Abstract
Liver X receptor α (LXRα) plays an important role in reverse cholesterol transport (RCT), and activation of LXRα could reduce atherosclerosis. In the present study, we developed a screening method to identify new potential LXRα agonists using an LXRα-GAL4 chimera reporter assay. A novel analogue of N,N-disubstituted 2,8-diazaspiro[4.5]decane, IMB-151, was identified as an LXRα agonist by using this method. IMB-151 showed a significant activation effect on LXRα, with an EC50 value of 1.47 µM. IMB-151 also increased the expression of ATP-binding cassette transporter A1 (ABCA1) and G1 (ABCG1) in RAW264.7 macrophages. The upregulating effects of IMB-151 on ABCA1 and ABCG1 markedly decreased when coincubated with geranylgeranyl pyrophosphate (GGPP) ammonium salt or LXRα small interfering RNA (siRNA). Our data indicated that the upregulation of ABCA1 and ABCG1 by IMB-151 depended on activation of LXRα. Moreover, IMB-151 significantly reduced cellular lipid accumulation and increased cholesterol efflux in RAW264.7 macrophages. Interestingly, IMB-151 slightly increased sterol response element binding protein 1c (SREBP-1c) protein expression levels in HepG2 cells compared with TO901317, and this indicated that IMB-151 might have less lipogenesis side effect in vivo. These results suggested that IMB-151 was identified as a selective agonist for LXRα by using a screening method and could be used as a potential antiatherosclerotic lead compound in the future.
Introduction
The liver X receptors (LXRα and LXRβ) are ligand-activated transcription factors that belong to the nuclear receptor superfamily.1,2 LXRα is abundantly expressed mainly in the liver, kidney, spleen, intestine, and adipose tissue, whereas LXRβ is expressed ubiquitously. 3 LXRs and retinoid X receptor form heterodimers that bind to specific response elements in the regulatory region of the target genes to regulate their expression. 4 Adenosine triphosphate (ATP)–binding cassette transporter A1 (ABCA1) and G1 (ABCG1) are regulated by LXRs via functional LXREs found in their genes, which play important roles in cholesterol efflux.5–7 ABCA1 can transfer both cholesterol and phospholipids to lipid-free apolipoprotein A-I (apoA-I), and ABCG1 can transfer cholesterol to high-density lipoproteins (HDLs).7,8
Excessive absorption of lipoproteins in macrophages causes foam cell formation within arterial walls, and these cells subsequently rupture and promote early atherosclerotic plaque formation.9,10 The efflux of excess cellular cholesterol from peripheral tissues and its return to the liver for excretion in the bile occurs by a process referred to as reverse cholesterol transport (RCT). 11 RCT is regarded as a major mechanism by which HDL protects against atherosclerotic cardiovascular disease. 11 LXRs play important roles in RCT by regulating cholesterol metabolism and transport, lipogenesis, and inflammation.9,12 Previous investigation showed that liver-specific LXRα overexpression reduced atherosclerosis. 13 Treatment of atherosclerotic mice with synthetic LXR ligands inhibited progression and promoted regression of atherosclerotic plaques.14,15 Furthermore, macrophage-specific deletion of LXRα in mice enhances atherogenesis. 16
Several LXRα ligands, such as endogenous ligand 22(R)-hydroxycholesterol 1 and synthetic agonists TO901317 and GW3965,17–19 have recently been reported. Once LXRα is activated by the ligand, conformational changes of LXRα cause cholesterol efflux by increasing the expression of ABCA1 and ABCG1.7,20 However, the known ligands have the undesirable side effect of inducing lipogenesis and hypertriglyceridemia. 12 Therefore, the identification of new LXRα agonists is a major challenge in pharmacology.
In this study, we discovered IMB-151 as a novel analogue of N,N-disubstituted 2,8-diazaspiro[4.5]decane with potential LXRα agonist activity using an LXRα-GAL4 chimera reporter assay. We then investigated the effect and mechanism of this compound on the target genes of LXRα and cholesterol efflux in murine macrophages.
