Abstract
To obtain natural or “me-better” antibodies (e.g., affinity-maturated antibodies), phage display libraries are widely used. However, the likelihood of obtaining satisfactory antibodies depends on the library content. Here, we used computer-aided design to model the use of the LoxP511 site as a linker between the heavy and light variable domains of an antibody for construction of a large single-chain fragment (scFv) antibody phage library by using the Cre/LoxP recombinant system. Then, we constructed two novel scFvs based on 2C4, namely, AH_scFv15 (15 amino acid [aa] linker; common [SG4]3 sequence) and AH_scFv21 (21-aa linker; LoxP511 sequence), to verify the use of the LoxP511 site as a linker. Our results indicate that LoxP511 could be used effectively for the construction of a large (e.g., 5 × 1012) phage display library of scFv antibodies from which it was possible to isolate an antibody with the same epitope as 2C4 but with higher affinity.
Introduction
An antibody’s potency depends on several factors, such as its affinity and stability, and an optimized antibody may demonstrate improved pharmacokinetics, a better safety profile, and lower toxicity as well as enable reduced dosing and incur a reduced cost of therapy. 1 In many well-studied cases, increased antibody affinity has been shown to correspond to improved biological efficacy. 2 However, human antibodies identified from antibody phage-display libraries are often unsatisfactory, as it is usually not possible to obtain large libraries. In addition, humanized antibodies from hybridomas often need to be optimized by phage display methods for therapeutic applications, for which large phage libraries are necessary. Large phage libraries require repeated cycles of cloning, which is tedious and a waste of time. Therefore, we hypothesized that aspects of the Cre/LoxP recombinant system, 3 in which DNA fragments located between LoxP and LoxP511 sites from two different vectors recombine to generate exponential numbers of new gene sequences, might be used to construct large phage libraries.
The single-chain variable fragment (scFv) is a small antibody fragment that contains a complete antigen-binding site and therefore maintains the binding specificity and affinity of the whole antibody. It consists of the covalently joined variable regions of the heavy and light chains (VH and VL, respectively) of immunoglobulins. To stabilize the association of VH and VL, a short polypeptide linker is present to connect the two domains between the carboxyl terminus of one domain and the amino terminus of the other domain. 4 The linker peptide generally contains glycine and serine residues to provide flexibility and protease resistance. When the linker is longer than 12 residues, VH and VL are able to form an scFv, which usually possesses an affinity similar to that of the parent antibody. 5 The small size of the linkers, together with their ability to produce functional scFvs in bacteria, has made them an interesting tool for protein engineering, with potential therapeutic or immunodiagnostic use. Since Yu et al. 6 successfully constructed a large (1.35 × 1010) scFv library by using a 21-amino acid (aa) linker, it should be feasible to use a 21-aa linker when constructing a large phage display library.
The HER family of receptor tyrosine kinases consists of four members: EGFR/HER1/ErbB-1, HER2/ErbB-2/Neu, HER3/ErbB-3, and HER4/ErbB-4. They contain four extracellular domains (ECDs): domain I, II, III, and IV. Binding of ligands leads to the homo- or heterodimer formation of the receptor tyrosine kinases, 7 which is essential for HER pathway activation via downstream-signaling molecules such as mitogen-activated protein kinase and phosphatidylinositol 3-kinase (PI3K)/AKT.8,9
HER2 overexpression in breast cancer is correlated with increased tumor growth rates, high metastatic potential, and poor long-term survival rates for the patient. 10 A number of therapeutic approaches are being developed to block the effects of HER2 overexpression, 11 including the use of the humanized monoclonal antibody pertuzumab (2C4). 12 2C4 is a therapeutic inhibitor of HER dimerization that acts by targeting the dimerization site in domain II, thus preventing activation of the HER signaling pathways, which mediate cancer cell proliferation and survival.13,14 Moreover, it is logical to target the dimerization domain of HER2 because HER2 is the most common receptor found in HER dimer combinations. 9
In this study, we examined, both theoretically and experimentally, the use of the Loxp511 site as a linker between VH and VL. Computer-aided design was used to model two scFv antibodies, namely, AH_scFv15 and AH_scFv21, that were based on the anti-Her2 therapeutic antibody 2C4 but with the commonly used (SG4)3 linker (15 aa long) and the LoxP511 site (21 aa) as the linker sequence, respectively. In addition, a series of HER2 ECD mutants expressed in 293T cells was used to analyze epitope binding of both scFvs by confocal microscopy and flow cytometry. Then, a large phage library based on the sequence of 2C4 was constructed by LoxP/LoxP511 recombination, from which an optimized antibody was isolated. Our data indicate that the LoxP511 site can be used effectively as a linker when constructing large scFv libraries for easier selection of better/optimized antibodies using the Cre/LoxP recombinant system.
