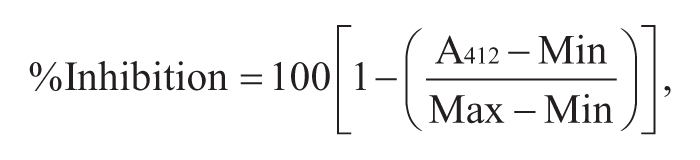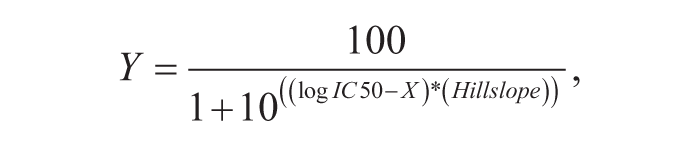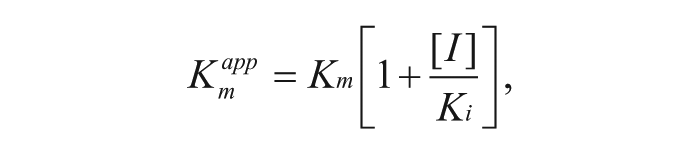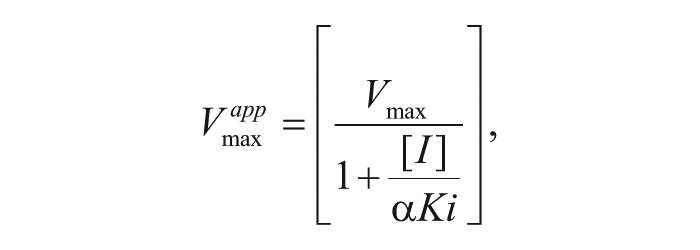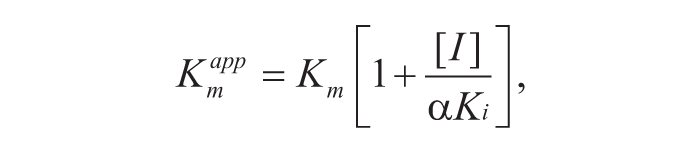Abstract
Secondary metabolite of Aspergillus terreus, terreic acid, is a reported potent antibacterial that was identified more than 60 years ago, but its cellular target(s) are still unknown. Here we screen its activity against the acetyltransferase domain of a bifunctional enzyme, Escherichia coli N-acetylglucosamine-1-phosphate-uridyltransferase/glucosamine-1-phosphate-acetyltransferase (GlmU). An absorbance-based assay was used to screen terreic acid against the acetyltransferase activity of E. coli GlmU. Terreic acid was found to inhibit the acetyltransferase domain of E. coli GlmU with an IC50 of 44.24 ± 1.85 µM. Mode of inhibition studies revealed that terreic acid was competitive with AcCoA and uncompetitive with GlcN-1-P. It also exhibited concentration-dependent killing of E. coli ATCC 25922 up to 4× minimum inhibitory concentration and inhibited the growth of biofilms generated by E. coli. Characterization of resistant mutants established mutation in the acetyltransferase domain of GlmU. Terreic acid was also found to be metabolically stable in the in vitro incubations with rat liver microsome in the presence of a NADPH regenerating system. The studies reported here suggest that terreic acid is a potent antimicrobial agent and support that E. coli GlmU acetyltransferase is a molecular target of terreic acid, resulting in its antibacterial activity.
Introduction
Gram-negative infections have emerged as one of the world’s greatest threats, causing significant mortality and morbidity. Antibiotic resistance in Gram-negative bacteria is increasing at an alarming rate, and multidrug resistance has now become a serious threat to public health and patient safety mainly because of the unavailability of antibiotics to treat infections caused by these organisms. 1 Furthermore, emergence of extended-spectrum β lactamase (ESBL)–producing organisms has also raised concern to work on new targets and methods for developing improved antibacterial drugs. The emergence of the antibiotic-resistant pathogens has outpaced the available treatment options, making it indispensable to work on new strategies for developing new and improved antibiotics. 1 There is, therefore, an urgent need to implement diverse compound screening and target identification strategies for discovering new and improved antibacterial agents to address the issue of bacterial resistance. Inhibiting cell wall biosynthesis has been proven to be attractive for antibacterial drug discovery, and the potency of antimicrobial agents targeting the cell wall of the bacteria recommends that the enzymes involved in the cell wall biosynthetic pathway can be exploited to find targets for antibacterial drug discovery. 2 Hence, enzymes of cell wall synthesis can be targeted to develop antibacterial molecules. Escherichia coli N-acetylglucosamine-1-phosphate-uridyltransferase/glucosamine-1-phosphate-acetyltransferase (GlmU) is one such enzyme that is involved in the biosynthesis of peptidoglycan and lipopolysaccharide in Gram-positive and Gram-negative bacteria, respectively. GlmU-catalyzed reactions lead to the formation of UDP-N-acetylglucosamine-1-P from glucosamine-1-P (UDP-GlcNAc) in a two-step reaction of acetylation and uridylation.3,4
UDP-GlcNAc, a cytoplasmic cell wall precursor in bacteria, is formed from fructose-6-P in a four-step reaction involving three enzymes—namely, glucosamine-6-phosphate synthase (GlmS), GlmM mutase, and GlmU ( Fig. 1 ).3,4 GlmU catalyzes the last two steps, acetylation and uridylation of glucosamine-1-phosphate, yielding UDP-GlcNAc, which is then used by the cell in the biosynthetic pathways of peptidoglycan and polysaccharide.3,4 In eukaryotes, the UDP-GlcNAc biosynthesis is a four-step pathway catalyzed by different enzymes ( Fig. 1 ). 5 The reaction catalyzed by the acetyltransferase domain of GlmU is unique to bacteria and does not have any homology with the eukaryotic counterpart, which makes it a more preferred target for the development of antibiotics specific to the bacterial enzymes.3,5 In addition, the acetyltransferase domain of GlmU has also been validated in vitro as an antibacterial target in Haemophilus influenzae, making it a suitable target for drug development. 6
