Abstract
Antiproliferative potential of nonsteroidal anti-inflammatory drugs (NSAIDs) has generated an immense interest among the scientific fraternity to assess its broader role in the chemoprevention of colon cancer. Due to serious adverse events associated with the chemotherapy, NSAIDs have been exploited as adjuvants to synergize the cytotoxic potential of conventional chemotherapeutic agents at low-dose regimens. The present investigation has been focused on in silico model generation for in vitro cytotoxicity activity of the clinically active NSAIDs using self-organizing molecular field analysis (SOMFA) studies. A statistically validated robust model for a diverse group of NSAIDs having flexibility in structure and cytotoxicity activity was obtained using SOMFA. The statistical measures having good cross-validated correlation coefficient r 2 cv (.8291), noncross-validated correlation coefficient r 2 values (.8686), and high F test value (41.8722) proved significance in the generated model. Analysis of 3-dimensional quantitative structure activity relationship (3D-QSAR) models through electrostatic and shape grids provided additional valuable information regarding shape and electrostatic potential influence on in vitro cytotoxicity profile. The analysis of SOMFA results provided a better insight about the generation of molecular architecture of novel NSAIDs yet to be synthesized, with optimum in vitro cytotoxicity activity and improved therapeutic profile.
Introduction
Colorectal cancer is the third most commonly diagnosed cancer in males and the second in females, with over 1.2 million new cancer cases and 608 700 deaths estimated to have occurred in 2008. 1 Both environmental (diet and physical activity) and genetic (family history, mutations, and polymorphisms) factors are related to colon cancer risk. 2 Nonsteroidal anti-inflammatory drugs (NSAIDs) have been an area of prime interest in achieving the antitumor response in various types of cancers including colon carcinogenesis. The NSAID-induced colon cancer chemoprevention is the specific area, which has been widely implicated by the scientific community. Recent studies showed that prediagnostic NSAID use is associated with improved colorectal cancer survival among women and have a function in altering colorectal cancer mortality. 3 Largest randomized trials conducted by the American Cancer Society showed that the use of NSAIDs has led to a 40% reduced risk of colon cancer. 4 Epidemiologic, preclinical, and clinical studies all suggested that NSAIDs are potent preventive agents for the therapeutic management of colon cancer. 5 Further, recent clinical studies have suggested the long-term benefits of regular use of low-dose aspirin in reducing the lower incidence of adenomatous polyps and lower incidences of deaths from colorectal cancer compared with nonusers. 6,7 Most epidemiologic studies (case–control and cohort) have advocated the long-term use of low-dose aspirin as suitable prospective to decrease the morbidity and increased survival rate in colon cancer. 8 –14 The entire class of NSAIDs appears to be effective in preventing carcinogen-induced colon cancer in animal models. Clinical trials using the NSAID, sulindac, have shown dramatic regression of colonic adenomas in patients with familial polyposis. 15,16 The clinical approach for the management of colon cancer has been flexibly broadened by the oncologists. The NSAIDs are being employed clinically as adjuvant in chemotherapy to counter the severity of adverse effects associated with conventional chemotherapeutic regimens. 17
Structural and shape variability are generally employed to probe biological systems and potentially discover novel compounds with selective target activity. Structure-based drug discovery relies on the fact that molecules with similar chemical features will elicit similar activity in biological assays. 18,19 In silico techniques offer highly robust models and tools to optimize the drug properties through quantitative structure activity relationship. Chemical similarity can also be used to construct diverse sets of probe molecules in order to find as many different molecular structures/scaffolds as possible that generate a similar biological response. 20 Cell viability assay for the inhibition of tumor cells are considered as the major determinant for the assessment of antiproliferative activity of a potential candidate in tumor cell’s biology. In this assay, concentration of the particular test compound or drug is quantified when growth of the cancer cells inhibits to 50% that is GI50 (median growth inhibition). The cytotoxic evaluations of the drugs through quantitative cell-based assays provide better understanding of the antitumorigenic activity and information about drug mechanism. Investigations of mechanism-based biological response have established that some drug classes, not necessarily with the same structure, cause a specific growth inhibition pattern across these tumor cell lines that can be related, by secondary testing, to a precise biological effect. 21 –25 Thus, it could be assumed from these findings that such screening of the biological activity can also be exploited to design novel compounds that elicit a desired mechanism of action, based solely on analogy to characteristic growth inhibition patterns. Further, 3-dimensional quantitative structure activity relationship (3D-QSAR) approach based on self-organizing molecular field analysis (SOMFA) could be used as a chemometric tool for deselecting newly screened compounds if their responses correspond to a previously well-exploited drug activity. The present investigation is an attempt to obtain a statistically validated model for in silico screening and prediction of the cytotoxicity activity (or GI50) of the clinically active NSAIDs using SOMFA studies in HT-29 colon cancer cells.