Materials and Methods
Materials
The pET-32a(+) vector was obtained from Promega (Madison, WI); isopropyl-β-
Cell Lines and Culture
The human ovarian cancer cell line SKOV3 (which express HER2) and the human renal epithelial cell line 293T were obtained from ATCC. The cells were cultivated in DMEM (high glucose) supplemented with 100 U/mL penicillin, 100 U/mL streptomycin, and 10% fetal bovine serum (FBS). All of the cells were incubated in a humidified incubator (Thermo, Waltham, MA) at 37 °C with 5% CO2.
Computer-Guided Homology Modeling and Molecular Docking
The model of the interaction between the HER2 ectodomain and 2C4 was designed using the three-dimensional (3D) crystal structures deposited in the Protein Data Bank (PDB codes 1S78 and 1L7I, respectively). 15 Under CVFF and CHARMM force fields, the structure was relaxed for 40,000 steps using the steepest descent minimization algorithm.
Construction of Expression Plasmids
DNA encoding the 2C4 VH and VL was synthesized using specific primers and overlap PCR. Then AH_scFv15 and AH_scFv21 were synthesized via overlap PCR using (SG4)3 (15 aa) and LoxP511 (21 aa; used in the Cre-LoxP system for recombination; aa sequence: SGGSTITSYNVYYTKLSSSGT)3,16 as the linker sequences, respectively. The PCR products were electrophoresed using 1.2% agarose gels and then subcloned into the double-digested pET-32a(+) vector to construct the expression plasmids AH_scFv15_P32 and AH_scFv21_P32.
To generate the HER2 ECD mutants, four point-mutated (alanine replacement) HER2 genes were subcloned into double-digested pEGFP-N1 vector by overlap PCR, enzymatic digestion, and ligation. The resulting vectors were then expressed in 293T cells.
Prokaryotic Expression and Purification of Recombinant Proteins
Escherichia coli Rosetta cells were transfected with AH_scFv15_P32 or AH_scFv21_P32 and cultured in 2-YT medium with 100 µg/mL ampicillin at 37 °C to an OD600 value of 0.5 to 1.0, followed by another 12 h of shaking at 25 °C after adding 0.05 mM IPTG. Cells were collected by centrifugation, resuspended, and lysed by sonication at 4 °C in binding buffer (20 mM phosphate buffer [PB] supplemented with 150 mM NaCl, pH 8.0). After centrifugation, the supernatant was immediately loaded onto an Ni2+–nitrilotriacetic acid sepharose affinity column. After being washed with five volumes of binding buffer, the column was loaded with diluted imidazole (10 mM, 30 mM, 50 mM, 200 mM, and 500 mM) in 20 mM PB, pH 8.0. The fusion proteins were collected and desalted using a Millipore column (MWCO, 10000). The concentrations of the purified proteins were measured using an ultraviolet spectrophotometer.
SDS-PAGE and Western Blot Analysis
Purified proteins were mixed with loading buffer to perform sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) using a 12% gel and detected with Coomassie Blue or transferred from the gel onto a nitrocellulose membrane. The membrane was blocked with nonfat 5% milk for 1 h at room temperature (RT) and then incubated with an HRP-conjugated anti-His antibody for 1 h at RT. After being washed three times, the membrane was detected by ECL and autoradiography.
Flow Cytometry
Centrifugation was used to harvest 5 × 105 cells per sample. The cells were then washed once with cold phosphate-buffered saline (pH 7.4) containing 2% FBS and incubated with purified protein at different concentrations for 30 min on ice; next, PE-conjugated anti-His antibody was added. Cells were then analyzed using a FACScan cytometer (Becton Dickinson, Franklin Lakes, NJ).
Confocal Microscopy
Cells were grown on glass-bottom dishes (3 × 105 cells/well) in a complete medium for 24 h at 37 °C and treated with purified scFvs for 0.5 h at RT, which was followed by a 15-min incubation with PE-conjugated anti-His antibody. Samples were observed and photographed with a laser scanning confocal microscope using the Zeiss LSM 410 confocal laser system connected to a Zeiss Axiovert 135M microscope with a 40_/1.2 C-Apochromat water immersion lens (Zeiss, Jena, Germany). Green fluorescent protein was imaged using an argon laser (488 nm for excitation with a 510 nm pass barrier filter), and the red fluorescence of PE was imaged using 488 nm for excitation and 670 nm pass barrier filter. Similar results were obtained in three independent experiments.
Phage Library Construction
Using the computer-aided analysis results, phage library primers were designed to introduce site mutations into the antibody as well as the LoxP511 linker. Using overlap PCR, ScFv sequences were synthesized, purified, and then cloned into the BssHII–NheI (New England Biolabs, Ipswich, MA) sites in the vector pDF (a kind present from Professor Yan Wang, Navy General Hospital). The inserted genes were under the control of a prokaryotic operator (lac operator) and a PelB leading sequence. The vector was derived from pDNA5 and thus was available for both scFv and Fab library construction.
