Abstract
Oral agents targeting Janus-associated kinases (JAKs) are promising new agents in clinical development. To better understand the relationship between JAK inhibition and biological outcome, compounds targeting JAKs were evaluated in peripheral human whole blood. To date, these analyses are low throughput and costly. Here, we developed a robust 384-well, high-throughput flow-based assay approach to screen small molecules for JAK/STAT signaling inhibition in human whole blood. This assay platform provides a highly sensitive analysis of signaling events in blood and facilitates measurement of target engagement. Further, the automation technologies and process optimizations developed here overcame sample integrity, handling, and multiparametric data analysis bottlenecks without affecting assay performance. Together these efforts dramatically increased sample throughput compared to conventional manual flow cytometric approaches and enabled development of novel JAK/STAT inhibitors.
Introduction
Human immunology is a complex network of intra- and extracellular interactions involving cytokines, chemokines, cell surface receptors, intracellular signaling molecules, and transcription factors in multiple cell types. These aforementioned cellular components are often altered in response to disease onset and therapy. In immune-based drug discovery, multiple high-throughput assay approaches are utilized to interrogate complex inflammatory signaling networks. Typically, first-tier screening assays include reporter systems in immortalized cell lines which are inexpensive, high in throughput, easy to use, reliable, and reproducible. However, these model systems, albeit amenable for high-throughput screening (HTS), may not always truly represent a natural biological setting and thus may fail to provide drug in vivo connectivity. In contrast, cells contained in human primary whole blood provide a physiologically relevant setting as well as facilitate assessment of synthetic small-molecule protein binding characteristics and the amount of free drug available for action on its target. Yet, assays typically utilized in whole blood analyses, such as enzyme-linked immunosorbent assays (ELISAs), are limited in throughput, multiplexing capabilities, and sample size requirements.
For several decades, flow cytometry has been the gold standard investigative tool for the identification of responsive immune cell subpopulations in the context of multiple disease states. It is powerful, providing simultaneous in-depth multiparameter analyses of individual cells or particles found in heterogeneous sample populations. Flow cytometry was initially applied characterizing cell surface proteins using selective antibodies, and now this technology has evolved to where the phosphorylation status of intracellular protein targets in individual cells can be monitored. Development of high-throughput phosphorylation-based flow cytometry was first reported in 2002 using phosphorylation epitope-specific antibodies, 1 and it made possible the measurement of cellular signaling protein phosphorylation status and its association with functional activation. Consequently, simultaneous measurement of phospho-protein signaling in single or multiple pathways using this technology is ideal for assessing the therapeutic modulation of activated cell surface receptors or their associated intracellular signal transduction cascades in heterogeneous cellular populations.
Historically for sequential analyses of discrete cell suspensions, flow cytometry was limited by its maximum processing speed of six samples per minute even in the most sophisticated platforms. 2 However, the emergence of high-throughput flow cytometry (HTFC) and advances in digital signal processing and detection capabilities extended cell-based screening capacities to processing samples contained in 384-well plates. Even though these enhancements enabled measurement of more than 10,000 particles per second, 3 they cannot alone facilitate the rigors of large-scale automated screening. From this perspective, incorporating HTS flow cytometry into streamlined, robotic processes requires measuring representative populations contained in a minimum number of collected events per well, and integrating automated whole blood sampling, handling, and multiparameter data analyses (gate parameters and population identification). Accordingly, these challenges are largely the reason for the absence of flow cytometry–based whole blood screening efforts in drug discovery.
Here, we established a first-in-class, multiparameter, 384-well phospho-protein signaling platform to evaluate treatment effects of the JAK family of nonreceptor tyrosine kinase inhibitors on ex vivo stimulated human cells contained in peripheral whole blood samples. In addition to better understanding the relationship between cellular JAK inhibition and activity in translational models, we optimized automation of several assay parameters to significantly reduce the whole blood sample size and enhance data capture. Collectively these efforts led to a sensitive translational assay platform reducing sample processing time and increasing compound testing capacity. Moreover, utilization of this platform allows for target identification, pharmacodynamic validation, lead assessment and optimization, and in-depth studies for compound mechanisms of action in primary cells.
