Abstract
Cadmium is an environmental pollutant well known for its nephrotoxic effects. Nevertheless, mechanisms underlying nephrotoxicity continue to be elucidated. MicroRNAs (miRNAs) have emerged in recent years as modulators of xenobiotic-induced toxicity. In this context, our study aimed at elucidating whether miRNAs are involved in renal proximal tubular toxicity induced by cadmium exposure. We showed that cadmium exposure, in 2 distinct renal proximal tubular cell models (renal proximal tubular epithelial cell [RPTEC]/human telomerase reverse transcriptase [hTERT] and human kidney-2), resulted in cytotoxicity associated with morphological changes, overexpression of renal injury markers, and induction of apoptosis and inflammation processes. Cadmium exposure also resulted in miRNA modulation, including the significant upregulation of 38 miRNAs in RPTEC/hTERT cells. Most of these miRNAs are known to target genes whose coding proteins are involved in oxidative stress, inflammation, and apoptosis, leading to tissue remodeling. In conclusion, this study provides a list of dysregulated miRNAs which may play a role in the pathophysiology of cadmium-induced kidney damages and highlights promising cadmium molecular biomarkers that warrants to be further evaluated.
Keywords
Introduction
Environmental chemical exposure remains a major public health matter worldwide. In particular, cadmium is a serious environmental pollutant emanating from industrial and agricultural sources. 1 Cadmium is a natural earth’s crust element, usually found as a mineral, and widely found as a byproduct of mining; smelting; and refining zinc, lead, and copper ores. 2 For the last 2 decades, despite a decreased production and use of cadmium, it is still found in nickel-cadmium batteries, cadmium telluride photovoltaics, plastic stabilizers, or paint pigments. 3 The general population exposure to cadmium predominantly results thus from air pollution, consumption of cadmium-contaminated foods and water, and also from smoking. 4
Cadmium, which is poorly metabolized, is filtered and reabsorbed in the renal proximal tubules, resulting in its bioaccumulation in target organs with a half-life of about 10 to 30 years. 5,6 Intrarenal cadmium concentration increases over time until it reaches a cellular threshold associated with progressive renal cell injury. 7 Cadmium is thus known to induce kidney injury in the general population as well as in particular occupational populations. 8 -10 Tubular proteinuria, the first clinical sign of cadmium-induced kidney injury, occurs even after a low exposure to cadmium. For example, Jarup 11 in 2000 reported, in a population of workers and people environmentally or occupationally exposed to low cadmium levels, a 10% increase prevalence of tubular proteinuria. At a later stage, tubular dysfunction has been shown to progress with nonspecific glomerular damages, and finally to a reduced glomerular filtration rate. 12 Furthermore, for women with type 2 diabetes, cadmium exposure was reported to be associated with a larger amount of albuminuria, suggesting that glomerular integrity of this subgroup of patients is particularly vulnerable to cadmium exposure. 13 Epidemiological studies have also reported an association between cadmium exposure and chronic kidney diseases 14 -16 or end-stage kidney diseases. 17 Cadmium has also been implicated in renal carcinogenesis 18 and has been classified as a category I carcinogen by the International Agency for Research on Cancer. 19
The mechanisms underlying renal cadmium toxicity are quite well understood. Briefly, at the cellular level, cadmium increases the production of reactive oxidative species leading to oxidative stress, DNA damage, lipid peroxidation, and apoptosis. 7,20 Cadmium has also been shown to activate several proto-oncogenes or genes associated with cell proliferation, such as c-myc or c-jun, both in vitro and in animal models. 21,22 However, the elucidation of novel regulatory mechanisms underlying cadmium injury remains a priority issue to allow better assessment of the risk associated with this common environmental contaminant and to develop novel pertinent biomarkers of cadmium exposure. In particular, the recently discovered importance of microRNAs (miRNAs) during kidney development 23 or in renal pathogenesis 24 -28 emphasizes the key role played by miRNAs in renal injury. 29 MicroRNAs are small endogenous, single-stranded, RNA molecules without protein-coding capacity, which act as powerful negative regulators of gene expression at the post-transcriptional level. 30 Generally, the inhibition of target gene expression by miRNAs requires the binding of miRNA to the 3′-untranslated region of its target genes, resulting in messenger RNA (mRNA) cleavage or suppressed translation. 31 Since their first discovery in Caenorhabditis elegans, 32 miRNAs have gained significant attention these last years with the understanding of their important modulatory role in cellular processes, such as proliferation, differentiation, or apoptosis. 33,34 In addition, the remarkable stability of miRNAs in biofluids has underscored their powerful value as a new class of noninvasive biomarkers for diagnosis, prognosis, or disease activity. 35,36
Thus, the present work aimed at identifying the miRNAs differentially expressed in renal proximal tubular cells in response to cadmium exposure. To establish a list of confident miRNAs that were modulated upon cadmium exposure, 2 cell models were used: (1) the renal proximal tubular epithelial cell (RPTEC)/human telomerase reverse transcriptase (hTERT) cells as a “discovery” cell model and (2) the human kidney-2 (HK-2) cells as a “validation” cell model.
Material and Methods
Cell Culture
Primary human RPTECs immortalized with pLXSN-hTERT retroviral transfection (RPTEC/hTERT, CRL-4031, ATCC [American Type Culture Collection], Manassas, VA) were cultured supplemented with 1% penicillin/streptomycin, 5 pmol/L triiodo-
Cadmium Exposure
Twenty-four hours after seeding in 96-well plates (20,000 cells/well), RPTEC/hTERT and HK-2 cells were exposed to 0 to 1,000 μmol/L cadmium chloride (CdCl2; Sigma) for 24 hours. Cell viability was evaluated using the CellTiterGlo kit (Promega, Madison, WI), based on ATP quantitation, to determine subsequent exposure treatment. Subconfluent cells (200,000 cells seeded in 6-well plates) were then subjected either to vehicle (untreated cells) or CdCl2 (20 µmol/L for HK-2 and 10 µmol/L for RPTEC/hTERT, corresponding to the estimated concentration required to inhibit cell growth by 20% [IC20]) for 24 hours.
RNA Extraction
Total RNAs were extracted from 2 wells of a 6-well plate, using the miRNeasy Mini kit (Qiagen, Redwood City, CA), according to the supplier’s recommendations. RNAs integrity was evaluated using an Experion (BioRad, Hercules, CA) automat.