Computational Analysis
Data Set
Data set of 23 clinically active molecules belonging to a diverse group of NSAIDs were taken from the literature 26 –33 and used for 3D-QSAR study. 34,35 The diverse group of NSAIDs employed in the present 3D-QSAR study using SOMFA approach along with their observed and actual GI50 in HT-29 colon cancer cells has been illustrated in Table 1.
In Vitro Cytotoxicity Assay of NSAIDs in HT-29 Colon Cancer Cells
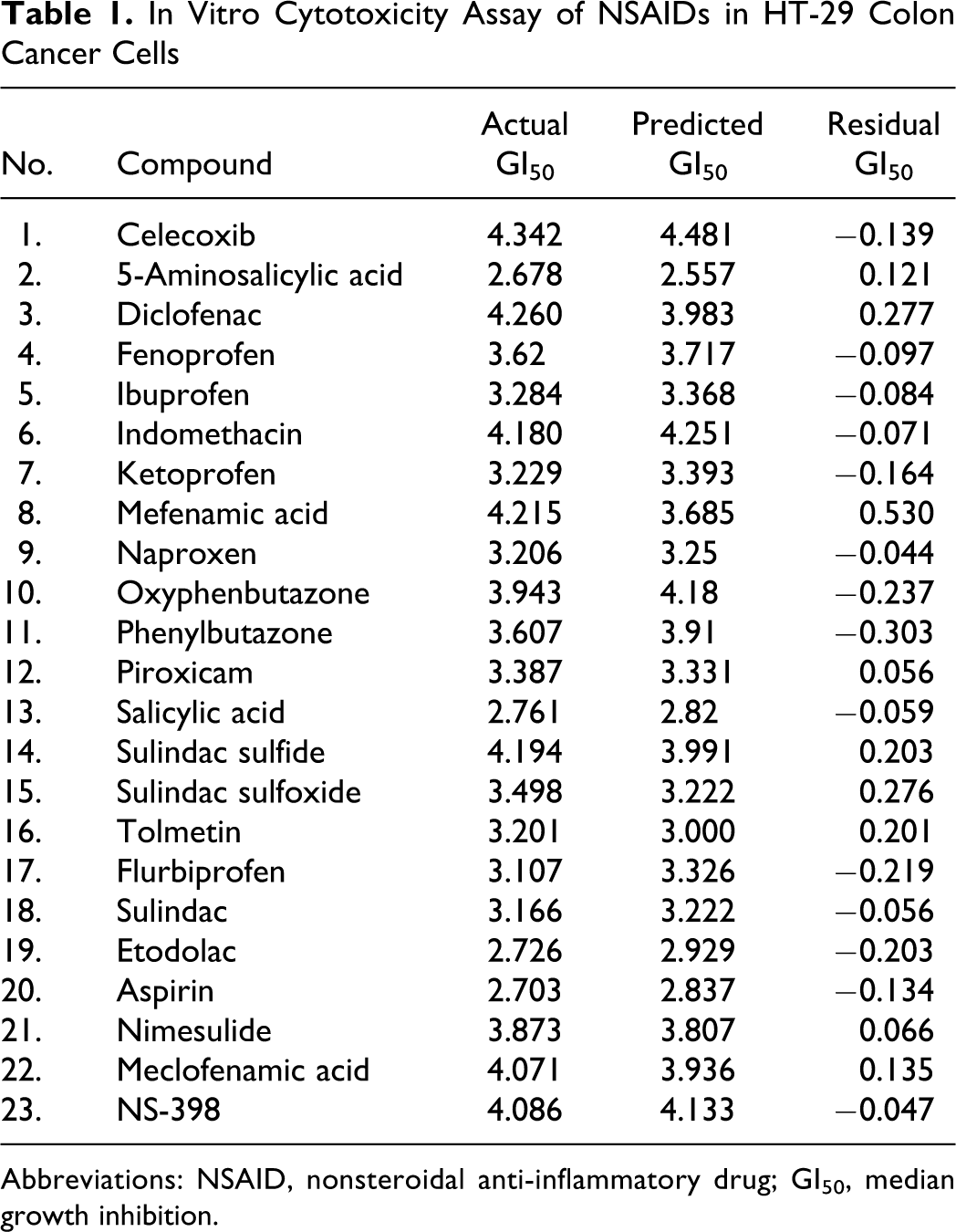
Abbreviations: NSAID, nonsteroidal anti-inflammatory drug; GI50, median growth inhibition.