Materials and Methods
Healthy Donor Peripheral Whole Blood Sample Acquisition
All patient-related procedures and protocols were performed in accordance with Bristol-Myers Squibb (BMS) Occupational Safety and Health Administration protocols. Whole blood samples were collected from healthy donors by venipuncture and stored in appropriate vacutainer tubes (Becton-Dickinson [BD], Franklin Lakes, NJ) at ambient temperature 1 h (or less) prior to experiment initiation.
Compounds
All compounds were synthesized at BMS and stock solutions (10 mM) were prepared using DMSO. Compounds were delivered manually to individual wells of 96-well plates (5 µL per well) or distributed to separate wells of 384-well plates using ECHO acoustic liquid handling technology (10 nL per well). Compound inhibitory activities were determined in biochemical kinase assays using human recombinant JAK enzymes as described. 4
Peripheral Whole Blood Assays
Compound cellular activity was monitored in human whole blood ex vivo using flow cytometry and cytokine stimulation. Briefly, blood samples were added to individual wells of 96-well (100 µL per well) or 384-well (10 µL per well) plates containing varying concentrations of compound and incubated for 1 h (37 °C) prior to interleukin 15 (IL15) cytokine treatment (Peprotech, Rocky Hill, NJ; 25 ng/mL). After 15 min, cellular stimulation was terminated by the addition of a lysis and fixation buffer (BD Phosflow Lyse/Fix buffer; following BD Biosciences protocols) to remove red blood cells and fix the remaining cells for cellular staining. Optimized concentrations of surface antibodies (anti-CD3 FITC [BD 555916], anti-CD8 PE [BD 555635]) were added to the cell suspensions and incubated for 1 h (37 °C) before being removed by two washing steps using phosphate-buffered saline (PBS) containing 0.1% bovine serum albumin (50 µL per wash per sample). Cells were subsequently permeabilized by the addition of a permeabilization buffer (BD Perm III buffer; following manufacturer protocols) and stained for 1 h using a phospho-STAT antibody (pY694, Alexa647-conjugated, BD 612599). Samples were analyzed using either a FACS Canto II or Intellicyte iQue flow cytometer (Albuquerque, NM).
Flow Cytometry and Data Analysis
96-well assay
Cellular data were exported from the BD Canto II instrument and analyzed using FlowJo analysis software (version 9.3; Ashland, OR). The amount of pSTAT5 expression in the CD3/CD8 positive population was measured by median fluorescence intensity (MFI). Percent induction was calculated comparing values between IL15-stimulated and unstimulated groups. The percent inhibition equation was applied as follows: [1 − (sample value − average vehicle only control)/(average IL15 stimulation only control − average vehicle only control)] × 100. Compound concentration–response curves were determined utilizing a BMS four-parameter logistic nonlinear regression model. Inhibitory concentration (IC50) determinations were calculated based on 50% inhibition on the fold induction between cytokine and unstimulated treatments. In all instances, dead cells and doublets were excluded prior to any data analyses. The Z′ value as a criterion for data quality was calculated according to Zhang et al. 5 All statistical analyses utilized GraphPad software tool sets (San Diego, CA).
High-throughput flow cytometry (384-well format)
Samples were suspended in wash buffer containing 8% pluronic acid, a nonionic surfactant (Gibco, Waltham, MA), upon completion of the intracellular staining step (70 µL per sample); samples are stable for up to 24 h following this step. The Intellicyt iQue flow cytometry platform was employed to evaluate samples from individual wells containing agitated cells (5–7 s acquisition time per well), and the probe was cleaned using PBS after every 12 well intervals. Data were examined using Forecyt analysis software (version 4.1; Albuquerque, NM). Levels of pSTAT5 expression in CD3/CD8 positive cells were measured by MFI. Percent induction, IC50 values, and concentration–response curves were determined as described above.
