Abstract
To develop new technology for detecting methicillin-resistant Staphylococcus aureus (MRSA), a novel fluorescent biosensor based on Klenow fragment (KF)–assisted target recycling amplification and synchronous fluorescence analysis was created. Carboxy-fluorescein (FAM)–labeled single-stranded DNA (ssDNA) containing a capture probe and a signal probe was adsorbed onto the surface of graphene oxide (GO) via π-stacking interactions, resulting in the fluorescence quenching of the dye. When target and primer were introduced, the fluorescence was restored due to P0 being completely released from the surface of the GO. Meanwhile, by using the KF and exploiting the synergistic effect of FAM and the double-stranded DNA (dsDNA)–SYBR Green I duplex structure, the fluorescence in this detection system was considerably amplified and the sensitivity was improved. The proposed strategy for mecA gene analysis showed a good linear range from 1 to 40 nmol/L, with a lower limit of detection of 0.5 nmol/L. In addition, a bacterial sample harboring the mecA gene was also detected, and its lower detection limit was up to 300 colony-forming units (CFU)/mL. Accordingly, this biosensor exhibits high sensitivity and selectivity and has great potential for early clinical diagnosis and treatment.
Introduction
Methicillin-resistant Staphylococcus aureus (MRSA), a potentially life-threatening pathogen, is a major cause of skin infection, food poisoning, gastroenteritis, pneumonia, and other diseases, which have a great impact on the health of both humans and animals.1–3 Bacteria that are resistant to β-lactam antibiotics are easily killed by methicillin. However, MRSA, first discovered by Jevons in 1961, was confirmed to be resistant not only to methicillin but also to many other antibiotics, which has led to great difficulties in the clinical treatment.4–6 MRSA has spread globally since its discovery, making it one of the most important pathogens in hospital and community infections, especially in the groups with weakened immune systems.7–9 Penicillin-binding protein 2a (PBP 2a), encoded by the mecA gene, has low affinity to methicillin, which helps MRSA avoid the fatal effects of various antibiotics, including methicillin; this avoidance results in fever, diarrhea, emesis, and multiple organ dysfunction syndrome.10,11 Sequence alignment analyses have failed to detect the mecA gene in other species besides Staphylococcus; this gene also shows good correlation with the drug resistance of MRSA. Hence, it is effective as a biomarker as it can not only be detected but also used to study the mechanism of drug resistance.
The conventional approaches for detecting MRSA mainly focus on culturing methods, enzyme-linked immunoabsorbent assay (ELISA), and PCR. However, all of these methods have particular disadvantages. Culturing methods are sensitive and cost-effective, but time-consuming and laborious. They also require a long time to monitor the bacterial colonies with the naked eye.12,13 In contrast, ELISA is a rapid and sensitive method, but is expensive and has a tendency to produce false-positive results due to cross-reactions.14,15 PCR is also available and has the advantages of high sensitivity and high specificity, but it requires expensive equipment and a complex pretreatment process. 16 Alternatives to these options have emerged in recent years, such as the use of electrochemical sensors. They were found to be superior to the conventional approaches, but require complicated electrode modification. 17 There is thus an urgent need to develop an alternative method for detecting MRSA that is rapid and efficient.
Graphene oxide (GO), a two-dimensional layered carbon material, has attracted attention all over the world due to its unique physical, chemical, thermal, and mechanical properties.18–20 It has an ultrahigh specific surface area for loading a large number of molecules, including proteins, nucleic acids, and certain drugs, via π-stacking interactions or chemical coupling.21,22 In addition, GO can act as a superb energy acceptor in fluorescence resonance energy transfer (FRET) as a result of its broad absorption spectrum. 23 Moreover, GO can protect the molecules bound to its surface from degradation or enzymolysis in the biological environments. 24 These three advantages have led to GO being widely applied in chemical sensors and biosensors. Using GO-based sensing platforms, some biosensors have already been designed to detect ATP, thrombin, and other small heavy metal ions.25–27 However, few studies have focused on the development of such sensors for detecting drug-resistant strains of bacteria. In order to detect such strains, there is a need to construct platforms based on GO that are sensitive and selective.
