Abstract
Alphaviruses are a prominent class of reemergent pathogens due to their globally expanding ranges, potential for lethality, and possible use as bioweapons. The absence of effective treatments for alphaviruses highlights the need for innovative strategies to identify antiviral agents. Primary screens that use noninfectious self-replicating RNAs, termed replicons, have been used to identify potential antiviral compounds for alphaviruses. Only inhibitors of viral genome replication, however, will be identified using replicons, which excludes many other druggable steps in the viral life cycle. To address this limitation, we developed a western equine encephalitis virus pseudoinfectious particle system that reproduces several crucial viral life cycle steps in addition to genome replication. We used this system to screen a library containing ~26,000 extracts derived from marine microbes, and we identified multiple bacterial strains that produce compounds with potential antiviral activity. We subsequently used pseudoinfectious particle and replicon assays in parallel to counterscreen candidate extracts, and followed antiviral activity during biochemical fractionation and purification to differentiate between inhibitors of viral entry and genome replication. This novel process led to the isolation of a known alphavirus entry inhibitor, bafilomycin, thereby validating the approach for the screening and identification of potential antiviral compounds.
Introduction
Mosquito-borne viruses (arboviruses) exemplify an important class of reemergent pathogens. Approximately 10 arboviruses belong to the Alphavirus genus within the Togaviridae family and cause disease in humans and animals, 1 several of which are listed by the National Institute of Allergy and Infectious Diseases as priority pathogens due to their reemerging status and potential use as biological weapons. These alphaviruses include chikungunya virus and western, eastern, and Venezuelan equine encephalitis viruses (WEEV, EEEV, and VEEV, respectively). WEEV, EEEV, and VEEV infect neurons and cause severe inflammation of the CNS, with mortality rates as high as 70% and long-term neurological sequelae often occurring in survivors.
The lack of approved therapeutics for potentially devastating alphavirus infections highlights the need for innovative approaches that reach beyond the constricted chemical boundaries of small-molecule libraries to discover new antiviral agents. Libraries consisting of natural product extracts take advantage of the diverse and complex biosynthetic pathways of living organisms, which can produce virtually limitless structural diversity that greatly exceeds the diversity in even the largest small-molecule libraries. 2 Equally important as library selection is the development of robust assays that are amenable to high-throughput screening (HTS) and duplicate most of the viral life cycle, but do not produce the secondary infectious virus particles that would require enhanced biocontainment facilities. Furthermore, the use of innovative counterscreening strategies that focus on specific biological activities can increase the probability of successful discovery and development of antivirals against alphaviruses.
Plasmid-based viral replicon reporter systems have been used effectively to screen libraries and identify candidate antivirals that inhibit viral genome replication. Indeed, we have used WEEV replicons for the discovery of several viral RNA replication inhibitors.3–5 Despite these successes, there are disadvantages to plasmid-based replicon screening systems, including labor-intensive procedures, transfection variability, and, most importantly, the limited scope of discovery to viral genome replication inhibitors, because replicons do not reproduce the entire viral life cycle. To address these limitations, pseudoinfectious particle (PIP) systems have been developed for antiviral screening. 6 PIPs are replicons packaged into viral particles that can infect and initiate replication in target cells, but do not produce secondary infectious particles. Thus, in contrast to plasmid-based replicons, reporter gene expression with PIPs is a function of successful viral binding, entry, envelope fusion, and genome replication.
We have developed and optimized a novel chimeric WEEV–Sindbis virus (WEEV-SINV) PIP system, and in this article we describe the use of this system to screen an extensive library of marine microbe–derived extracts to identify multiple bacterial strains that produce compounds with antiviral activity. We used PIPs in parallel with a plasmid-based WEEV replicon assay to counterscreen extracts and guide fractionation, thus differentiating potential viral entry inhibitors from genome replication inhibitors. This process led to the isolation of bafilomycin, a known inhibitor of alphavirus entry, 7 as an active molecule in an extract derived from a novel Streptomyces species recovered from off the coast of Costa Rica. These results provide proof-of-concept that PIP-based primary screens followed by parallel PIP–replicon counterscreens can effectively identify and differentiate between candidate antiviral compounds that target defined steps in the viral life cycle.
Materials and Methods
Plasmids and Inhibitors
The WEEV replicon plasmid, pWR-LUC, has been previously described. 4 Mycophenolic acid (MPA) and bafilomycin were purchased from Sigma-Aldrich (St. Louis, MO).
Cell Culture
Baby hamster kidney (BHK-21) cells were purchased from the American Type Culture Collection (Manassas, VA). BSR-T7 and HEK293 cells have been previously described.4,5 Cells were cultured in complete Dulbecco’s Modified Eagle Medium (cDMEM) containing 5% bovine growth serum or fetal bovine serum, 1% sodium pyruvate, 0.1 mM non-essential amino acids, 10 U/ml penicillin, and 10 µg/ml streptomycin.