Results and Discussion
Successful drug development requires large-scale compound screening platforms allowing for lead molecule identification and optimization in physiologically relevant cells. In the present work, we established a high-throughput, 384-well flow cytometry workflow for detecting STAT5 phosphorylation in human leukocyte subsets from whole blood. Initially our efforts focused on optimizing steps (cytokine treatment, red blood cell lysis, cell fixation/permeabilization, multifluorochrome staining, compound treatment) of a 96-well assay using a BD FACS Canto II instrument.6,7 Hence, IL15 treatment conditions were adjusted to facilitate JAK kinase activation and tyrosine phosphorylation of STAT transcription factors2,8 in CD3/CD8 positive lymphocytes (average = 12%–15%;
Fig. 1A
). These treatments induced p-STAT5 (phospho-STAT5 protein) activity greater than 10-fold in these T cells as measured by the median fluorescent intensity (
Fig. 1B
). Next, a reference compound was used to validate the assay and determine its sensitivity. In these studies, whole blood samples (100 µL) were treated with the reference compound (3–1000 nM) for 1 h prior to IL15 stimulation (15 min) and p-STAT5 activity assessment. Results showed reference compound treatment reduced IL15-induced STAT activation in a dose-dependent manner (
Fig. 1C
,
D
) with IC50 values ranging from 85 to 100 nM across independent blood donors (
Fig. 1D
). Most importantly, these data correlate with those reported for a similar inhibitor.
9
Additionally, reference compound treatment did not attenuate the percentage of lymphocytes isolated from blood, nor did it affect isolation of CD3/CD8 positive T cells, and DMSO (0.3%) treatment did not affect baseline p-STAT signaling (
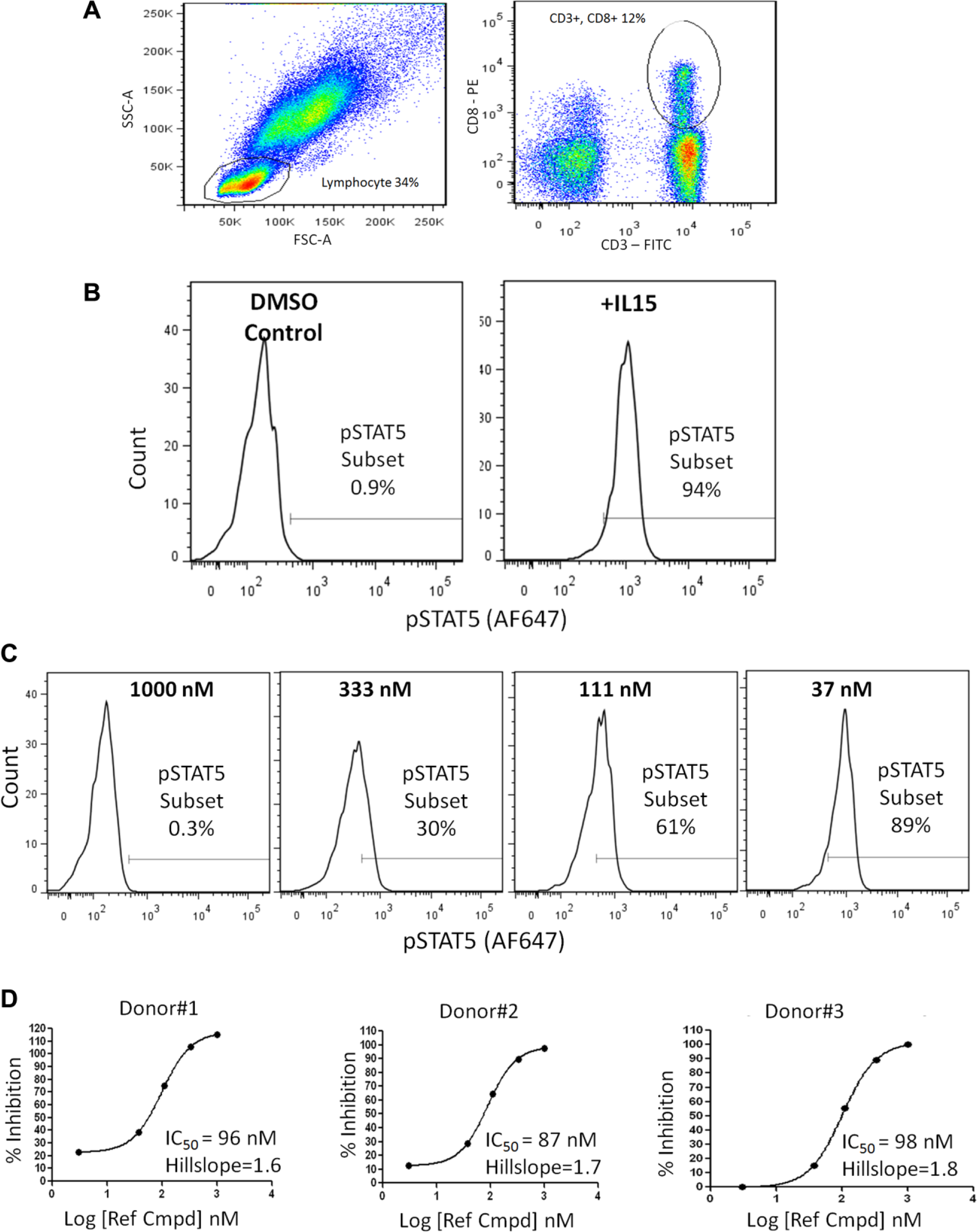
