Abstract
Circulating microRNAs (miRNAs) could represent sensitive and specific biomarkers for tissue injury. However, their utility as biomarkers in nonclinical toxicological studies using nonhuman primates is limited by a lack of information on their organ specificity and circulating levels under resting condition of the animals. Herein, liver, heart, and skeletal muscle-specific expression patterns of miRNAs were determined in 27 tissues/organs from male and female monkeys (n =2/sex) by next-generation sequencing (NGS) analysis. This analysis revealed organ-specific miRNAs in the liver (miR-122), heart (miR-208a and miR-499a), and skeletal muscle (miR-206). Next, plasma was collected from conscious-naive male and female cynomolgus monkeys (n = 25/sex) to better understand the expressions of organ-specific circulating miRNAs. The absolute values of circulating miRNAs were quantified using a Taqman microRNA assay. MiR-1, miR-133a, and miR-208b showed preferential expression in the heart and skeletal muscles, whereas miR-192 was abundant in the liver, stomach, small intestine, and kidney. These miRNAs had identical sequences to their human counterparts. Six organ-specific miRNAs (miR-1, miR-122, miR-133a, miR-192, miR-206, and miR-499a) could be evaluated quantitatively by quantitative real-time reverse transcription polymerase chain reaction with or without preamplification. No significant sex differences were noted for these circulating miRNAs. For their circulation levels, miR-133a showed more than 900-fold interindividual variation, whereas miR-122 showed only a 20-fold variation. In conclusion, we profiled circulating organ-specific miRNAs for the liver, heart, and skeletal muscle of cynomolgus monkeys.
Keywords
Introduction
MicroRNAs (miRNAs) are small noncoding RNA molecules that play crucial roles in diverse biological processes. 1 MiRNAs have been found in extracellular body fluids, and an increasing number of studies have explored miRNAs as biomarkers for cancers and other diseases in nonclinical and clinical studies. 2,3 The expression patterns of miRNAs are informative as biomarkers because, uniquely, some miRNAs in body fluids originate from tissues or cells where miRNAs are highly or specifically expressed. 4
MiR-122 was identified as a liver-specific miRNA that is released into blood during liver injury in rodents and humans. 5,6 Furthermore, miR-1 and miR-133a are expressed abundantly in the heart, and their levels were higher in blood obtained from patients with acute myocardial infraction or rats treated with a cardiotoxicant. 7,8 Although the mechanism underlying this elevation in circulating miRNAs remains to be elucidated, circulating miRNAs could leak from the cytoplasm of damaged cells as by-products at variable levels based on the observation that extracellular miRNA levels well correlated with the extent of cell death. 9 Therefore, organ-specific circulating miRNAs represent potential safety biomarkers for organ toxicity.
MiRNAs are also recognized as promising candidates for translational biomarkers because the sequences of most miRNAs are conserved among different animal species. 10 The high degree of sequence homology raises the possibility that their distribution and roles would be similar across species, which suggests that the release of plasma miRNAs from injured cells would have a common mechanism. However, miR-208a has been reported to be present in rat plasma but not in human plasma, possibly reflecting a difference in expression levels. 11 –13 This result indicated that species differences in circulating miRNAs should be taken into account for toxicological evaluation of drugs.
The cynomolgus monkey (Macaca fascicularis) is used commonly as a nonhuman primate, especially in the field of toxicology. They are considered to be anatomically, physiologically, and genetically closer to humans compared to other laboratory animals, such as rats and mice. 14,15 However, the latest version of miRBase (version 21) contains mature miRNAs only from closely related species, such as Rhesus macaque (Macaca mulatta) or humans. Recently, next-generation sequencing (NGS) has emerged as a powerful tool to reveal novel or homologous sequences of unknown miRNAs. Using this technology, mature miRNAs in the kidney 16 or testis 17 of cynomolgus monkeys were identified and shown to have high homology to those from Rhesus macaque and humans. To identify other organ-specific miRNAs in this species, preferentially expressed miRNAs from different organs should also be evaluated.
Quantitative real-time reverse transcription polymerase chain reaction (RT-qPCR) is one of the most common methods used to assess individual target miRNAs. 18 In general, miRNAs are measured by relative expression relative to a reference sample. This approach is appropriate for most purposes to examine physiological changes in target gene expression levels. However, absolute quantification is preferred for the comprehensive assessment of basal expression level, with a better statistical analysis.
In this study, we focused on 8 miRNAs that have been proposed as potential safety biomarkers associated with tissue injury to the liver (miR-122 and miR-192), heart (miR-1, miR-208a, miR-208b, and miR-499a), and skeletal muscle (miR-133a and miR-206) in rodents and in human. 11,19 Their organ specificity and absolute circulating levels in cynomolgus monkeys were identified using NGS and RT-qPCR, respectively.
Materials and Methods
Next-Generation Sequencing Analysis
To identify organ-specific miRNAs from the liver, heart, and skeletal muscle, a previously published NGS data set was used to determine testicular-specific miRNAs in our laboratory. 17 Briefly, the NGS data for 27 tissues and organs, including the adrenal gland, aorta, colon, duodenum, eye, heart (left ventricular wall and papillary muscle), jejunum (including the ileum), kidney (cortex, medulla, and papilla), liver, lung, ovary (from females), plasma, pancreas, rectum, sciatic nerve, skeletal muscle (rectus femoris and soleus), skin, spleen, stomach, testis (from males), trachea, urinary bladder, and uterus (from females) were employed. The expression levels in each tissue or organ were normalized from the raw count data to reads per million (rpm). The percentage of miRNA expression in the target organ was calculated by dividing the sum of the counts (rpm) in all organs by the counts (rpm) in each tissue or organ. To assess the organ specificity of each miRNA, the counts for organs in which different parts were sectioned, such as the heart and kidney, were expressed as their average.