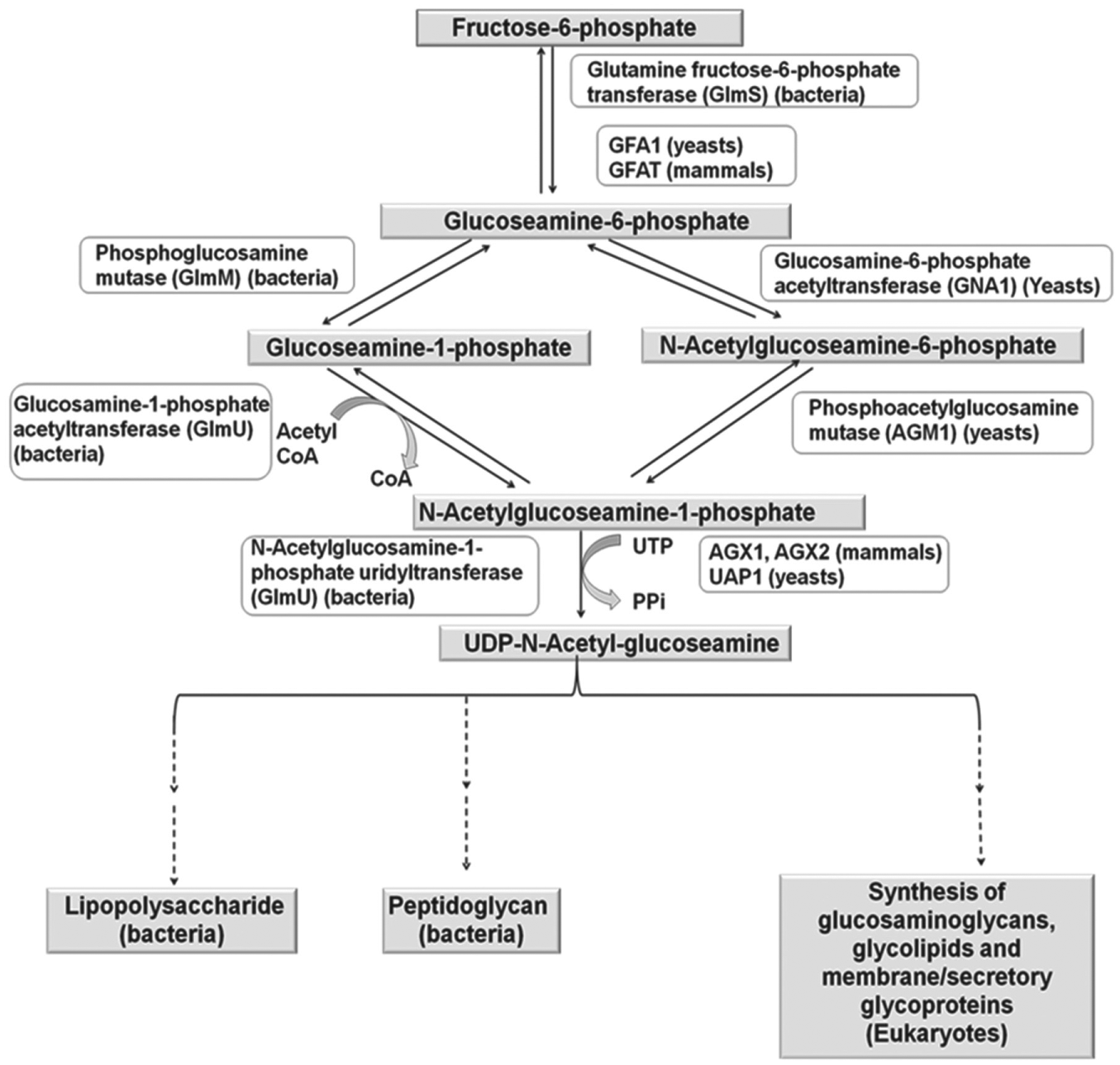
Biosynthetic pathway and metabolism of UDP-N-acetylglucosamine in prokaryotes and eukaryotes.
The structure of GlmU enzyme and its mechanism has been revealed from the crystallographic studies of the GlmU enzyme with its substrates and inhibitors. The N-terminal domain (residues Asn3–Arg229) adopts an α/β fold similar to the dinucleotide binding Rossmann fold, whereas the C-terminal domain (residues Phe260–Ala437) catalyzes the acetyltransferase activity and adopts a left-handed parallel β-helix (LβH) structure homologous to bacterial acetyltransferases (
Terreic acid is a reported antibacterial inhibitor, but its cellular and molecular targets are still unknown. 15 It was initially reported as an inhibitor of MurA, 16 an enzyme involved in the lipopolysaccharide (LPS) biosynthetic pathway, but later studies revealed that MurA is not the molecular target of terreic acid, and a different mechanism is responsible for the antibacterial activity of terreic acid. 17 In this study, terreic acid was tested for its target engagement with acetyltransferase activity of E. coli GlmU. The acetyltransferase activity of terreic acid was monitored using Ellman’s reagent, and its IC50 was 44.24 ± 1.85 µM. Terreic acid was also found to be specific to the acetyltransferase domain of E. coli GlmU, which was determined by using the GlmU variant lacking acetyltransferase activity. Molecular docking studies and the characterization of resistant mutant strains further characterized E. coli GlmU as a molecular target of terreic acid.
Materials and Methods
Isolation and Maintenance of Aspergillus terreus
The fungal strain MRCJ-356, used in the present study, was isolated from a soil sample collected from the Shiwalik region situated at a height of about 900 meters above sea level, having geographical location 33.38°N 74.38°E. Phylogenetic analysis of the isolate was carried out by the acquisition of the ITS1-5.8S-ITS2 ribosomal gene sequencing. The sequence was submitted to the GenBank and the strain was deposited to the institutional repository under the strain identifier MRCJ-356. During the present study,the isolate was stored and maintained on Sabouraud dextrose agar (SDA) at 4 °C.
Cultivation Conditions
Preseed inoculum was prepared in a 250-mL Erlenmeyer flask containing 50 mL Sabouraud dextrose broth (SDB) medium inoculated with a loop full of culture. The culture was grown at 28 °C with shaking at 200 rpm. Five percent of 2-day preseed culture was further transferred to a 1-L Erlenmeyer flask containing 250 mL SDB medium. The seed culture was allowed to grow at 28 °C with shaking at 200 rpm for the next 2 days. Batch fermentation was carried out in a 7-L fermentor (BioFlo 110; New Brunswick Scientific, Enfield, CT) with 5 L SDB medium under the following conditions: temperature 28 °C, agitation speed 200 rpm, and aeration 0.5 vvm. Fermentation was carried out for 10 days, and the batch was terminated and processed for isolation of the desired compound.
Extraction Procedure and Isolation of Terreic Acid
Solvents were distilled before use. 1H and 13C nuclear magnetic resonance (NMR) spectra were recorded on 400-MHz spectrometers (model no. D 205/52-2382, Avance 500 [Billerica, MA, USA]) using tetramethylsilane (TMS) as the internal standard. Chemical shifts are expressed in parts per million (δ ppm). Silica gel–coated aluminum plates were used for TLC (thin layer chromatography). The product was purified by column chromatography on silica gel (60–120/100–200 mesh) using petroleum ether–ethyl acetate as the eluent to obtain the pure product. Elemental analyses were performed on the Elementar. Liquid chromatography/mass spectrometry (LC/MS) spectra (model no. Symapt MS; Waters, Milford, MA, USA) and optical rotation (model No. 241; PerkinElmer Waltham, Massachusetts, USA) were recorded. The tested compound yielded data consistent with 95% purity measured by high-performance liquid chromatography (HPLC) (Agilent 1260 Affinity; Agilent, Santa Clara, CA, USA).