This article presents a rapid, sensitive, and selective fluorescent biosensor for the identification of MRSA carrying the mecA gene based on the noncovalent assembly of a probe labeled with a carboxy-fluorescein (FAM) fluorophore and GO. This probe consists of regions I and II, representing a capture probe and a signal probe, respectively. Region I is a complementary sequence that can specifically bind to the target DNA. Region II is used to recognize the primer sequence that amplifies fluorescence signals by displacing the target DNA and forming an SYBR Green I–double-stranded DNA (dsDNA) structure. In addition, the maximum excitation wavelength and maximum emission wavelength of FAM and SYBR Green I are both very close to each other in this analytical system; in fact, the fluorescence peaks of these two dyes overlap completely. By using the synergistic effect of FAM and SYBR Green I, the sensitivity of this detection method is significantly improved. Thus, a rapid, sensitive, and selective biosensor was obtained for assaying MRSA, which functions in the detection of drug-resistant strains, clinical diagnosis, and treatment.
Materials and Methods
Materials
All DNA oligonucleotides used in this study were synthesized by Shanghai Sangon Biological Engineering Technology & Services Co., Ltd. (Shanghai, China) (
Methods
Preparation of FAM-labeled probe/GO complex
GO powder was dissolved in Milli-Q water and sonicated for 3 h in an ice bath before being used to obtain a homogeneous black-yellow solution. The probe and GO solution were incubated in Tris-HCl buffer (25 mmol/L Tris-HCl, 5 mmol/L MgCl2, pH 7.4) at room temperature before the target was added. In order to optimize the ratio of probe to GO, seven different concentrations of GO ranging from 0 to 0.24 mg/mL were incubated with 50 nmol/L probe. Fluorescence was recorded at 514 nm.
Sensitivity and selectivity assays
The probe was mixed with 20 μL of GO (1 mg/mL) solution in Tris-HCl buffer (25 mmol/L Tris-HCl, 5 mmol/L MgCl2, pH 7.4) for 15 min. Then, the primer (50 nmol/L) and target DNA with different concentrations ranging from 0 to 50 nmol/L were added to the probe/GO reactive solution for another 40 min at 37 °C. A total of 2 μL of KF (1 U/μL) and 0.5 μL of 2 mmol/L dNTPs was subsequently added and incubated for 10 min; then, the reaction was stopped by keeping it at 75 °C for 10 min. Next, 2 μL of working solution of SYBR Green I was injected into the resulting solution for 10 min. Fluorescence intensity was recorded at 514 nm with an excitation wavelength of 480 nm in a quartz cuvette. In addition, specificity was tested using the target DNA and three different types of mismatch DNA. The concentration of the target DNA was set at 10 nmol/L, while that of the three types of mismatch DNA was set at 20 nmol/L. The protocol was the same as that described above. All experiments were repeated three times.
Bacterial sample analysis
MRSA cultured in nutrient medium (1% tryptone, 0.27% beef extract, 0.5% NaCl, pH 7.2) for 24 h at 37 °C was used as the target bacterium. After the number of colony-forming units (CFU) per milliliter had been determined by using the plate calculation method, bacteria at different concentrations from 3 × 102 to 108 CFU/mL were then heated to 95 °C for 15 min, after which they were immediately placed in ice water for 10 min to obtain their corresponding single-stranded DNA (ssDNA). Then, the primer (50 nmol/L) and the released ssDNA were added to the probe/GO complex. After incubation at 37 °C for 40 min, 2 μL of KF (1 U/μL) and 0.5 μL of 2 mmol/L dNTPs were successively added for 10 min to produce a polymerization reaction, which was then terminated by heating at 75 °C for 10 min. Next, 2 μL of working solution of SYBR Green I was injected into the resulting solution, followed by 10 min of incubation. Fluorescence intensity was then measured in a quartz cuvette at 514 nm with an excitation wavelength of 480 nm. Furthermore, six different bacteria containing S. aureus, S. epidermidis, S. saprophyticus, S. typhimurium, E. coli, and S. dysenteriae were examined to demonstrate the specificity of this biosensor in the bacterial samples. All the bacteria were kept at a concentration of 106 CFU/mL. The detection protocol was the same as previously described. All experiments were repeated three times.