Based on the Cre/LoxP system, a large-content synthetic phage antibody library for affinity maturation of 2C4 was successfully constructed. 17 Briefly, after transferring the ligation products into XL1-Blue, the primary library was generated. By infecting the phagemid into the Cre+ bacteria BS1365 (genotype: F′kan recA1 endA1 gryA96 thi21ΔlacU169 supE44 hsdR17 (λ imm 434 X12-cre) at high multiplicity of infection (MOI), the VH genes were recombined to create a large phage antibody library. Then the working repertoire was obtained by infecting XL1-BLUE cells at a low MOI.
Cell-Based Clone Screening
A 2 mL aliquot of the library was incubated with SKOV3 cells for 2 h at RT. After centrifugation, the pellet was resuspended in 200 µL of 0.1 M citric acid, pH 3.0, after which XL1-Blue cells were added to enable phage proliferation. After three cycles of screening, positive clones were cultivated on LB plates containing both ampicillin and kanamycin and selected for sequencing and further identification by flow cytometry.
Results
General Features of the 3D Structures of AH_scFv15 and AH_scFv21
Using computer-guided homology modeling, the 3D structures of AH_scFv15 (15 aa linker; common [SG4]3 sequence) and AH_scFv21 (21 aa linker; LoxP511 site sequence) were designed, as shown in Figure 1 . The pocket conformations formed by the six complementarity determining regions (CDRs) of AH_scFv15 and AH_scFv21 are similar. The root mean square distance of the main chain carbon atoms in the CDRs in AH_scFv15 and AH_scFv21 is 0.015 nm. The results indicate that the two scFvs possess the same function.