Spectral Analysis of Terreic Acid
Pale yellow needles, m.p. 126–127 °C. UV spectra (MeOH) λmax (log ε) 213 (4.03) and 314 (3.88). [α]28.6D25 (c, 0.046 in MeOH). Molecular formula: C7H6O4. HR-MS: [M+H]+ m/z 155.0358. Data 1H NMR (400 MHz; CDCl3): 1.93 (s, 3H); 3.86 (d, 1H, J = 3.7 Hz); 3.89 (d, 1H, J = 3.7 Hz); 6.87 (s,1H) and 13C NMR (100 MHz; CDCl3): 8.8, 51.6, 53.8, 120.4, 151.9, 187.5, 190.8.
Bacterial Strains and Culture Conditions
The bacterial strains E. coli ATCC 25922, Pseudomonas aeruginosa ATCC 29212, and Klebsiella pneumoniae ATCC 700603 were obtained from ATCC (American Type Culture Collection, Manassas, VA). Metallo-β-lactamase (NDM1) clinical isolates, K. pneumoniae, and Acinetobacter baumannii, obtained from hospital settings, were also included in the study. Cation-adjusted Mueller-Hinton broth (MHB; Becton-Dickinson, Cockeysville, MD) was used for minimum inhibitory concentration (MIC) determination. Mueller-Hinton agar (MHA; Becton-Dickinson) was used for minimum bactericidal concentration (MBC) studies. Trypticase soy agar (TSA; Becton-Dickinson) was used for culturing of bacteria and colony counts.
Construction of Expression Vectors
Standard procedures for molecular cloning and E. coli cell transformation were used. The E. coli glmU gene was amplified by PCR using genomic DNA isolated from E. coli ATCC 25922 as the template and the following oligonucleotides as primers: 5′-AAA
Similarly, DNA fragment encoding C-terminally truncated enzyme (GlmU-Tr331) lacking acetyltransferase activity was generated by PCR using the following oligonucleotides as primers: 5′-AAA
Both the cloned genes were confirmed by sequencing (Chromus biotech pvt. Ltd., Bengaluru, India). The nucleotide sequence was analyzed by basic local alignment search tool (BLAST) available at the National Center for Biotechnology Information (NCBI; http://www.ncbi.nlm.nih.gov/). pBA750, 6 an expression vector expressing 1.7 kb H. influenzae glmU, was generously gifted by Dr. Ed T. Buurman (AstraZeneca, Boston, MA).
Protein Expression and Purification
Standard procedures for protein expression and purification were used. To express E. coli GlmU, the expression vector pET28a harboring the glmU gene was transformed in E. coli BL21 (DE3) by electroporation (25 µF capacitance, 2.5 kV and 200 Ω) using Bio-Rad Gene Pulser X cell (Bio-Rad Laboratories) and plated on Luria-Bertani (LB) agar containing 50 µg/mL kanamycin. After overnight growth at 37 °C, 1% inoculum was inoculated into 200 mL LB broth containing 50 µg/mL kanamycin, and the cells were grown to an optical density (600 nm) of 0.5. These cells were then induced with 0.2 mM IPTG, allowed to grow for 12 to 14 h at 18 °C, and harvested by centrifugation at 5000 rpm at 15 °C for 15 min. Cells were resuspended in a 1:8 ratio of lysis buffer (1× phosphate-buffered saline [PBS] buffer, pH 7.3, 5% glycerol) with 1 mM DL–dithiothreitol (DTT) and 1 mM phenylmethylsulfonyl fluoride (PMSF) and were then lysed by sonication (12–15 cycles, 30-s pulse and 55-s halt). Cell debris was pelleted by centrifugation at 10,000 rpm for 30 min. The supernatant and pellet were analyzed by sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) for the presence of protein. Clarified supernatant containing the protein of interest was loaded onto a preequilibrated Ni-NTA column (Qiagen). The column was then washed with 10 mL wash buffer (lysis buffer with 500 mM NaCl, pH 7.3). The protein was eluted using a linear gradient of imidazole from 50 to 250 mM in 25 mM Tris-HCl buffer (pH 8.0) containing 140 mM NaCl with 1 mM PMSF and 1 mM DTT. E. coli GlmU-Tr331 was also expressed and purified using the same protocol.
Recombinant vector pBA750 was transformed in BL21 (DE3) electrocompetent cells, and H. influenzae GlmU was expressed and purified using the above protocol except for the IPTG, which was added to a final concentration of 1 mM during induction.
Acetyltransferase Assay
For the acetyltransferase activity, the activity was monitored in an assay by quantification of CoA produced. CoA reacts with 5,5′-dithiobis-(2-nitrobenzoic acid) (DTNB) to release 4-nitrothiophenolate, which can be quantified at 412 nm. 6 Enzyme reactions were performed in a 50-µL volume containing 200 µM acetyl CoA (Sigma, St. Louis, MO) and 200 µM D-glucosamine-1-phosphate (Sigma) in assay buffer containing 50 mM Tris-HCl and 5 mM MgCl2. Reaction was initiated with the addition of 19.49 nM E. coli GlmU or 0.20 µM H. influenzae GlmU, incubated at 37 °C for 30 min, and stopped by adding 50 µL stop solution (50 mM Tris-HCl [pH 7.27] and 6 M guanidine hydrochloride). To the terminated reaction, 50 µL DTNB solution (50 mM Tris-HCl [pH 7.6], 1 mM EDTA, and 0.2 mM DTNB) was added and the absorbance was determined at 412 nm using a microplate reader (Infinite PRO 200; Tecan, Männedorf, Switzerland). Unit activity of the acetyltransferase domain of the enzyme was defined as the µM of the product formed per minute per milligram of the enzyme. Primary screening of terreic acid against acetyltransferase domain of E. coli GlmU and H. influenzae GlmU was done at a 100-µM concentration. Neat DMSO was added to both positive (minimum inhibition) and negative controls (maximum inhibition). The maximum (100%) inhibition was measured for reactions inhibited by preincubation of enzyme with 1 µM of an already reported inhibitor, 1-[2,4-Dimethoxy-5-(10H-phenoxazin-10-ylsulfonyl) phenylamino]-2-(-4-pyridyl)-1-ethanone, from AstraZeneca (Boston, MA, USA). 6
The IC50 for the inhibition of acetyltransferase activity of E. coli GlmU and H. influenzae GlmU by terreic acid was determined by serially diluting terreic acid in DMSO in the concentration range starting from 0.39 to 100 µM and preincubating with 19.49 nM E. coli GlmU or 0.20 µM H. influenzae GlmU prior to the start of the reaction with AcCoA and GlcN-1-P.
Percent inhibition was calculated using the following equation:
where A412 is the absorbance in the test well, Min is the absorbance of the fully inhibited reaction, and Max is the absorbance of the uninhibited reaction. A log substrate concentration versus percent inhibition (XY) graph was created using GraphPad Prism 6 (GraphPad Software, La Jolla, CA), and the compound concentration resulting in 50% inhibition (IC50) was calculated by nonlinear least squares regression and equation solving for IC50:
where Y is % inhibition and X is log inhibitor concentration.
