Abstract
Most noncytoplasmic bacterial proteins are exported through the SecYEG channel in the cytoplasmic membrane. This channel and its associated proteins, collectively referred to as the Sec pathway, have strong appeal as a possible antibiotic drug target, yet progress toward new drugs targeting this pathway has been slow, perhaps due partly to many researchers’ focus on a single component, the SecA ATPase. Here we report on a pathway-based screen in which beta-galactosidase (β-gal) activity is trapped in the cytoplasm of Escherichia coli cells if translocation through SecYEG is impaired. Several hit compounds passed a counterscreen distinguishing between β-gal overexpression and impaired β-gal export. However, the most extensively characterized hit gave limited E. coli growth inhibition (EC50 ≥ 400 µM), and growth inhibition could not be unambiguously linked to the compound’s effect on the Sec pathway. Our study and others underscore the challenges of finding potent druglike hits against this otherwise promising drug target.
Introduction
Bacteria of clinical concern continue to develop resistance to most antibiotics. Meanwhile, the drug development pipeline includes few antibiotic candidates with novel mechanisms of action. Particularly worrisome is our limited arsenal against the highly antibiotic-resistant ESKAPE pathogens (Enterococcus faecium, Staphylococcus aureus, Klebsiella pneumoniae, Acinetobacter baumannii, Pseudomonas aeruginosa, and Enterobacter species). 1
In searching for new drug targets, many researchers have become interested in the SecA-dependent pathway of protein export.2–4 This Sec pathway sends proteins through a cytoplasmic membrane channel—made up of subunits SecY, SecE, and SecG—in a process that involves several additional proteins (SecA, SecB, SecD, SecF, and YajC). 5 Several of the Sec proteins are essential for survival, as demonstrated by genetic knockout studies. 6 Moreover, inhibition of the Sec pathway could limit pathogens’ export of β-lactamases (which degrade penicillins and cephalosporins) and toxins (responsible for pathogenesis). Additionally, jamming of the SecYEG channel appears to cause rapid destruction of the SecY and SecE components, 7 which should exacerbate any problems in SecYEG-mediated transport and hasten cell death.
Despite these and other reasons why the Sec pathway might be an excellent drug target, 3 progress toward clinical candidates has been modest. The limited progress might be partly due to many researchers’ focus on the SecA protein 4 rather than the pathway as a whole. While SecA is an essential enzyme (an ATPase) and therefore a convenient target to study, it is not necessarily the most druggable component of the pathway. We have taken a more inclusive approach to Sec pathway drug development by developing a screening assay that should identify inhibitors of protein export by the Sec pathway as a whole, rather than inhibitors of SecA alone. 8 Our assay uses an Escherichia coli strain (EC626) that expresses a recombinant beta-galactosidase (β-gal) with a signal peptide that sends it through the SecYEG channel to the periplasm, where the oxidizing environment does not allow conformational maturation to an active enzyme. Thus, any β-gal activity seen in these cells is cytoplasmic and indicates an impairment of export through SecYEG.
Here we report on a screen of ~57,000 chemical samples and our follow-up studies of selected hits.