Animals
All experiments were approved by the Institutional Animal Care and Use Committee of Daiichi Sankyo Co, Ltd (Tokyo, Japan). In total, 25 male and 25 female cynomolgus monkeys originating from the Philippines were obtained from the Oriental Yeast Co, Ltd (Tokyo, Japan) and used to measure the levels of selected circulating miRNAs. The animals were 3 to 4 years of age and weighed between 4.25 and 6.90 kg. The animals were housed individually in stainless steel cages (W 594 × D 870 × H 1,015 mm) in an animal study room with a controlled temperature of 24°C and humidity of 60%; the lighting was on a 12-hour light–dark cycle. The monkeys were fed with commercial pellets (100 g/day/animal; PS-A, Oriental Yeast) and were allowed free access to chlorinated tap water.
Sample Preparation
Whole blood samples (1 mL per sampling point) were collected from the femoral vein into tubes containing EDTA-2K (MICROTAINER; Japan Becton Dickinson company, Ltd, Tokyo, Japan). The obtained samples were immediately placed on ice and processed for plasma isolation within 2 hours postcollection, according to a previously reported method. 20 The samples were centrifuged at 10 000×g at 4°C for 5 minutes using a fixed-angle rotor (CF15R; Hitachi Koki, Tokyo, Japan). The supernatant was inspected visually for hemolysis or discoloration and then transferred into a 2-mL microtube (Eppendorf Japan, Tokyo, Japan), followed by centrifugation at 16 000×g at 4°C for 5 minutes to remove cell debris. 21 The supernatant was aliquoted into a new microtube.
RNA Isolation
A fixed volume of each sample was used as described previously. 20,22 Total RNA was extracted using a commercially available column-based system: the miRNeasy Mini kit (QIAGEN, Tokyo, Japan), according to the manufacturer’s instructions with some modifications. Briefly, 200 μL samples of plasma were prepared and 1,000 μL of QIAzol lysis reagent was added to each sample, and the mixture was shaken vigorously. Following shaking, 5 μL of 5 nmol/L synthetic Caenorhabditis elegans miRNA (Syn-cel-miR-238-3p, miScript miRNA Mimic; QIAGEN) was added, and each sample was immediately mixed with 200 μL of chloroform (Wako Pure Chemical Industries, Osaka, Japan). The samples were centrifuged at 12 000×g at 4°C for 15 minutes. The aqueous phase was transferred to a new microtube, 1.5 volumes of ethanol (Wako Pure Chemical) were added to the tube, and the samples were applied to an RNeasy Mini Spin Column with SC adaptors (Taigen Bioscience Corporation, Taipei, Taiwan), followed by vacuum drying for 2 minutes using a VacEzor system (Taigen Bioscience Corporation). A further vacuum-rinsing step was conducted on the system as follows: 200 μL of ethanol for 1 minute, 800 μL of RWT Buffer for 2 minutes, repeated 800 μL of RPE buffer for 2 minutes, and 300 μL of ethanol for 1 minute. The column was then placed onto a new microtube and centrifuged at 12 000×g at room temperature for 1 minute. The total RNA was eluted from the column by 2 repeated elutions with 50 μL of RNase-free water (QIAGEN) to increase the amount of miRNAs obtained.
Complementary DNA Synthesis
The isolated RNA was reverse transcribed using a TaqMan MicroRNA Reverse Transcription kit (Thermo Fisher Scientific, Tokyo, Japan) combined with a multiplexed stem-loop primer pool from the TaqMan MicroRNA Assays (Thermo Fisher Scientific). The reaction components, including the multiple miRNA reverse transcriptase (RT) primer, were the same as those detailed by Roberts et al. 23 Briefly, 5 μL of total RNA was added to each RT reaction mix, which comprised 3 μL of the RT primer pool, 0.15 μL of 100 mmol/L dNTPs with dTTP, 1 μL of MultiScribe RT (50 U/μL), 1.5 μL of 10 × RT buffer, 0.19 μL of RNase inhibitor (20 U/μL), and 4.16 μL of nuclease-free water in a total reaction volume of 15 μL. Reverse transcription was performed at 16°C for 30 minutes, 42°C for 30 minutes, followed by a final reverse transcriptase inactivation step at 85°C for 5 minutes on a GeneAmp PCR System 9700 (Thermo Fisher Scientific). Standard curves for absolute quantification were generated using a QuantiGene miRNA control (Affymetrix, Tokyo, Japan) or commercially available synthesized RNA oligonucleotides (Hokkaido System Science Co, Ltd, Hokkaido, Japan) that corresponded to known miRNAs. These controls were serially diluted and added to the RT reaction mix described earlier. For normalization, another dilution series was created with a random complementary DNA (cDNA) from our sample group to construct standard curves for cel-miR-238.
Preamplification
Among the 8 selected miRNAs (miR-1, miR-122, miR-133a, miR-192, miR-206, miR-208a, miR-208b, and miR-499a), 5 (miR-1, miR-206, miR-208a, miR-208b, and miR-499a) showed quantification cycle (Cq) values above 35 or were not detectable in samples by qPCR. The RT products for those 5 miRNAs were preamplified before the qPCR step to enhance the potential sensitivity, in combination with a multiplexed stem-loop primer pool. The primer pool comprised all primers for less abundant target miRNAs, with the final concentration of each RT primer being diluted 200-fold by Tris-EDTA buffer in a final volume of 1,000 μL. The preamplification step was carried out in a reaction comprising 12.5 μL of 2 × Taqman PreAmp Master Mix (Thermo Fisher Scientific), 3.75 μL of the RT primer pool, 6.25 μL of nuclease-free water, and 2.5 μL of the undiluted RT product in a total reaction volume of 25 μL. Each sample was thermocycled on a GeneAmp PCR System 9700 under the following cycling conditions: 95°C for 10 minutes, followed by 14 cycles of 95°C for 15 seconds, and 60°C for 4 minutes. Standard curves for preamplified miRNAs were also constructed using the RT products from the synthesized controls as mentioned in the previous section.