Uridyltransferase Assay
Uridyltransferase assay for determining the specificity of terreic acid to the acetyltransferase domain of E. Coli GlmU was performed in an 8-µL volume containing 25 mM HEPES buffer (pH 7.6), 10 mM MgCl2, 1 mM DTT, 0.5 mM GlcNAc-1-P, 1 mM UTP, 0.04 units of thermostable inorganic pyrophosphatase, and 0.17 µM E. coli GlmU-Tr331. 18 The reactions were incubated for 30 min at 30 °C and terminated by incubating at 65 °C for 30 min, and the product formed was estimated by a malachite green phosphate assay kit (Bioassay Systems, Hayward, CA) according to the manufacturer’s instructions. Unit activity of the uridyltransferase domain of the enzyme was defined as the µM of the product formed per minute per milligram of the enzyme.
Mode-of-Inhibition Studies
Mode-of-inhibition and Ki values were determined by simultaneously changing the terreic acid concentration and either the AcCoA or GlcN-1-P concentration (50–1600 µM). The resulting curves were fit by nonlinear least squares regression models describing competitive, uncompetitive, noncompetitive, or mixed inhibition using the enzyme kinetics module of GraphPad Prism 6 (GraphPad Software). Inhibition constant of a competitive inhibitor (Ki) can be determined by the following equation 19 :
where
The inhibition constant for uncompetitive inhibition (αKi) can be determined by the following equations 19 :
where
where
Modeling Studies
Molecular docking of terreic acid was carried out to analyze its binding interactions at GlmU acetyltransferase domain of E. coli. The molecular modeling studies were carried out using GLIDE v 6.2 implemented in Maestro v 9.7 (Schrodinger, LLC, New York, NY). 20 Ligand structures were drawn using a 2D sketcher, and further preparation of the ligands was performed using Ligprep, where their 3D states, possible stereoisomers, and ionization states at pH 7 ± 2 were generated. Docking was carried out on the homotrimer biological assembly of GlmU retrieved from PDB (PDB code: 4AA7), as the protein exists as a trimer, and the acetyltransferase site has a contribution from all the three subunits. Protein trimer was prepared, optimized, and minimized using the Protein Preparation Wizard. The 3D grid was generated around the co-crystallized inhibitor bound of the acetyltransferase domain. Docking was performed on this grid using the Extra Precision (XP) scoring function of GLIDE.
MIC and MBC determination of terreic acid against Gram-negative pathogens
The MIC of terreic acid was determined as per the guidelines of Clinical and Laboratory Standards Institute (formerly, the National Committee for Clinical Laboratory Standards). 21 Bacterial suspensions were prepared by suspending 24-h grown culture in sterile normal saline and adjusting the turbidity to 0.5 McFarland standard, which is equivalent to 1.5 × 108 colony-forming units (CFU)/mL. Twofold serial dilutions of terreic acid were prepared in the 100-µL volume of Mueller-Hinton broth (MHB; BD Biosciences, San Jose, CA) in 96-well U-bottom microtiter plates (Tarson, Mumbai, India). The final concentrations of terreic acid ranged from 2.87 to 368 µg/mL. The above-mentioned bacterial suspensions were further diluted in MHB, and a 100-µL volume of these diluted inoculums was added to each well of the respective plates, resulting in a final inoculum of 5 × 106 CFU/mL in the well. After incubation at 37 °C for 24 h, the plates were read visually, and the minimum concentration of the compounds showing no turbidity was recorded as the MIC. The MBC was determined by spreading a 100-µL volume on Mueller-Hinton agar plates, and the bacterial cells were enumerated after incubation at 37 °C for 24 h. MBC was defined as the lowest concentration of the compound at which more than 99.9% of the cells were killed compared with a nontreated control.
Time-Kill Studies against E. coli ATCC 25922
E. coli ATCC 25922 was grown in MHB at 37 °C for 24 h. The turbidity of bacterial suspension was adjusted to a McFarland standard of 0.5 in sterile normal saline. A total of 200 µL of this suspension was used to inoculate 20 mL MHB containing increasing concentrations of terreic acid ranging from 23 to 184 µg/mL. DMSO control was also included in the study. Suspensions were incubated at 37 °C, and the number of CFU was determined on MHA using a serial dilution method at various time points. 22
Biofilm Susceptibility Assay
The effect of terreic acid on biofilm formation by E. coli ATCC 25922 was examined by the microdilution method. 23 This method was similar to the MIC assay for planktonic cells. The bacterial suspensions were prepared from the overnight-grown culture, and the turbidity of the suspension was adjusted to an optical density at 600 nm (OD600) of 0.05 (1 × 109 CFU/mL). Twofold serial dilutions of terreic acid were prepared in MHB in the wells of a 96-well flat-bottom polystyrene tissue culture plate (Tarson) containing MHB in a volume of 100 µL/well. Then, 40 µL fresh MHB was added to each well, followed by the addition of 60 µL of the above-mentioned suspension to each well of the plate. This resulted in the final inoculum of 6 × 107 CFU/mL in each well; the final concentrations of terreic acid ranged from 2.875 to 368 µg/mL. After incubation at 37 °C for 24 h, absorbance at 595 nm was recorded to assess the culture growth. The culture supernatant from each well was decanted, and planktonic cells were removed by washing the wells with PBS (pH 7.2). The biofilm was fixed with methanol for 15 min and then air dried at room temperature. The wells of the dried plate were stained with 0.1% (wt/vol) crystal violet (Sigma) for 10 min and rinsed thoroughly with water until the negative control wells appeared colorless. Biofilm formation was quantified by the addition of 200 µL of 95% ethanol to the crystal violet–stained wells and recording the absorbance at 595 nm (A595) using a microplate reader (Infinite PRO 200, Tecan, Mannedorf, Switzerland).
Combination Studies of Terreic Acid with Antibiotic against E. coli ATCC 25922
The potentiating effects of terreic acid on the MICs of antibiotics (ciprofloxacin, ceftriaxone, carbenicillin, tetracycline, gentamycin; Sigma) were determined by the checkerboard microdilution method as described previously. 22 The experiment was performed in 96-well U-bottomed plates (Tarson). Ten 2-fold serial dilutions of antibiotics, ranging from MIC to below MIC, were prepared in the presence of increasing concentrations of terreic acid (0.35–184 µg/mL), in such a way that row B of the microtiter plate received the lowest concentration of terreic acid and row H received the highest. Bacteria grown overnight on MHA plates were suspended in normal saline (0.85%) and the turbidity was adjusted so that it was equivalent to that of a 0.5 McFarland standard, corresponding to 1.5 × 108 CFU/mL. Further dilution of the inoculum in MHB was done in such a manner that each well contained 5 × 105 CFU/mL as final bacterial inoculums and the plates were then incubated at 37 °C for 18 h.