Results and Discussion
A GO-based sensing platform based on strand-displacement polymerization recycling and synchronous fluorescent signal amplification was designed for the detection of MRSA carrying the mecA gene. As shown in Figure 1 , in the absence of the target, the FAM-labeled probe is adsorbed onto the surface of GO via π-π stacking and the fluorescence is quenched by GO as a result of FRET. However, when the target DNA and primer are introduced, the probe completely dissociates from the GO due to the specific recognition between complementary nucleic acid sequences. Upon addition of KF and dNTPs, the polymerization reaction that replaces the target sequence for the next round occurs, creating a large amount of dsDNA. Subsequently, SYBR Green I is added and forms a DNA–SYBR Green I duplex structure that produces significant fluorescence. When target DNA is sensed by synchronous fluorescence analysis, the fluorescence peaks of FAM and SYBR Green I overlap completely, which substantially enhances fluorescence in this detection system. The synergistic effect of FAM and SYBR Green I on the fluorescence intensity will considerably improve the sensitivity of this method. Moreover, the method of using the probe/GO was confirmed to be rapid, inexpensive, and not involve complex operations, making it particularly suitable for use in developing countries.
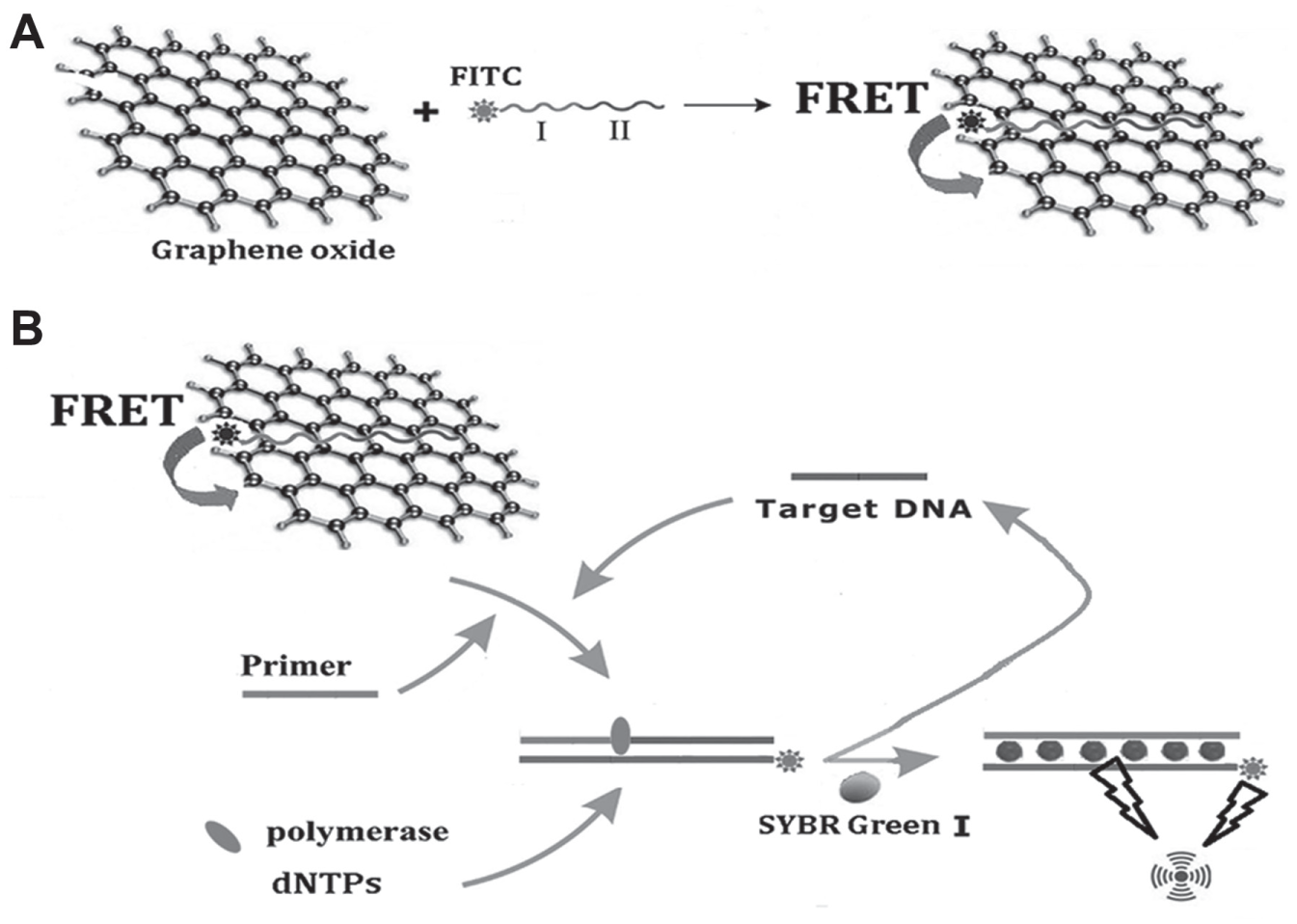
Schematic representation of the GO/P0 architecture platform for MRSA detection. (
To confirm the feasibility of using the proposed biosensor, fluorescence emission spectra of the probe/GO complex were studied under various conditions. As presented in
It has been reported that the optimal ratio of nanomaterial to biomolecule is a critical factor for the efficient detection of target. The presence of excess FAM-labeled probes or GO in a solution often produces false-positive signals or false-negative signals. Consequently, it is necessary to determine how many FAM-labeled probes could actually be loaded onto GO.
