two-step activation. To elucidate the mode of action of chemicals, effects on gene expression in the model can be assessed. 21
Supplemental Material
Supplemental Material - Evaluation of Pharmaceuticals for DNA Damage in the Chicken Egg Genotoxicity Assay (CEGA)
Supplemental Material for Evaluation of Pharmaceuticals for DNA Damage in the Chicken Egg Genotoxicity Assay (CEGA) by Tetyana Kobets, Jian-Dong Duan, Esther Vock, Ulrich Deschl, and Gary M. Williams in International Journal of Toxicology
Footnotes
Acknowledgments
The authors would like to acknowledge our colleague for many years, Klaus D. Brunnemann, B.S., now deceased, for his help with experimental procedures.
Author Contributions
Kobets, Tetyana contributed to conception and design, contributed to acquisition, analysis, and interpretation, drafted manuscript, and critically revised manuscript; Duan, Jian-Dong contributed to acquisition and critically revised manuscript; Vock, Esther contributed to conception and critically revised manuscript; Deschl, Ulrich contributed to conception and critically revised manuscript; Williams, Gary M. contributed to conception and design, contributed to analysis and interpretation, and critically revised manuscript. All authors gave final approval and agree to be accountable for all aspects of work ensuring integrity and accuracy.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by funding form Boehringer Ingelheim Pharma GmbH&Co. KG, Biberach an der Riss, Germany.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