Ex vivo treatment of human whole blood with a reference compound (Ref Cmpd) inhibits STAT5 phosphorylation in CD3/CD8 positive cells as measured using a 96-well formatted flow cytometry assay. (
The initial 96-well screening approach exhibited capacity to discern potent, concentration-dependent inhibition of STAT phosphorylation (
Fig. 1
). Nonetheless, its application for large-scale screening and identifying structure–activity relationships (SARs) was limited by cell collection time requirements, being labor-intensive, and requiring large blood volume sample sizes (100 µL). This blood volume requirement, in turn, also necessitated increased quantities of IL15, test compounds, antibodies, and amounts of assay reagents for red blood cell lysis, fixation and permeabilization. Thus, further optimization took place to accommodate a 384-well format. Our first step was reducing the whole blood sample volume while maintaining representative extracellular (CD3/CD8 positive) and intracellular (phospho-STAT) profiles and retaining acceptable variation values (well-to-well or plate-to-plate). In view of this, the liquid handling parameters (such as monitoring the tips for air gaps, positioning the 384-well head, fine-tuning the z height, and tip touch) on the automated Hamilton Star liquid handling workstation were adjusted to avoid blood clotting and to ensure simultaneous, uniform, low-volume (10 µL) delivery of blood across all wells of each 384-well destination plate. We then focused our attention on optimizing delivery of test compounds to blood samples. In our initial studies, the compounds were manually added to individual wells containing the whole blood samples and each required an intermediate dilution step. The disadvantages of this workflow were that it required sizable compound quantities that may be in limited supply and each dilution step