Isolation of Resistant Mutants In Vitro
For the confirmation of GlmU as the molecular target of terreic acid, resistant mutants were generated against it. Single-step resistance mutant generation method was used for raising mutants using a previously described method. 24 A bacterial suspension containing 109 CFU/mL (100 µL) was plated on MHA containing terreic acid at concentrations equal to 2×, 4×, and 8× MIC. Mutation frequency was calculated by counting the total number of colonies appearing after 24 h of incubation at 37 °C on the terreic acid–containing plate and by dividing the number by the total number of CFU plated. Isolated colonies were picked and retested for growth on terreic acid–containing agar plates to confirm their resistance. The MIC of the mutant strains was then determined as per the above discussed protocol, and the strain with increased MIC was taken for further analysis through sequencing of the glmU PCR product.
In Vitro Metabolism
Terreic acid at 0.5 µM was incubated with 0.32 mg/mL of rat liver microsomes (RLM) at 37 °C for 30 min in phosphate buffer (pH 7.4) with an NADPH regeneration system (0.8 mM NADP+, 2.5 mM G6P, 1 U/mL G-6-PDHase). 25 Verapamil was used as a positive control in the assay. Upon terminating the incubations by the addition of acetonitrile, the extract was analyzed by LC/MS/MS to determine the remaining concentration of terreic acid. For LC/MS/MS analysis, an HPLC-30AD (Shimadzu, Kyoto, Japan) connected to an LCMS-8030 mass spectrometer was used. A Chromolith RP 18 (Merck Millipore, Germany) column (4.6 × 100 mm) was eluted with a mobile phase of water/acetonitrile (20–80 in an 8.4-min elution time) containing 0.1% formic acid at a flow rate of 0.5 mL/min. The MS detector was run at electrospray ionization (ESI) negative mode at m/z 153.2. Verapamil as internal standard was monitored at SIM of m/z 455.4.
Results
Extraction and Isolation of Terreic Acid
Batch fermentation of A. terreus in 5 L production medium for 10 days under specified conditions resulted in about 86 g/L wet biomass. After termination of fermentation, the fermented broth was extracted with EtOAc. About 100 mg crude extract was obtained from the fermented broth of A. terreus and subjected to column chromatography on silica gel 60 using a stepwise gradient from n-hexane/EtOAc 90:10 v/v (18 mg). The molecular weight of the compound was established as 154 by ESI-MS, having the corresponding molecular formula C7H6O4 and 5 DBE. The 1H NMR spectrum (
GlmU Acetyltransferase Activity and IC50 of Terreic Acid
The recombinant E. coli glmU was cloned, expressed, and purified in the E. coli system. The activity of terreic acid against the acetyltransferase domain of E. coli GlmU was evaluated by a colorimetric assay as described above, where the assay conditions were standardized using an already reported inhibitor, 1-[2,4-Dimethoxy-5-(10H-phenoxazin-10-ylsulfonyl) phenylamino]-2-(-4-pyridyl)-1-ethanone, from AstraZeneca.
6
Primary screening of terreic acid against the acetyltransferase domain of E. coli GlmU was performed at a 100-µM concentration where terreic acid was found active, showing percent inhibition of 84% ± 0.69%. IC50 value of terreic acid was calculated by the above discussed protocol, and IC50 of 44.24 ± 1.85 µM against acetyltransferase activity of E. coli GlmU was seen (
Uridyltransferase assay, using E. coli GlmU-Tr331, was performed to validate the specificity of terreic acid to the acetyltransferase domain of E. coli GlmU. Terreic acid was not found to be active against the uridyltransferase domain (with IC50 > 100 µM), which thereby confirms that terreic acid is indeed specific to the acetyltransferase domain of E. coli GlmU.
Mode-of-Inhibition Studies of Terreic Acid
The mode of inhibition for terreic acid with the acetyltransferase activity of GlmU was tested to determine how the inhibitor binds to the enzyme. Terreic acid showed uncompetitive inhibition, with GlcN-1-P having an uncompetitive inhibition constant value (αKi) of 41.94 ± 4.99 µM, and competitive inhibition was observed with AcCoA having a competitive inhibition constant (Ki) value of 40.29 ± 3.54 µM ( Fig. 2 ).
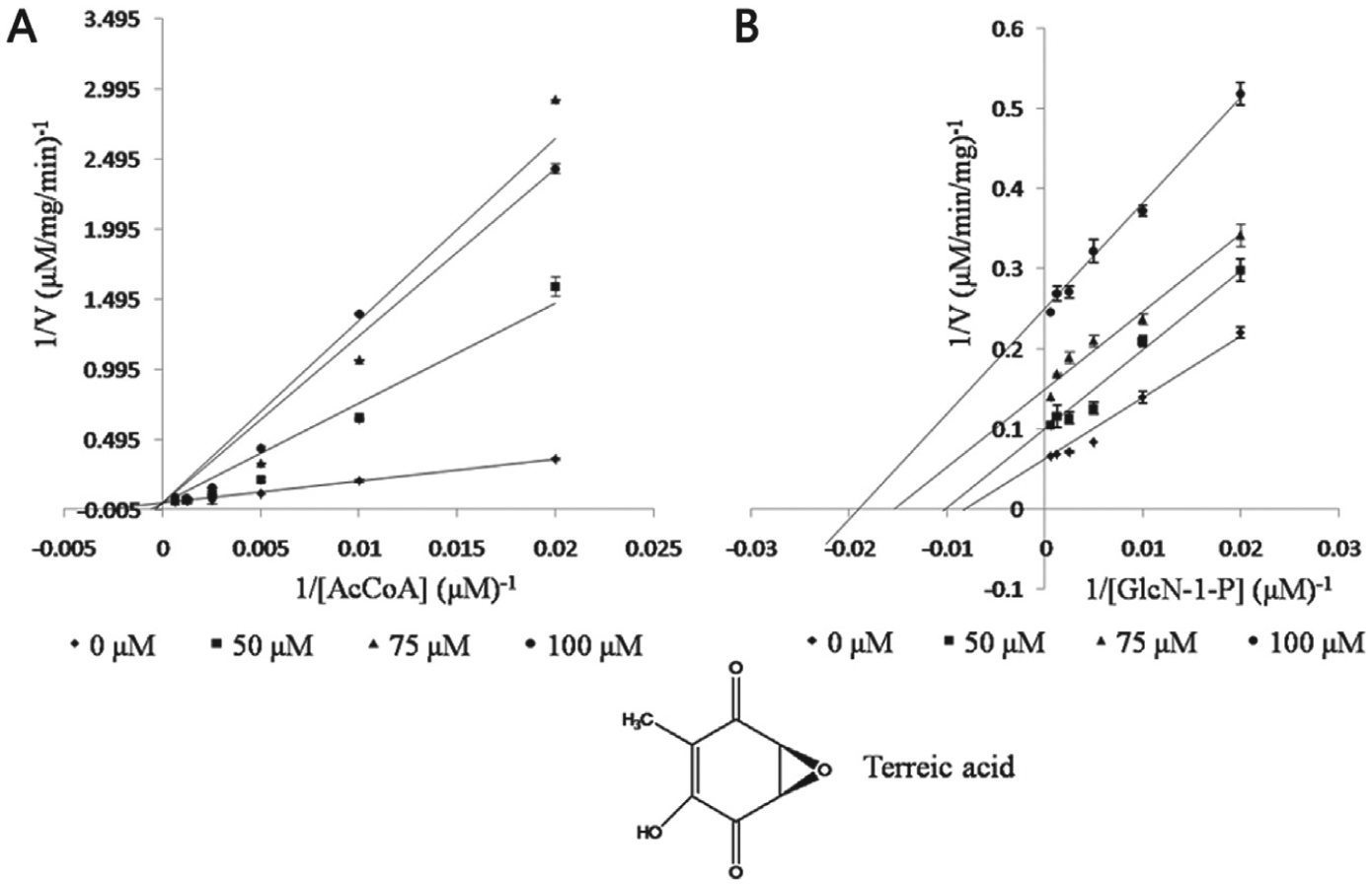
Lineweaver-Burk plots of inhibition studies of terreic acid against Escherichia coli GlmU acetyltransferase activity. Increasing concentrations of compound were used and either AcCoA (
Modeling Studies
Analysis of the docked protein–terreic acid complexes revealed the important residues interacting with the compound with the GLIDE docking score of −1.854 ( Fig. 3 ). GLIDE score represents the protein-ligand binding affinity. Residues of the acetyltransferase domain binding pocket present within 4Å of the docked ligand poses were analyzed. Residues A:ILE383, A:CYS385, A:TYR387, A:ASP388, A:GLN408, and A:THR426 were identified as the essential residues that are present in the binding pocket ( Fig. 3A ). Binding pose of the compound showed that its O– atom has the tendency to form hydrogen bonding interactions with the S-H of A:CYS385, where this residue is a hydrogen bond donor. The docked pose of terreic acid at the acetyltransferase site of E. coli GlmU shows the presence of A:ALA411 and A:ILE383 having hydrophobic groups as well as A:GLN408 having hydrogen bonding interactions. A:PHE393 provides the hydrophobic environment in the binding pocket of acetyltransferase site of GlmU.