To confirm the ability of the proposed method to quantify a target, different concentrations of the mecA gene were analyzed. Figure 2 shows that the fluorescence was apparently restored by the addition of target DNA in the concentration range of 0.5–40 nmol/L, and the recovered fluorescence increased linearly in the range of 1–40 nmol/L (R2 = 0.9993). Moreover, the detection limit peaked at 0.5 nmol/L. In addition, the target DNA and mismatch DNA were used to investigate the selectivity of this approach under the same experimental conditions. As shown in Figure 3 , remarkable recovery of fluorescence was observed after the addition of the target DNA; nevertheless, the signals of the control groups were much lower than that of the experimental group even when the concentrations of the control groups were twice that of the experimental group. Furthermore, although the sequences differed by only one base between the target DNA and one mismatch DNA, the S/N of the former was 3.4 times higher than that of the latter. This clearly indicates that the proposed platform could detect the target DNA specifically without any cross-reaction or background interference. Hence, we considered that this novel fluorescence biosensor is promising for distinguishing the target from composite samples.
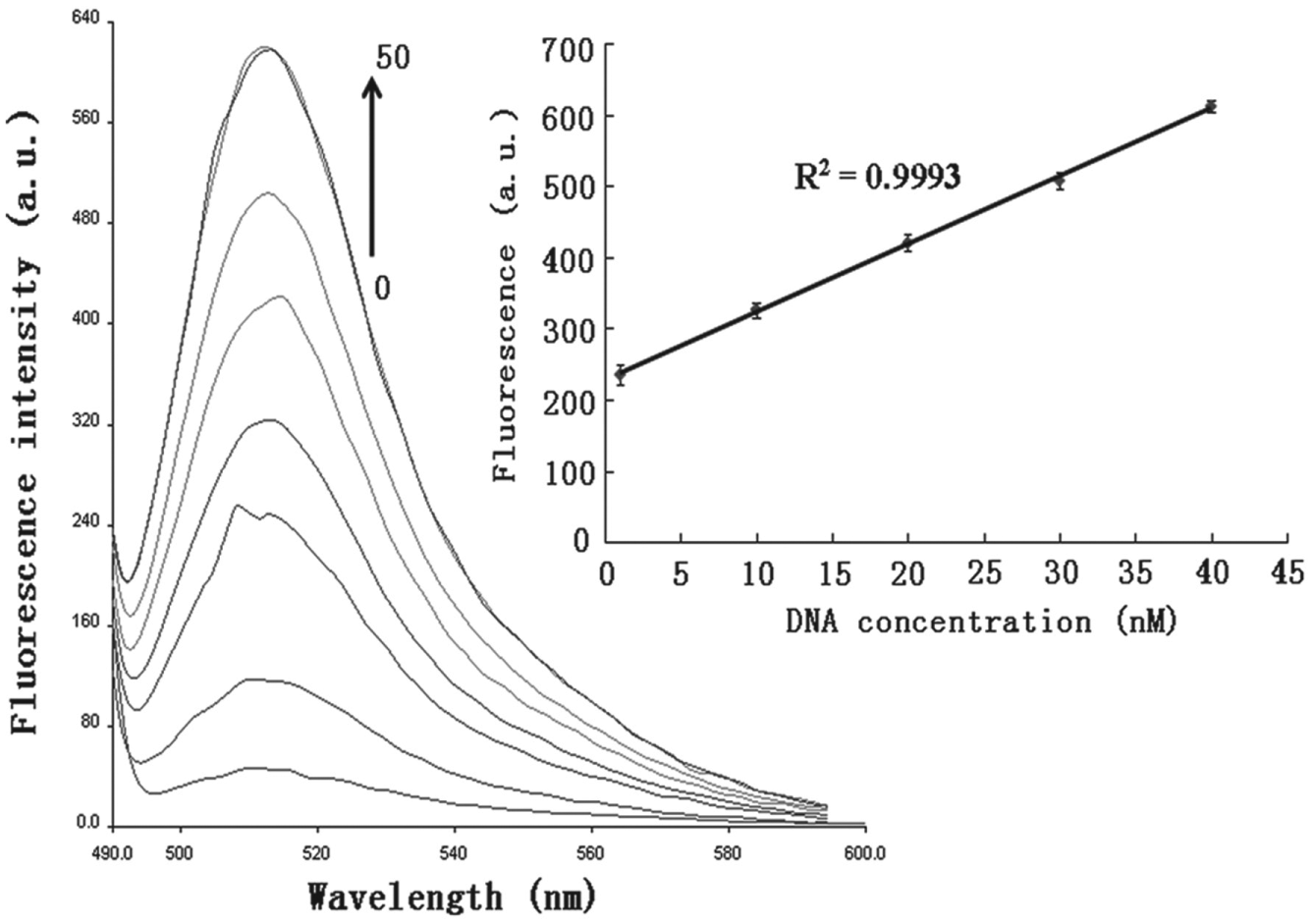