In Vitro Cytotoxic Activity
In order to correlate the data linear with the free energy change, 36 the negative logarithm of the quantified GI50 M (molar) against HT-29 colon cancer cell line as pGI50 (log 1/GI50 or actual activity) was used as the dependent variable. Variation in structures and in vitro cytotoxic activity of drug molecules qualified are included in the present QSAR study. The structure of the drug molecules have been presented in Figure 1.
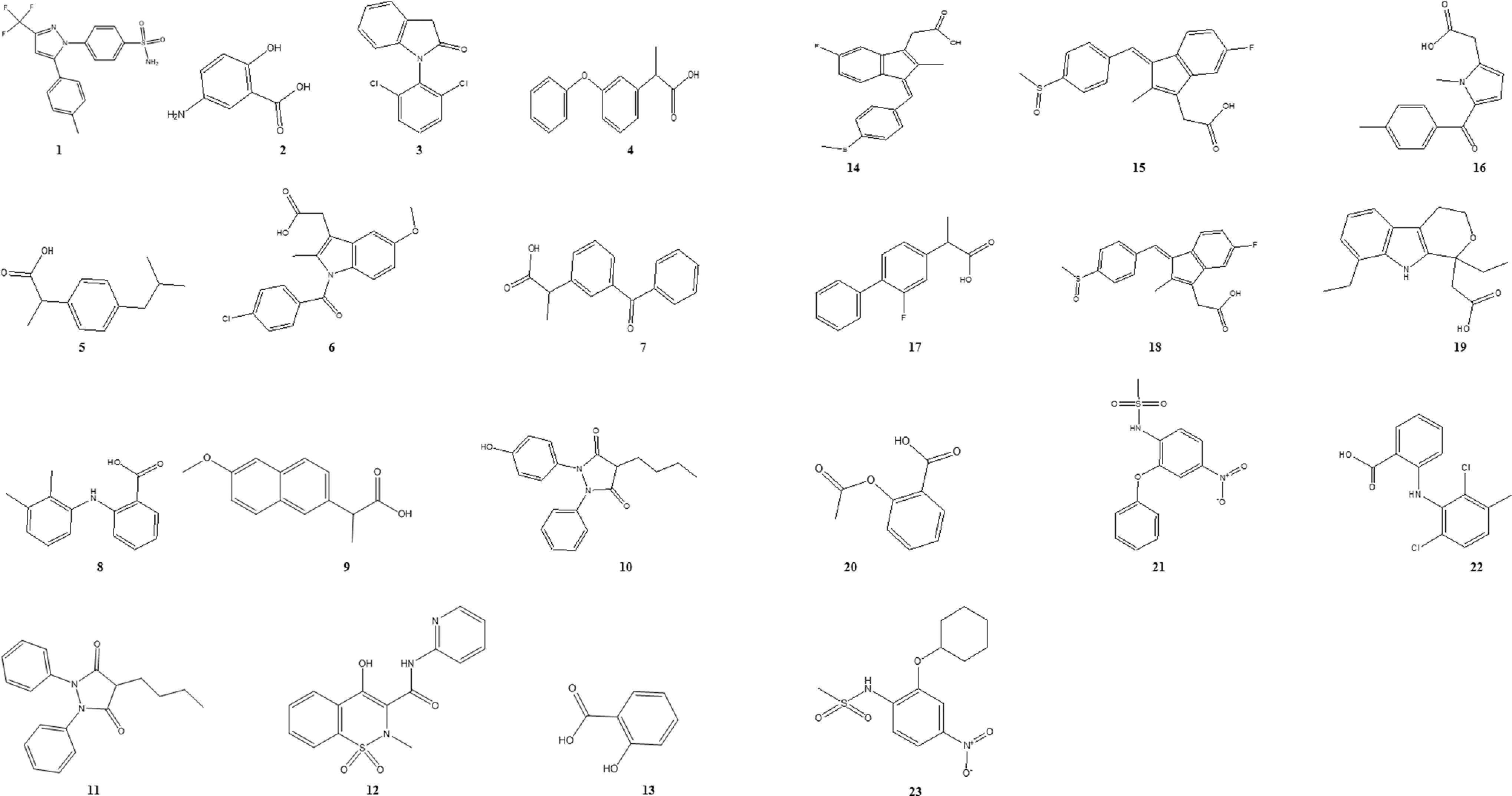
Structures of diverse set of NSAID molecules for SOMFA analysis.
Molecular Modeling
Using Chemdraw Ultra 8.0 running on Intel core (TM)2 Duo CPU T7500 @ 2.2 GHz with a 32-bit operating system/Windows7 Ultimate platform, all the 3D structures of the NSAIDs were constructed and were subjected to energy minimization using molecular mechanics (MM2). The minimization is continued until the root mean square (RMS) gradient value reaches a value smaller than 0.001 kcal/mol Å. The Hamiltonian approximations Austin model 1 (AM1) method 34 available in the MOPAC module 35 of Chem3D is adopted for reoptimization until the RMS gradient attains a value smaller than 0.001 kcal/mol Å. Further, all parameters were kept default unless stated. The physicochemical properties such as molar refractivity (MR), log P (partition coefficient), and total dipole moment were calculated using the “Analyze” option of the Chem3D 37 in order to obtain lowest energy structure of the compounds in the series.
SOMFA 3D-QSAR Models
Resolutions of 0.5 Å and 1 Å were spawned around the aligned molecules 38 from a 40 × 40 × 40 Å grid originating at (−20, −20, −20). The best model obtained using 0.5 Å grid resolution under exploration has been depicted in Table 2. The partial least square (PLS) algorithm was used in conjugation with leave one out (LOO) cross-validation to develop the final model. The following statistical parameters were considered to compare the robustness of generated QSAR models: correlation coefficient r, r 2, percentage contribution of each component (C), F test, and internal predictive power by cross-validated coefficient (r 2 cv).