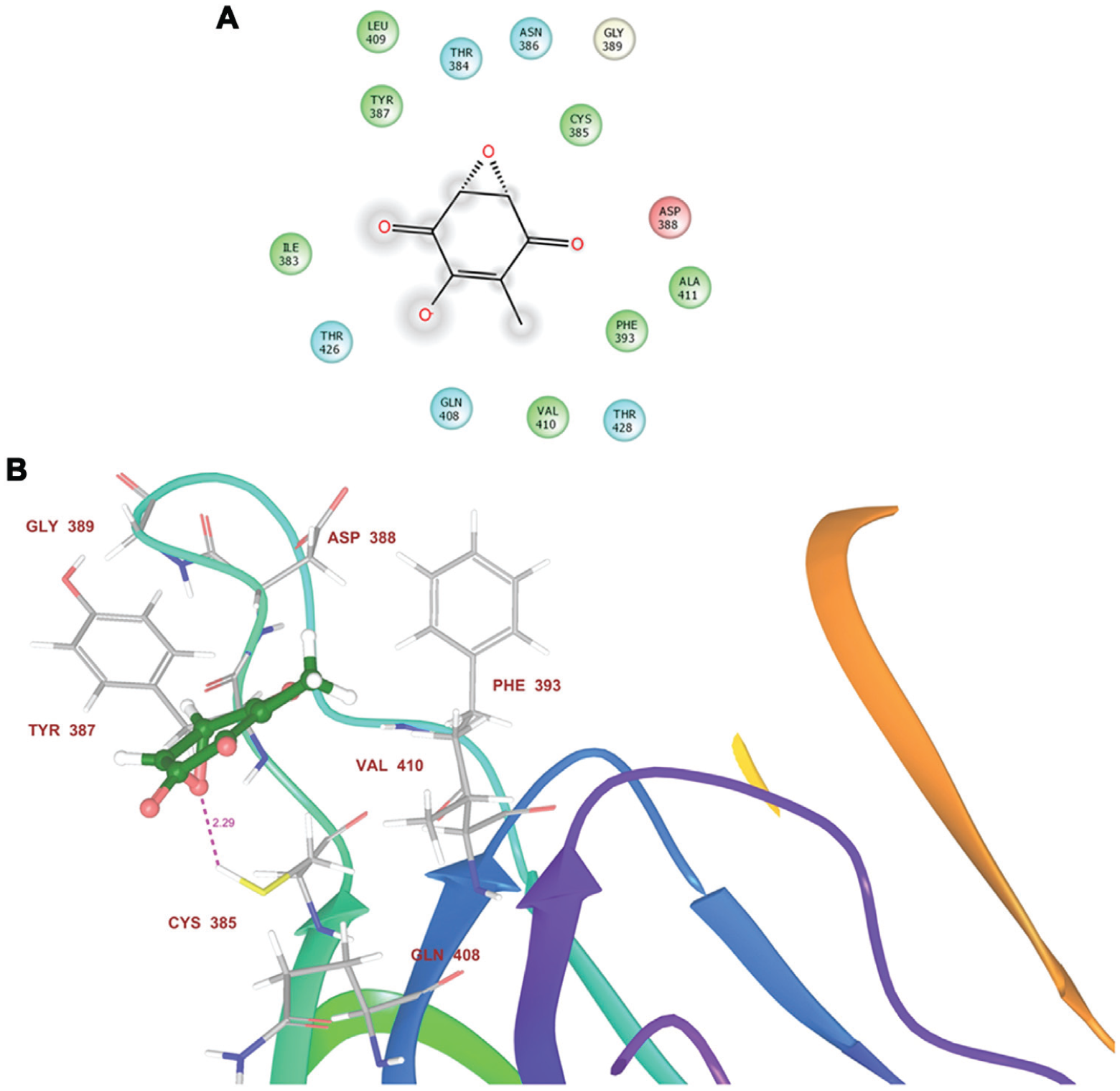
(
MIC and MBC of Terreic Acid against Gram-Negative Pathogens
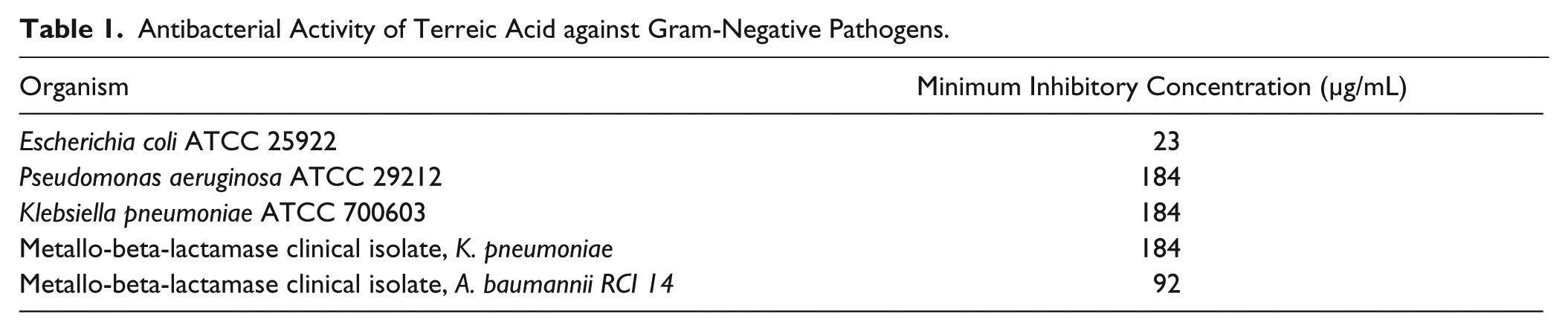
To evaluate the antimicrobial activity of terreic acid against Gram-negative microorganisms, the MICs and the MBCs were determined, and the results are shown in Table 1 . Terreic acid exhibited an MIC of 23 µg/mL (150 µM) against E. coli ATCC 25922, whereas the MBC was 2-fold greater than the inhibitory concentration ( Table 1 ). Terreic acid also exhibited antibacterial activity against other Gram-negative bacteria tested (P. aeruginosa ATCC 29212, K. pneumoniae ATCC 700603, and clinical isolates: A. baumannii and K. pneumoniae) ( Table 1 ).