had the potential to introduce assay result variation due to the physical properties of each compound (e.g., solubility). Since now the volume of each sample was reduced 10-fold, we leveraged use of ECHO acoustic liquid handling technology that dispenses nanoliter quantities of compound (10 nL), eliminating the intermediate dilution step, and took advantage of the Formulatrix Tempest liquid handing technology to reduce antibody and IL15 ligand volumes to 200 nL. Together, these automated liquid handling procedures increased assay reproducibility and substantially reduced manual assay operation time. Additionally, miniaturizing this workflow resulted in a significant reduction of reagents, thereby reducing overall assay costs by more than 15-fold. Lastly, often tedious steps of flow-based assays are the sequential wash and centrifugation procedures after cell fixation, lysing, permeabilization, and antibody staining. These procedures, when run manually over large sets of plates, require significant processing time and contribute to data variability. To this end, we integrated use of a Biotek Elx405 384 well plate washer (Winooski, VT), Velocity 11 VSpin microplate centrifuge (Palo Alto, CA), plate stacker, Thermo Scientific Multidrop Combi (for high-volume wash buffer addition; Waltham, MA), and Liconic automated incubator (Woburn, MA). Consequently, these integrated devices increased consistency between the numerous required wash steps (
HTFC screening systems, such as those from IntelliCyt, facilitate implementation of flow cytometry assays for large-scale screening due to their capacity to analyze samples contained in small volumes (microliters) in a large number of plates and their capabilities to simultaneously assess different cellular parameters from a single well. Here we adjusted the HTFC Intellicyt iQue