Models Generated by Multilinear Regression Analysis and Statistical Measures Indicating Significance of Models With LOO Cross-Validation and Randomized Biological Activities
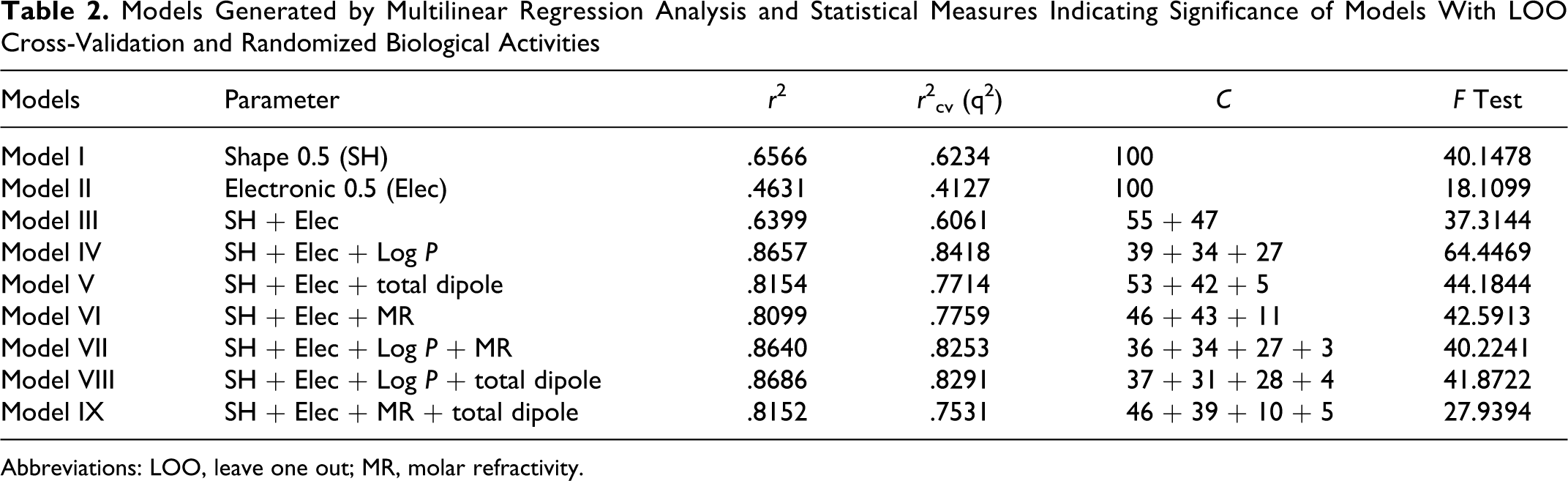
Abbreviations: LOO, leave one out; MR, molar refractivity.
The overall objective of a QSAR procedure is to derive a model that is optimally predictive. A model that does a good job of predicting the activities of molecules on which it is based must be tested to see whether any of the the test data affect the model excessively. This is done using the QSAR validation procedure. It is useful to assess the reliability and significance of QSAR models. Validation can be done by using cross-validation test, the bootstrap test, and by randomization test. The cross-validation process repeats regression many times on subsets of data. Usually each molecule is left out in turn, and the r
2 value is computed using the predicted values of the missing molecules (the cross-validated r
2). If molecules are removed N at a time from a total set of M, then N*M regressions are performed. In LOO, 1 compound was dropped in turn and a model was generated from the remaining molecules. This model was then used to predict the activity of the dropped compound and the whole method was reiterated until all the molecules were predicted. This PLS analysis offered the optimum number of components that was used to generate the final models without cross-validation. The result from a cross-validation analysis was expressed as r
2
cv (q
2
) value, which is defined as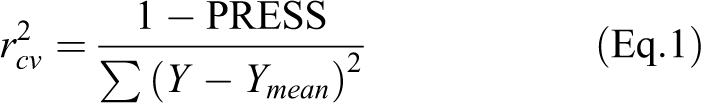
Where, PRESS = ∑Y − Y pred. The acronyms Y, Y pred, and Y mean are actual, predicted, and mean values. PRESS is an acronym for prediction sum of squares which is used to validate a regression model with regard to predictability. The PRESS value above can be used to compute an r 2 cv statistic, called r 2 cross-validated, which reflects the prediction ability of the model without selecting another sample or splitting the data. The result from a cross-validation analysis that is r 2 cv value, which can take up values in the range from 1, suggesting a perfect model, to less than 0 where errors of prediction are greater than the error from assigning each compound mean activity of the model. 39 Fischer statistics (F test) is the ratio between explained and unexplained variance for a given number of degrees of freedom. The larger is the value of F, the greater the probability that the QSAR models will be statistically signifcant. 40 Since the final equations were not very useful to represent efficiently the SOMFA models, 3D master grid maps of the best models were displayed by Grid-Visualizer program.