Open Array Analysis
Three independent exposures (10 μmol/L CdCl2) were used for miRNA profiling in RPTEC/hTERT cells with the QuantStudio 12 K Flex Open Array Real-Time PCR system (Thermofisher). The OpenArray reverse transcription reaction was performed, according to the manufacturer’s protocol, using Megaplex RT Primers Pool A v2.1 and Pool B v3.0 following preamplification reaction. Then, TaqMan Open Array Real Time PCR was performed on a QuantStudio 12K Flex Real-Time PCR System (Thermofisher) according to the supplier’s recommendations.
Relative quantification was evaluated using the Thermofisher Cloud Data Analysis Apps (Thermofisher) and the comparative threshold cycle method (Crt); 754 miRNAs and 4 internal controls were analyzed. Low expressed miRNAs (Crt > 26) and miRNAs with poor amplification quality (<1.24) were excluded from the study. The expression of each miRNA was calculated after normalization to the levels of small nuclear RNA U6 (U6 snRNA) and finally expressed as fold change as compared to the control samples.
Pathway Analysis
Enriched Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis was performed using DNA Intelligent Analysis (DIANA)-miRPath v3.0 software (http://diana.cslab.ece.ntua.gr), 38 based on predicted targets by DIANA-microT-CDS. Targets of miRNAs with a score of more than 0.8 were selected. Only KEGG pathways with a P value <0.05 and a false discovery rate (FDR) <0.05 were retained.
Quantitative Real-Time Polymerase Chain Reaction
MicroRNAs retrotranscription was performed using Taqman microRNA Reverse Transcription Kit (Thermofisher). Quantitative polymerase chain reaction (PCR) was performed using a StepOnePlus Real-Time PCR System (Thermofisher) with Taqman Universal Master Mix II, no UNG (Thermofisher) and with the following Taqman assays: miR-21-5p (assay ID 000397), miR-29c-3p (assay ID 000587), miR-193a-5p (assay ID 002281), miR-27a-3p (assay ID 000408), miR-30b-5p (assay ID 000602), and miR-30c-5p (assay ID 00019); U6 snRNA (assay ID 001973) was used as a normalizer.
Messenger RNAs retrotranscription was performed using the high-capacity complementary DNA reverse transcription kit (Thermofisher) according to manufacturer’s instructions. Quantitative PCR was performed on a StepOnePlus Real-Time PCR System (Thermofisher) with Taqman Universal Master Mix II, no UNG (Thermofisher) and with the following Taqman assays: neutrophil gelatinase-associated lipocalin (NGAL; assay ID hs01008571), heme oxygenase 1 (HO-1; assay ID hs01110250), and interleukin 6 (IL-6; assay ID hs00174131). PPIA (cyclophilin A; assay ID hs99999904) was used as a normalizer. Relative expression levels of mRNAs and miRNAs were assessed using the comparative threshold cycle method (2−ΔΔCT). 39
Western Blot
Total proteins, from 3 wells of a 6-well plate, were lysed in radioimmunoprecipitation assay buffer (Sigma) containing protease and phosphatase inhibitors cocktail (Roche, Meylan, France) and were sonicated for 20 seconds. Lysates were quantified for protein concentration using the BCA protein assay kit (Pierce, Waltham, MA). Twenty μg of proteins were separated by electrophoresis on a 4% to 12% BisTris NuPAGE gel (Thermofisher) or 7% Tris acetate NuPAGE gel (Thermofisher) and were transferred onto nitrocellulose membrane using the Trans-Blot Turbo Transfer System (Biorad). After blocking, the membranes were incubated with their respective primary antibodies (anti-caspase 3 [sc7148; Santa Cruz, Dallas, TX; 1/1,000], anti-cleaved caspase 3 [9661; Cell Signaling (Danvers, MA); 1/1,000], anti-Poly(ADP-Ribose) Polymerase 1 [PARP-1, 9542P; Cell Signaling; 1/1,000], anti-HO-1 [5853 S, Cell Signaling; 1/1,000], and anti-HSP60 [sc-1052; Santa Cruz; 1/4,000]) overnight at 4°C. Membranes were further incubated with horseradish peroxidase-conjugated secondary antibody (Sigma, Saint Louis, MO; 1/10,000) for 45 minutes at room temperature. Protein bands were visualized with the Amersham ECL Western blotting detection reagents (GE Healthcare, Chicago, IL) using a Fusion FX Spectra imager (Vilber Lourmat, Collégien, France). Image analysis was performed using ImageJ.
Statistical Analysis
Statistical analyses were performed with GraphPad Prism version 6.00 for Windows (GraphPad Software). Results are given as mean ± standard error of the mean. Two-tailed student t test was used for single comparisons. One-way analysis of variance was used for multicomparison following by Bonferroni post hoc test. P value < 0.05 was considered statistically significant.
Results
Cadmium Affects Renal Cell Survival and Induces Cell Toxicity
We first assessed cadmium-induced toxicity in vitro using immortalized proximal tubular epithelial cells (RPTEC/hTERT). As expected, cadmium exposure induced RPTEC/hTERT toxicity in a dose-response manner (half maximal inhibitory concentration [IC50]: = 26.01 ± 6.38 µmol/L; Figure 1A). Dose-dependent cadmium-induced cell injury was also confirmed by microscopy. Indeed, exposure to CdCl2 for 24 hours caused RPTEC/hTERT cells to separate from each other and switch from epithelioid to rounded shape (Figure 1B). Subsequent CdCl2 exposure was then conducted using a mild 10 µmol/L CdCl2 concentration.