Materials and Methods
Reagents
The compound IMB-151 was purchased from WuXi AppTec Company (Shanghai, China). Geranylgeranyl pyrophosphate (GGPP) ammonium salt, TO901317, and Red Oil O stain were purchased from Sigma (St. Louis, MO). Fetal bovine serum (FBS) and Opti-MEM Reduced Serum Medium used for transfection were obtained from Gibco (Invitrogen, Carlsbad, CA). RAW264.7 macrophages, HepG2 cells, and HEK293T cells were obtained from the Cell Center of PUMC (Beijing, China). Modified Eagle’s medium (MEM) and Dulbecco’s modified Eagle’s medium (DMEM) were purchased from Hyclone (Thermo Scientific, Rockford, IL). Oxidized low-density lipoprotein (ox-LDL) and HDL were purchased from Union-Biology Company (Beijing, China). 22-NBD-cholesterol was purchased from Invitrogen.
Cell Culture and Construction of Reporter Plasmid
RAW264.7 macrophages (ATCC, TIB-71) and HEK293T cells (ATCC, CRL-11268) were cultured in DMEM containing 10% FBS. HepG2 cells (ATCC, HB-8065) were grown in MEM containing 10% FBS at 37 °C in 5% CO2.
Total RNA was isolated from HepG2 cells using TRIzol reagent (Invitrogen), and first-strand complementary DNA (cDNA) was synthesized using a reverse transcriptional kit (Fermentas, Vilnius, Lithuania). The forward and reverse primers used for amplifying the human LXRα and LXRβ ligand-binding domain (LBD) fragments were listed as follows: LXRα forward primer, 5′-CGGGATCCTGGGCA-TGATCGAGAAGCT-3′; LXRα reverse primer, 5′-GGGGTACCTCATT CGTGCACATCCC-3′; LXRβ forward primer, 5′-ATTCGGGATCCCAGCGGCTCAA-3′; and LXRβ reverse primer, 5′-TGGGGTACCTCACTCG-TGGACGT-3′. To construct the pBIND-LXRα -LBD/pBIND-LXRβ-LBD plasmid, LXRα and LXRβ LBD PCR products were digested with BamH I and Kpn I and then were ligated into the pBIND vector (Promega, Madison, WI) respectively, which included the GAL4 DNA-binding domain (GAL4-DBD). GAL4-pGL4-luc plasmid was constructed by inserting the 5× GAL4 response elements into the promoter region of the pGL4.17 reporter vector (Promega).
LXRα-GAL4 Chimera Reporter Assay and Screening
A special synthetic compound library with 3000 drug-like structures from WuXi Apptec (Shanghai, China) was used for screening in this present study. These compounds were stocked in 96-well plates at 10 mg/mL in 100% DMSO. Compounds occupied 80 wells in each plate, and the remaining 16 wells contained DMSO only as negative controls.
HEK293T cells were seeded in 96-well plates at 2 × 104 cells/well in 100 µL DMEM containing 10% FBS. After incubation for 12 h, the cells at 90% confluence were washed once with phosphate-buffered saline (PBS), then transfected with GAL4-pGL4-luc reporter plasmids (180 ng/well) and pBIND-LXRα-LBD (or pBIND-LXRβ-LBD) expression plasmids (18 ng/well) using Lipofectamine 2000 (Invitrogen; 0.5 µL/well). After 6 h, the transfected cells were washed twice with PBS and replaced with 200 µL DMEM, containing indicated screening compounds respectively (10 µg/mL of various screening samples, 1 µM TO901317 as a positive control, and 0.1% DMSO as a negative control). For primary screening, each compound was assayed only in singlet. For rescreening, each primary positive compound was tested in triplicate. For IMB-151 activity assaying, the cells were treated with various concentrations of IMB-151 in serum-free DMEM. After 18 h, the cells were washed with PBS once and lysed with 20 µL 1× CCLR (Promega) per well. The luciferase activity was detected as relative luminescence units (RLUs) in a final volume of 60 µL with the Luciferase Assay System (Promega) on a microplate reader (PerkinElmer, Waltham, MA). The compounds with LXRα agonistic activity were identified using LXRα-GAL4 chimera reporter assay through screening.