Materials and Methods
High-Throughput Screen
We conducted a high-throughput screen at the National Screening Laboratory for the Regional Centers of Excellence in Biodefense and Emerging Infectious Diseases (NSRB) located on the Harvard Medical School campus in Boston, MA. Our protocol was essentially as described previously
8
aside from adaptations to the high-throughput screening environment. In brief, EC626 cells suspended in 3 mL of Miller’s Lysogeny broth (LB), with 0.2% glucose (w/v) added to suppress LacZ expression, were grown overnight in 15 mL round-bottom tubes at 37 °C under vigorous aeration. The next morning, cells were diluted 1:100 into LB with 0.2% glucose (w/v) and grown in a 500 mL flask for 4 h at 37 °C with vigorous aeration. Each well of a 384-well plate then received 16 µL of washed cells resuspended in phosphate-buffered saline (PBS) and 16 µL of LB (no glucose). Each small-molecule sample (100 nL) was pin-transferred to columns 1–22 of each plate using Epson robotic machinery. The final concentrations of these samples were 16.67 µg/mL for pure compounds and 50 µg/mL for natural product extracts. Column 23 served as a negative control, receiving no compounds, while column 24 contained the positive control of sodium azide, an established inhibitor of SecA (final concentration 0.25 mM). Following the addition of test samples, plates were incubated for 30 min at 37 °C, after which each well received 8 μL of ZOB buffer (59 mM monobasic sodium phosphate, 101 mM dibasic sodium phosphate, 1.6 mM magnesium sulfate, 320 mg/L hexadecyltrimethylammonium bromide, 160 mg/L sodium deoxycholate, 1.6 mg/mL o-nitrophenyl-β-
Screening Statistics
The Z′ factor, a measure of the separation of positive and negative controls, was highly variable among the plates, ranging from 0.74 (excellent) to below 0 (very poor). To address this variability in our data, we defined compounds as hits if and only if they appeared exceptional in both duplicates of a plate, according to the following analysis. For each plate, a median absolute deviation (MAD) was calculated from A405 values for all experimental wells (not positive or negative control wells), and each well’s A405 was converted to a Z score representing the number of MADs above or below the mean. These data exhibited edge effects, with hits most likely to come from the leftmost column and the top and bottom rows of each plate. Edge effects were corrected with free HTS Helper software provided by Vladimir Makarenkov et al. 9 Well correction (WC)–adjusted Z scores were then used to rate compounds as strong hits (duplicate A405’s > 4 MADs above the average of the plate’s experimental wells), medium hits (duplicate A405’s > 3 MADs above average), weak hits (duplicate A405’s > 2.5 MADs above average), or nonhits.
Counterscreen for LacZ Overexpression
This assay was essentially identical to the screening assay except for the use of EC627 rather than EC626. EC627 is an E. coli strain identical to EC626 except that its β-gal lacks an export signal. 8 β-gal activity in EC627 thus serves as a readout of LacZ expression independent of protein export.
Growth and Cytotoxicity Assays
We tested whether the growth of a strain of K12 E. coli, MC4100, could be impaired by selected compounds. MC4100 is the parent strain of EC626 and EC627; it lacks the LamB-LacZ fusion gene possessed by EC626 and EC627. We grew cultures of MC4100 overnight, then tracked the growth of the diluted, resuspended cultures in 96-well plates over 4–6 h using a BioTek Synergy plate reader. Growth rate was quantified as the change in absorbance at 600 nm (A600) per unit time during log-phase growth. Cytotoxicity assays were also performed with human B lymphocytes (CRL-8155), as previously described. 10
Detection of MBP Export via Western Blot
Proteins exported by the SecA-mediated translocation system are recognized by a leader sequence that is cleaved from the precursor protein as it reaches the periplasmic space. Thus, the cytoplasmic precursor and the mature periplasmic product can be distinguished by their difference in molecular mass. Strain MC4100 carrying plasmid pMAL-p5X (New England BioLabs, Ipswitch, MA) was used as an indicator strain, designated EC749, to evaluate SecA-mediated export of its lactose-inducible maltose binding protein (MBP).
An overnight culture of EC749 in trypticase soy broth was diluted 10-fold into fresh medium supplemented with 0.4% glucose (w/v). After approximately 2 h (A600 = 0.5–0.8), 0.75 mL of the expanded culture was added to microcentrifuge tubes containing an equal volume of 37 °C trypticase soy broth supplemented with the test compound. After an additional 30 min of tumbling at 37 °C, proteins were precipitated by addition of 300 µL of cold 30% trichloracetic acid (w/v) and placed in an ice bath for 5 min. Precipitated protein was collected by centrifugation (2 min, 17,000g, 4 °C) and pellets were rinsed with 1 mL of cold acetone. The rinse (900 µL) was aspirated and the remainder allowed to evaporate overnight at 4 °C. Dried pellets were dissolved in electrophoresis sample buffer at a concentration equivalent to A600 = 5 (based on the starting culture) and warmed for 5 min at 70 °C prior to electrophoresis.
Proteins equivalent to an A600 of 0.01 (e.g., 2 µL of a sample with A600 = 5) were separated on 8% bis-tris plus polyacrylamide (w/v) gels (Life Technologies, Carlsbad, CA) and transferred to PVDF membranes (iBlot, Life Technologies). The recombinant MBP polypeptides were detected by Western analysis using rabbit anti-MBP (Pierce/Thermo PA1-989, 1:2000) and chromogenic detection (Life Technologies) according to the suppliers’ protocols.