Quantitative Real-Time PCR
The qPCR reactions were performed using a 7900HT Fast Real Time PCR System (Thermo Fisher Scientific) using the TaqMan MicroRNA Assays (Thermo Fisher Scientific) and the TaqMan Fast Advanced Master Mix (Thermo Fisher Scientific), according to the manufacturer’s instructions. First, the samples were diluted by adding 20 μL of water to 5 μL of either the preamplification or the RT product. Then 2 μL of the diluted sample was added into the reaction premix aliquots, which comprised 10 μL of 2 × TaqMan Fast Advanced Master Mix, 1 μL of miRNA-specific primer/probe, and 7 μL of nuclease-free water in a total reaction volume of 20 μL in an optical 96-well plate. The plate was then sealed with ABI MicroAmp Optical Adhesive Film (Thermo Fisher Scientific). Cycling conditions were as follows: 50°C for 2 minutes and 95°C for 20 seconds followed by 45 cycles of 95°C for 1 second and 60°C for 20 seconds.
Data Analysis
Data were analyzed with SDS Software version 2.3 (Thermo Fisher Scientific). The threshold line was set manually at “1.0” in all plates in the study to confirm the reproducibility by comparison with their Cq values. A copy number of each sample was calculated from the standard curve for absolute quantification and the Cq values of the samples, followed by further normalization by the relative values from cel-miR-238. The efficiencies of PCR amplification were calculated using the formula E =(10 −1/slope −1) × 100%, using the slope of the plot of the Cq values for each serial dilution against the log cDNA input level. The cutoff level was set at Cq > 40 cycles. When the data were observed to be skewed positively, a logarithmic transformation was applied to make the distribution close to normal. 24 The means and standard deviations were transformed back into the original scale. The values for 2 standard deviations below and above the mean were defined as the “dynamic range” in the present study.
Statistics
To determine statistical significance of the differences, the expression levels of miRNAs were analyzed using the Wilcoxon rank-sum test (for comparison of 2 nonpaired groups) or a paired t test (for intraindividual variations). These analyses were performed using Microsoft Office Excel 2010 (Microsoft Corporation, Washington) and the SAS System Release 9.2 (SAS Institute Inc, Cary, North Carolina). A P value of less than .05 was considered statistically significant.