RNA Isolation and Real-Time Quantitative PCR
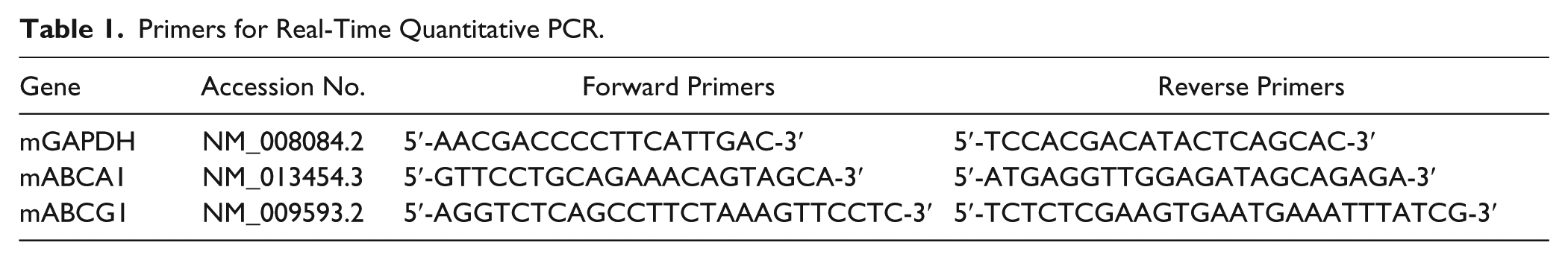
RAW264.7 macrophages were seeded in 6-well plates at 6 × 104 cells/well in DMEM containing 10% FBS. Total RNA was extracted from the cells using TRIzol reagent according to the manufacturer’s protocol. First-strand cDNA was synthesized from the total RNA in a 20-µL system using a reverse transcriptional kit. Real-time quantitative PCR with SYBR Green (Roche Diagnostics, Lewes, UK) detection chemistry was performed on a CFX96 Real-Time PCR Detection System (Bio-Rad, Hercules, CA). The sequences of the primers are listed in Table 1 . Melting curves were recorded, and the specificity of PCR products was checked on agarose gel. The messenger RNA (mRNA) levels of all genes were normalized for glyceraldehyde-phosphate dehydrogenase (GAPDH) levels, and the quantitative measurements were carried out by the ΔΔCt method.
Primers for Real-Time Quantitative PCR.

Western Blot Analysis
RAW264.7 macrophages and HepG2 cells were seeded on 6-well plates at 6 × 104 cells/well and 4 × 104 cells/well, respectively. After cell attachment (24 h), IMB-151 at various concentrations was added. The cells were harvested after 18 h, and protein extracts were prepared as previously described. Equal amounts of proteins (40 µg per lane) were electrophoresed by 10% sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) and electroblotted onto a 0.45-µm polyvinylidene fluoride membrane (Millipore, Bedford, MA). After the blots were blocked with 5% (w/v) skimmed milk in Tris-buffered saline containing 0.2% Tween-20 (TBST) for 1 h, they were incubated with the following primary antibodies, which were diluted in 5% (w/v) skimmed milk in TBST: mouse anti-GAPDH (1:1000; Santa Cruz Biotechnology, Santa Cruz, CA), rabbit anti-LXRα (1:1000; Sigma), anti-ABCA1 (1:1000; Novus, Littleton, CO), anti-ABCG1 (1:500; Abcam, Cambridge, MA), and anti–sterol response element binding protein 1c (SREBP-1c) (1:1000; Novus) for 4 °C overnight. Then the membranes were washed with TBST three times, followed by incubation with horseradish peroxidase–conjugated secondary antibodies: anti–mouse and anti–rabbit immunoglobulin G (IgG) antibodies (1:5000; Novus) for 2 h in temperature. After being washed with TBST three times, the protein bands were detected with an Enhanced Chemiluminescence (ECL) reaction kit (Millipore) and quantified by Quantity One Software (Bio-Rad). All the proteins were normalized to GAPDH.
Oil Red O Staining
Lipid accumulation in RAW264.7 macrophages was evaluated by means of Oil Red O staining. The cells were seeded in 96-well plates at 6 × 104 cells/well, and 60 µg/mL ox-LDL was added after cell attachment. The next day, when the cells were grown to 90% to 95% confluence, they were treated with IMB-151 at various concentrations. After 18 h, the samples were fixed with 4% (v/v) paraformaldehyde (15 µL/well) for 10 min and washed with ddH2O twice. Then the samples were added with 60% (v/v) islpropanol (150 µL/well) for 5 min, then stained with 0.5% Oil Red O (150 µL/well) for 1 h. After being washed with 60% (v/v) islpropanol (150 µL/well) three times and ddH2O twice, the samples were observed by light microscopy.