platform and its automated sampling system3,10 to accurately process samples directly from multiwell plates and permit investigation of numerous extracellular and intracellular markers. Figure 2A shows an example of a typical p-STAT5 analysis of CD3/CD8 positive cells over time. In these studies, HTFC effectively identified CD3/CD8 positive cell populations and provided the cellular activities of 32 independent compounds tested at 11 different concentrations in these subpopulations. Moreover, since the HTFC autosampler aspirates multiple cellular samples through a single tubing line directly into the instrument, data from an entire compound plate can be effectively acquired at rates of less than 60 min per 384-well plate ( Fig. 2A ) versus the 120 min per 96-well plate using the BD FACS Canto II instrument. However, after handling an extensive number of 384-well plates, it became apparent that not all wells were properly defined as a result of the instrument software improperly producing well identifiers from individual, time-resolved histograms ( Fig. 2B ). Additionally, incomplete red blood cell lysis, sample debris, and cellular aggregation contributed to noteworthy well-to-well variation and inaccurate MFI values ( Fig. 2B ). Overall, these factors affected HTFC data analytics and statistical significance calculations, so samples were subsequently processed in buffers supplemented with the nonionic surfactant, pluronic acid. This surfactant treatment facilitated efficient automated population gating and accurate well identifier assignments and decreased sample acquisition volumes from 50 µL to less than 12 μL ( Fig. 2B ). Now this assay is able to reproducibly identify populations of interest (after manually setting individual cell gates), evaluate the cellular activities of individual compounds, and automatically generate statistically fortified data sets for each population and parameter of interest across a large number of plates.