Antibacterial Activity of Terreic Acid against Gram-Negative Pathogens.

Time-Kill Studies
The time-kill studies were performed against E. coli ATCC 25922, and the results are shown in Figure 4 . The 2×MIC and 4×MIC of terreic acid showed the reduction in the CFU count of E. coli ATCC 25922 below the detection limit (50 CFU/mL) in 6 h and 3 h, respectively. The kill kinetics study, hence, showed that terreic acid exhibits a concentration-dependent killing effect against E. coli ATCC 25922.
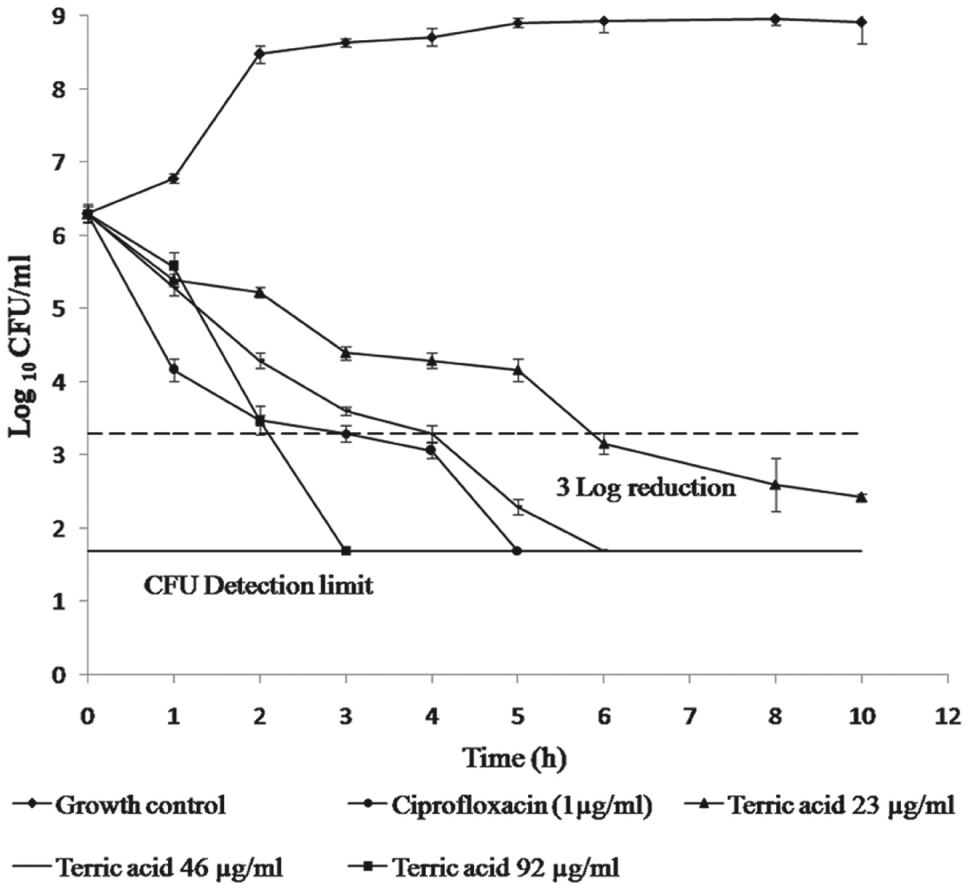
Effect of terreic acid on the cell growth of Escherichia coli ATCC 25922 at different concentrations. Values are means ± SD from three independent determinations.
Biofilm Inhibition
Terreic acid exhibited an inhibitory effect on the formation of biofilm generated by E. coli ATCC 25922, with a minimum biofilm inhibition concentration (MBIC) of 46 µg/mL for E. coli ATCC 25922, which is 2-fold higher than its MIC.
Synergistic Studies
Cell wall inhibitors may facilitate the entry of other antibiotics and act synergistically with them. Hence, the checkerboard microdilution method was used to study the effect of terreic acid on the MICs of antibiotics for in vitro combination studies. The MIC of terreic acid was determined in the presence of other antibiotics (ciprofloxacin, ceftriaxone, carbenicillin, tetracycline, gentamycin); no significant synergistic effects were detected with these antibiotics.
Characterization of Resistant Mutants In Vitro
Five strains of E. coli ATCC 25922 resistant to terreic acid were isolated with a mutation frequency of 10–7 to 10–8 at a compound concentration equal to the 2× MIC value (46 µg/mL). No mutants could be recovered at the higher concentrations of terreic acid (4× MIC, 8× MIC). MIC determination of the identified stains against terreic acid exhibited a fold increase in the MIC of one out of the five mutant strains tested. To characterize the identified mutant strain, DNA was isolated from the mutant strain and used as template to perform PCR as described above. DNA from E. coli ATCC 25922 was used as a positive control. The purified PCR product of the glmU gene was sequenced and analyzed by using protein-protein blast (blastp) at NCBI, which revealed a single mutation (F393L) in the resistant strain located in the acetyltransferase domain (
Microsomal Stability of Terreic Acid
The samples incubated with RLM were analyzed as per the method defined above. Peak area at the 0-min sample was taken as 100% of terreic acid present in the sample. Reduction in area of peak of interest was observed after 30 min. Percent drug remaining after 30 min was calculated. The metabolic stability of terreic acid was thus established in RLM, which was found to be experimentally 99.9% of drug remaining in RLM, indicating that terreic acid is not rapidly metabolized by liver microsomes.
Discussion
To tackle the threat of drug resistance in Gram-negative bacteria along with the drying antibiotic pipeline, we must look into new targets and strategies for developing new drugs for the treatment of increasing Gram-negative infections.1,2 N-acetylglucosamine-1-phosphate-uridyltransferase/glucosamine-1-phosphate-acetyltransferase (GlmU) is one such target that is involved in the cell wall biosynthesis in Gram-negative and Gram-positive bacteria, making it an attractive target for developing antibacterials.3,4 GlmU has drawn attention as a novel drug target in the recent time, and many GlmU inhibitors have been identified. Recent studies have led to the identification of nonspecific thiol reactive agents, 2-phenylbenzofurans, arylamines, and arylsulfonamides, as potent inhibitors of the acetyltransferase domain of GlmU.