Cell Viability and Virus Replication Assays
Cell viability was determined by 3-[4,5-dimethylthiazol-2-yl]-2,5-diphenyltetrazolium bromide (MTT) assay as previously described.4,5 Firefly luciferase (fLUC) accumulation was measured as a surrogate marker for viral RNA replication as previously described. 4
Pseudoinfectious Particle Production
We generated chimeric WEEV-SINV PIPs in BHK-21 cells using a three-component system as previously described. 3
High-Throughput Screening
Subconfluent BHK-21 cells grown in 10 cm2 tissue culture dishes were detached with 0.5% trypsin, then diluted to a concentration of 4×105 cells per milliliter, and 25 µl of cell suspension per well was dispensed into 384-well plates. Individual bacterial extracts (0.5% v/v in DMSO) were added to single wells with a Biomek Pintool (Beckman Coulter, Indianapolis, IN), and all plates contained a series of 32 wells each of negative and positive controls, which consisted of vehicle (DMSO) and 50 µM MPA, respectively. We added 25 µl of predetermined optimal PIP dilution in cDMEM per well, and plates were cultured at 37 °C in 5% CO2. After 24 h incubation, 40 µl media was removed and replaced with 10 µl per well of Steady-Lite luciferase reagent (PerkinElmer, Waltham, MA), and luminescence was read on a PHERAstar multimode plate reader (BMG Labtech, Ortenberg, Germany).
Marine Microbe Culture, Fermentation, Extract Preparation, and Fractionation
The extract library was derived from a collection of Actinomycetes isolated from marine sediments. Isolation was performed by sprinkling sediment onto a mixed cellulose filter (Millipore HAWP0900; Millipore, Billerica, MA) on top of a Bennett’s agar plate containing 25 µg/ml of cycloheximide and 25 µg/ml of nalidixic acid. The plate was incubated at 28 °C for 3 days before the filter was removed, after which the plate was incubated for another 7–10 days until colonies were visible; the colonies were picked off the plate and streaked onto International Streptomyces Project Medium 2 (ISP2) agar until pure. Extracts were prepared for HTS by starting seed cultures in 17 ml dual position cap tubes containing 2 ml of ISP2 media and grown for 4 days on a rotary shaker at 200 rpm. The seed culture was then poured into a 250 ml baffled flask containing 100 ml of ISP2 and grown for 18 days on a rotary shaker at 200 rpm. The culture was centrifuged at 4000 rpm for 10 min to remove cells, and 2 g of Amberlite XAD16 resin (Sigma-Aldrich) contained within a polypropylene mesh bag was subsequently added to the broth and incubated overnight on the rotary shaker. The resin bag was removed and placed into 10 ml of methanol, followed sequentially by equal volumes of acetone and ethyl acetate. Each of the three individual prefractionated crude extracts was evaporated and reconstituted to a final concentration of 15 mg/ml in DMSO.
For confirmation assays, an oatmeal plate was streaked from a glycerol spore stock of the active Actinomycetes strain and incubated for 5 days. Seed cultures of 3 ml of ISP2 media were inoculated with a loop full of vegetative cells from oatmeal plate cultures and incubated with shaking (200 rpm) at 28 °C for 5 days, then transferred to 100 ml cultures incubated under the same conditions for an additional 7–28 days, based on the original incubation period for the HTS library sample. After incubation, bacteria were pelleted by centrifugation, and the resulting cell-free broth was subjected to solid phase extraction using 2 g of XAD-16 resin. The resin was subsequently separated by filtration and subjected to organic extraction using methanol and methanol–ethyl acetate (1:1). Extracts were concentrated under vacuum and dissolved in DMSO for validation in virus replication assays.