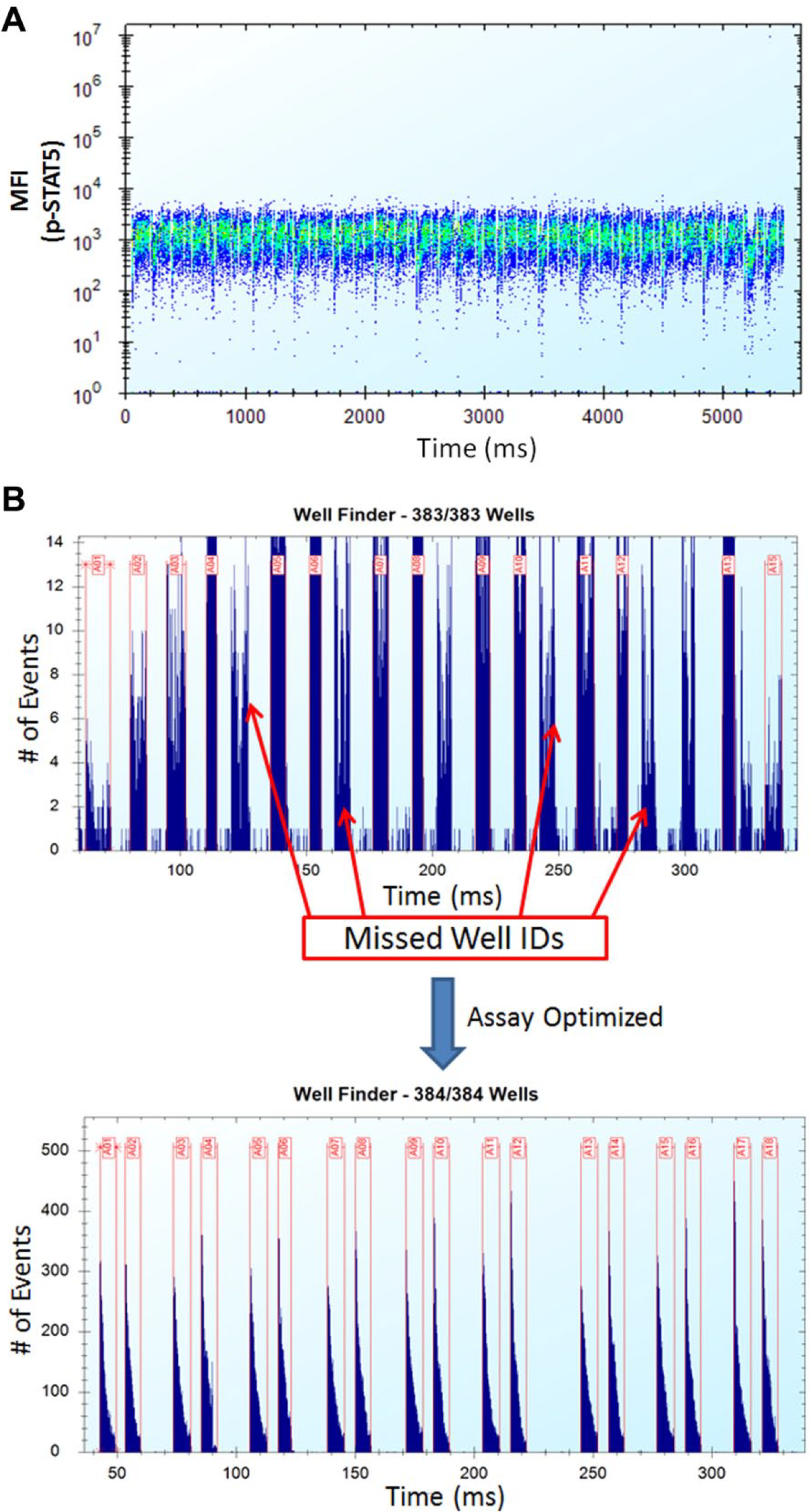
HTFC effectively identifies CD3/CD8 positive cells from whole blood samples and permits compound testing for cellular activity. (
The robust 96-well assay described here was hindered for large-scale screening by its manual cell gating requirement for all samples. A feature of the IntelliCyt iQue HTFC platform is the convenience of using its proprietary template-driven acquisition and analysis tools to automate gating, compensation, and statistical analyses for entire studies. In our hands, these tools supported programmed multiparameter gating and compensation across all 384 samples (
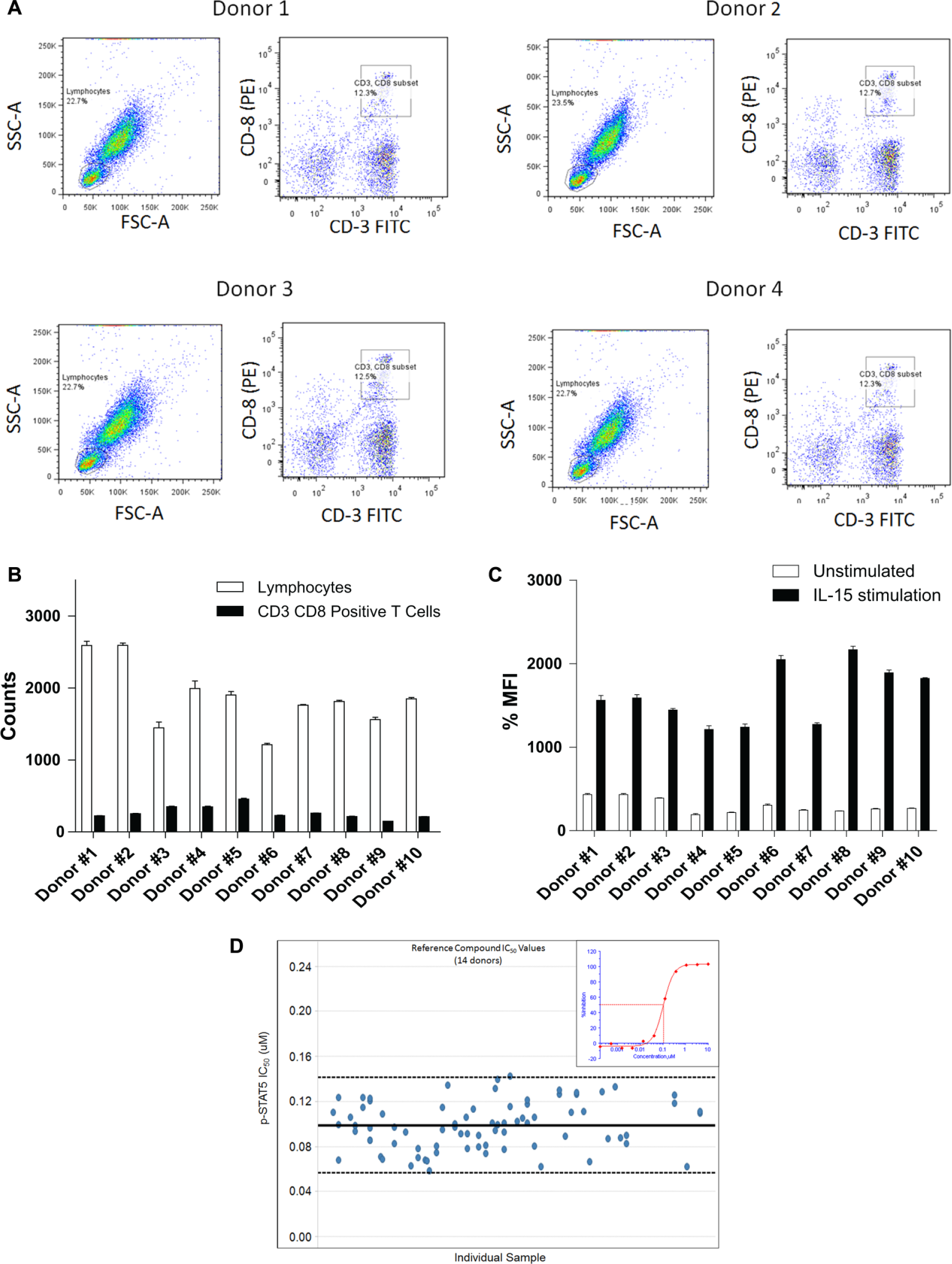
Ex vivo treatment of human whole blood inhibits STAT5 phosphorylation in CD3/CD8 positive cells as measured using HTFC. (
A large-scale study evaluating the cellular activities of selective, p-STAT small-molecule inhibitors was executed using the manual 96-well and automated 384-well assays described in this report. As shown in
Figure 4A
, individual compound inhibitory activities against p-STAT5 are comparable regardless of the assay used (Pearson correlation coefficient value = 0.932). As a result, 4000 compounds were screened for their activity in whole blood obtained from 30 independent donors using our validated HTFC approach. Screening was completed in less than 180 days, and
Figure 4B
shows representative compound activity profiles, of which many potently reduce pSTAT5 signaling in CD3/CD8 positive cells (IC50 < 3 nM, lowest concentration tested). A handful of these are JAK1 and JAK3 inhibitors whose IC50 values range in this assay from 177 to 270 nM (
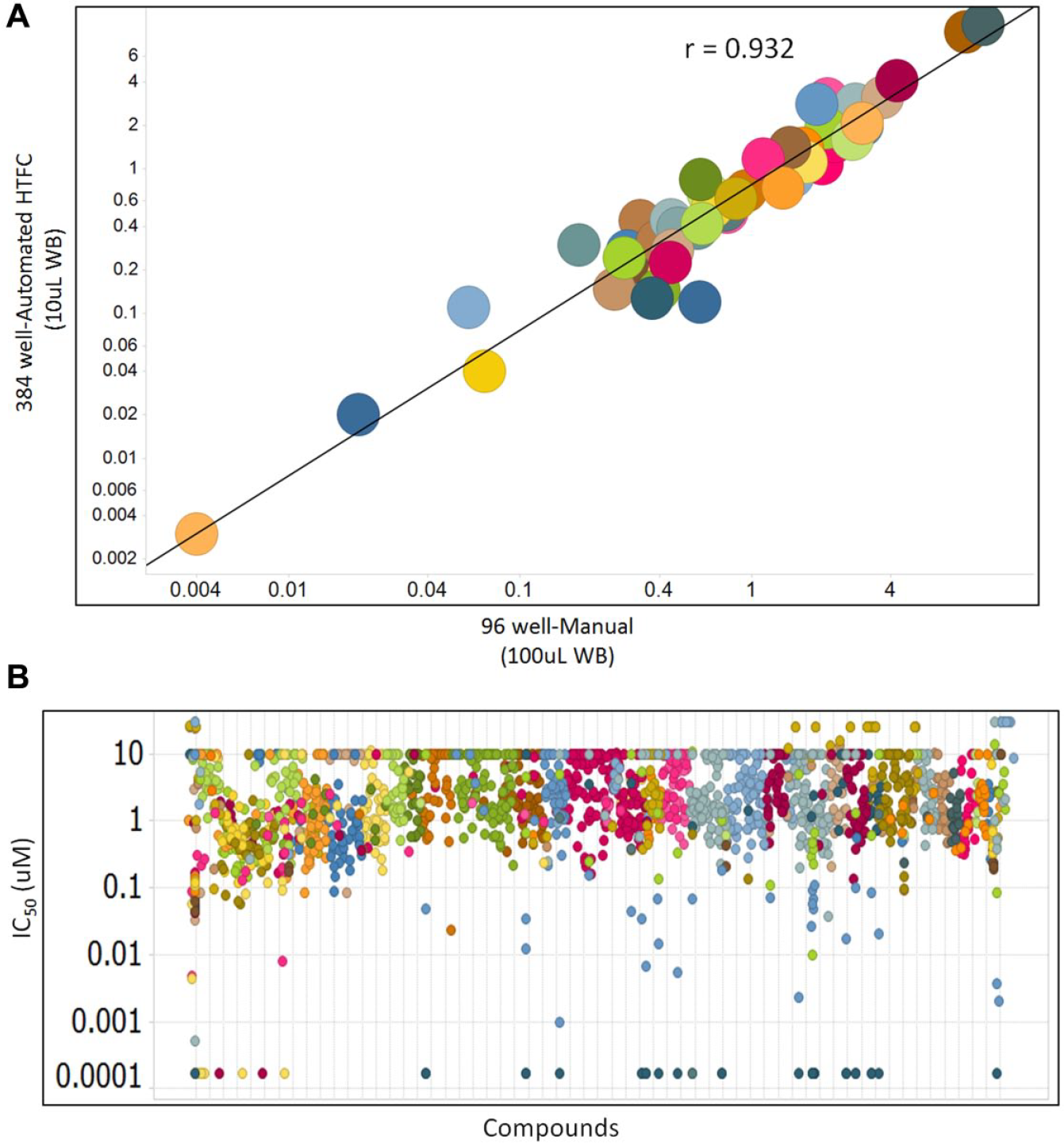