Assays of Cytoplasmic and Periplasmic Marker Enzymes (G6PD and β-Lactamase)
As an alternative means of assessing export via the Sec pathway, we measured the activity of a periplasmic enzyme, β-lactamase, as described previously. 11 Activity of a cytoplasmic enzyme, glucose 6-phosphate dehydrogenase (G6PD), was quantified as a control. 11 Cultures of EC749 were grown overnight, and then diluted and regrown for 4 h in the presence of isopropyl β-D-1-thiogalactopyranoside (IPTG) (80 µM) and either DMSO alone or test compound dissolved in DMSO. Periplasmic contents were released by incubating cells with 0.2 M guanidine (which disrupts the outer membrane of Gram-negative bacteria), while cytoplasmic contents were released by incubating cells with a combination of 0.05% Triton X-100 (which disrupts the cytoplasmic membrane; v/v) and 0.2 M guanidine. These incubation periods lasted 20–30 h at 4 °C. β-Lactamase activity was then monitored via increases in A490, reflecting hydrolysis of the substrate nitrocefin. G6PD activity was tracked via increases in A340, reflecting reduction of the substrate NADP+ to NADPH. Enzyme activities were normalized to cell density readings (A600).
Thermal Shift Assays
Thermal shift assays, which detect ligand binding to a protein as an increase in the protein’s melting temperature (Tm), were performed as described previously. 12 In the presence of possible ligands, samples of purified SecA (100 µg/mL) in 96-well plates were heated from 20 to 90 °C over ~55 min by a DNA Engine Opticon 2 (manufactured by MJ Research, now part of Bio-Rad, Hercules, CA). The fluorescent probe SYPRO Orange (Invitrogen, Grand Island, NY; excitation at 530 ± 30 nm, emission at 575 ± 20 nm) was used to detect unfolding of the protein’s hydrophobic regions. Melting temperature (Tm) was defined as the temperature at which the increase in fluorescence was the steepest.
Results and Discussion
High-Throughput Screen and Hit Confirmation
One hundred seventy 384-well plates were screened in duplicate. These included 56,768 duplicate experimental wells (45,856 pure compounds and 10,912 natural product extracts). Each plate’s experimental wells’ absorbance readings at 405 nm (A405) were used to calculate a median absolute deviation (MAD) for that plate. After correction for edge effects, 9 a total of 367 wells were considered hits. These included 108 strong hits (duplicate A405’s > 4 MADs above the average of the plate’s experimental wells), 105 medium hits (duplicate A405’s > 3 MADs above average), and 154 weak hits (duplicate A405’s > 2.5 MADs above average).
We selected 211 of the 367 initial hits for confirmation assays back at the University of Washington. These included all 69 natural product extracts considered strong hits, plus all 142 pure compounds that were strong, medium, or weak hits and were considered to have acceptable medicinal chemistry potential. Ten of the 211 samples shipped from the screening center were yellow in color, thus interfering with our readout of A405, and were not studied further. When the remaining 201 samples were retested in the assay used in the primary screen, 67 of these gave A405 values well above baseline (>3 standard deviations above the negative control wells). This hit confirmation rate of 33% is within the normal range for real-life experiments. 13
Counterscreen for LacZ Overexpression
We next determined whether any of the 67 confirmed hits increased A405 simply by increasing expression of the LacZ gene and thus jamming the SecYEG channels with excess β-gal, giving higher β-gal activity. Upregulation of LacZ expression would not be detrimental to E. coli growth and thus would not be relevant to drug development or of interest to us. To see whether any of the hits cause LacZ overexpression, we tested them in a modified screening assay with strain EC627, which is identical to EC626 except that its β-gal does not include a signal sequence and thus is not exported to the periplasm. Most of the 67 samples did increase A405 in EC627, indicating an undesirable increase in LacZ expression. Five pure compounds and six natural products did not increase A405 in EC627, establishing themselves as the most promising hits ( Table 1 ).
Summary of High-Throughput Screen and Initial Follow-Up.