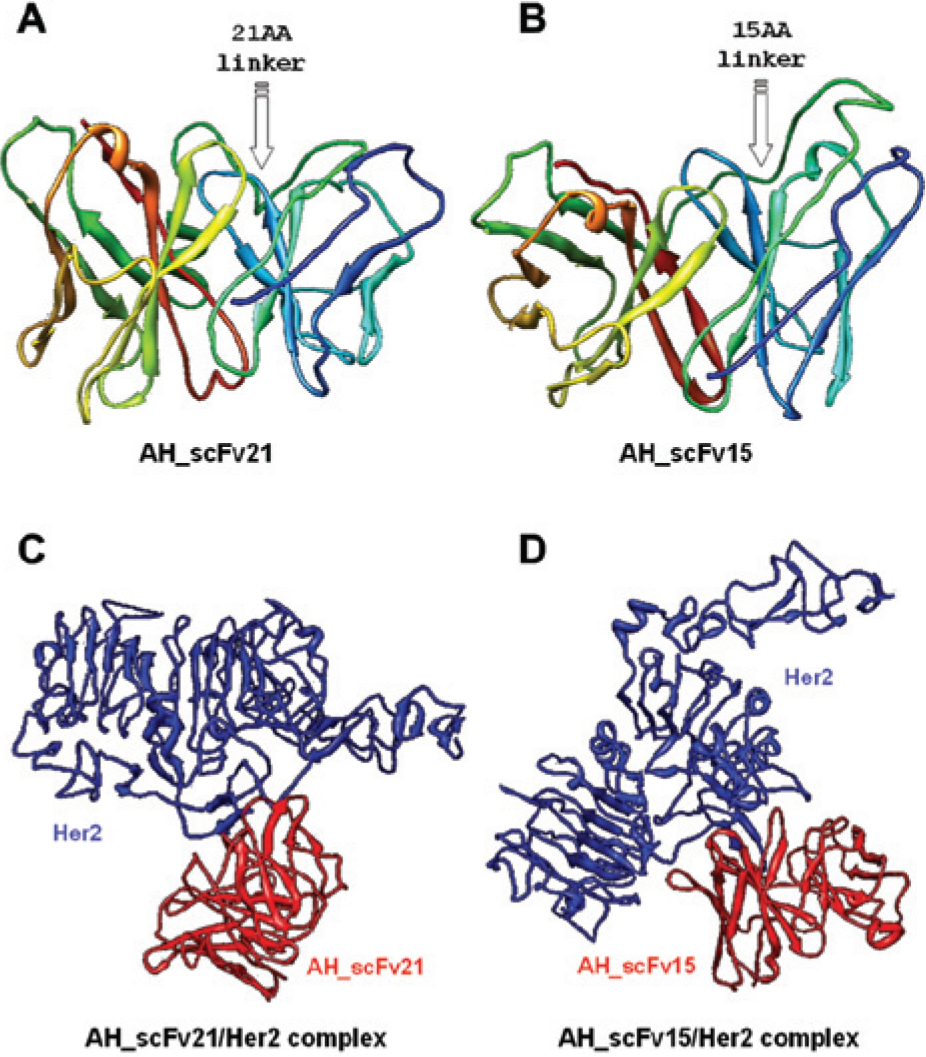
Three-dimensional structures of the two single-chain variable fragments (scFvs) binding to the extracellular domain of HER2 (HER2_ECD), derived by computer-aided analysis. The optimized three-dimensional structures of (
Theoretical Structure of the Complex Formed by AH_scFv15 and AH_scFv21 with the ECD of HER2
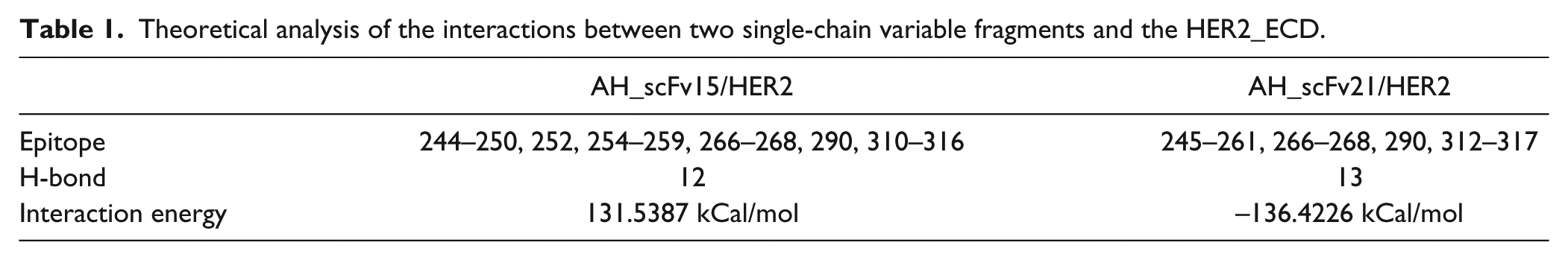
Based on the 3D structures of AH_scFv15 and AH_scFv21 and the crystal structure of HER2, the 3D structures of the scFv-HER2 complexes were constructed using the molecular docking method and optimized using the molecular dynamics method. The 3D minimized structure of the scFv and HER2 complexes is shown in Figure 2 . Using distance geometry, intermolecular hydrogen bond forming theory, and computer graphics technology, the interaction between the two scFvs and HER2 was predicted, as shown in Table 1 . The results show that the identified key domain, the number of intermolecular H-bonds, and the interaction energy of the two complexes are predicted to be similar.
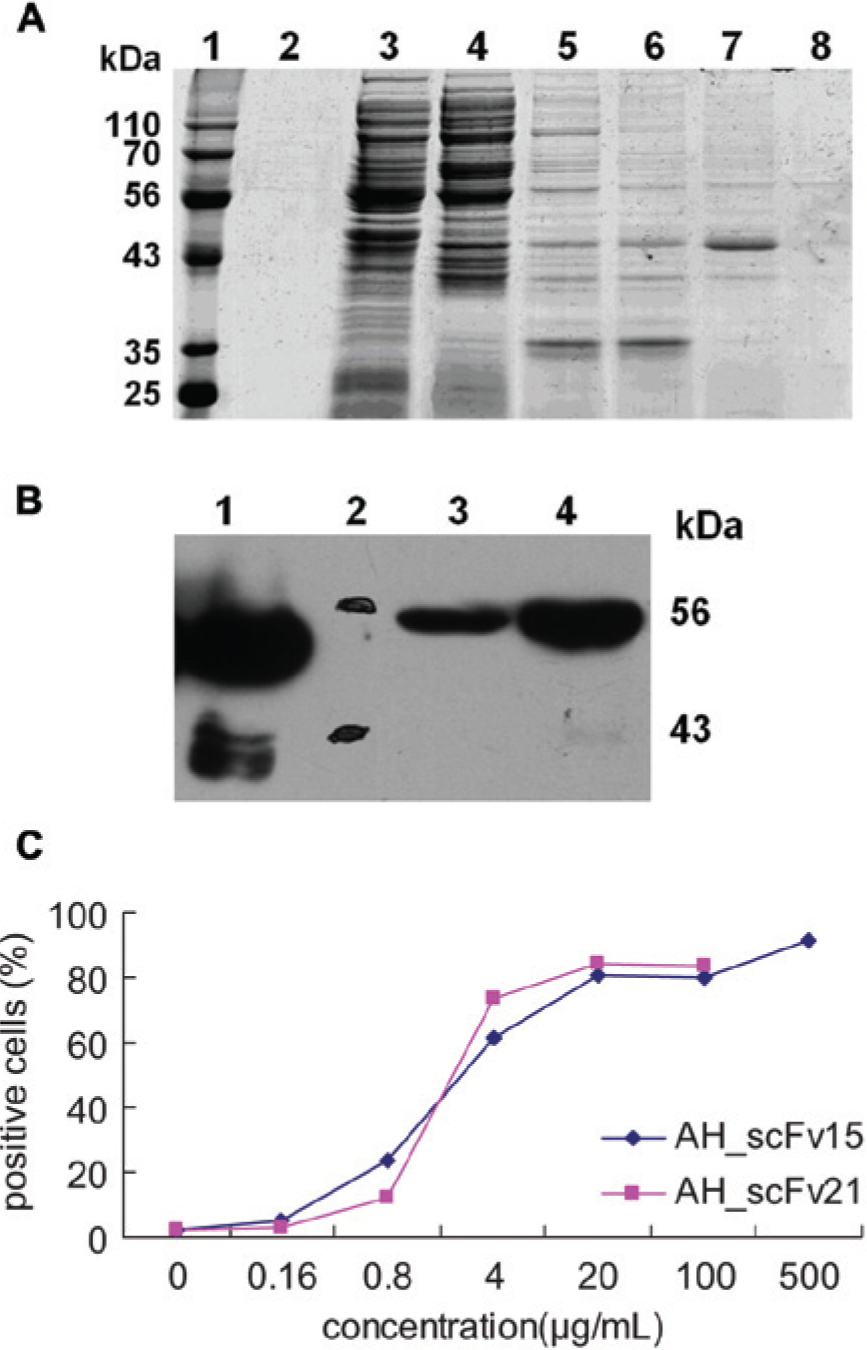
Purification and membrane antigen binding of the two single-chain variable fragments (scFvs). (