Small Interfering RNA against LXRα
We employed siRNA against mouse LXRα (sc-38829; Santa Cruz Biotechnology) and scrambled siRNA for the negative control (sc-37007; Santa Cruz Biotechnology). RAW264.7 macrophages were seeded at 6 × 104 cells/well in the 6-well formats. Briefly, 50 nM siRNA against LXRα or 50 nM scrambled siRNA per well (final concentration) was transfected into RAW264.7 cells using Lipofectamine RNAiMAX transfection reagent (Invitrogen; 5 µL/well) at a cell density of 30% to 50% confluence. After 48 h of the transfection, in comparison to the control siRNA, the siRNA of LXRα suppressed the expression of LXRα mRNA by 75% according to real-time quantitative PCR assays.
To examine whether the effect of IMB-151 on ABCA1 and ABCG1 was affected by LXRα knockdown, cells were treated with serum-free DMEM for 18 h in the absence or presence of IMB-151 (10.0 µM) after LXRα siRNA transfection for 48 h. Then Western blotting assays were performed as described above.
Cellular Cholesterol Efflux Experiments
Cellular cholesterol efflux experiments were performed using 22-NBD-cholesterol in RAW264.7 macrophages. 21 The cells were plated in 96-well clear-bottom black plates (Costar, Corning Inc., NY) at 4 × 104 cells/well. Six hours later, the medium was removed and the cells were labeled with 22-NBD-cholesterol (2.0 µM at the final concentration) in DMEM containing 0.2% (w/v) bovine serum albumin (BSA) (Sigma) (medium A) for 24 h in a 37 °C, 5% CO2 incubator. After 24 h of labeling, cells were washed twice with PBS and incubated with 100 µL medium A containing IMB-151 (0, 1.25, 2.5, 5.0, and 10.0 µM) for an additional 18 h. HDL (50 µg/mL as the final concentration) was added to start the efflux experiment at 37 °C for 6 h. Then the amounts of cholesterol in medium and cells were measured using a microplate reader, respectively (PerkinElmer; excitation 485 nm, emission 535 nm). The percentage of 22-NBD-cholesterol efflux was calculated as (medium)/(medium + cell) × 100%. Each efflux assay was performed in duplicate in three experiments.
Data and Statistical Analysis
The signal-to-background (S/B) ratio, signal-to-noise (S/N) ratio, and Z′ value of 96 well-plates were calculated according to the method described by Zhang et al. 22 The activity of negative control 0.1% DMSO was defined as the basis for 100% activity. The regulatory activity of tested samples was defined as (RLUtest compound/RLUnegative control) × 100%. The compound was considered a primary hit when the activity was ≥150%. The positive sample was further validated in triplicate reassays, and the validated hit was defined as a sample for which reassay showed ≥150% regulatory activity at a 95% confidence interval.
Statistics and best-fit curves were calculated with GraphPad Prism 5.0 software (GraphPad Software, San Diego, CA). The data are expressed as means ± SEM. Results were analyzed by Student t test and one-way analysis of variance by SPSS version 11.0 (SPSS, Inc., an IBM Company, Chicago, IL).
Results
Cell-Based Assay Optimization
To assess an assay system, the reproducibility and signal variation at the activity range must be evaluated. In our screening system, 1 µM TO901317 was the positive control, and four assay parameters influencing the signal and noise of the cellular reaction were taken into consideration, including the following:
DMSO concentration: 0.1% (
Ratio between the reporter plasmid and the expression plasmid: 10:1 (
Cell number: 2 × 104 cells/well (
Incubation time: 18 h (
S/N, S/B, percent coefficient of variation (CV%), and Z′ factor are classic and scientific indices for the comparison and evaluation of the quality of assays and can be used in assay validation and optimization.
22
According to our evaluation, this transient transfected system can be used for screening (
IMB-151 Has LXRα Agonist Activity
We identified that IMB-151, a structural analogue of N,N-disubstituted 2,8-diazaspiro[4.5]decane ( Fig. 1A ), had LXRα agonistic activity via the LXRα-GAL4 chimera reporter screening assay as described in the Materials and Methods. IMB-151 showed maximal activity of approximately 2.2-fold on LXRα, and a plot of LXRα activity versus concentration exhibited a clear dose-response relationship with an EC50 of 1.47 µM ( Fig. 1B ). However, it is interesting that IMB-151 had almost no activation on LXRβ compared with LXRα ( Fig. 1C ). In contrast, TO901317 exhibited approximately 3.3-fold LXRα activation and 2.8-fold LXRβ activation with an EC50 of 0.05 and 0.06 µM, respectively ( Fig. 1D , E ).