For large-scale fermentation of Streptomyces strain 7756-H5, a 25 ml seed culture was transferred to a 2.8 L Fernbach flask containing 1.5 L of ISP2 medium and incubated on a rotary shaker (200 rpm) at 28 °C for 21 days. The culture was harvested by centrifugation, and the resulting cell-free broth was subjected to solid phase extraction as described above using 20 g resin per liter of culture. The resin was subsequently separated by filtration and subjected to methanol extraction twice and methanol–ethyl acetate (1:1) once to yield 5 g of dried crude extract. The crude extract was dissolved in 500 ml of water and was subjected to organic phase separation using 500 ml ethyl acetate three times. The organic phase extract (~600 mg dried material) was selected for further purification based on antiviral activity, and it was initially separated on a Phenomenex Luna C18 high-performance liquid chromatography (HPLC) column using an acetonitrile–water gradient (10:90 → 100:0) to isolate six fractions (A–F). Fraction D was selected based on antiviral activity for further separation using HPLC with a Waters XBridge Column (Waters, Milford, MA) and acetonitrile–water (77:23) elution to isolate seven fractions (D1–D8). Fraction D6 (3.6 mg) was highly active in the WEEV-SINV PIP assay, contained a pure compound based on liquid chromatography–mass spectrometry (LC-MS) analysis, and was selected for molecular structure elucidation by nuclear magnetic resonance (NMR), as described. 5
Results and Discussion
Development of WEEV Pseudoinfectious Particles (PIPs) as a Platform for HTS and Validation of Antiviral Compounds Derived from Marine Microbes
We previously developed and used a phenotypic cell-based WEEV replicon assay to successfully identify inhibitors of viral genome replication from both small-molecule and natural-product libraries.4,5 The WEEV genome, like all alphaviruses, consists of a positive-strand (+) RNA that serves directly as the viral mRNA for host translation of the viral nonstructural proteins, which are subsequently responsible for viral genome replication. Viral replication produces a full-length antigenomic negative-strand (−) RNA intermediate that serves as a template for the production of the full-length genomic (+) RNA, in addition to a subgenomic RNA that encodes the structural capsid protein and envelope glycoproteins. The structural proteins are dispensable for genome replication, and we previously replaced the WEEV structural genes with fLUC to generate the replicon, pWR-LUC
4
(
To reduce the potential variability associated with replicon transfections, reproduce steps in the WEEV life cycle in addition to genome replication, and maintain lower biosafety level requirements than those required for fully infectious encephalitic alphaviruses, we developed and validated WEEV PIPs.
3
Whereas previous groups have reported the production of alphavirus PIPs for SINV
8
and VEEV,
9
these studies have primarily explored basic virology questions or the development of gene vectors for next-generation vaccines, whereas our focus is on drug discovery. We generated a three-vector system using the SINV capsid and envelope proteins encoded on separate plasmids to produce chimeric WEEV-SINV PIPs when mRNAs derived from these plasmids were co-transfected with the WEEV LUC replicon into producer BHK-21 cells (
We optimized the WEEV-SINV PIP system with BHK-21 cells in a 384-well format (Z’ = 0.55), and we subsequently performed an HTS assay using an extensive collection of ~26,000 extracts derived from marine microbes recovered from diverse geographical locations. Secreted metabolites were adsorbed onto resins, and sequential organic solvent extraction was used to prepare two or three prefractionated crude extracts from individual microbial isolates, as previously described. 11 Table 1 provides a composite overview of the experimental systems, assay formats, criteria, and results from the initial HTS and subsequent primary and secondary validation steps. For the HTS, we eliminated extracts without notable suppression of fLUC activity compared to DMSO controls. We also excluded extracts suspected to contain direct luciferase inhibitors based on previous fLUC-based HTS assays performed at the University of Michigan Center for Chemical Genomics to limit false-positive results. For primary validation, we repeated WEEV-SINV PIP assays on 1245 extracts and also examined cell viability to exclude cytotoxic extracts. Furthermore, we selected only those strains with at least two active extracts, and eventually identified 334 extracts derived from 140 individual strains with antiviral activity against PIPs. For secondary validation, we repeated WEEV-SINV PIP and viability assays, but also included a WEEV replicon assay in parallel to help differentiate between inhibitors of viral entry and genome replication. This novel approach resulted in the identification of 214 active extracts from 88 strains, of which 31 strains suppressed fLUC activity in both PIP and replicon assays, potentially representing viral genome replication inhibition, whereas 57 strains suppressed fLUC activity only in the PIP assay, potentially representing viral binding or entry inhibition.
Identification and Validation Steps in the Discovery of Marine Microbe–Derived Extracts Containing Antiviral Activity.