Theoretical analysis of the interactions between two single-chain variable fragments and the HER2_ECD.
Prokaryotic Expression of AH_scFv15 and AH_scFv21
To evaluate the theoretical prediction and design results, the expression plasmids AH_scFv15_P32 and AH_scFv21_P32 were transfected into E. coli Rosetta cells, and the resulting anti-HER2 scFvs were purified using an Ni2+–nitrilotriacetic acid sepharose affinity column. SDS-PAGE analysis indicated that the target proteins could be diluted by 200 mM imidazole in 20 mM PB, pH 8.0 ( Fig. 2A ). Western blot showed a specific band for the scFvs, with a molecular weight of approximately 49 kDa, as predicted ( Fig. 2B ).
AH_scFv21 Is Able to Bind the Same Epitope as 2C4
Daudi cells (105) were incubated with the diluted AH_scFv21 and AH_scFv15, respectively, for 30 min on ice, followed by incubation with PE_anti-His for another 20 min. Cells were then analyzed by flow cytometry to evaluate the binding activities of the two scFvs. As shown in Figure 2C , the percentage of AH_scFv21–positive cells is similar to that of AH_scFv15–positive cells, indicating that using the Loxp511 site as a linker does not affect the binding ability of the scFv, and this result is consistent with the theoretical prediction.
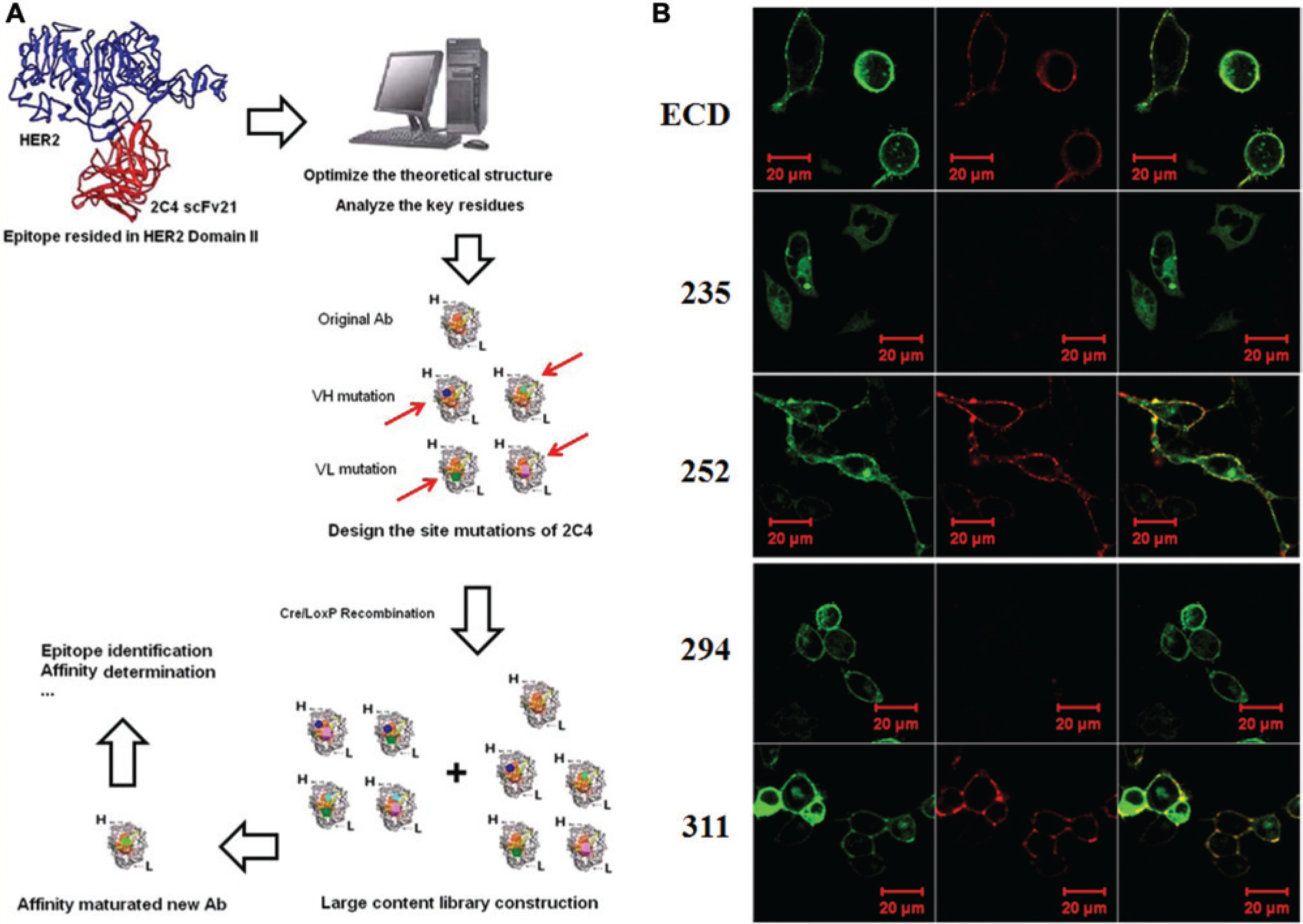