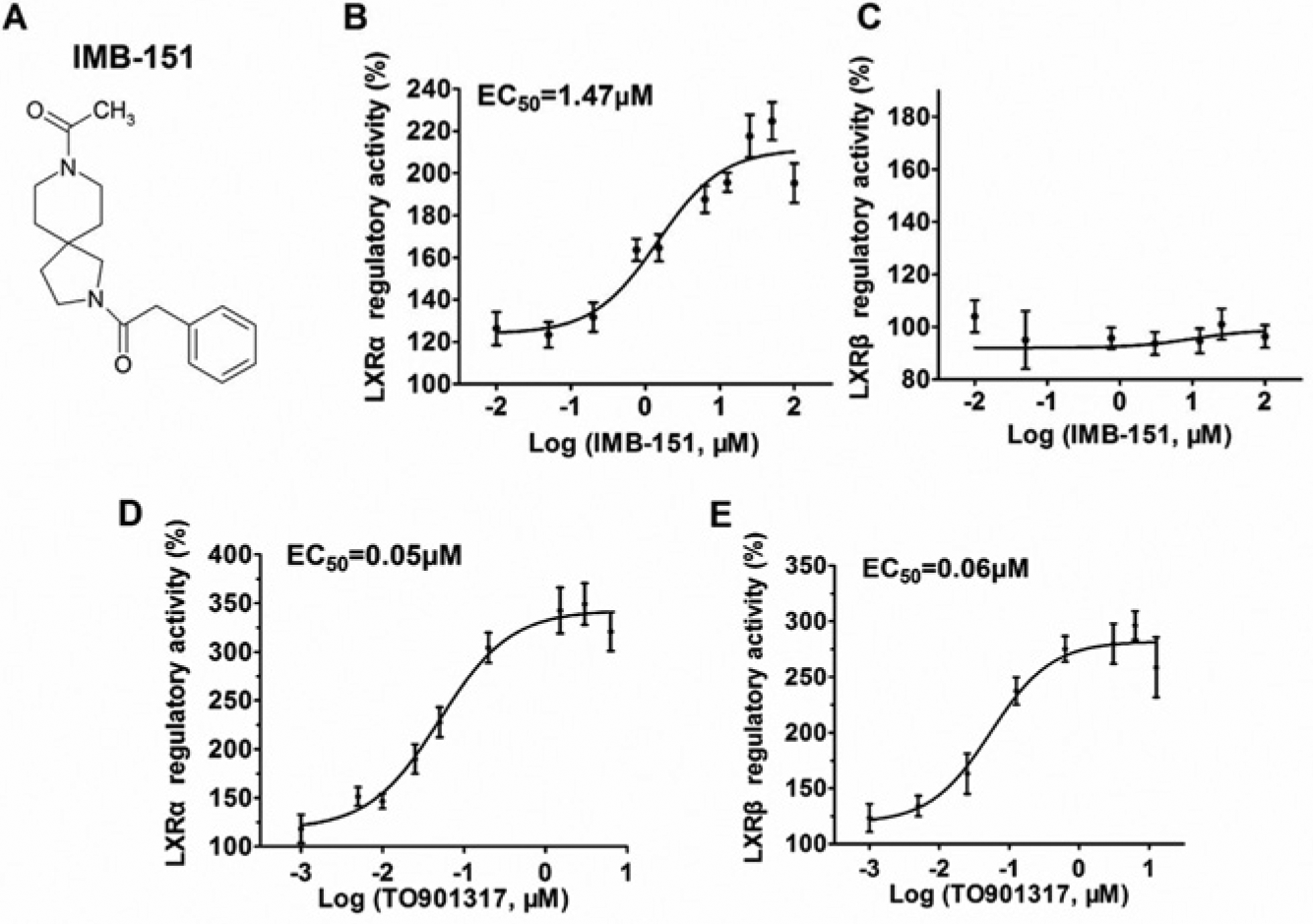
IMB-151 regulates liver X receptor α (LXRα). (
IMB-151 Induces LXRα, ABCA1, and ABCG1 Expression In Vitro
We first examined the effect of IMB-151 on the expression of ABCA1 and ABCG1 in RAW264.7 macrophages by real-time quantitative PCR and Western blotting assays. In RAW264.7 cells, IMB-151 significantly increased the expression of ABCA1 and ABCG1 at both protein ( Fig. 2A , B ) and mRNA ( Fig. 2C ) levels. It has been reported that LXRα agonists not only activate LXRα but also promote the expression of LXRα mRNA and protein. 23 Meanwhile, IMB-151 dose-dependently induced LXRα expression at the translational level ( Fig. 2D ) in HepG2 cells.