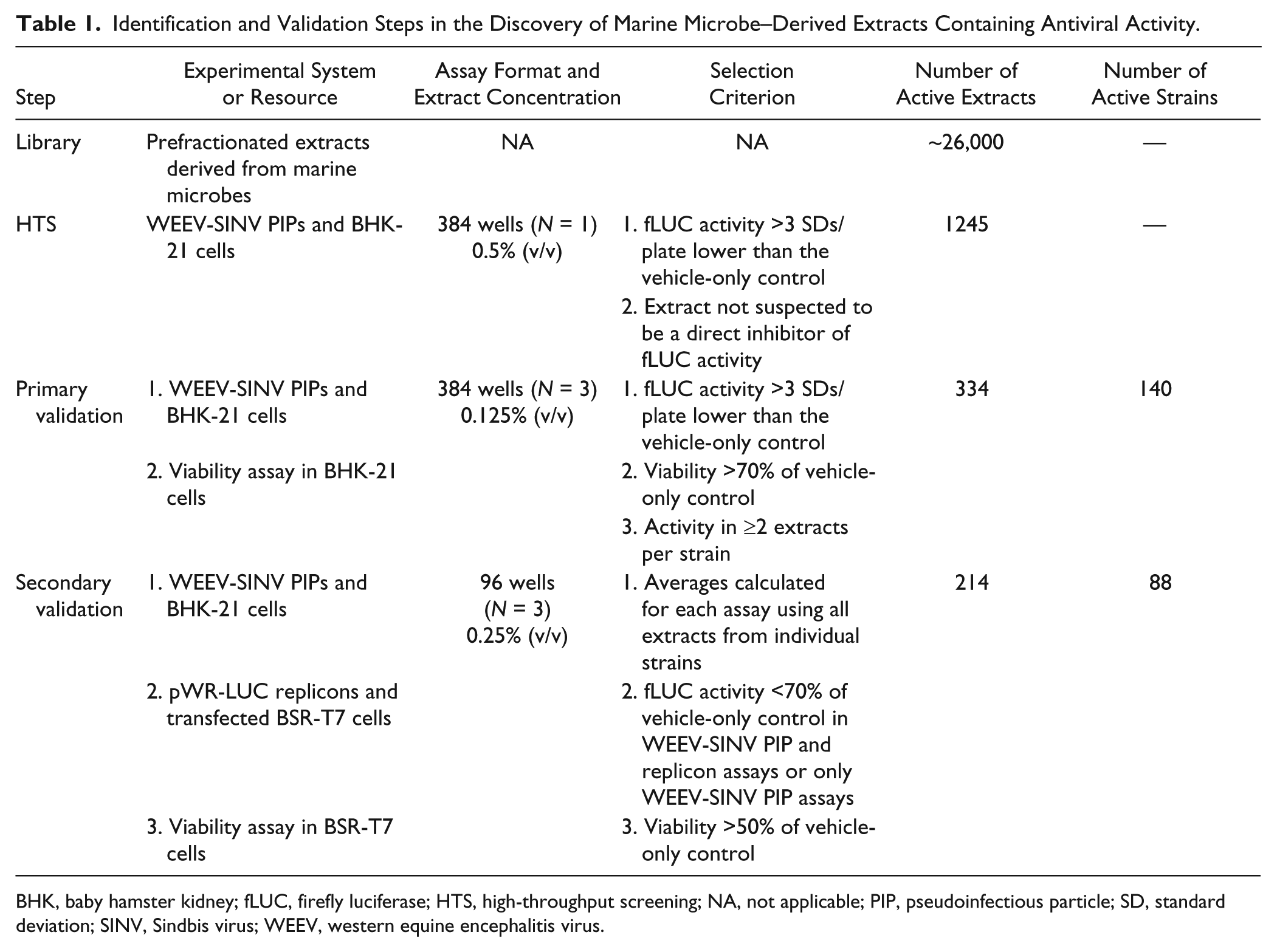
BHK, baby hamster kidney; fLUC, firefly luciferase; HTS, high-throughput screening; NA, not applicable; PIP, pseudoinfectious particle; SD, standard deviation; SINV, Sindbis virus; WEEV, western equine encephalitis virus.
We initially chose 11 microbial strains whose extracts displayed inhibitory activity in both PIP and replicon assays for repeat culture, extract preparation, and tertiary validation to ensure the antiviral activity of their metabolites. Weight-based concentrations that produced a 50% inhibition of WEEV replicons or PIPs (IC50), 50% cytotoxicity (CC50), and selectivity indices (CC50/IC50) for the extracts derived from these 11 strains are shown in
Table 2
. The extracts from four strains showed minimal toxicity, in which three retained potent activity against both replicons and PIPs (7756-H5, 32231-A2, and 24835-H1), whereas one retained activity against PIPs only (07-353-H2). The extracts from the remaining seven strains either showed substantial toxicity or had minimal activity [selectivity index (SI) <10] in replicon or PIP assays. We chose strain 7756-H5 for the subsequent analyses reported herein, because the extract from this strain had low toxicity, had the most potent antiviral activity in WEEV-SINV PIP assays, and also maintained activity in WEEV replicon assays. Strain 7756-H5 was obtained from marine sediment off the coast of Playa Grande in Costa Rica, and on the basis of 16S rRNA sequence and phylogenetic analysis, it is in the Streptomyces genus and similar to three known strains: Streptomyces spp. VAN21, DS3024, and DS21 (
Tertiary Validation of Marine Microbe–Derived Extract Antiviral Activity in WEEV Replicon and PIP Assays.