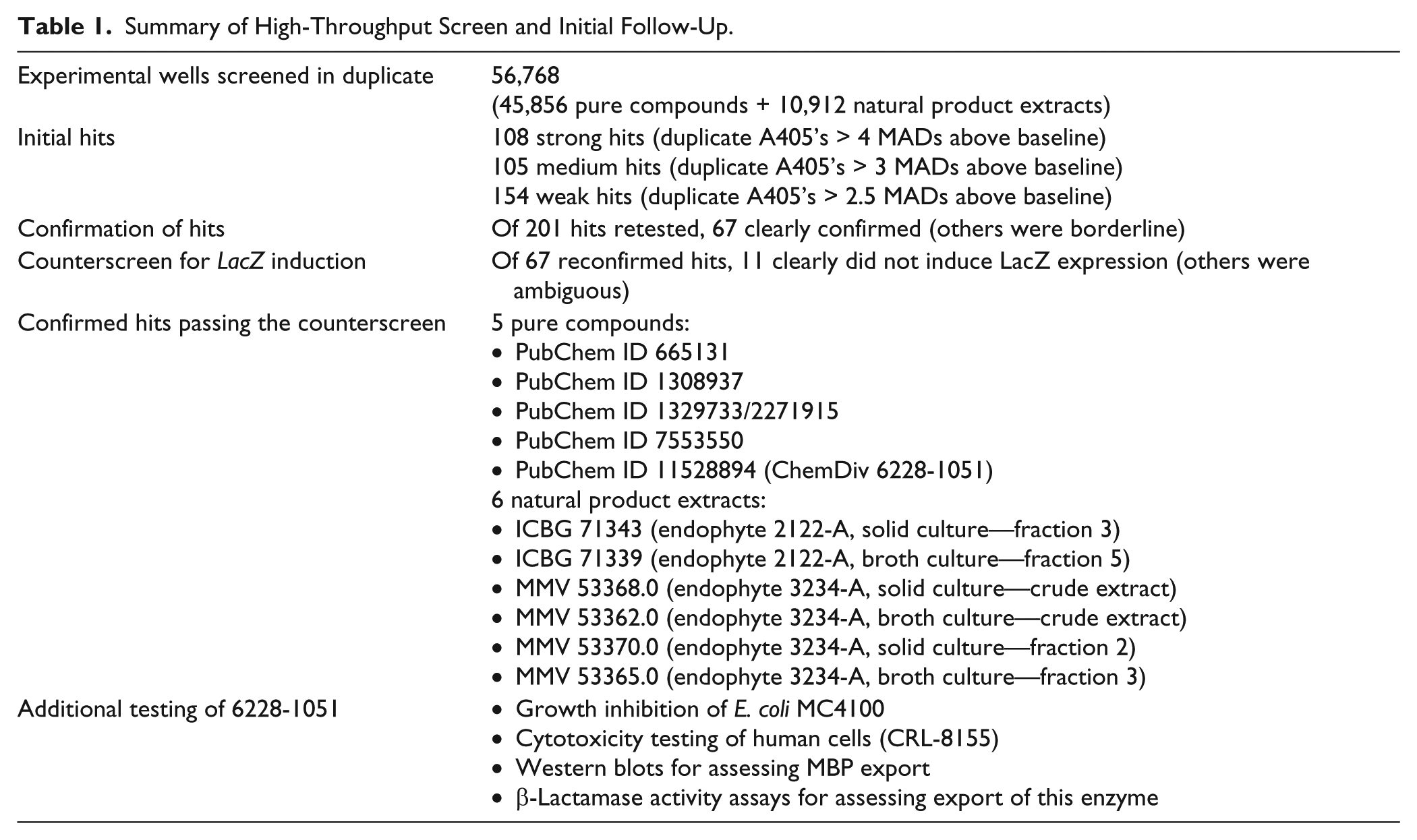

The natural product extracts were derived from two endophytes, isolate 3234-A (obtained from the petiole of a Heliconia species) and isolate 2122-A (obtained from the roots of a Rivina species). Unfortunately, these extracts—originally derived from species native to Costa Rica—were in extremely limited supply at the screening center and could not be purchased or regenerated cheaply, precluding further study.