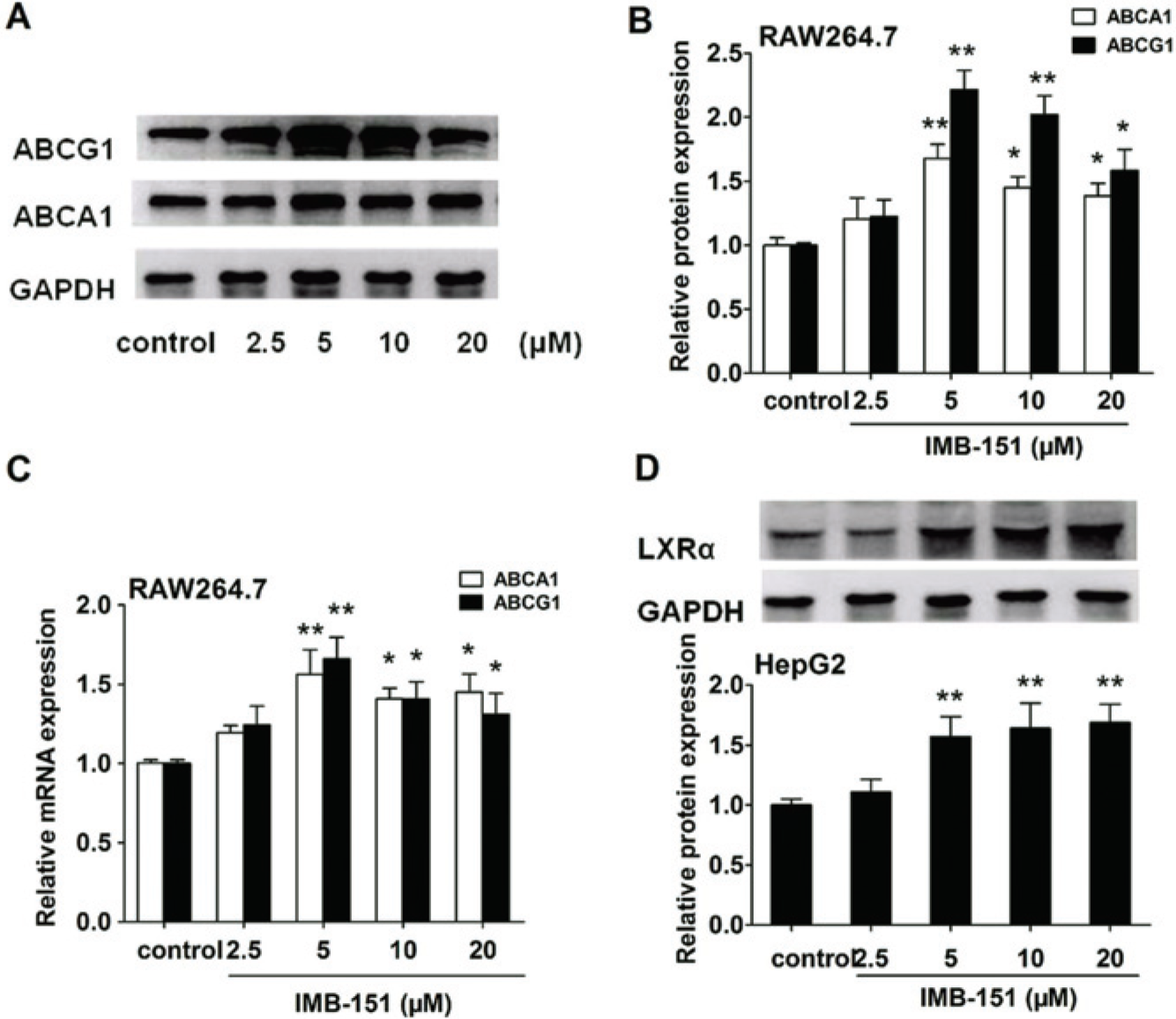
Effect of IMB-151 on liver X receptor α (LXRα), ABCA1, and ABCG1 expression. (
IMB-151 Reduces Cellular Lipid Accumulation in RAW264.7 Macrophages
To investigate the potential role of the active compound IMB-151 in modulating lipid efflux from mouse peritoneal macrophages, foam cell assay was performed using RAW264.7 cells. As shown in Figure 3C–E , treatment of RAW264.7 cells with IMB-151 significantly reduced lipid accumulations compared with ox-LDL alone ( Fig. 3B ). In addition, when the cells were treated with both antagonists (GGPP and IMB-151), two concentrations of the IMB-151 lipid efflux effect reduced ( Fig. 3G–H ).