Results
Next-Generation Sequencing Analysis of the Organ Specificity of miRNAs in Cynomolgus Monkeys
More than 1,000 mature miRNAs were filtered using the criteria that the expression level of the miRNA in a target organ exceeded both 90% and 500 rpm. The expression levels of each miRNA with the top 5 most abundant tissues or organs are shown in Figure 1. As a result, miR-122, miR-208a and miR-499a, and miR-206 were observed to be highly expressed in the liver, heart, and skeletal muscle, respectively (Figure 1). Furthermore, miR-1, miR-133a, and miR-208b were abundant in the heart and skeletal muscle compared to their levels in other organs. By contrast, miR-192 was abundant not only in the liver but also in the stomach, intestines (duodenum, jejunum, and colon), and kidney. Most of the highly expressed organ-specific miRNAs exhibited 100% sequence homology with the respective human mature miRNAs (Table 1); therefore, the primers and probes for the human miRNAs were considered to be available for subsequent RT-qPCR assay.
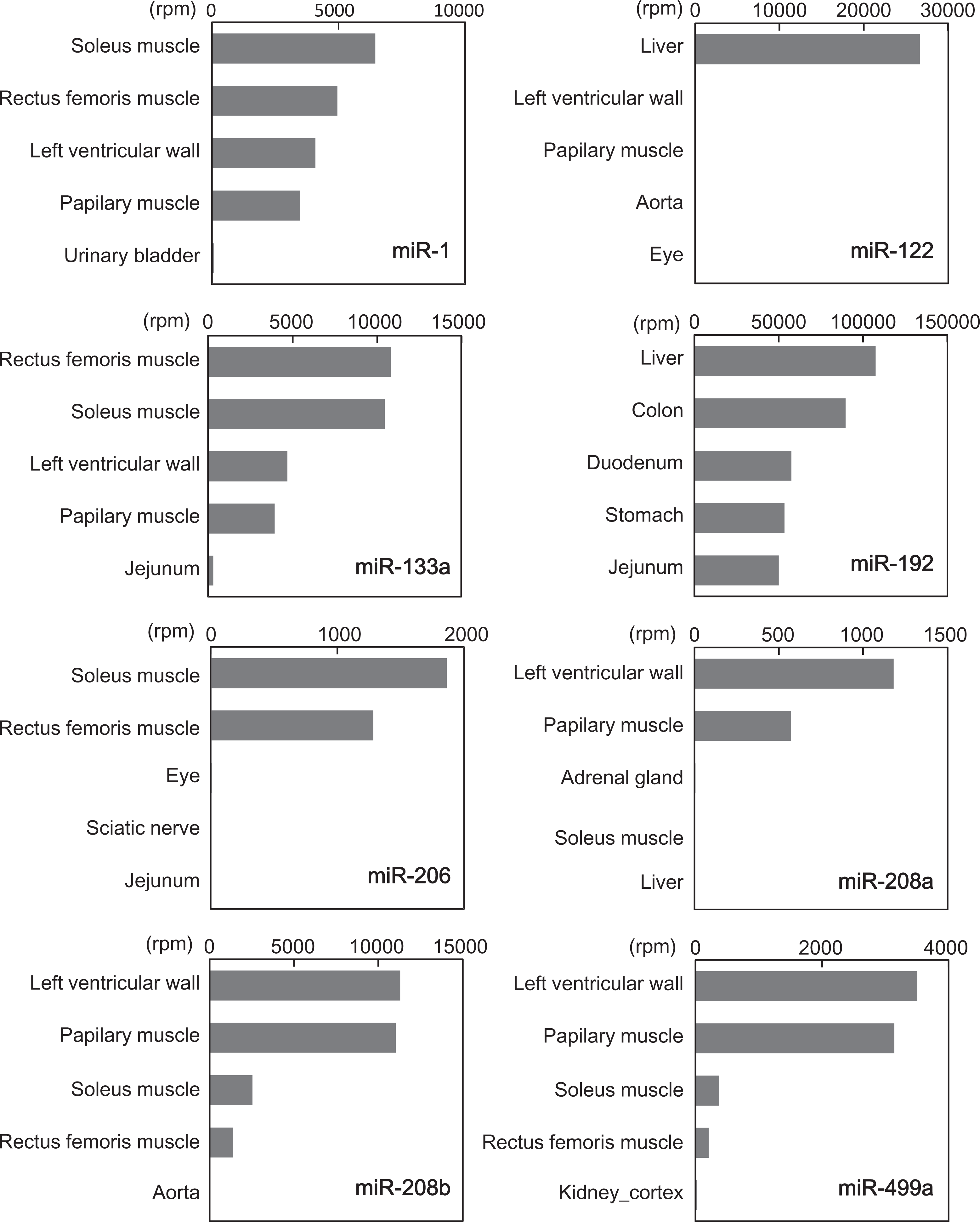
Expression profiles of selected microRNAs (miRNAs) in cynomolgus monkeys identified by next-generation sequencing (NGS). The x-axis represents reads per million.
Sequences of Organ-Specific microRNAs (miRNAs) Identified by Next-Generation Sequencing (NGS).a
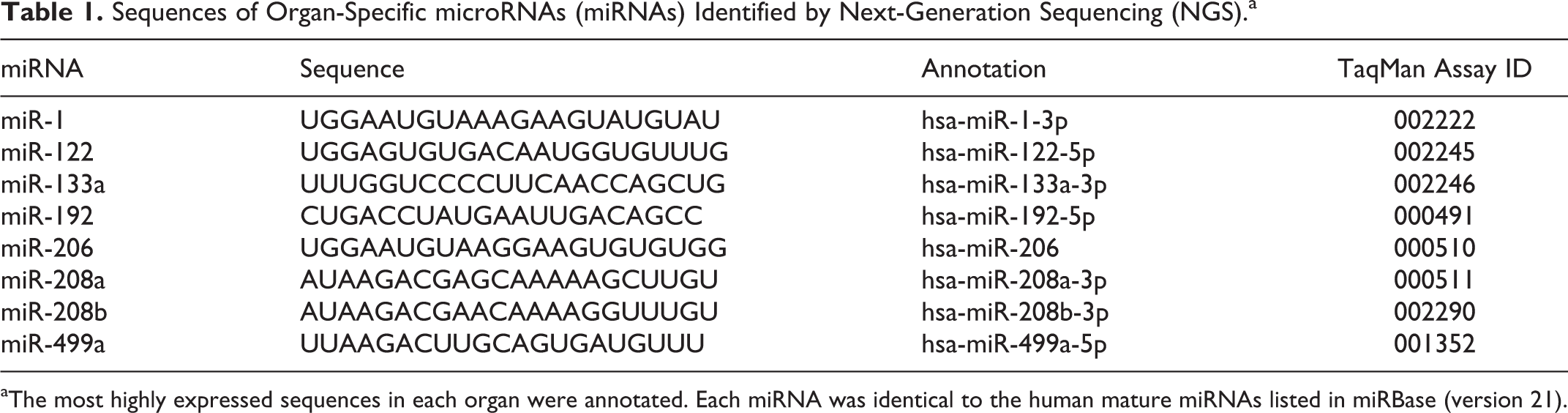

aThe most highly expressed sequences in each organ were annotated. Each miRNA was identical to the human mature miRNAs listed in miRBase (version 21).
Quality Assessment of Extraction and RT-qPCR Performance
To confirm the efficiency of sample processing, the Cq values of the spiked-in cel-miR-238 were compared in each assay. The difference in the Cq values was less than 2.5-fold (Table 2) even in cases where the cel-miR-238 was diluted through the preamplification step. A 10-fold serial dilution series of a synthetic oligonucleotide revealed that the lower limit of quantification (LLOQ) was 102 copy/μL for most of the miRNAs. Only miR-1 showed a higher LLOQ (103 copy/μL). The average amplification efficiency was approximately 90%, with the correlation coefficient (R 2) of the standard curves ranging from .998 to 1.000 (data not shown).