Here in this study, terreic acid isolated from A. terreus has been studied for its activity against acetyltransferase domain of E. coli GlmU. Terreic acid exhibited an IC50 of 44.24 ± 1.85 µM against the acetyltransferase domain of E. coli GlmU. Since E. coli GlmU enzyme shows an identity of 69% and reveals striking similarities both at the structural and kinetic levels with a homologue sequence of H. influenzae, 28 terreic acid was screened for its potency against H. influenzae GlmU as well to find the spectrum of its activity. It was found to be less active against H. influenzae GlmU (IC50 97.56 ± 1.59µM). Since GlmU is a bifunctional enzyme showing both acetyltransferase and uridyltransferase activities, it is important to see the specificity of terreic acid to the acetyltransferase domain of E. coli GlmU. 4 The specificity of terreic acid to the acetyltransferase domain was determined using an uridyltransferase assay, where the end point was estimated using a malachite green assay. 18 C-terminally truncated enzyme (E. coli GlmU-Tr331) lacking acetyltransferase activity was used in the assay to completely target the uridyltransferase activity. Terreic acid was found to be inactive against the uridyltransferase domain of E. coli GlmU, confirming its specificity to the acetyltransferase domain of GlmU.
For understanding the basic mechanism of the inhibition of terreic acid to the acetyltransferase domain of E. coli, mode-of-inhibition studies were done. They showed competitive inhibition with acetyl coenzyme A with a competitive inhibition constant (Ki) of 40.29 ± 3.54 µM. A competitive inhibitor reversibly binds to the same site as the substrate, so its inhibition can be entirely overcome by using a very high concentration of substrate. The Vmax does not change, and the effective Km increases. 19 Terreic acid also showed uncompetitive inhibition with glucosamine-1-phosphate with uncompetitive inhibition constant (αKi) values of 41.94 ± 4.99 µM, respectively. An uncompetitive inhibitor binds to the enzyme-substrate complex but not to the free enzyme, and both the effective Vmax and the effective Km get reduced. Mode-of-inhibition studies suggest inhibition of E. coli GlmU in an ordered manner where Glc-1-P binds first, then terreic acid.
Molecular docking of the identified hits at the acetyltransferase domain of GlmU identified residues A:ILE383, A:CYS385, A:TYR387, A:ASP388, A:GLN408, and THR426 as the essential residues for providing desired configuration for terreic acid binding at the acetyltransferase site of E. coli GlmU. Docking studies show a GLIDE score of −1.854, which is the approximation of the energetic of the conformational, orientational, and positional space of the docked ligand, that is, terreic acid in the binding site of protein (E. coli GlmU). Hydrogen bond distances can be clearly seen in Figure 3B between the –S-H group of A:CYS385 and O of terreic acid with bond length of 2.29 Å. The observed hydrogen bond distance confirms the proposed mechanism, which is in agreement with the previous studies. 8
The MIC of terreic acid against E. coli ATCC 25922 was found to be approximately three times higher than its IC50 against the acetyltransferase domain of E. coli GlmU. The correlation of MIC and IC50 suggested the possibility that the acetyltransferase activity of E. coli GlmU is the cellular target of terreic acid, resulting in its antibacterial activity. To further validate that E. coli GlmU is a molecular target of terreic acid, resistant mutants were isolated and characterized, which revealed a single base pair substitution resulting in Phe-393 to Leu in the acetyltransferase domain of GlmU. Phe-393 is an important residue present in the acetyltransferase domain of E. coli GlmU as reported in literature. 6 The substitution of this important residue (F393L) must have been responsible for the conformational changes, resulting in the weak interaction with the ligand, which may have resulted in the resistance of mutant E. coli against terreic acid ( Fig. 3B ).
Terreic acid also displayed antimicrobial activities against K. pneumoniae ATCC 700603, P. aeruginosa ATCC 29212, and metallo-beta-lactamase (NDM1) clinical isolates K. pneumonia and A. baumannii. The sequence of the enzyme E. coli GlmU when aligned to the homologue sequences of K. pneumoniae, P. aeruginosa, and A. baumannii revealed sequence identities of 89%, 56%, and 55%, respectively, with E. coli GlmU (
The antibiofilm activity of terreic acid was also evaluated against biofilm generated by E. coli ATCC 25922, and it was seen that terreic acid exhibited an inhibitory effect on biofilms by E. coli. This finding co-relates well with already reported nonspecific GlmU inhibitors such as iodoacetamide, N-ethyl maleimide (NEM), and NEM analogues that have been reported to be effective against biofilm formation in Gram-negative and Gram-positive bacteria. 10 In addition to antimicrobial activity of terreic acid against Gram-negative pathogens, the microsomal stability studies predicted good stability of the compound. In literature, there are various reports describing potent E. coli GlmU inhibitors, but they do not exhibit simultaneous antimicrobial activity in Gram-negative wild-type strains. Terreic acid, reported here, exhibits dual activity, thereby providing an ideal scenario for the development of a novel antimicrobial agent. There are also a few reports of GlmU inhibitors from natural sources out of many synthetic small-molecule inhibitors of GlmU reported. One such inhibitor is 2-phenylbenzofuran, isolated from Aspergillus flavipes, which upon suitable modifications yielded a compound with IC50 of 1 µM against acetyltransferase activity of GlmU. 9 From the work presented here, terreic acid, a secondary metabolite of A. terreus, was identified as an inhibitor of acetyltransferase activity of E. coli GlmU. Further characterization of this compound revealed it to be antibacterial and metabolically stable. Chemically, terreic acid is quinone epoxide. There are reports where epoxide containing natural products such as fosfomycin produced by Streptomyces species and fumagillin from Aspergillus fumigates have shown antimicrobial activity.29,30 These structures are chemically stable and have shown stability under the performed biological assays conditions. The structure offers plenty of medicinal chemistry opportunities for optimization of biological activity by structure-activity relationship studies for the design of potential drug candidate.
Footnotes
Acknowledgements
We are grateful to Dr. Ed T. Buurman (AstraZeneca R&D, Boston, MA) for providing the expression vector pBA750 expressing 1.7 kb H. influenzae glmU and the inhibitor, 1-[2, 4-Dimethoxy-5-(10H-phenoxazin-10-ylsulfonyl)phenylamino]-2-(-4-pyridyl)-1-ethanone, for standardizing the GlmU assay. We thank Mr. Chand Raina for his help in carrying out fermentation experiments.
Declaration of Conflicting Interest
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was funded by Council of Scientific and Industrial Research Indian Council, (CSIR) New Delhi, India (Grant no. BSC0205). Authors (RS, MRL, CR) were provided Junior and Senior Research Fellowship by CSIR and UGC, India.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.