Results
The correlation between calculated descriptors as independent variable and in vitro cytotoxicity as response variable was calculated using PLS analysis. In the present QSAR study, 3D-SOMFA model was used to assess the data set composed of clinically used 23 NSAIDs whose experimental GI50 were reported in the literature, in order to predict the optimal molecular and steric features responsible for in vitro cytotoxic activity and antiproliferative activity. Statistical results of SOMFA models obtained by PLS analysis that is cross-validated r 2 cv, noncross-validated r 2, F test value served as quantitative measure of the predictability of the SOMFA. During SOMFA studies, grid spacing of 1.0 Å and 0.5 Å was investigated for both GI50 calculations. The different models generated for better correlation by statistical analysis have been given in Table 2. The best model given in equation (model VIII) exhibits good internal predictivity as established by the cross-validation r 2 cv value (.8291) of the model and also good external predictivity indicated by r 2 (.8686). The plot of observed versus predicted activities and residual values have been given in Figures 2 and 3, respectively. The descriptors in the best model indicated the contributions of steric, electronic, log P, total dipole moment, and MR on the in vitro cytotoxic activity (Table 2).
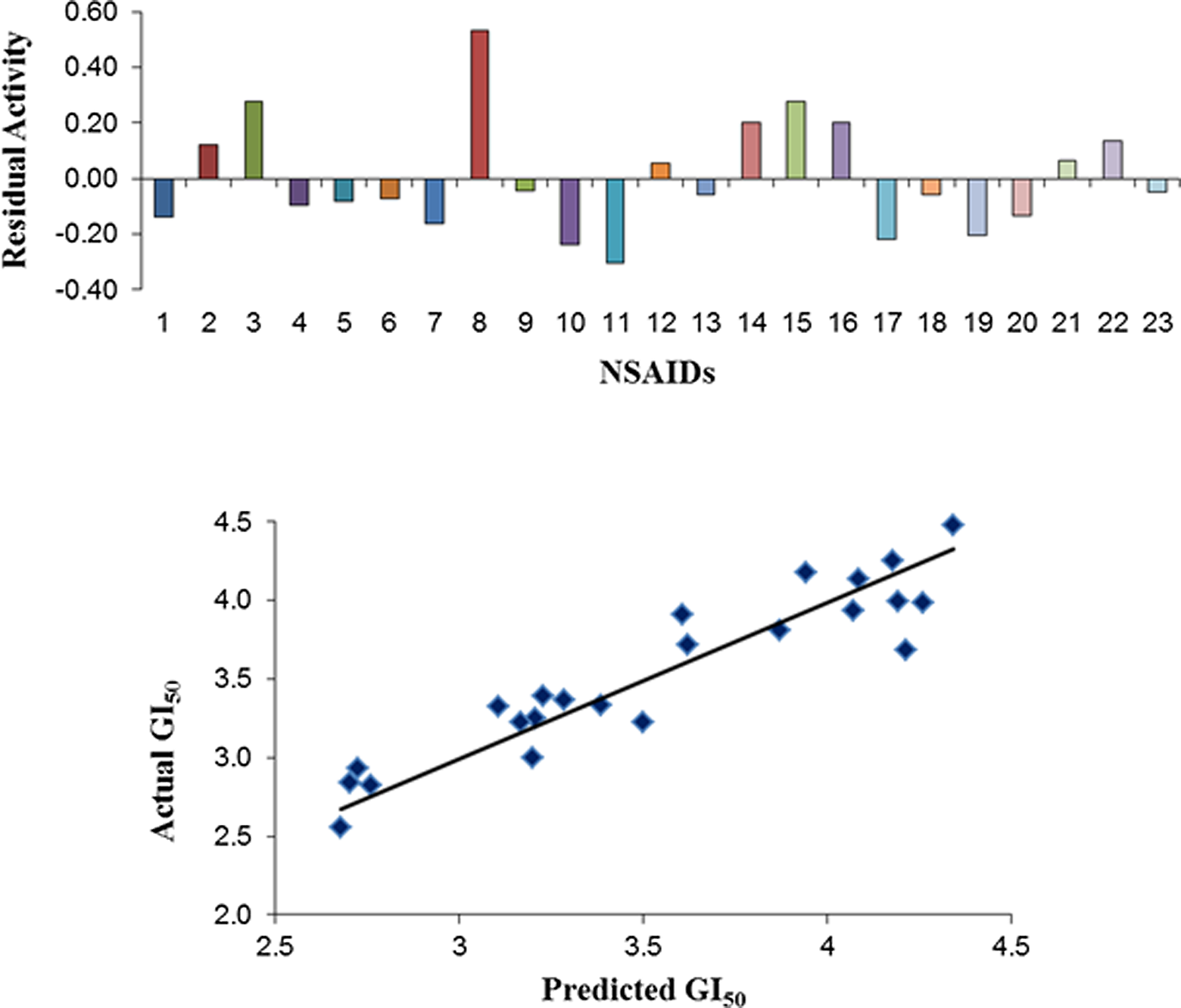
Comparative residual and predictive activities (GI50) using the best SOMFA model representation of NSAIDs. NSAID indicates nonsteroidal anti-inflammatory drug; SOMFA, self-organizing molecular field analysis; GI50, median growth inhibition.