Detection of target DNA at different concentrations. The concentrations of DNA are varied from 0 to 50 nmol/L from the bottom to the top; fluorescence curve in the bottom represents the background signal of this system. (The inserted figure shows the increases in fluorescence are proportional to the concentration of the target DNA, ranging from 1 to 40 nmol/L.) Error bars SD, n = 3.
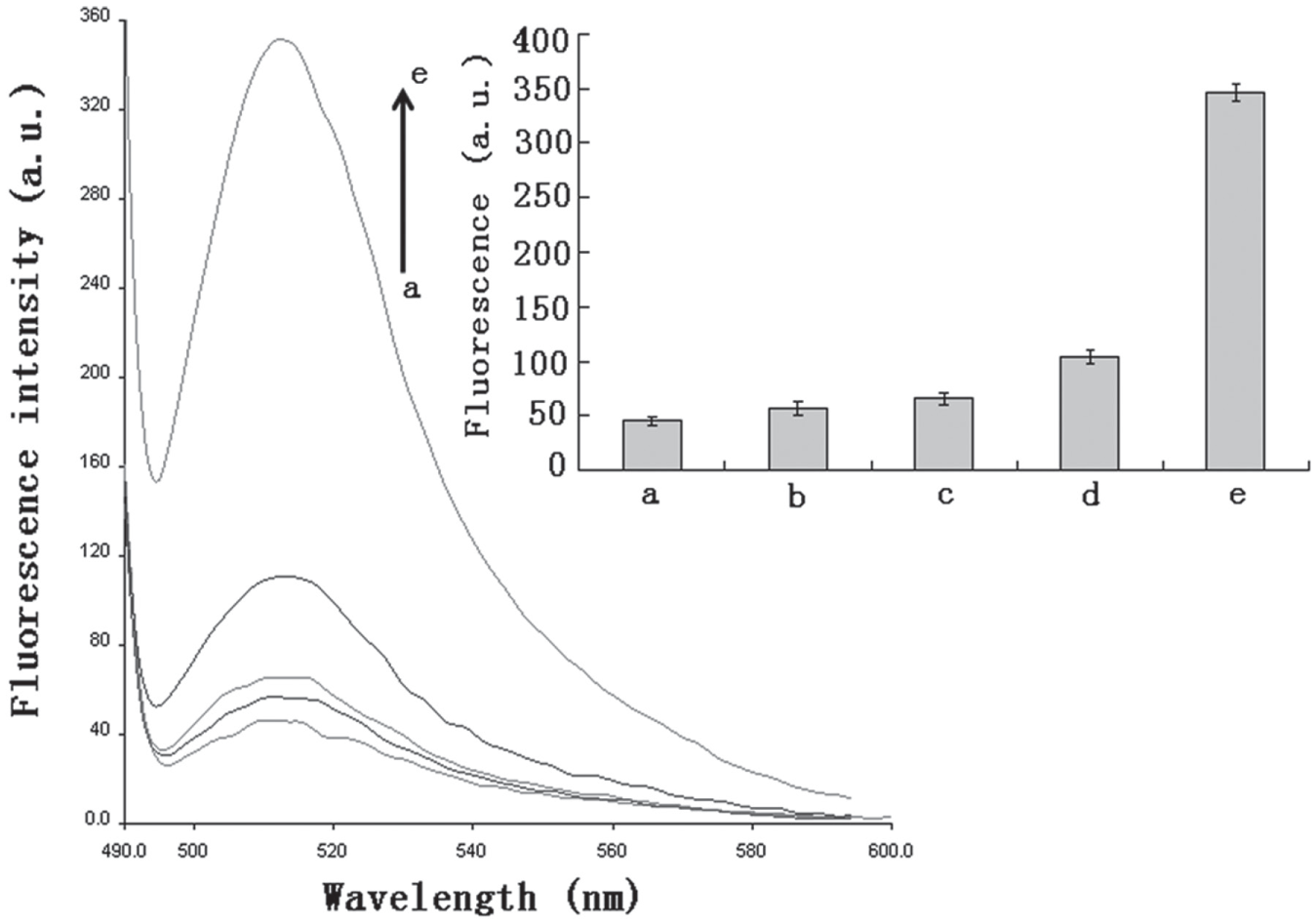
Fluorescence signals triggered by target DNA and mismatch DNA: (
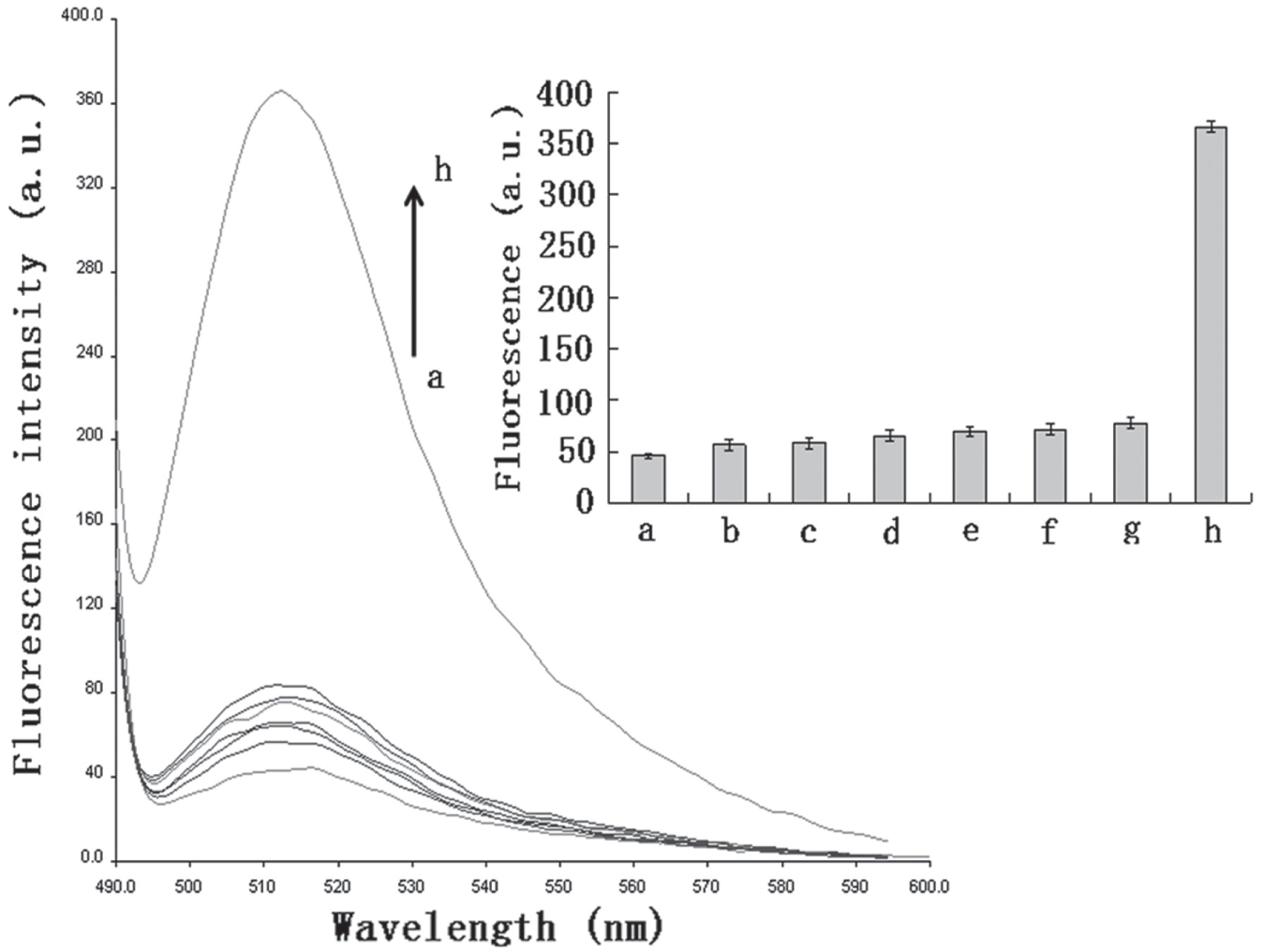
To detect MRSA carrying mecA gene by this method, the pretreated bacteria ranging from 3 × 102 to 108 CFU/mL were added to the FAM-labeled probe/GO complexes. As shown in Figure 4 , the background fluorescence signal was low before the DNA was injected due to the high efficiency of GO quenching; however, when the bacteria were added at increasing concentrations, the fluorescence gradually increased, which occurred linearly from 103 to 107 CFU/mL (R2 = 0.9954). Importantly, the lower detection limit reached 3 × 102 CFU/mL, which is superior to those of surface plasmon resonance (SPR), colorimetric, and fluorescence biosensors.28–30 The specificity of this designed platform was also estimated by surveying the fluorescence of target bacteria and six other bacteria: S. aureus, S. epidermidis, S. saprophyticus, S. typhimurium, E. coli, and S. dysenteriae. Figure 5 shows a comparison among the fluorescent intensities in response to MRSA and other nontarget bacteria. The results indicate that