According to our computer-aided modeling results, the two scFvs could identify an epitope in domain II of HER2. To test this, a series of HER2 mutants (HER235, HER252, HER 294, and HER 311) was designed, as shown in Figure 3A , in which the indicated residues were replaced by alanine. The mutants were expressed in the 293T cells, which were then used for epitope determination. As shown in Figure 3B , herceptin is able to bind all the HER2 mutants, because its epitope resides in domain IV of HER2. In contrast, although both anti-HER2 scFvs could identify HER2_ECD (wild-type), HER252, and HER311, they had no or only a weak ability to bind to HER235 or HER294. Flow cytometry analysis similarly showed that residues 235–236/245–248 and 294–297 of HER2 could interact with the two scFvs directly, consistent with the theoretical prediction.
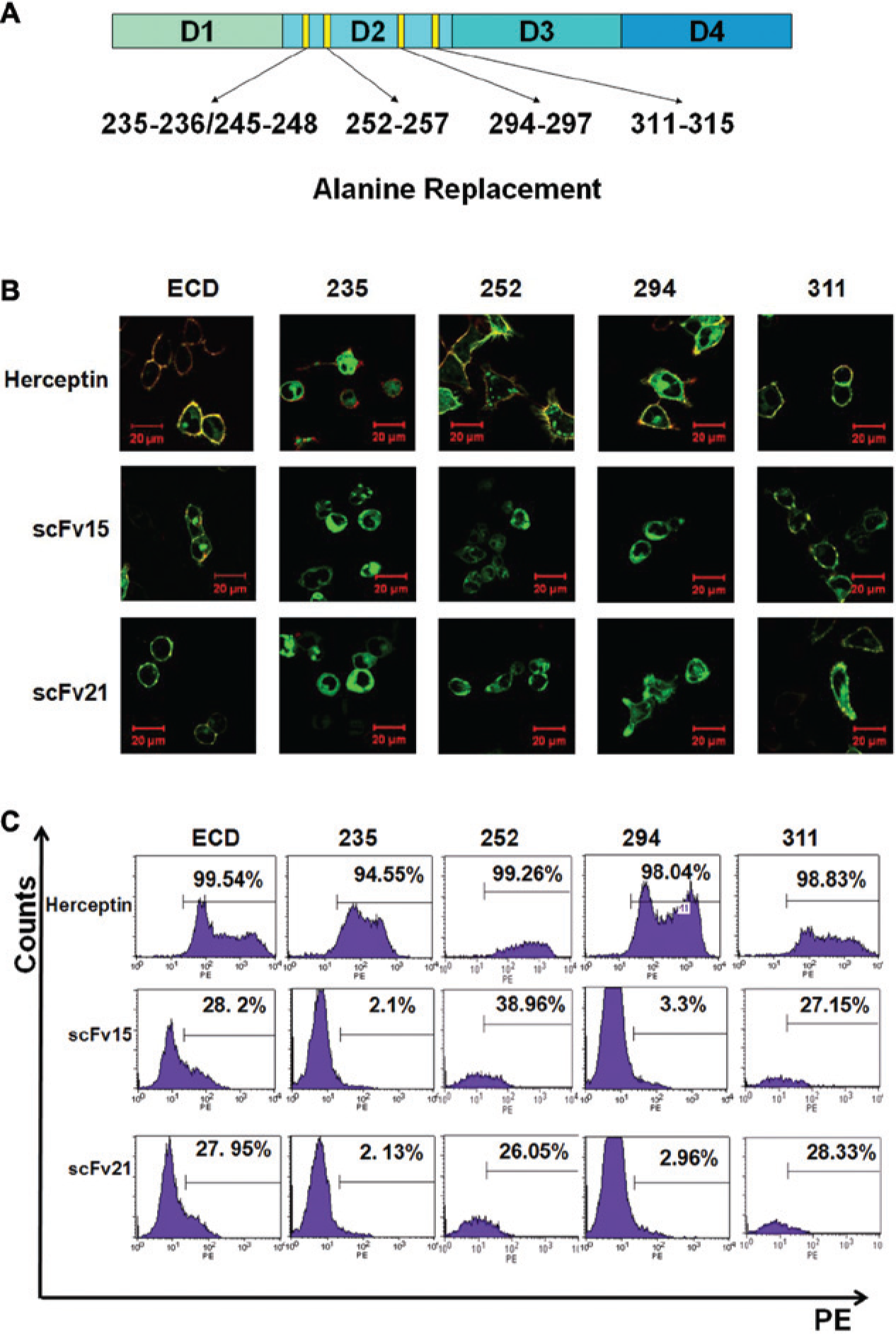
Epitope identification of the two single-chain variable fragment (scFv) antibodies. (
Construction of a Large Phage Library by Recombination Using AH_scFv21
Next, we used AH_scFv21 to construct a large phage library. The pDF vector has a prokaryotic operator (lac operator) and a PelB leading sequence but was derived from pDNA5 and thus could be used for both scFv and Fab library construction. Based on the LoxP/LoxP511 system, by transforming the recombinant AH_scFv21 into BS1365 (a Cre+ E. coli strain), it was possible to form a large working library for antibody screening; that is, the scFv fragments between LoxP and the Loxp511 linker sites could be reciprocally interchanged, thereby increasing