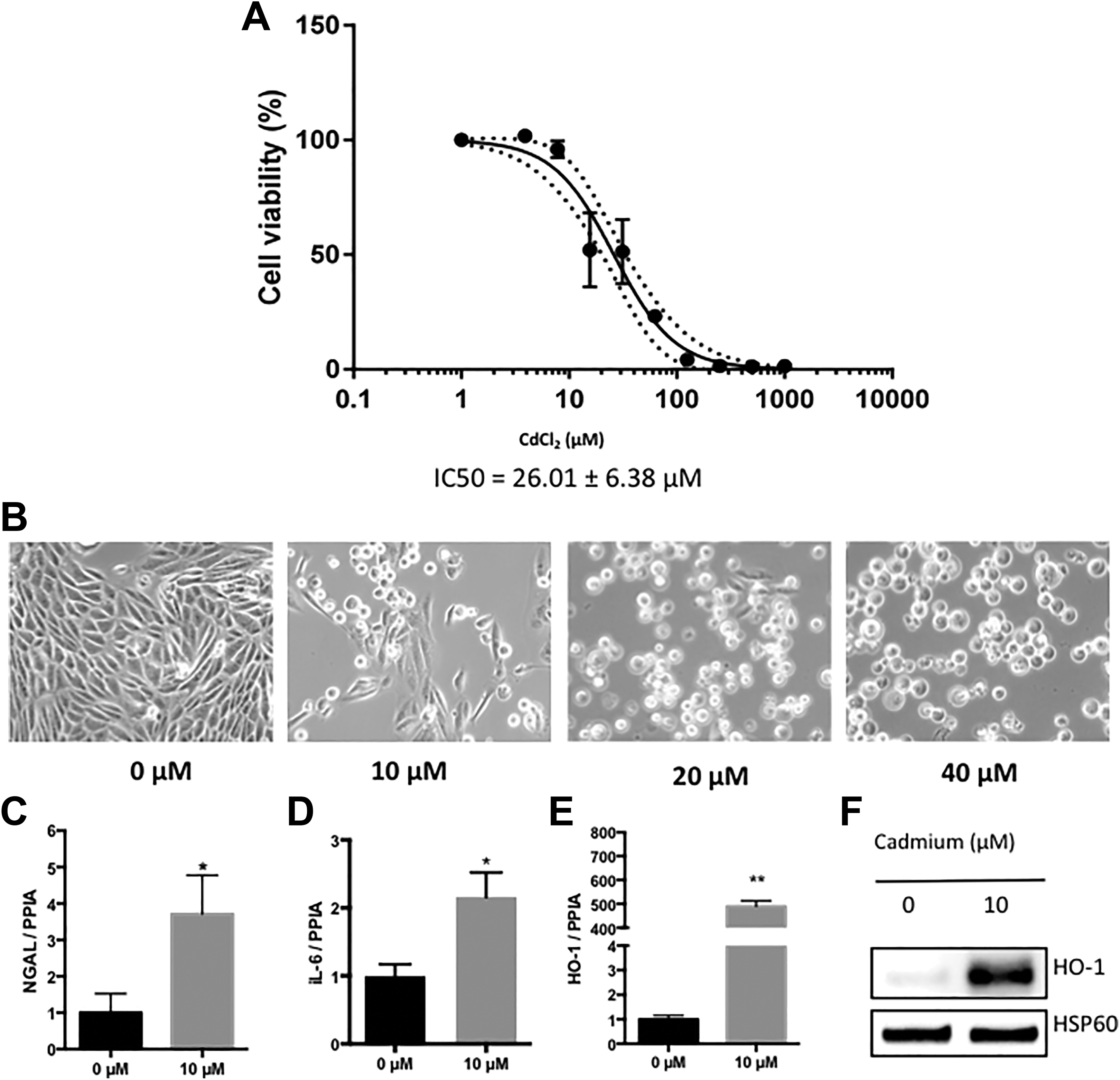
Cell death and cytotoxicity induced by cadmium in RPTEC/hTERT cells. A, Cell viability was measured after treatment with CdCl2 (0 to 1,000 µmol/L) for 24 hours using the CellTiterGlo kit (Promega). Values are the means ± SEM of 4 independent experiments. P value <0.0001. B, Representative bright field microscopy under 40× magnification. Cells were treated for 24 hours with 0 to 40 µmol/L CdCl2. C-E, Relative expression of neutrophil gelatinase-associated lipocalin (NGAL), IL-6, and heme oxygenase 1 (HO-1). Values are the means ± SEM of 5 independent experiments. *P value<0.05; **P value<0.01. PPIA was used as an internal control. F, Representative Western blot of the 3 independent experiments. Cells exposed to 10 μmol/L CdCl2 versus untreated cells. Blots were incubated with antibodies against HO-1. The HSP60 protein was used as an internal control. CdCl2 indicates cadmium chloride; hTERT, human telomerase reverse transcriptase; RPTEC, renal proximal tubular epithelial cell; SEM, standard error of the mean.
In addition, we confirmed that the relative expression of NGAL, a renal injury marker, 40 was increased following cadmium exposure (Figure 1C). Furthermore, cadmium increased the expression of inflammatory genes such as IL-6 (Figure 1D). Finally, we observed a significant induction of heme oxygenase 1 (HO-1), a well-recognized biomarker for oxidative stress, 41 at mRNA (Figure 1E) and protein levels (Figure 1F).
Cadmium Exposure Triggers a Modulation of miRNA Expression in Renal Cells
As previously described, proximal tubular epithelial cells, such as RPTEC/hTERT, display high metabolism and xenobiotic disposition capacity 42 and are among the most relevant in vitro models to evaluate xenobiotic deleterious effects. 28,43,44 As described above, RPTEC/hTERT cell model closely recapitulates cadmium toxicity in vitro. We then aimed to identify key miRNAs potentially involved in cadmium deleterious effects.
A miRNA was considered as nonexpressed when its adjusted Crt value was higher than 26, according to the supplier’s recommendations. Using this criterion, of the 754 miRNAs that we studied, 150 miRNAs were regarded as expressed in RPTEC/hTERT cells. Among them, the most expressed miRNAs were miR-222-3p, miR-21-5p, miR-146a-5p, miR-30c-5p, and miR-30b-5p. In addition, cadmium induced the significant upregulation of 38 miRNAs (Figure 2).