the enhancement of fluorescence intensity as soon as the MRSA DNA was added was much greater than that induced by other control bacteria. The results indicate that the other bacteria besides MRSA almost failed to generate a response to the biosensor. Hence, this method apparently has excellent specificity and sensitivity for MRSA detection, which would enable its wide application in the detection of pathogenic microorganisms, and the clinical diagnosis and treatment of their infections.
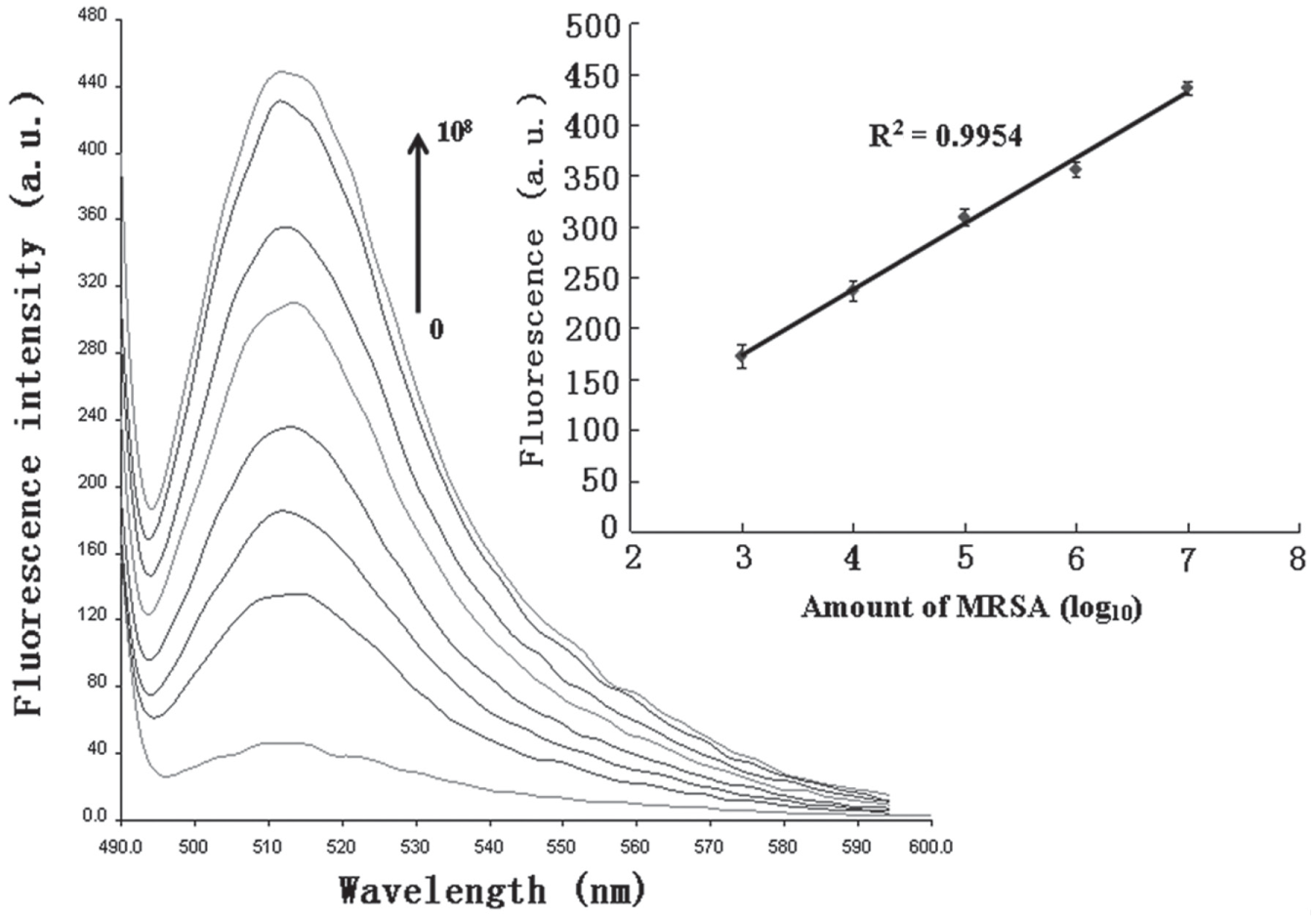
Sensitive detection of the target bacteria carrying mecA gene with different concentrations. The concentrations of target bacteria ranged from 0 to 108 CFU/mL from the bottom to the top; fluorescence curve in the bottom represents the background signal of this system. (The inserted figure shows the fluorescence enhancements are proportional to the concentration of the target bacteria, ranging from 103 to 107 CFU/mL.) Error bars SD, n = 3.
Specific detection of the target bacteria compared to other bacteria strains: (
In summary, we have successfully constructed an amplified GO-based fluorescence biosensor for detecting the mecA gene and MRSA carrying this gene. This sensor is based on strand-displacement polymerization recycling and synchronous fluorescent signal analysis. This detection system has been shown to be a simple, cost-effective, sensitive, and specific method due to its use of