Dose-Response Study of 6228-1051
One of the pure compounds (PubChem ID 11528894, aka ChemDiv 6228-1051) was obtained from a commercial vendor and studied in greater detail. A half-maximal effect of 6228-1051 on cytoplasmic β-gal accumulation in EC626 is evident at a concentration of ~25 μM. The compound also inhibits growth of MC4100, EC626’s parent strain, with an EC50 of ≥400 μM ( Fig. 1 ). Our screening assay cannot quantify percent reductions in β-gal export because it only reports the β-gal that is retained in the cytoplasm, not the β-gal that is successfully exported. The growth rate starts to decline at the approximate concentration that the β-gal activity is maximal. This suggests that if 6228-1051 kills E. coli cells through its effect on the Sec pathway, near-total inhibition of the pathway is needed to slow growth.
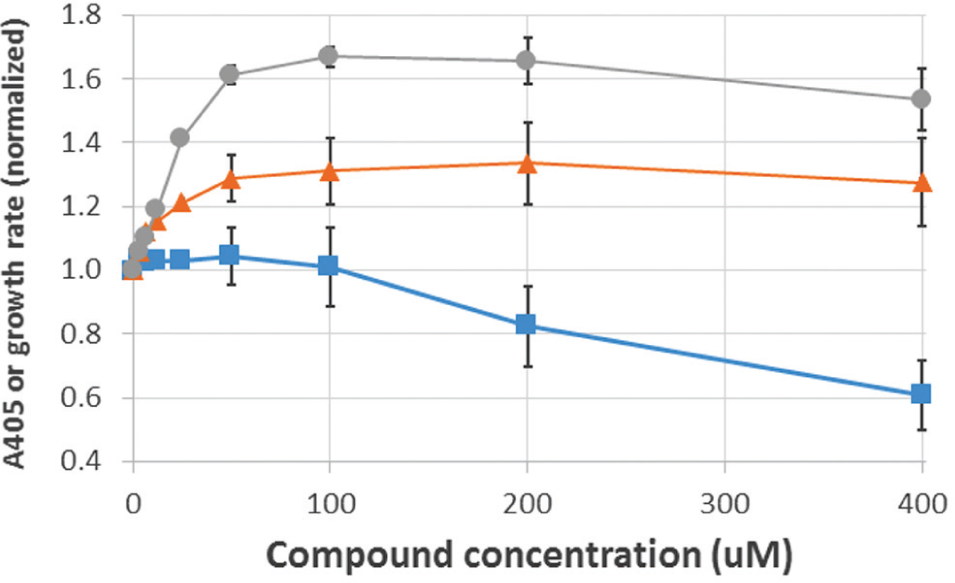
Effects of 6228-1051 on β-gal export in strain EC626 (circles) and cell growth of strain MC4100 (squares), and effects of related compound 025610 on β-gal export in strain EC626 (triangles). Each curve shows averages of 8–12 experiments, each performed on a separate day. Error bars (some omitted for clarity) represent SEMs. A405 values and growth rates (quantified as increases in A600 per unit time) are expressed relative to DMSO-only controls, which were set to 1.0.
Preliminary Hit Expansion around 6228-1051
Several close relatives of 6228-1051 are commercially available ( Fig. 2 ). We tested several of them for inhibition of β-gal export in our screening assay. To our disappointment, none was more potent than 6228-1051 in inhibiting β-gal export. Compound 025610 inhibited export somewhat less strongly than 6228-1051 ( Fig. 1 ); compounds 6228-1061, 6228-0867, and 020083 gave equivocal or no inhibition. In cytotoxicity assays of human CRL-8155 cells, 6228-1051 and 025610 gave EC50’s of 101 and 216 μM, respectively. These values, compared with the growth inhibition data in Figure 1 , do not indicate selectivity for E. coli over human cells.