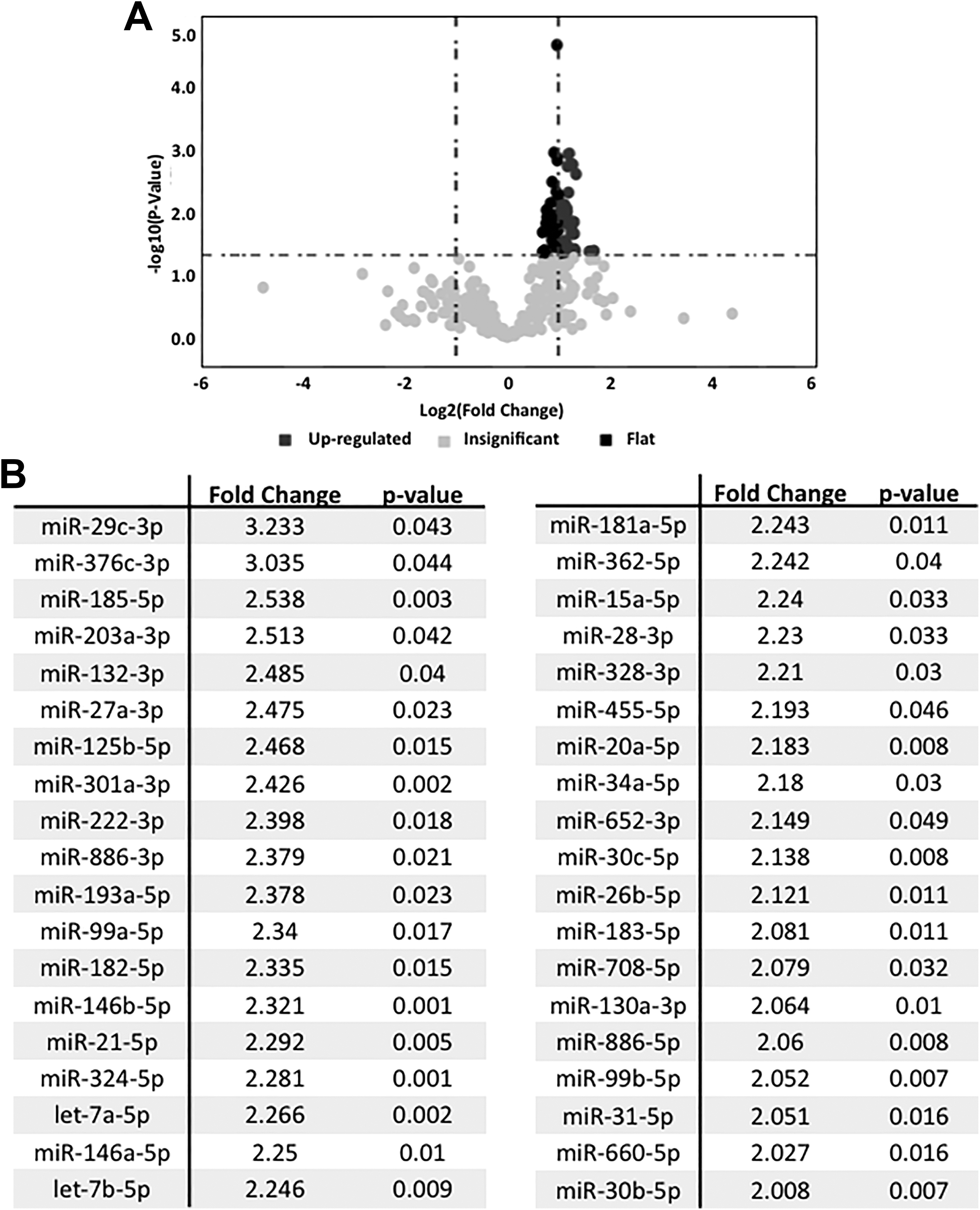
Significantly deregulated miRNAs in RPTEC/hTERT cell model of cadmium-induced cytotoxicity. A, Volcano plot showing the 38 significantly differentially expressed miRNAs in RPTEC/hTERT cells exposed to 10 µmol/L CdCl2 versus untreated cells (n = 3 independent experiments). B, List of the miRNAs significantly modulated by cadmium treatment. CdCl2 indicates cadmium chloride; miRNA, microRNA; hTERT, human telomerase reverse transcriptase; RPTEC, renal proximal tubular epithelial cell.
Of note, none of the studied miRNAs was found to be significantly downregulated in cells exposed to cadmium. These data were validated for 5 of the 6 miRNAs that were selected according to their basal expression, upregulation level, and P value (miR-21-5p, miR-27a-3p, miR-29c-3p, miR-30b-5p, and miR-30c-5p) in independent RPTEC/hTERT cell samples (Figure 3).
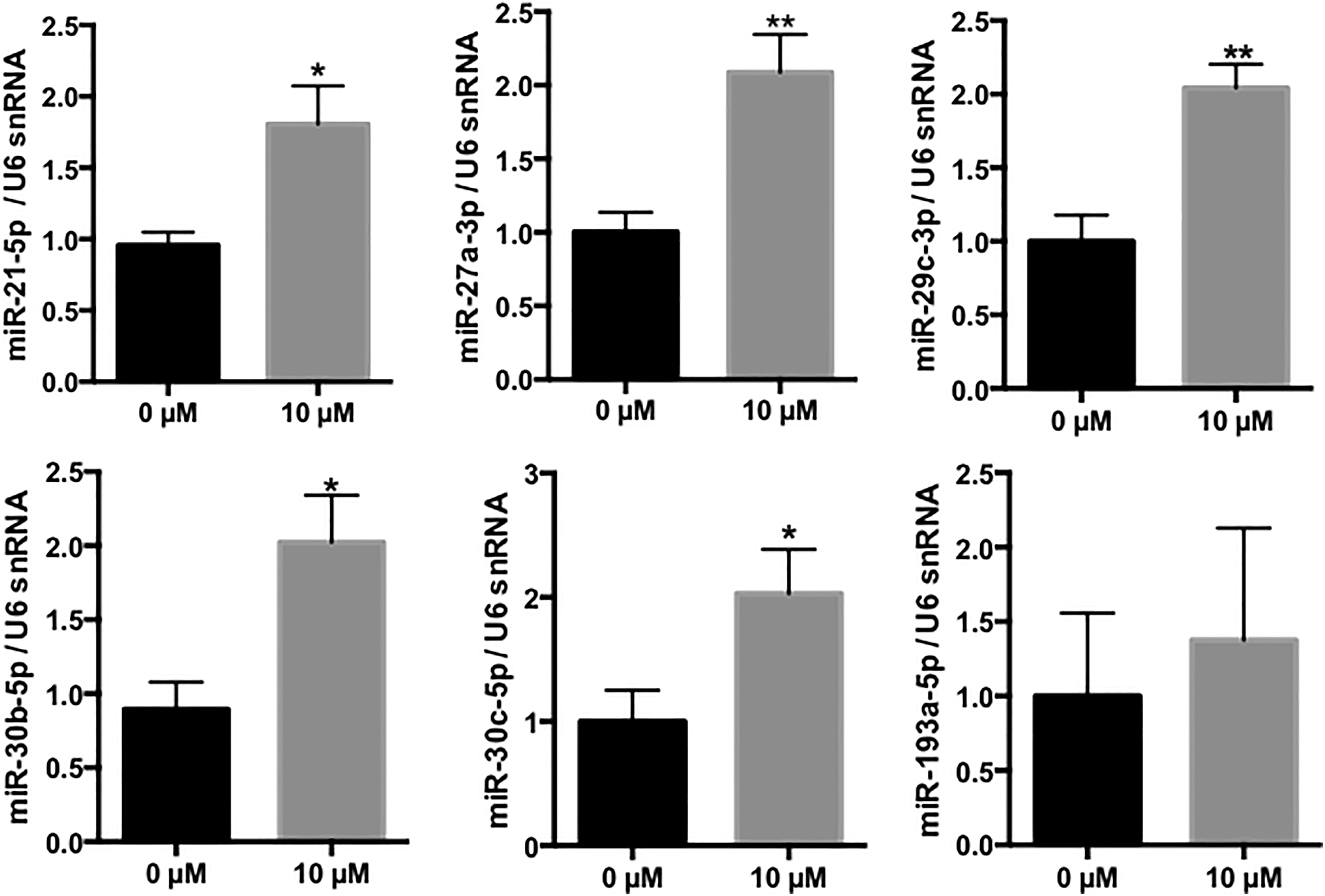
Validation of the cadmium-induced upregulation of miR-21-5p, miR-27a-3p, miR-29c-3p, miR-30b-5p, miR-30c-5p, and miR-193a-5p in RPTEC/hTERT cells. Relative expression of miRNAs. Values are the means ± SEM of 5 independent experiments. *P value<0.05; **P value<0.01. U6 snRNA was used as an internal control. miRNA indicates microRNA; hTERT, human telomerase reverse transcriptase; RPTEC, renal proximal tubular epithelial cell; SEM, standard error of the mean.
Validation of miRNA Modulation Induced by Cadmium in HK-2 Cell Line