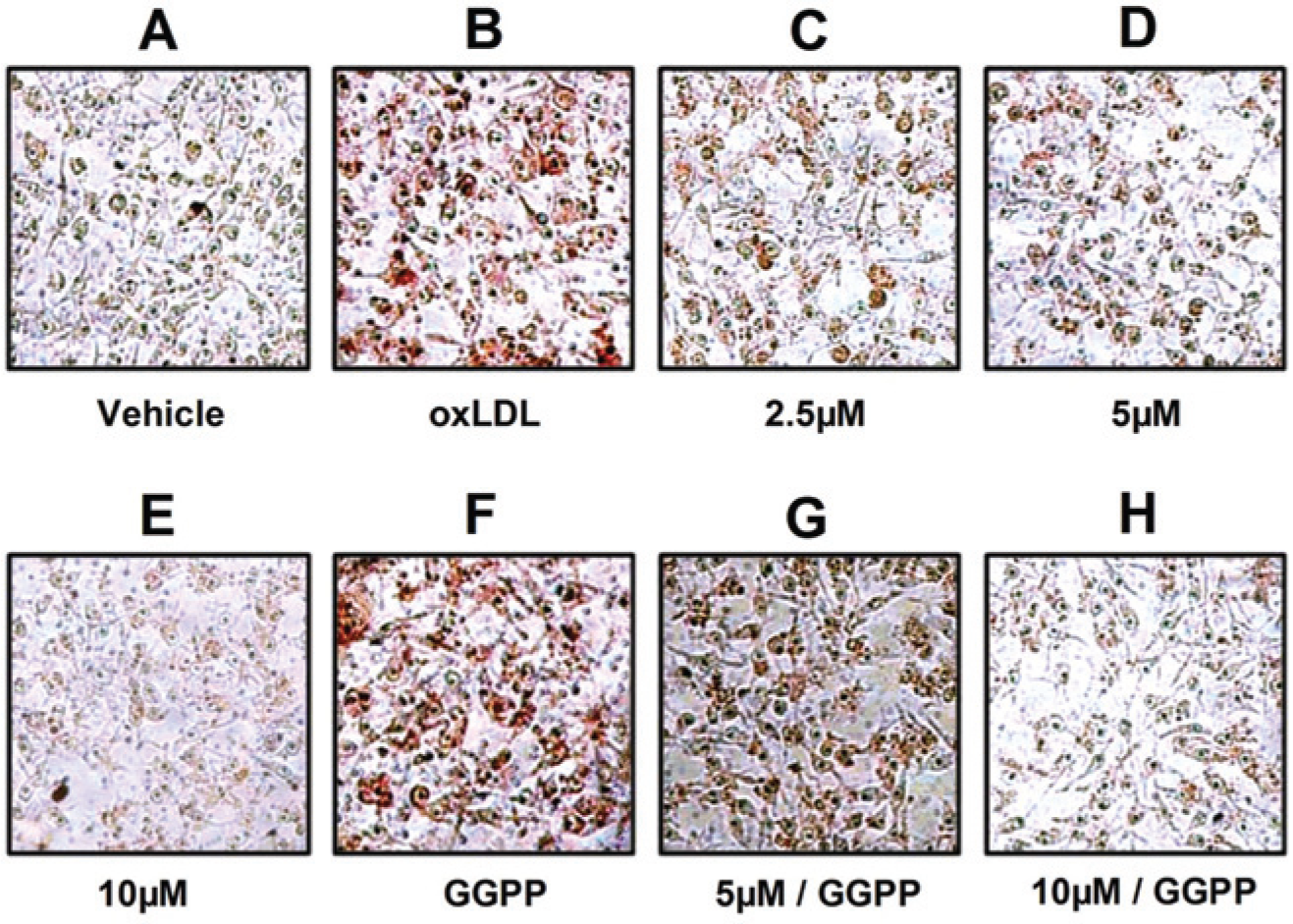
IMB-151 reduces oxidized low-density lipoprotein (ox-LDL)–induced lipid accumulation in RAW264.7 macrophages. RAW264.7 macrophages were preincubated with (
IMB-151 Promotes Cholesterol Efflux in RAW264.7 Macrophages
We next investigated the effect of IMB-151 treatment on cholesterol efflux in RAW264.7 macrophages. IMB-151 increased cholesterol efflux to HDL and reduced cellular cholesterol concentration in a dose-dependent manner in RAW264.7 macrophages ( Fig. 4A ). These promoting cholesterol efflux effects of IMB-151 were significant compared with the control at concentrations of 5.0 and 10.0 µM.