Large-scale HTFC study data evaluating the cellular activities of selective, p-STAT small-molecule inhibitors in human whole blood (WB). (
Over the past decade, flow cytometry has developed into a critical tool for examining mechanisms of action, monitoring intracellular signaling pathways and low-throughput screening.
13
Typically these studies use immortalized cells expressing transgenes of interest, require sizable sample quantities and several manual manipulations of samples and plates. In our case, analyzing compound activities in primary T cells isolated from peripheral whole blood, each study initially required a 10-fold blood volume excess and aggressive mixing to lyse red blood cells as well as reagent- and labor-intensive cell fixation, permeabilization staining steps all of which hampered HTS platform development. We overcame these limitations, by optimizing and chemically validating a fully automated HTFC platform capable of processing miniaturized human peripheral whole blood samples in a resource- and cost-effective manner (
Another key factor to successful drug development is measuring compound activities in nonhuman animal models. Our human whole blood HTFC assay has the potential to process blood samples as low as 5 μL per well so it could be adapted to test compound activities in other animal species (such as mouse, rat, dog and monkey) and accordingly could preserve animals and significant veterinary sciences staff investment. Collectively, our HTFC platform can potentially provide a translational understanding of complex physiological cellular networks involved in disease processes at a significantly earlier stage of drug discovery which enables development of efficacious and potent therapies sooner.
Footnotes
Declaration of Conflicting Interests
The authors declare no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