Quantification Cycle (Cq) Values of the Spiked-In Control miR-238 in Each Assay.a
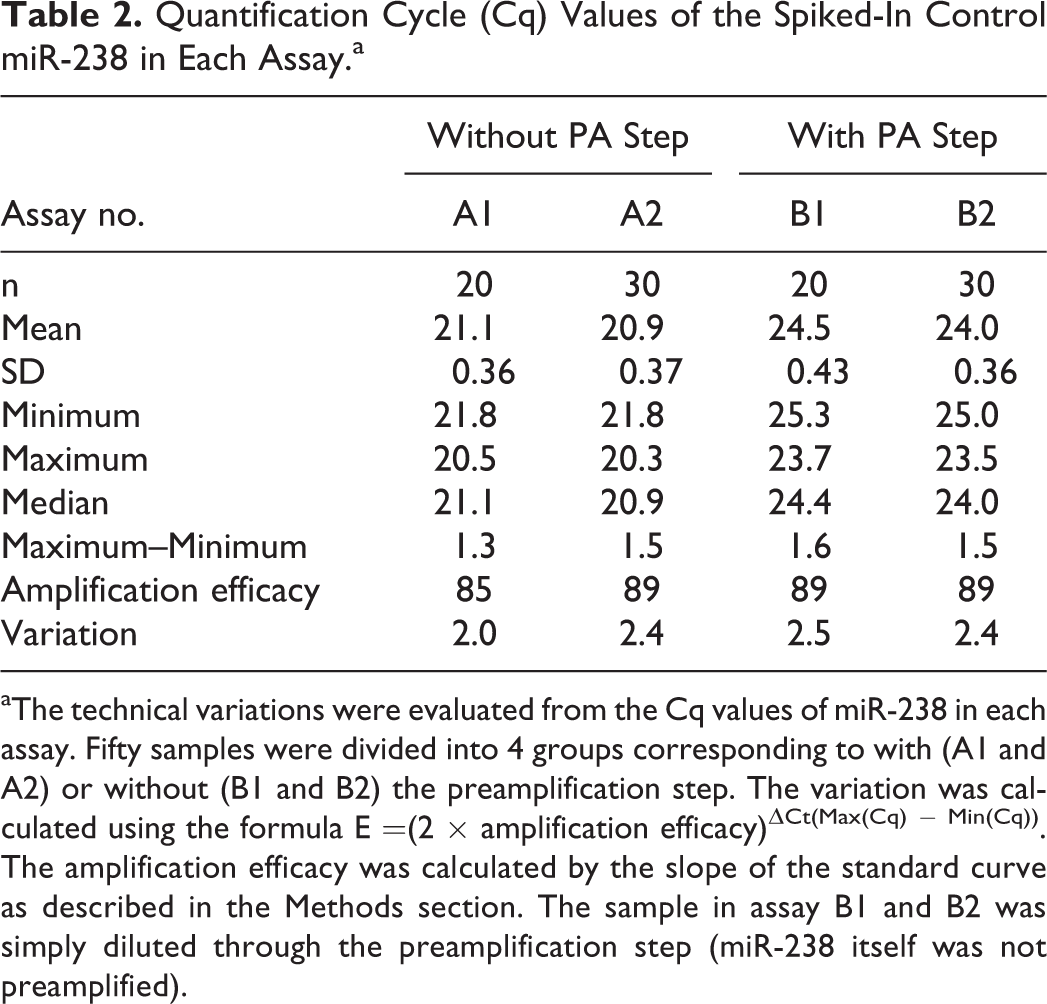
aThe technical variations were evaluated from the Cq values of miR-238 in each assay. Fifty samples were divided into 4 groups corresponding to with (A1 and A2) or without (B1 and B2) the preamplification step. The variation was calculated using the formula E =(2 × amplification efficacy)ΔCt(Max(Cq) − Min(Cq)). The amplification efficacy was calculated by the slope of the standard curve as described in the Methods section. The sample in assay B1 and B2 was simply diluted through the preamplification step (miR-238 itself was not preamplified).
Real-Time qPCR Profiling of Circulating miRNAs
Using the established assay platform, plasma levels of 8 miRNAs (miR-1, miR-122, miR-133a, miR-192, miR-206, miR-208a, miR-208b, and miR-499a) from 50 cynomolgus monkeys were comprehensively analyzed. In the assay, miR-122, miR-133a, and miR-192 were detectable without preamplification, whereas miR-1, miR-206, and miR-499a required preamplification because of their low expression levels. However, neither miR-208a nor miR-208b was detectable even using preamplification.
To confirm any sex difference in the circulating miRNAs, each detectable miRNA was compared between 25 males and 25 females. No sex differences were observed for any of the miRNAs (Figure 2; Table 3). The data were not normally distributed; therefore, a logarithmic transformation was performed. Among the miRNAs examined, miR-122 showed the highest mean plasma level (5.71 × 104 copy/μL), with a small dynamic range (20-fold). Conversely, a large dynamic range was observed for miR-1 (581-fold), miR-133a (971-fold), and miR-206 (426-fold). The small dynamic range of miR-499a (38-fold) indicated that preamplification had no negative effects on the dynamic range. Intraindividual variations in the expression of each circulating miRNA were also evaluated; however, these variations were almost within the reference ranges, except for 1 animal that showed high miR-133a expression (Figure 3).