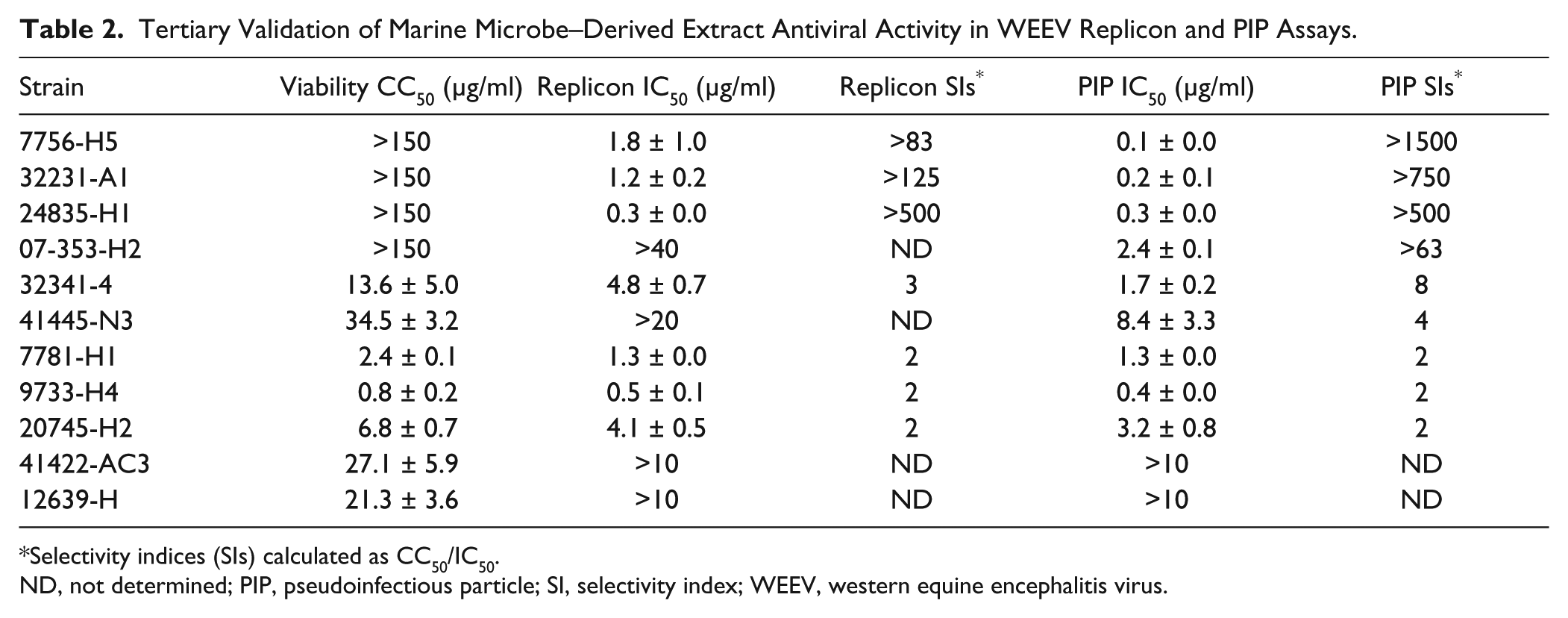
Selectivity indices (SIs) calculated as CC50/IC50.
ND, not determined; PIP, pseudoinfectious particle; SI, selectivity index; WEEV, western equine encephalitis virus.
Isolation of a Purified Antiviral Compound Derived from the Extract of Strain 7756-H5
The crude natural product extract derived from Streptomyces strain 7756-H5 was likely a complex mixture of secreted microbial secondary metabolites comprising an unknown number of diverse chemical entities, including compounds potentially active against distinct steps in the viral life cycle. The use of microbial extracts as source material for drug discovery has a potentially high yield due to the vast chemical space that can be explored with natural products, 2 although it does add additional levels of complexity to drug discovery and development. To partially mitigate this complexity, we used both PIP and replicon assays in parallel to differentiate antiviral activity that likely targeted viral genome replication (in which inhibition would be evident in both assays) and antiviral activity that targeted virus attachment and entry (in which inhibition would be evident with PIPs only). This led to the development of an empiric, three-step separation procedure that was guided by antiviral activity in both WEEV-SINV PIP and WEEV replicon assays at each fractionation step to isolate the active antiviral compounds from the crude extract of strain 7756-H5. Furthermore, we used weight-based concentrations and completed dose–titration curves at each step to follow potency.