We also aimed to further assess the relevance of the cadmium-induced miRNA dysregulation that we observed in RPTEC/hTERT cells in the commonly used HK-2 cell line. 45 -53 The HK-2 cells also exhibited typical features of cadmium toxicity, including a decreased cell viability and a morphological modification. However, HK-2 cells were less sensitive to cadmium than RPTEC/hTERT, as suggested by a higher IC50 (IC50 = 36.42 ± 7.10 µmol/L versus IC50 = 26.01 ± 6.38 µmol/L; Supplemental Figure 1).
Expression of the previously selected miRNAs (miR-21-5p, miR-27a-3p, miR-29c-3p, miR-30b-5p, miR-30c-5p, and miR-193a-5p) was further evaluated following cadmium exposure (IC20 = 20 µmol/L). We observed a similar miRNA response to cadmium with the significantly upregulation of 5 of the 6 selected miRNAs in this cell line (Figure 4).
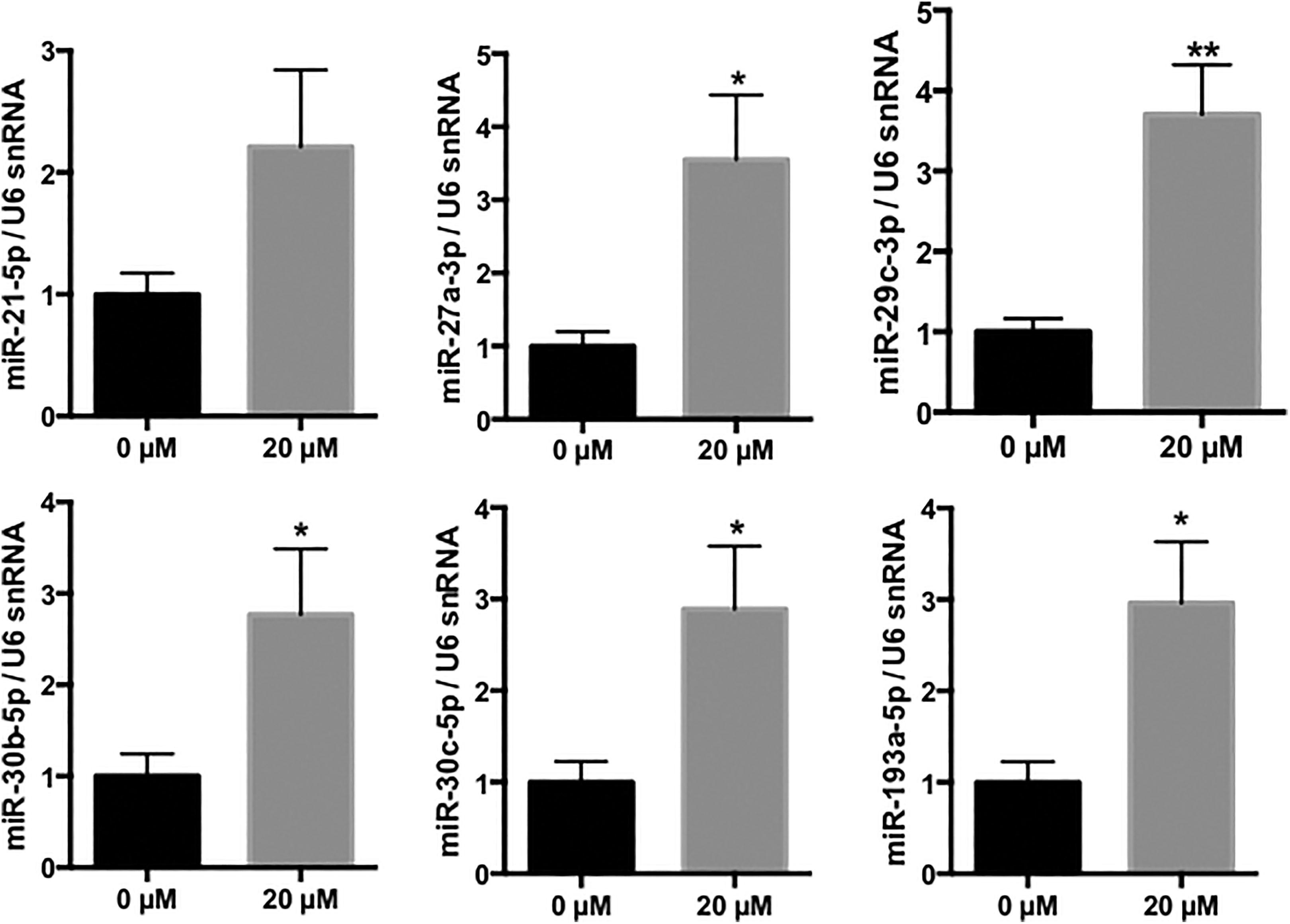
Validation of the cadmium-induced upregulation of miR-21-5p, miR-27a-3p, miR-29c-3p, miR-30b-5p, miR-30c-5p, and miR-193a-5p in HK-2 cells. Relative expression of miRNAs following CdCl2 exposure. Values are the means ± SEM of 4 independent experiments. *P value<0.05; **P value<0.01. U6 snRNA was used as an internal control. CdCl2 indicates cadmium chloride; HK-2, human kidney-2; miRNA, microRNA; SEM, standard error of the mean;
Prediction of miRNA Targeted Genes and Pathways
A thorough review of the literature was in favor of the involvement of these upregulated miRNAs in key cellular processes such as apoptosis and cancer (Supplemental Table 1). Moreover, a bioinformatic analysis revealed that about 80% of the upregulated miRNAs have been previously experimentally shown to target genes involved in the apoptotic process (Table 1).