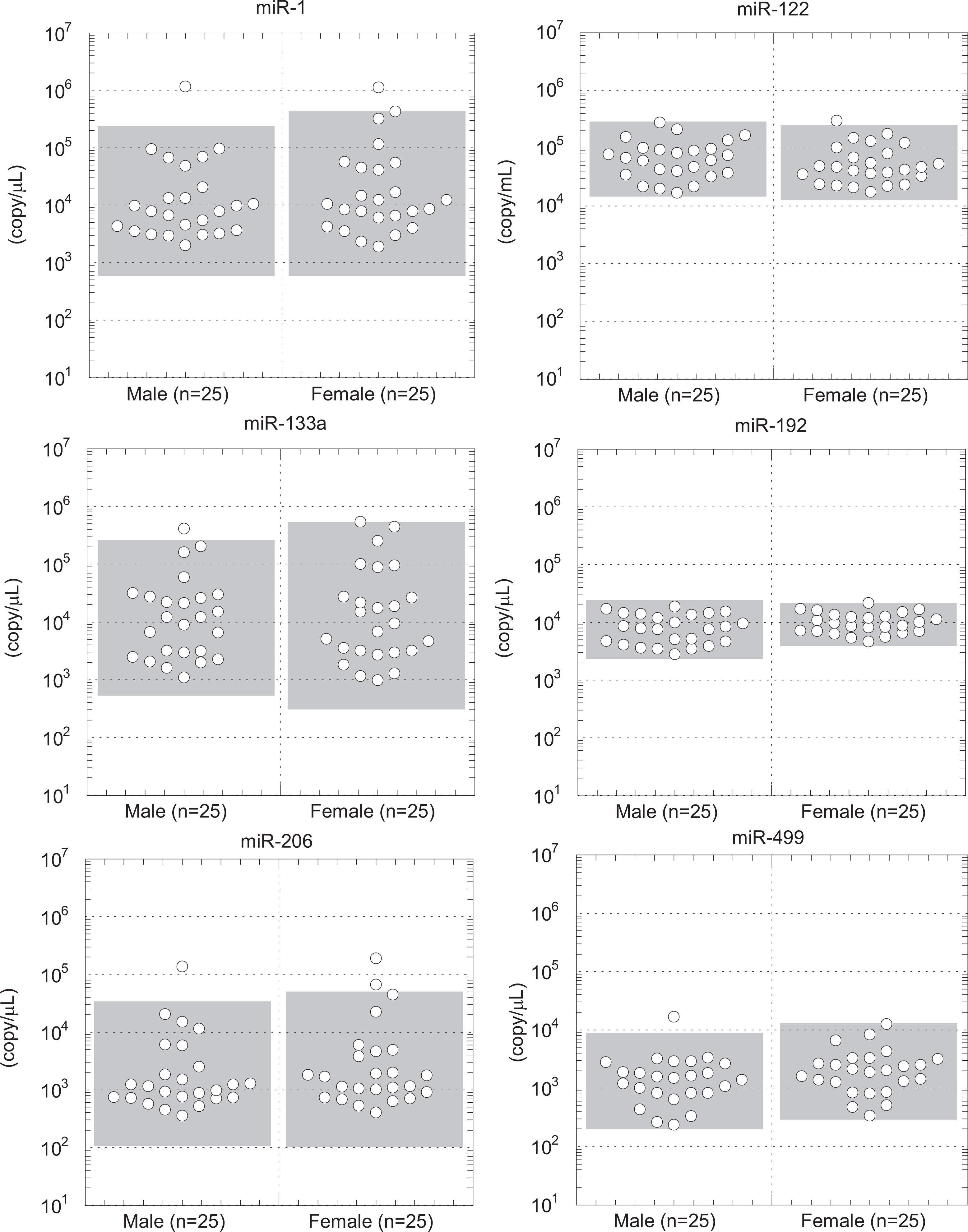
Circulating levels of organ-specific microRNAs (miRNAs) in male and female monkeys. The expression levels of the miRNAs are represented by dot plots. Differences were analyzed by the Mann-Whitney U test. The range of the mean ± 2 standard deviation (SD) from the male or female data are shown in gray boxes.
Statistical Values for the Plasma Levels of Organ-Specific MicroRNAs (miRNAs).a
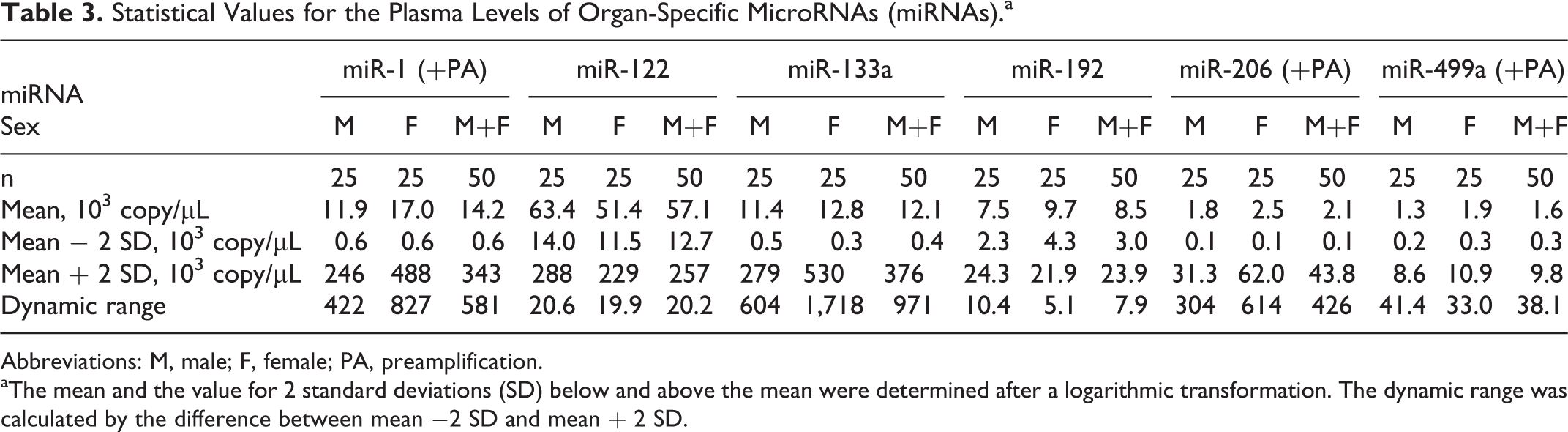
Abbreviations: M, male; F, female; PA, preamplification.