The first separation step was an organic solvent extraction, in which the majority of antiviral activity with both assays separated into the organic phase ( Fig. 1A ). The organic extract was subsequently separated by C18 HPLC in the second step to yield six fractions, termed A through F, in which fraction D exhibited the most activity against both WEEV-SINV PIPs and WEEV replicons. We chose fraction D for further separation because it still contained potent inhibitory activity in both assays. The third and final step was an additional HPLC-based separation that yielded eight fractions, termed D1 through D8. Fraction D6 contained a pure compound that retained potent antiviral activity in the WEEV-SINV PIP assay ( Fig. 1A ), but had no significant activity against WEEV replicons ( Fig. 1B ). Efforts to isolate additional compounds with retained inhibitory activity in the replicon assay at the third separation step were unsuccessful (data not shown). When we calculated replicon and PIP IC50 values for all separation procedure steps, there was an approximate 40-fold increase in potency in the PIP assay from crude extract to fraction D. Unexpectedly, there was a small decrease in potency at the last separation step, when the pure compound in fraction D6 had a small but statistically significant (P < 0.01) increase in IC50 value compared to the less pure fraction D ( Fig. 1A ), suggesting possible chemical instability with the purified compound. We also conducted confirmatory tests in HEK293 cells and found that fraction D6 retained antiviral activity against WEEV-SINV PIPs in human cells, albeit with a fivefold increase in IC50 value (0.029 ± 0.001 µg/ml) compared to its activity in BHK-21 cells (see Fig. 1A ). Nevertheless, the robust nature of using parallel PIP and replicon assays during the bioassay-guided fractionation process, and the retention of potent activity against WEEV-SINV PIPs but not WEEV replicons in fraction D6, suggested that we had successfully isolated an antiviral compound produced by Streptomyces strain 7756-H5 that targeted viral binding, entry, or fusion.
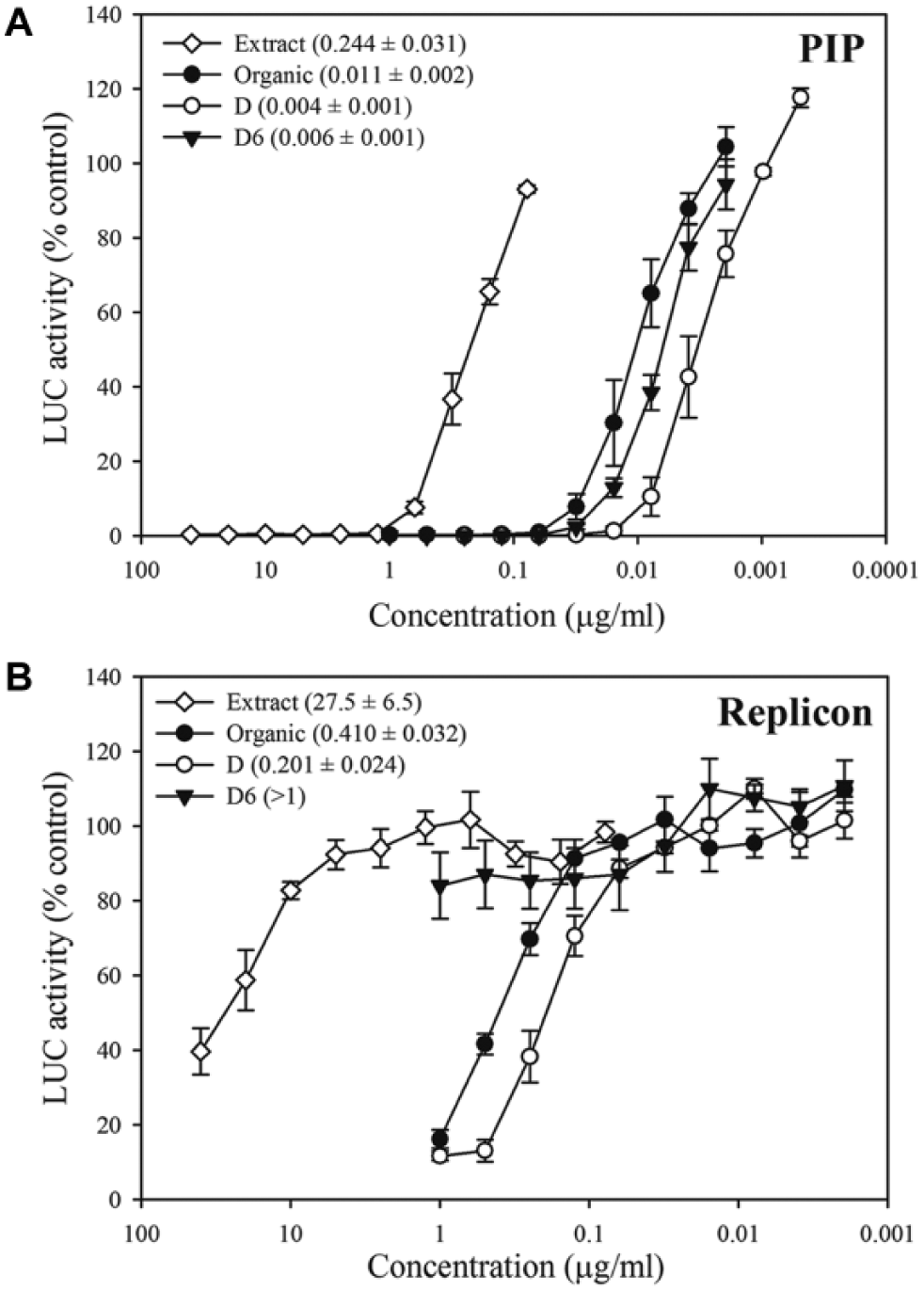
Sequential fractionation and purification of an antiviral compound derived from Streptomyces strain 7756-H5. (
Antiviral Compound Isolated from Streptomyces Strain 7756-H5 Is a Bafilomycin Analog
Fraction D6 was pure based on high-resolution LC-MS analyses, in which the primary component had a molecular weight (MW) of 436.28 g/mol, which corresponded to a WEEV-SINV PIP assay IC50 of 14 nM. We used an array of 1D and 2D NMR techniques to determine the core structure of fraction D6, and we verified the primary component as a bafilomycin analog. Bafilomycins are a family of structurally related macrolide antibiotics initially isolated from Streptomyces griseus in the 1980s. 12 The core bafilomycin structure consists of a 16-membered lactone ring, in which different family members have varied R groups ( Fig. 2A ). Due to the absence of sufficient material and the presence of three unassigned signals in the proton and carbon NMR, we were unable to definitively identify the bafilomycin analog in fraction D6. However, the MW of the primary component in fraction D6 corresponded exactly to the bafilomycin core with a single hydrogen atom as the R group ( Fig. 2A ).