List of the Experimentally Validated Genes Related to Apoptosis Targeted by Cadmium-Responsive MiRNAs in RPTEC/hTERT Cells.

Abbreviations: hTERT, human telomerase reverse transcriptase; MiRNA, microRNA; RPTEC, renal proximal tubular epithelial cell.
The relevance of the cadmium-induced miRNA pattern was further assessed. As expected, apoptosis was observed in cadmium-treated cells, as shown by the cleavage of caspase 3 and PARP-1, 2 key proteins of the apoptotic process (Figure 5).
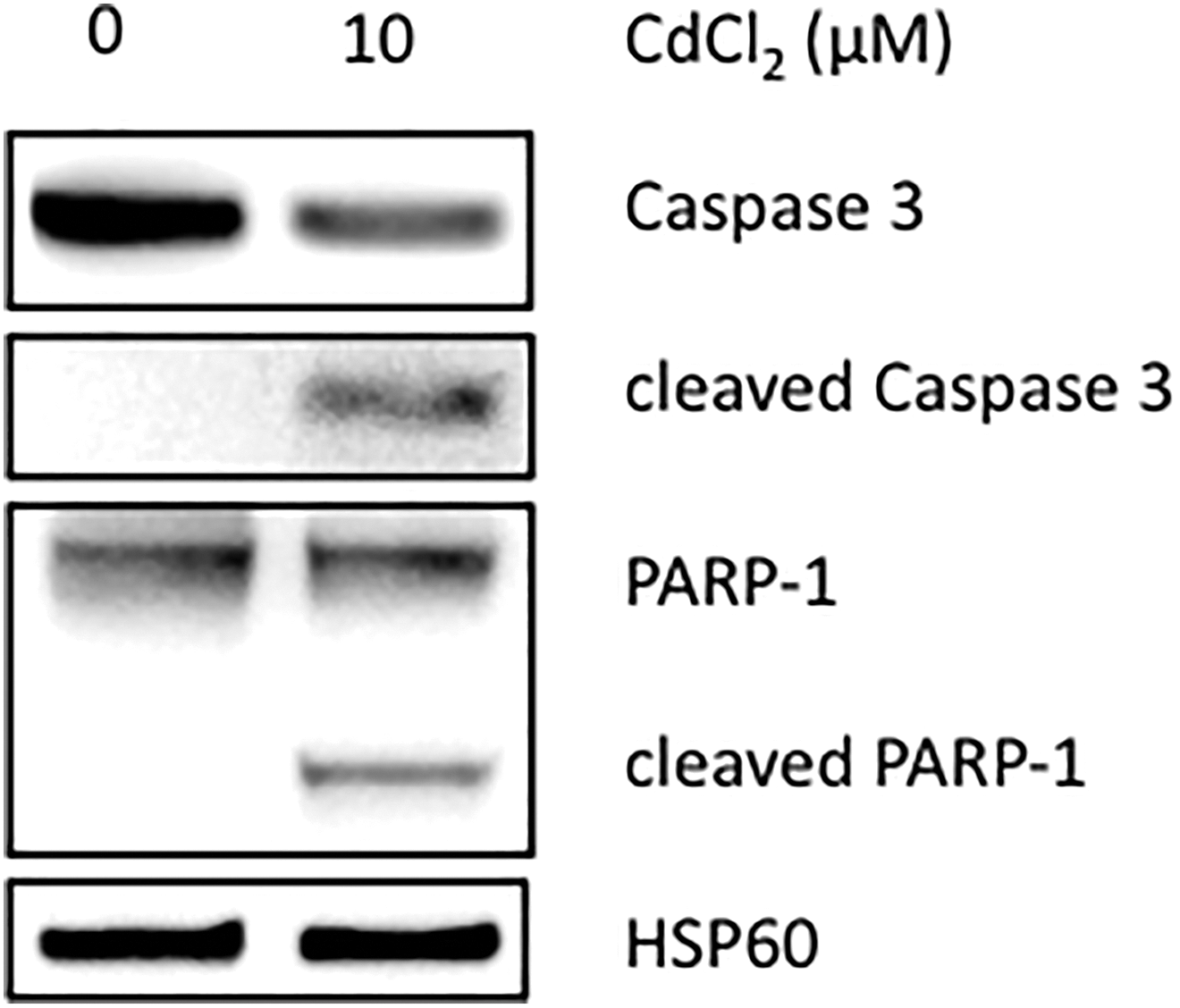
Apoptosis induced by cadmium in RPTEC/hTERT cell model. Representative Western blot of the 3 independent experiments for cells exposed to 10 μmol/L CdCl2 versus untreated cells. Blots were incubated with antibodies against cleaved caspase 3 and PARP-1. The HSP60 protein was used as an internal control. CdCl2 indicates cadmium chloride; hTERT, human telomerase reverse transcriptase; RPTEC, renal proximal tubular epithelial cell.
Moreover, the miR-Path analysis tool highlighted that the direct targets of the upregulated genes were enriched in KEGG network processes involved in tissue remodeling, including fibrogenesis and cancer. These pathways included phosphoinositide 3-kinase-protein kinase B (PI3K-AKT), mitogen-activated protein kinase (MAPK), Ras, Rap1, and Hippo signaling pathways (Figure 6).