aThe mean and the value for 2 standard deviations (SD) below and above the mean were determined after a logarithmic transformation. The dynamic range was calculated by the difference between mean
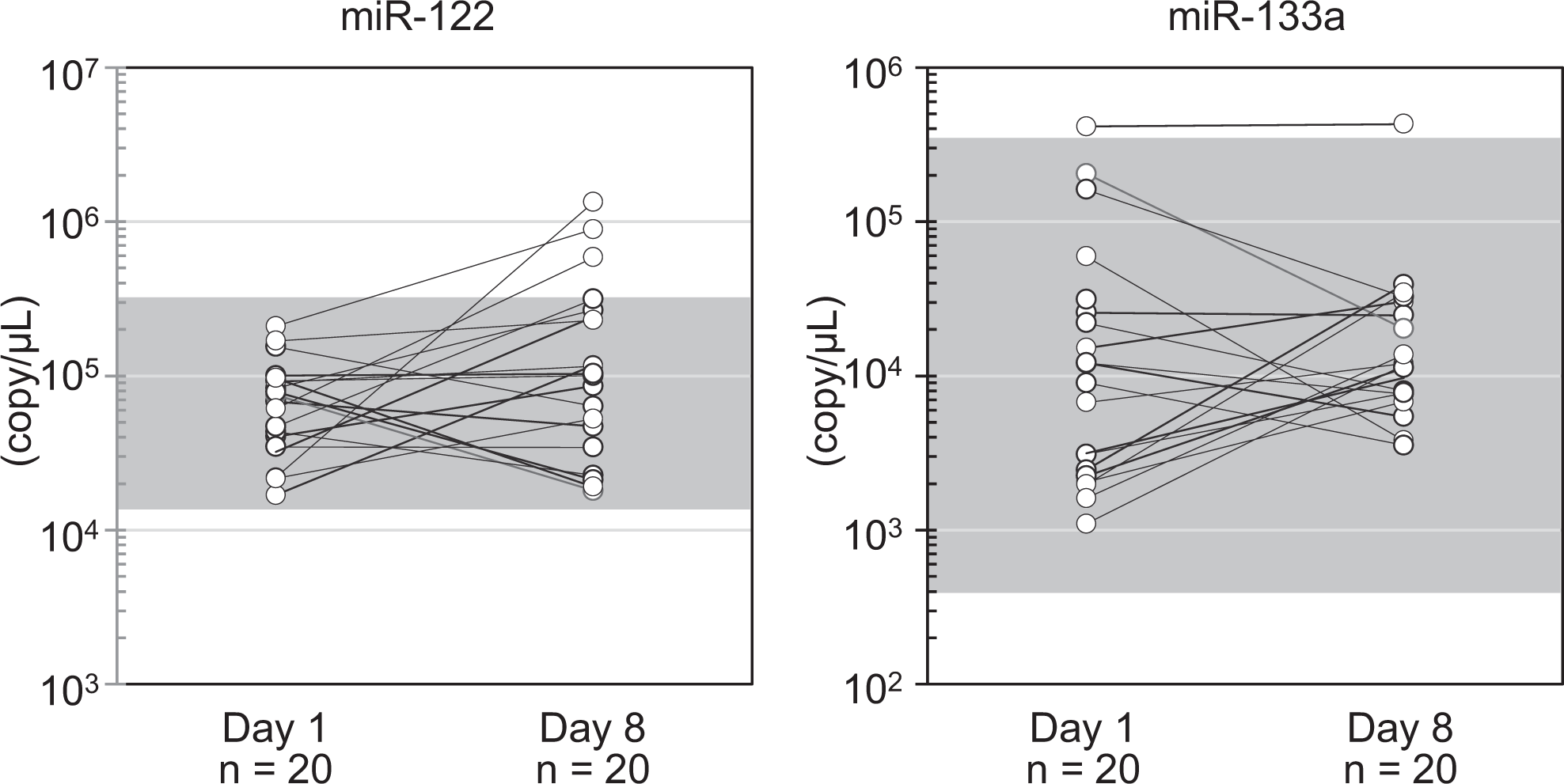
Intraindividual differences in microRNAs (miRNAs). The plasma level of the miRNAs between different sampling days (day 1 and day 8) in the same animal are shown. The dynamic ranges (mean ± 2 standard deviation [SD]) calculated from 50 cynomolgus monkeys are shown in gray boxes. No significant changes were observed between the sampling days using a paired t-test.
Discussion
Among more than 1,000 mature miRNAs identified in the 27 tissues and organs of the cynomolgus monkey by NGS analysis, we focused on only 8 miRNAs that are thought to be associated with tissue injury to the liver (miR-122 and miR-192), heart, and/or skeletal muscle (miR-1, miR-133a, miR-206, miR-208a, miR-208b, and miR-499a) in rodents and in humans. The top hit sequence of each miRNA had completely identical sequences to its human miRNA counterpart. Recently, Veeranagouda et al 16 identified 913 miRNAs in the cortex of the kidney of cynomolgus monkeys. Their annotations of these miRNAs were almost identical to those of the present results, except for miR-133a, which was annotated differently as mfa-miR-133c. In our current NGS data, hsa-miR-133a (mfa-miR-133c) was more abundant in the heart and skeletal muscles compared to mfa-miR-133a.
The NGS analysis revealed that miR-122 was a liver-specific miRNA. In contrast, miR-192, which was suggested as a potential biomarker for liver injury in rodents and humans, 5,6 was abundant not only in the liver but also in the stomach, intestines, and kidney. This expression pattern is not specific to cynomolgus monkeys because miR-192 was also identified in multiple organs in rats and dogs. 25,26 MiR-192 was reported to increase as a result of renal ischemia–reperfusion injury in rats 3 ; therefore, the source of miR-192 should be interpreted carefully when its circulating level is elevated. Muscle-specific miRNAs, referred to as “myomirs,” are a group of miRNAs that are highly enriched in cardiac and/or skeletal muscles. 27 Based on the present study, certain myomirs showed exclusive expression in the heart and/or skeletal muscle. Notably, myomir miR-208b showed the highest expression in the heart of cynomolgus monkeys, which is in line with data from dogs. 25 By contrast, miR-208b was reported to be expressed predominantly in the soleus muscle in mice. 28 Among species, the predominant miR-208 subtype in the heart could vary in parallel with α-major histocompatibility complex (α-MHC) and β-MHC expression levels. In the heart, α-MHC is highly expressed in rodents, whereas β-MHC is the predominant myosin in large animals, such as dogs and humans. 29 The miR-208a gene is located in the MYH6 gene that encodes α-MHC, whereas the miR-208b resides in the MYH7 gene that encodes β-MHC. 30 Although their precise mechanism of regulation remains to be investigated, miR-208a and miR-208b appear to be coexpressed with their host myosin.