GO as an efficient fluorescence quencher. This novel biosensor can also detect the mecA gene with a linear range from 1 to 40 nmol/L and has a lower detection limit of 0.5 nmol/L. In addition, a much higher S/N of the target than that of the nontarget was observed when this system was used for mismatch DNA analysis, which implied that the biosensor has high specificity for the mecA gene. Importantly, prominent fluorescence signals were obtained when this system was used for assaying bacteria samples. The lower detection limit of bacteria was 3 × 102 CFU/mL, with a linear range from 103 to 107 CFU/mL. The platform also showed high specificity to MRSA, indicating that the FAM-labeled probe/GO architecture could specifically identify the mecA gene from target bacteria. Given the frequency of infection with MRSA both in hospitals and in the community, the system that we have established provides an efficient platform for MRSA detection, which offers a range of advantages over conventional methods. Its range of applications can also be extended to detecting proteins, cells, heavy ions, and some other targets, by substituting the probe for aptamers. Therefore, in future studies, we will focus on designing amplified GO-based fluorescence aptasensors for the identification of drug-resistant bacteria.
Footnotes
Acknowledgements
We would like to thank the National Natural Science Foundation of Hunan Province (2016JJ3098), Outstanding Youth Scientific Research Project funded by the Education Department of Hunan Province (15B169), National Natural Science Foundation (81202451), Science and Technology Innovation Team in Colleges and Universities in Hunan Province (Chinese traditional medicine for treatment of infectious diseases, no. 15, Grxjb-7), Doctor Start-Up Foundation of Hunan University of Chinese Medicine (9982-1001019), Key Subjects of Hunan University of Chinese Medicine (pathogenic biology, no. 1), and Project Funded by Hunan Provincial Level Course (immunology and pathogenic biology, no. 48) for their financial support
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: We would like to thank the National Natural Science Foundation of Hunan Province (2016JJ3098), Outstanding Youth Scientific Research Project funded by the Education Department of Hunan Province (15B169), National Natural Science Foundation (81202451), Science and Technology Innovation Team in Colleges and Universities in Hunan Province (Chinese traditional medicine for treatment of infectious diseases, no. 15, Grxjb-7), Doctor Start-Up Foundation of Hunan University of Chinese Medicine (9982-1001019), Key Subjects of Hunan University of Chinese Medicine (pathogenic biology, no. 1), and Project Funded by Hunan Provincial Level Course (immunology and pathogenic biology, no. 48) for their financial support.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