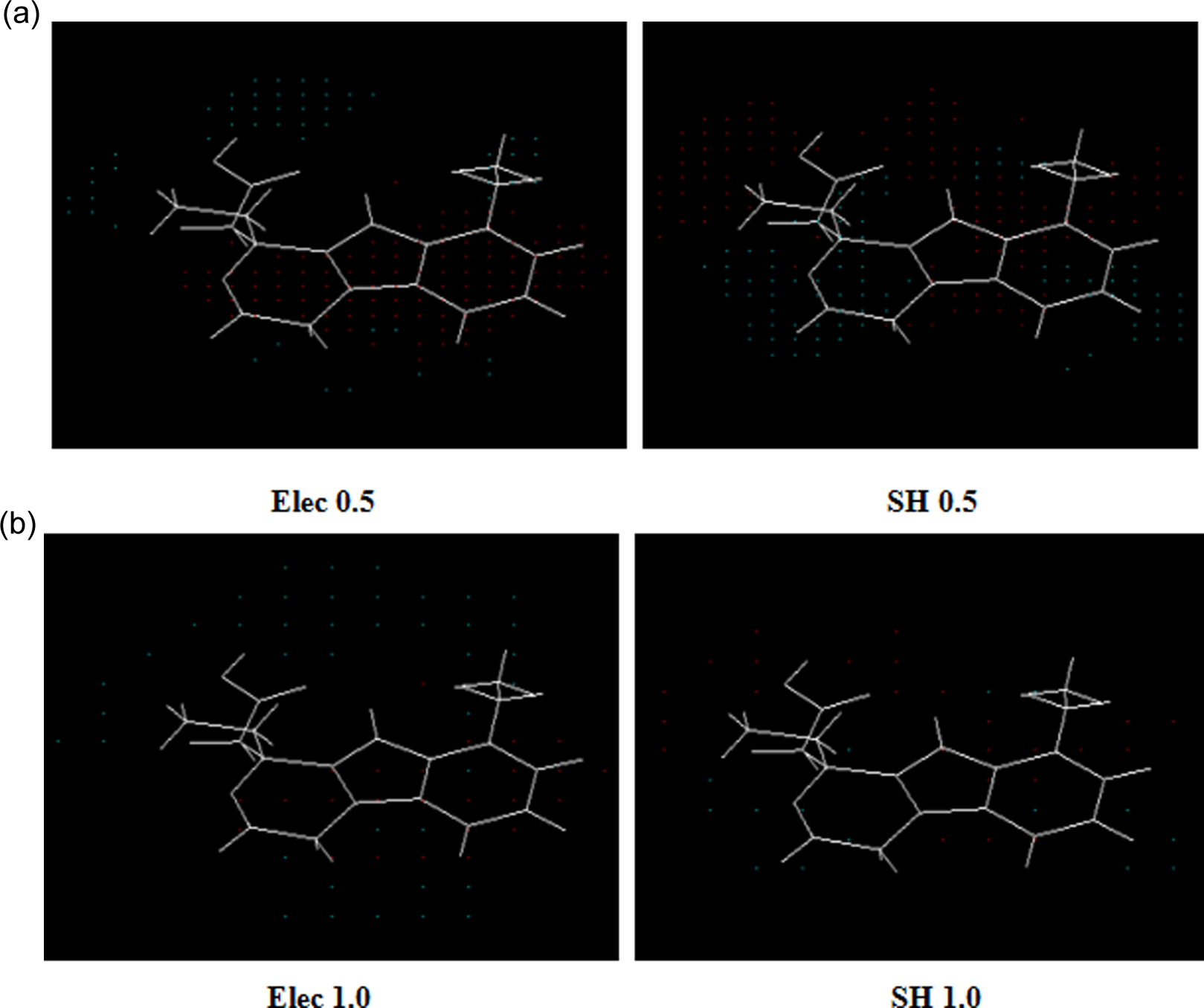
The SOMFA-derived shape grids illustrating most active compound having electronic arrangement for the GI50 (red and blue indicate regions with more steric bulk or less steric bulk, respectively) at different resolutions: (A) 0.5 Å and (B) 1 Å. SOMFA indicates self-organizing molecular field analysis; GI50, median growth inhibition.