Preamplification is useful to enhance the detection of poorly expressed miRNAs, without introducing a detection bias. 31,32 In the present study, the Cq values of the standard curves constructed using preamplified synthesized controls showed almost same values across the assays. This high reproducibility allowed us to perform absolute quantitation using preamplified samples, which resulted in the detection of additional miRNAs. Contrastingly, some miRNAs were not detectable even when combined with preamplification. Further experiments are necessary to verify whether the undetected miRNAs exist in circulating blood.
Our comprehensive assessment using samples from 50 animals also provided a more rigorous statistical analysis of the extent of the dynamic range. However, it should be noted that the data included biological variation as well as technical variation. The biological variation remains a challenge for the use of these data because a standard internal control has not yet been established. In this respect, external reference controls were proposed to compare the results of studies conducted in different laboratories incorporating different platforms. 33,34 In fact, almost all miRNA studies used exogenous controls for normalization routinely; however, the PCR efficiency for the data has rarely been addressed. To minimize these variations, the procedures in our method were kept consistent throughout the study, in accordance with the following points. First, we processed the plasma as quickly as possible (within 2 hours postcollection) to exclude the effect of the stability of miRNAs in plasma among different species. 35 Second, each plasma sample was inspected visually to avoid analytical bias from contamination caused by hemolysis, which represents one of the confounding factors that might affect the levels of circulating miRNAs. 36 Consequently, the technical variation in the expression levels of the external control was less than 2.5-fold.
In our RT-qPCR method, 6 (miR-1, miR-122, miR-133a, miR-192, miR-206, and miR-499a) of 8 miRNAs could be evaluated quantitatively in plasma from animals in the resting state. MiR-1, miR-133a, and miR-206 showed more than a 400-fold variation, whereas miR-122 and miR-192 showed only 20-fold and 8-fold variation, respectively. In another study using human serum or plasma from healthy participants (n =25), 5 the dynamic ranges of miR-1, miR-122, and miR-192 were estimated as 87-fold, 22-fold, and 15-fold, respectively. Despite conducting preamplification, neither miR-208a nor miR-208b could be detected in the monkey plasma. Circulating miR-208a was reported to be measured consistently in rats but not in humans. 8,11 –13 Furthermore, miR-499a was reported to increase in acute myocardial injury in humans 37 but not in rodents. These results suggested the expression levels of miR-208a and miR-499a in cynomolgus monkeys might be comparable to those in humans. However, it is difficult to compare the circulating levels of the miRNAs between cynomolgus monkeys and other species because all previously reported miRNAs were measured as relative values compared to a reference sample.
The extent of miRNA expression in their predominant organs was not reflected in the circulating miRNA levels. MiR-122 showed an approximately 8-fold higher mean plasma level compared to that for miR-192, whereas the counts in the liver for miR-122 were lower than those for miR-192. Similarly, miR-499a showed relatively fewer counts in the heart compared to miR-208b, but only miR-499a was detectable in the plasma. Such inconsistencies suggest that detectable miRNAs in plasma might be secreted actively via microvesicles, such as exosomes, to play a role in cell-to-cell communication. 38 In fact, hepatic and cardiac exosomes could serve as a source of organ-specific miRNAs that are involved in various function in the liver and heart, respectively. 39,40 Although the individual roles of the detected miRNAs remain incompletely understood, the observed high expression levels of the extracellular miRNAs might contribute to certain physiological and pathological activities of target cells. Further investigation is needed to clarify the roles of the circulating miRNAs.
In conclusion, we profiled circulating organ-specific miRNAs for the liver, heart, and skeletal muscle of cynomolgus monkeys using NGS and absolute RT-qPCR methods. Further study using liver, heart, or skeletal muscle injury models in cynomolgus monkeys should be undertaken to clarify whether these circulating miRNAs have potential use as translational safety biomarkers.
Footnotes
Acknowledgments
The authors thank Hidetoshi Ooshima, Isamu Kikuchi, Yu Yoshimatsu, and Shunsuke Takada for their excellent technical assistance in animal treatments and sample preparations which facilitated this work.
Author Contribution
Takuma Iguchi, Satoshi Tamai, Ken Sakurai, and Kazuhiko Mori contributed to conception and design, acquisition, analysis, and interpretation, drafted the article, critically revised the article, gave final approval, and agreed to be accountable for all aspects of work ensuring integrity and accuracy. Noriyo Niino contributed to measurement of miRNAs, analysis, and interpretation, drafted the article, critically revised the article, gave final approval, and agreed to be accountable for all aspects of work ensuring integrity and accuracy. Takuma Iguchi contributed to conception and design and contributed to acquisition, analysis, and interpretation; Noriyo Niino contributed to conception and design and contributed to acquisition, analysis, and interpretation; Satoshi Tamai contributed to conception and design and contributed to acquisition, analysis, and interpretation; Ken Sakurai contributed to conception and design and contributed to acquisition, analysis, and interpretation; Kazuhiko Mori contributed to conception and design and contributed to acquisition, analysis, and interpretation. All authors drafted and critically revised the manuscript, gave final approval, and agreed to be accountable for all aspects of work ensuring integrity and accuracy.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