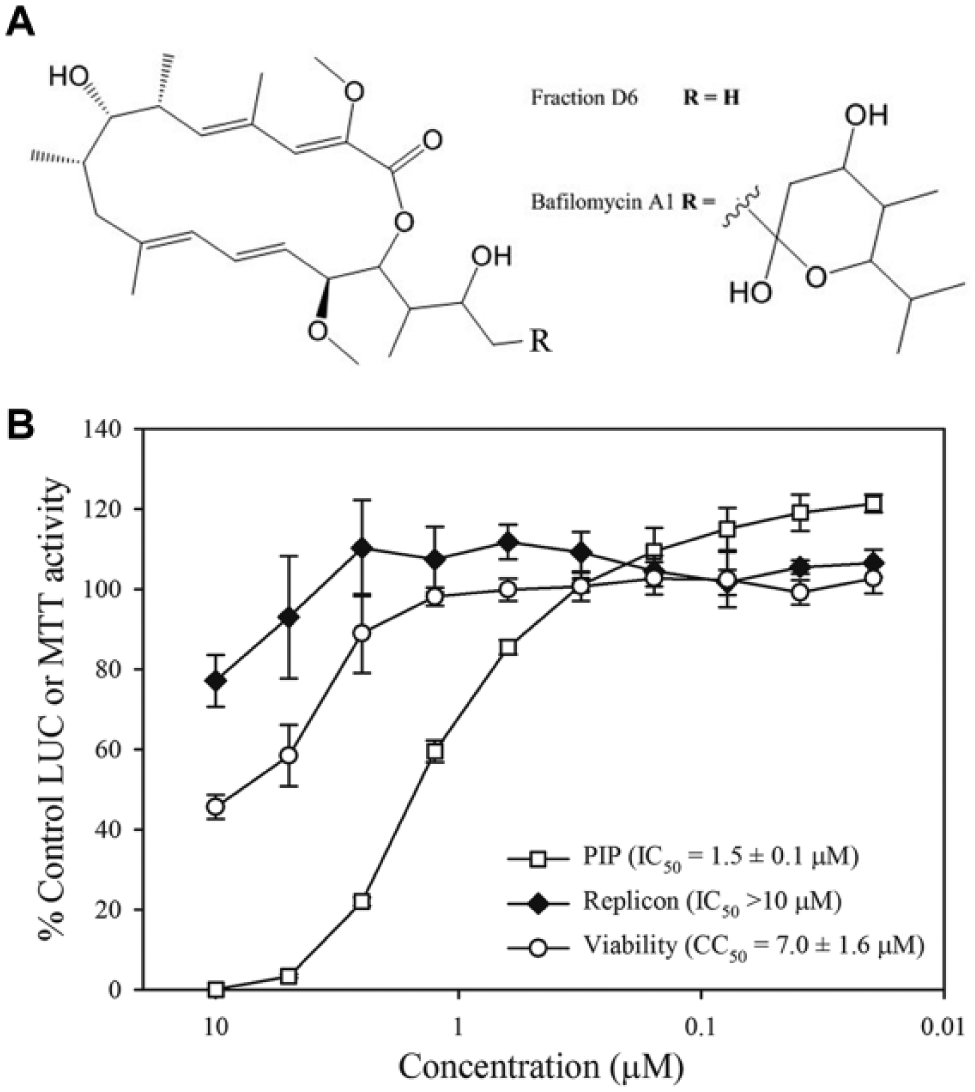
Commercial bafilomycin A1 inhibits western equine encephalitis virus–Sindbis virus (WEEV-SINV) pseudoinfectious particles (PIPs) but not WEEV replicons. (
Bafilomycins are potent and specific inhibitors of vacuolar-type ATPases in eukaryotic cells, and they produce an increased pH in intracellular organelles such as lysosomes and endosomes. 13 Alphaviruses contain a lipid envelope and require the low pH of the cellular endocytic pathway for fusion of the viral membrane with the host endosome or lysosome membranes to initiate genome replication. Consequently, bafilomycins have been shown to potently inhibit the replication of several alphaviruses by disrupting viral entry, and they are active against highly pathogenic encephalitic alphaviruses.7,14 To verify that bafilomycins inhibit WEEV entry but not genome replication in our system, we examined the activity of an authentic bafilomycin A1 standard (Sigma B1793; Sigma-Aldrich) in both WEEV-SINV PIP and WEEV replicon assays ( Fig. 2B ). Commercial bafilomycin A1 potently inhibited PIPs with an IC50 of 1.5 µM, but showed minimal activity in the replicon assay. The PIP assay IC50 for commercial bafilomycin A1 was substantially higher than the value obtained for strain 7756-H5 fraction D6 (14 nM). Because, however, we were able to elucidate only the core molecular structure of fraction D6, a full and direct comparison of potencies was not possible. Nevertheless, the fortuitous isolation of bafilomycin validated our innovative approach and indicated that a primary HTS with WEEV-SINV PIPs followed by parallel validation steps with PIPs and replicons can facilitate the identification of alphavirus inhibitors that disrupt viral binding, entry, or fusion.
Although the WEEV-SINV PIP system described in this report expands the viral life cycle steps that are interrogated compared to the plasmid-based replicon system, the PIP system does not reproduce every feature of alphavirus replication within cells. For example, the capsid protein is expressed in producer cells during PIP production, but the subgenomic mRNA encoding the structural proteins is not amplified in target cells, and therefore the overall level of capsid expression is likely negligible compared to that of cells infected with wild-type virus. This is important because the alphavirus capsid protein functions as an autocatalytic protease, encapsidates the viral genome, and, in encephalitic alphaviruses such as WEEV, also inhibits the host’s transcription. 