Discussion
SOMFA shape and electrostatic potential for the analysis have been presented as master grid (Figure 3) using 2 different resolutions of grid that is 1.0 Å and 0.5 Å. The actual and predicted activities of the molecules are reported in Table 1 using the best model VIII. Figure 2 shows a good linear correlation and moderate difference between actual and predicted values of molecules. The master grid maps derived were used to display the contribution of shape and electrostatic potential. The master grid maps gave a direct visual indication regarding the structural features responsible for differentiating the activities of the molecules in the study. The master grid also offered an interpretation as how to design and optimize novel molecules with much higher activities. Each master grid map was colored in 2 different colors for favorable and unfavorable effects. In other words, the electrostatic features were red (more positive charge increases activity or more negative charge decreases activity) and blue (more negative charge increases activity or more positive charge decreases activity), and the shape features are red (more steric bulk increases activity) and blue (more steric bulk decreases activity), respectively.
The SOMFA shape potential map showed some important features that mainly included a cluster of high-density blue points around pyranoindole ring indicative of unfavorable steric interaction, while the red points around carboxylic group of pyran ring and the alkyl moiety suggested a steric bulky substitution may be expected in this region to enhance inhibitory activities. Meanwhile, in electrostatic potential map, a high density of blue points around carboxylic moiety indicated the presence of electronegative groups for favorable inhibitory activities, while the presence of red points in the vicinity of pyranoindole ring indicated electropositive substituents are favorable for optimal activity. A combination of both steric and electrostatic potential map gives 3D insight into structural features of NSAIDs. In order to get further insight we have considered the effect of other parameters on the 3D features (steric and electrostatic). It is evident from the Table 2 that log P, total dipole moment, and MR contributed positively although to different extents indicated by models I to IX.
The contribution of shape and electrostatic resulted in moderate r 2 in models I to III. Introduction of log P into the conventional models resulted in significant improvement in model quality indicated by r 2 (.8657) for model IV. However, with the incorporation of total dipole moment and MR also resulted in improvement of r 2 but less as compared to the model IV. Addition of total dipole with log P resulted in model VIII with highest r 2 (.8686).
In conclusion, statistically validated robust 3D-QSAR models were developed for a diverse group of NSAIDs having antitumor activity against HT-29 colon cancer cell lines. The results obtained for the present series of NSAIDs revealed good correlation as evident from the best model with in vitro cytotoxicity. The prediction potential of the QSAR model was tested by LOO method, which provides a good internal predictivity with r 2 cv (.8291). The results of the QSAR study showed characteristic influence of various physicochemical parameters, which mainly include steric, electronic components, MR, total dipole moment, and log P of NSAIDs against HT-29 colon cancer cells. The correlations of 2-D parameters with 3-D parameters provide better understanding of the desired structural features required to optimize antitumor activity. This can further be exploited to design newer NSAID molecules with optimal in vitro cytotoxicity for better therapeutic and prophylactic management of colon cancer.
Footnotes
Acknowledgments
The authors gratefully acknowledge UGC (University Grants Commission) New Delhi, for UGC-RSFMS (Research Fellowship Scheme for Meritorious Students) fellowship (Scheme No. F.4-1/2006 (BSR)/5-89/2007 (BSR)) and Dr Daniel Robinson (Oxford University) for SOMFA software.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