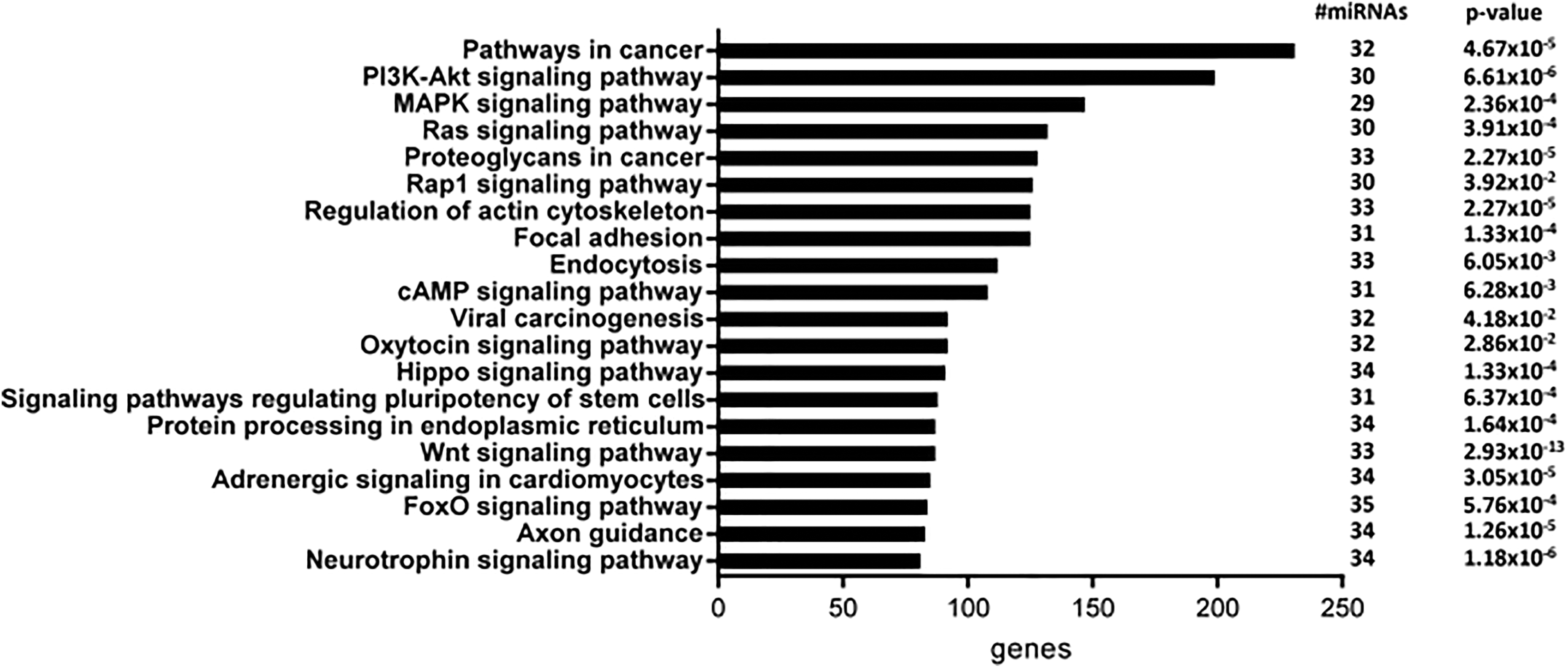
Pathway prediction for the differentially upregulated miRNAs in RPTEC/hTERT cells treated with cadmium versus untreated cells according to mirPath v.3 (Diana Tools). The number of miRNAs associated with each pathway and number of genes involved in each pathway are indicated. The top 20 pathways are listed (P ≤ 0.005). miRNA indicates microRNA; hTERT, human telomerase reverse transcriptase; RPTEC, renal proximal tubular epithelial cell.
Discussion
MicroRNAs have emerged in the past 2 decades as powerful gene repressors and are involved in various cellular and biological key processes. 54 As metals are widely present in the environment, they are prone to be absorbed, to accumulate and to interfere with cell fate. In particular, cadmium is a widespread environmental and industrial compound that induces various deleterious effects including nephrotoxicity. 55 To date, there are few reports describing miRNA expression changes due to other metal exposure. In particular, arsenic, one of the most common environmental carcinogenic pollutants, 56 has been shown to alter miRNA expression, leading to malignant transformation and tumorigenesis in human bronchial epithelial cells. 57
Cadmium exposure in human is then postulated to modify miRNA expression, leading to deregulation of gene expression and toxic effects. However, studies related to the modulation of miRNAs in response to cadmium exposure are lacking, especially in renal cells. Indeed, as environmental cadmium may accumulate in soils and water, previous studies have mainly focused on plants or aquatic organisms. 58 For example, the plant adaptive response to heavy metal stress has been demonstrated to be notably modulated by miRNAs in rice, soybean, or wheat. 59 -61
To our knowledge, our study allowed for the first time a comprehensive analysis of miRNA modulation following cadmium exposure in human RPTEC. Our data are in favor of an association between cadmium cytotoxicity and altered miRNA expression in human renal tubular proximal cells. In order to clarify mechanisms underlying miRNA relevance, we have first chosen to explore miRNA expression profile in RPTEC/hTERT, a cellular model that has been proven to be reliable in several nephrotoxicity studies. 28,53,44,62 In particular, this cell line may be viewed as an improved tool for in vitro nephrotoxicity assessments. 44 Indeed, by using only the catalytic subunit of the endogenous hTERT enzyme for immortalization, telomere length is stabilized, and then, the RPTEC/hTERT cell line does not undergo replicative senescence, 37 a well-known limitation of primary RPTECs. Moreover, the RPTEC/hTERT cell line exhibits a similar metallothioneins activity and stress-induced gene overexpression compared to healthy primary human kidney cells. 44,63 We used a CdCl2 concentration of 10 µmol/L, corresponding to the IC20 in the RPTEC/hTERT cell line, which is in line with previous studies. 64 -66
More precisely, we observed that cadmium treatment for 24 hour induces the upregulation of 38 miRNAs in RPTEC/hTERT cells. The fact that the modulation of miRNA expression is quite pronounced in our experimental conditions, when the cytotoxicity is mild (10 µmol/L CdCl2 that corresponds to about IC20), suggests that miRNA modulation could represent an early event in the pathophysiology of cadmium-induced renal injury. Similar early effect was recently reported in a rat subchronic model of cadmium exposure, in which animals were treated with subcutaneous injections of CdCl2 (0.6 mg/kg) 5 days a week for 12 weeks. MicroRNA high-throughput analysis allowed the identification of 44 miRNAs significantly upregulated after cadmium administration. 67 In particular, despite some expected discrepancies, 5 miRNAs (miR-222-3p, miR-146b-5p, miR-20a-5p, miR146a-5p, miR-21-5p, and miR-132-3p) are upregulated here and in Fay’s study. In addition, to assess the robustness of the data observed in RPTEC/hTERT cells and to further support the validity of our profiling approach, the miRNA deregulation of 6 miRNAs was evaluated in the widely used HK-2 cell line. 53 Interestingly, despite limited differences that may be caused by basal miRNA expression and cadmium concentrations, a similar miRNA upregulation was also globally observed. Taken together and with acknowledgement of intrinsic limitations of in vitro studies, these findings demonstrate that cadmium significantly modulates the miRNA expression profile in renal proximal tubular cells and suggest that miRNA dysregulation may participate in the pathogenesis of cadmium-induced kidney injury.