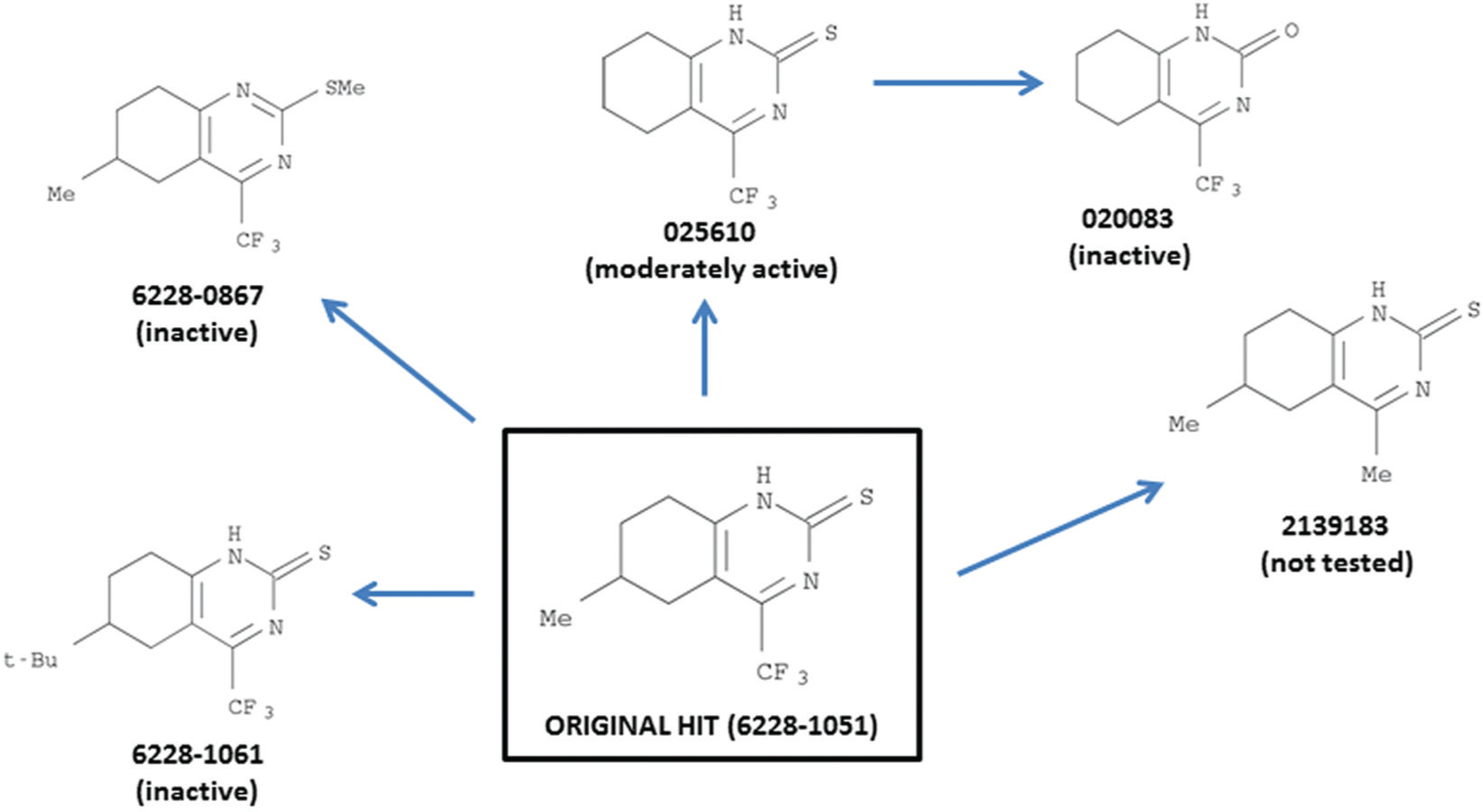
Commercially available compounds closely related to hit compound 6228-1051.
Further Exploration of 6228-1051’s Mechanism of Action
Our initial evidence that compounds like 6228-1051 target the Sec pathway was based on a readout of one particular genetically engineered protein (β-gal). To verify that 6228-1051 really does impair the Sec pathway, we tested its ability to block export of other proteins that also require the Sec pathway for proper localization in the cell.
We focused first on maltose binding protein (MBP, aka MalE). We ran Western blots of treated and untreated cell extracts and probed them with a commercially available anti-MBP antibody. Successfully exported MBP migrates faster than immature MBP (pre-MBP) due to extracytoplasmic cleavage of its 26–amino acid signal peptide, 14 a significant portion of its 396–amino acid length. We achieved reliable separation of mature MBP from pre-MBP, and the pre-MBP band appeared to intensify in cells grown in the presence of 67 µM 6228-1051 ( Fig. 3 ); however, differences were small and somewhat variable from experiment to experiment.