15 Consequently, several potentially druggable capsid-mediated events in the alphavirus life cycle are not evaluated with a WEEV-SINV PIP assay. Furthermore, the use of chimeric PIPs containing SINV envelope glycoproteins may lead to the identification of SINV-specific inhibitors. Multiple common replication mechanisms are shared by alphaviruses with low (e.g., SINV) and high (e.g., WEEV, VEEV, and EEEV) pathogenic potential, however, and the isolation of bafilomycin, which is active against both groups of alphaviruses,7,14 provides proof-of-concept that WEEV-SINV PIPs are a viable platform for the discovery of candidate antivirals against encephalitic alphaviruses.
The increase in viral life cycle steps examined with WEEV-SINV PIPs does present potential additional difficulties with downstream evaluation of molecular targets and modes of action, which often represents a formidable hurdle with phenotypic cell-based drug discovery platforms. One approach to narrow the spectrum of candidate modes of action is to use related assays in parallel to eliminate or highlight candidate compounds of interest. We used this approach with WEEV-SINV PIPs and WEEV replicons, for which parallel assays can potentially differentiate genome replication inhibitors, which would be active in both assays, from viral-binding, entry, or fusion inhibitors, which would be active with PIPs only. Indeed, after the initial HTS, primary, and secondary validation using a large and diverse natural product library as starting material, we identified 31 bacterial strains that produced putative genome replication inhibitors and 57 strains that produced putative entry inhibitors, and we successfully isolated and identified a known inhibitor of alphavirus entry, bafilomycin. Our development and validation of a WEEV-SINV PIP assay for drug discovery, along with the novel approach of using both PIP and replicon assay during validation screens, will now allow the further exploration of chemical libraries in the search for novel alphavirus inhibitors.
Footnotes
Acknowledgements
We thank Kenneth T. Krill for his helpful comments on the manuscript and Steven Swaney for technical assistance. We thank the Costa Rican government for the collection permit (R-CM-INBio-03-2006-OT) to obtain the environmental samples at the Tempisque Conservation Area.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to this research and article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was funded by the National Institute of Allergy and Infectious Diseases grants U54 AI057153 (Region V “Great Lakes” Regional Center of Excellence for Biodefense and Emerging Infectious Diseases Research Career Development and Developmental Project Awards; to DJM) and R21 AI093642 (to DJM), the International Cooperative Biodiversity Groups Initiative (U01 TW007404) at the Fogarty International Center (to DHS and GTC), and the Hans W. Vahlteich Professorship (to DHS). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