the size of the library ( Fig. 4A ). Here, the size of the preliminary library content is 4 × 107, whereas the size of the recombinant library is 5 × 1012. Then, the work library was established with the content of 2 × 1011 for further screening. After three cycles of cell screening, a new antibody with a theoretically designed higher affinity than that of 2C4 was identified and sequenced.
Screening and identification of an affinity-maturated antibody. (
Affinity Determination and Epitope Identification of the New Ab
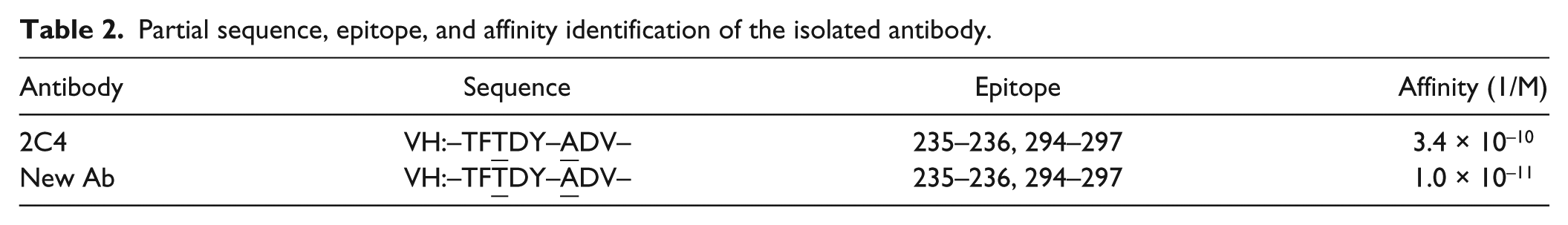
The binding kinetics of 2C4 or the new Ab with respect to HER2 were measured using Biocore, which showed that the new Ab possessed a higher affinity than 2C4, consistent with the theoretical prediction results. The KD values are 1.0 × 10−11 (the new Ab) and 3.4 × 10−10 (2C4), respectively ( Table 2 ).
Partial sequence, epitope, and affinity identification of the isolated antibody.

In addition, the confocal results confirmed that the new Ab binds to the same epitopes as its parent scFv. As shown in Figure 4B , it could identify only HER2_ECD, HER252, and HER311 but could not bind to HER235 or HER294, indicating that residues 235–236/245–248 and 294–297 were key sites recognized by the new Ab. These results indicate that it is possible to use a LoxP511-based recombination system to construct large phage libraries for “me-better” antibody screening.