Considering their known targets, our study pointed out that the significantly deregulated miRNAs mentioned above are involved in key cellular processes including oxidative stress, inflammation, and cell death (Supplemental Table 1). In particular, evidences favor a role for miR-182-5p and miR-27a-3p in oxidative stress. Indeed, miR-182-5p has been shown to repress oxidative stress in RAW264.7 cells, an atherosclerosis model, exposed to oxidized low-density lipoprotein. 68 In addition, miR-27a-3p has been recently described to potentially target nuclear factor erythroid 2-related factor 2 (NRF2) a master regulator of oxidative stress. 69 Moreover, in our study, a large proportion of the cadmium-modulated miRNAs target genes are related to apoptosis. Among them, miR-21-5p has been suggested to act as a negative regulator in the apoptosis of tubular epithelial cells. 70 MiR-30 family members have been also shown to alter the balance between cell proliferation and apoptosis in myocardial 71,72 and renal cells. 73
Altogether, these cellular processes promote tissue remodeling, including fibrosis and cancer (Supplemental Table 1). More precisely, it is well known that a large number of miRNAs may act as either oncogenes or tumor suppressors under certain conditions and that miRNA modulation contributes to cancer-associated processes such as proliferation, invasion, metabolic reprogramming, or resistance to cell death. 74 In particular, miR-27a-3p, which is one of the most deregulated miRNAs in the present study, has been previously demonstrated to be involved in malignant transformation in lung 75 and liver 76 after cadmium exposure. Interestingly, KEGG pathway analysis, based on miR-Path 3.0, revealed that of the 38 cadmium-responsive miRNAs we described, 34 are involved in cancer pathways.
The expression of several miRNAs, termed as “fibromiRs” 77 and whose involvement in kidney fibrosis has been extensively demonstrated, 25,29,78 is also significantly modulated by cadmium exposure. For example, among the miRNAs we have validated, the well-characterized miR-21-5p has been previously reported to be increased in several experimental models of renal fibrosis, such as folic acid-induced kidney injury, ischemia/reperfusion injury, unilateral ureteral obstruction, and Alport nephropathy models. 79,80 Translational relevance of miR-21-5p has been also confirmed in patients with chronic kidney diseases or exhibiting several renal fibrotic lesions. 24,26,81
The purpose of our work was to determine whether cadmium-induced nephrotoxicity was associated with aberrant miRNA expression in kidney to potentially highlight miRNAs as biomarkers of cadmium exposure. Indeed, upon tissue injury, miRNAs have been shown to be released into the bloodstream. 82 MicroRNA propensity to be highly stable in biofluids 83 -85 has allowed seminal studies to demonstrate their potential interest as new noninvasive biomarkers. Here, we highlighted several miRNAs of interest that deserve to be further evaluated. In particular, the versatile role of miR-21-5p has previously led to propose this miRNA as a valuable biomarker in the context of renal cancer 86 and kidney injury. 24,87
In conclusion, our findings demonstrate that cadmium profoundly alters miRNA expression of proximal tubular epithelial cells and suggests that dysregulated miRNA expression may play a role in the pathogenic molecular events occurring during cadmium-induced kidney injury. In the future, the characterization of miRNAs and mRNAs interactions will provide better insight into cadmium-induced mechanisms and could allow the identification of miRNAs as promising markers of environmental stress.
Supplemental Material
Supplemental_Material - Cadmium-Induced Renal Cell Toxicity Is Associated With MicroRNA Deregulation
Supplemental_Material for Cadmium-Induced Renal Cell Toxicity Is Associated With MicroRNA Deregulation by J. Lemaire, C. Van der Hauwaert, G. Savary, E. Dewaeles, M. Perrais, J. M. Lo Guidice, N. Pottier, F. Glowacki and C. Cauffiez in International Journal of Toxicology
Footnotes
Authors’ Note
J. Lemaire and C. Van der Hauwaert contributed equally.
Author Contributions
Lemaire, Julie contributed to acquisition and analysis, drafted manuscript, and critically revised manuscript; Van der Hauwaert, Cynthia contributed to conception and design, contributed to acquisition, analysis, and interpretation, drafted manuscript, and critically revised manuscript; Savary, Grégoire contributed to acquisition and analysis and critically revised manuscript; Dewaeles, Edmone contributed to acquisition and analysis and critically revised manuscript; Perrais, Michael contributed to analysis and interpretation and critically revised manuscript; Lo Guidice, Jean-Marc contributed to design, contributed to acquisition and analysis, and critically revised manuscript; Pottier, Nicolas contributed to conception and design, contributed to analysis and interpretation, drafted manuscript, and critically revised manuscript; Glowacki, François contributed to conception and design, contributed to interpretation, drafted manuscript, and critically revised manuscript; Cauffiez, Christelle contributed to conception and design, contributed to analysis and interpretation, drafted manuscript, and critically revised manuscript. All authors gave final approval and agree to be accountable for all aspects of work ensuring integrity and accuracy.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The authors would like to thank the “Conseil Régional des Hauts-de-France” for their valuable support.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