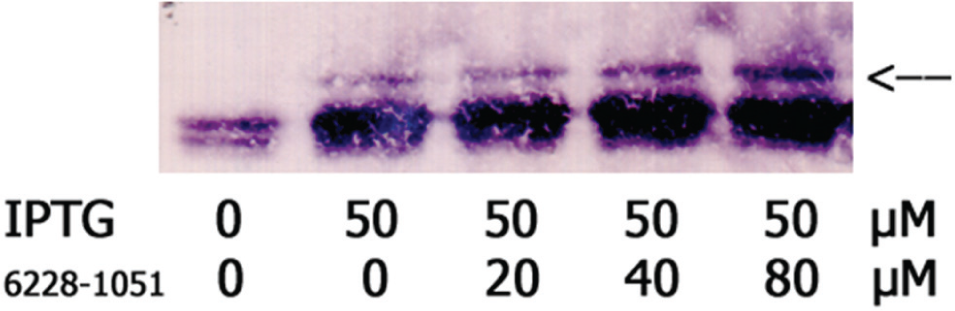
6228-1051 may impair export of MBP, according to Western blot analysis. When MBP production was induced with IPTG, a small amount of uncleaved MBP was retained in the cytoplasm. This “pre-MBP” was visible as a band of higher molecular weight than mature MBP (arrow). The fraction of cytoplasmic MBP appeared to increase in the presence of 40–80 µM 6228-1051.
As a further look at whether 6228-1051 acts specifically on the Sec pathway, we studied the periplasmic marker enzyme β-lactamase and the cytoplasmic marker enzyme glucose 6-phosphate dehydrogenase (G6PD). Our hypothesis was that, by disrupting the Sec pathway, 6228-1051 would reduce periplasmic β-lactamase activity while having no effect on cytoplasmic G6PD activity. Data on EC749 cells grown in the presence of 6228-1051 were in partial agreement with the hypothesis. Cells exposed to 67 µM 6228-1051 had only 66 ± 6% the periplasmic β-lactamase activity of control cells (mean ± standard deviation from four experiments on separate days). However, these cells also appeared to have reduced cytoplasmic G6PD activity (72% ± 16% of control, N = 4).
If 6228-1051 does indeed target the Sec pathway, its specific molecular target within that pathway remains unknown. One means of verifying compound–protein binding is via thermal shift assays. 12 6228-1051 (100 µM) did not increase the melting temperature of SecA purified from E. coli, whereas the positive control compound of ATP (1 mM) did increase SecA’s melting temperature (Tm) by 1.4–2 °C, consistent with weak binding of ATP to SecA. These data argue against the hypothesis that 6228-1051 binds to SecA.
Prospects for Drug Development
The approach reported here advances previous drug development efforts by screening for inhibitors at the level of the Sec pathway rather than the SecA protein in particular. However, the compound studied in greatest detail, 6228-1051, did not slow growth of E. coli cells except at concentrations of >100 µM, was not selective for E. coli over human cells, and operates via a still-uncertain mechanism of action, so it does not qualify as an exciting hit compound from a drug development standpoint.
Reports that severe reductions in SecY and SecE levels have only modest effects on protein export15,16 suggest that the Sec pathway’s capacity may greatly exceed cells’ needs, and that blockage of the Sec pathway must be nearly 100% for cell growth to be affected. This was supported by the fact that 6228-1051 did not reduce growth except at concentrations beyond those necessary to achieve maximal levels of β-gal cytoplasmic activity.
In retrospect, it is possible that the other 10 samples listed in Table 1 , and/or other medium and weak natural-product hits not covered by Table 1 , might have been our best hits from a drug development perspective, and that our focus on 6228-1051 was unfortunate. Alternatively, perhaps the components of the Sec pathway, like many other prospective drug targets, are difficult to perturb with small-molecule inhibitors.
Footnotes
Acknowledgements
Su Chiang and staff at the National Screening Laboratory for the Regional Centers of Excellence in Biodefense and Emerging Infectious Diseases at Harvard Medical School offered excellent support of the high-throughput screen. Thomas Silhavy and colleagues (Princeton University) provided the E. coli LamB-LacZfusion strains (EC626 and EC627). Konstantin Korotkov (University of Kentucky) cloned and purified SecA from E. coli. Ryan Choi performed growth assays with CRL-8155 cells. Jack Mo and Zhongsheng Zhang (University of Washington) assisted with biochemical assays. Lynn Barrett provided general laboratory and administrative support. Daniel Weaver provided programming assistance.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported in part by a pilot grant from the University of Washington Royalty Research Fund.