Discussion
An antibody should be humanized and/or optimized for better performance, including higher affinity, and so forth, for subsequent application. 18 Traditional optimization methods, including site mutation and CDR/CDR3 randomization, have disadvantages, and large phage libraries are necessary for quick and successful screening. 19 The use of a recombination system such as Cre-LoxP is a good method for increasing library size.
Many native Fv antibody fragments are not stable because the VH and VL domain can rapidly dissociate; therefore, different strategies have been developed to stabilize the association of these two domains by adding linkers such as the 15-aa linker (SG4)3 between VH and VL. We hypothesized that it could be possible to use the LoxP511 site as a linker to construct a large scFv phage library for obtaining “me-better” antibodies quickly.
HER2 overexpression is an easily recognizable marker that correlates with increased tumor growth rates, high metastatic potential, and poor long-term survival rates for the patient. Unlike HER1, HER3, and HER4, no direct ligand for HER2 has yet been discovered. HER2 assumes an open conformation, which is constitutively active, with its dimerization domain permanently exposed and ready for structural interaction with other ligand-activated HER receptors such as activated ligand-bound EGFR to form dimers. 20 2C4, a new anti-HER2 humanized antibody, inhibits the formation of HER homo- or heterodimerization by binding the dimerization site in HER2 domain II to downregulate the activation of HER signaling pathways that mediate cancer cell proliferation and survival.
To determine whether it would be possible to use LoxP511 as a linker to construct a large scFv phage library, we used computer-aided design to model two 2C4-derived scFvs with different linkers: AH_scFv21, with a 21-aa linker (LoxP511), and AH_scFv15, with a 15-aa linker ([SG4]3). The modeling results indicated that both of them could bind antigen HER2 at a similar epitope: 235–236/245–248 and 294–297 ( Table 1 ). Next, to validate the theoretical prediction, we constructed, bacterially expressed, and used an Ni2+–nitrilotriacetic acid sepharose affinity column to purify the two scFvs. SDS-PAGE and Western blot analysis show that the target protein has a molecular weight of approximately 49 kDa ( Fig. 2 ). Next, HER2 mutants (HER235, HER252, HER 294, and HER 311) were expressed in 293T cells to identify the epitopes of the two scFvs. According to confocal microscopy observation and flow cytometry analysis, the epitope of both scFvs resides in residues 235–236/245–248 and 294–297 of HER2, indicating that the linker size does not influence antigen binding ( Fig. 3 ). Then, we constructed a synthetic phage library with a size of 5 × 1012 for affinity maturation of 2C4 ( Fig. 4A ) using the Cre-LoxP recombinant system and isolated a new “me-better” antibody with higher affinity than 2C4. Theoretical prediction and flow cytometry analysis show that the new antibody binds to the same epitope as its parent scFv ( Table 2 ).
In this study, the 15-aa linker (SG4)3 was used the positive control, as it is a linker that is frequently used between VL and VH. Based on the theoretical results, the 21-aa linker did not influence the 3D structure of the scFv. Furthermore, according to the experimental results, the VL-21aa-VH protein had a similar epitope and similar antigen-binding capacity as VL-15aa-VH, which showed that it was possible to use LoxP511 as a linker. Normally, because of the constraints present within molecular biological techniques (e.g., enzyme restriction and ligation), the number of mutated sites that can be generated is limited. Here, as the LoxP511 sequence was used as the linker of scFv, the Cre-LoxP system allowed the construction of a large library by recombination, which permitted us to design and evaluate more site mutations that could produce antibodies with a potentially high affinity. In addition, by using recombination, the library could include not only the predicted single-site mutants but also combined candidates with mutation sites in both heavy and light chains.
In this way, using BS1365 (Cre+ E. coli strains) and the LoxP/LoxP511 recombination system, a synthetic phage library for 2C4 optimization was established, resulting in a large working library in which fragments between LoxP and the scFv Loxp511 linker were reciprocally interchanged between different phagemids, thereby increasing the library content ( Fig. 4A ). Finally, we used this library to isolate a new “me-better” Ab, which our computer platform predicted to possess higher affinity than 2C4. Biocore detection shows that it indeed has a lower KD value to bind the antigen ( Table 2 ), and confocal analysis shows that its epitope resides in 235–236/245–248 and 294–297 of HER2, which is the same as that of the parent scFv ( Fig. 4B ).
This system does have some disadvantages. LoxP511 could be used as the linker only between two/multi domains and could not be used inside a domain because it might change the domain’s 3D structure. Nonetheless, our study shows that LoxP511 is a good choice as the linker to construct large scFv phage libraries for use in quick and easy “me-better” antibody screening.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grants from National Sciences Fund (No. 31370938), National High Technology Research and Development Program (863 program, No. 2012AA02A302) and National Key Basic Research Development Program (973 program, No. 2010CB833604).