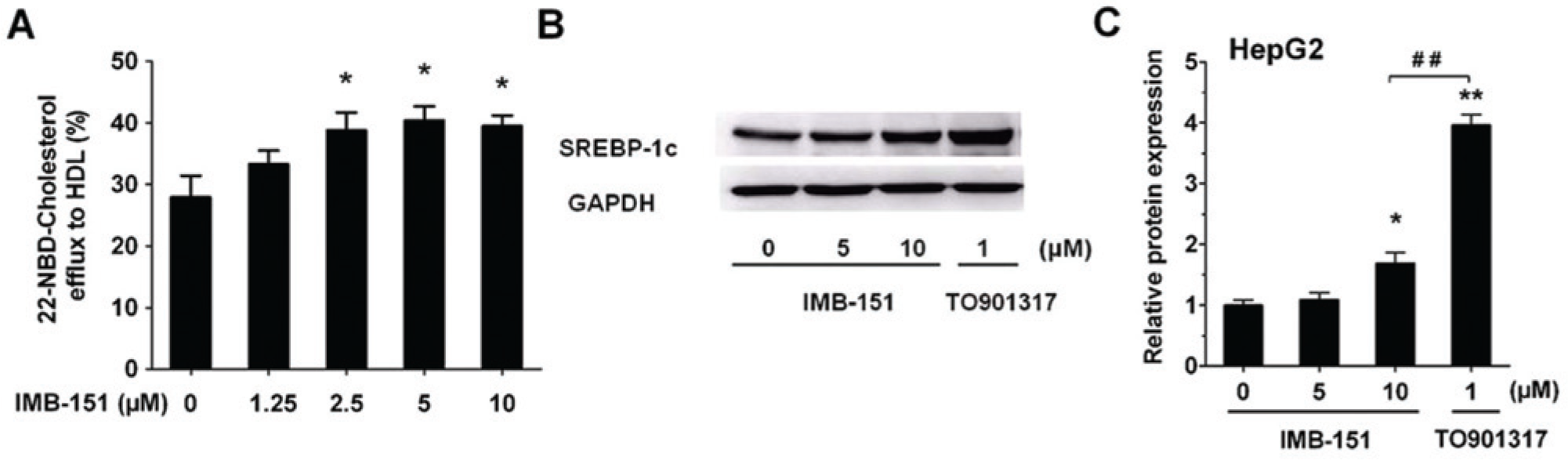
(
IMB-151 Slightly Induced SREBP-1c Expression
A potential side effect of LXRα agonists is increasing lipogenesis via upregulation of the SREBP-1c pathway in the liver. Therefore, the role of IMB-151 on SREBP-1c expression was also examined in this study. Our results showed that IMB-151 slightly increased SREBP-1c protein expression levels in HepG2 cells compared with TO901317 ( Fig. 4B , C ).
IMB-151-Induced Upregulation of ABCA1 and ABCG1 Depends on LXRα Activation
Previous work demonstrated that GGPP inhibits the activation of LXRα and the expression of its target genes. 24 Therefore, the effects of IMB-151 on ABCA1 and ABCG1 were examined when treated with GGPP simultaneously in RAW264.7 cells. Our data showed that the upregulating effects of IMB-151 on ABCA1 and ABCG1 were significantly reduced when GGPP was added ( Fig. 5A , B ), which indicated that IMB-151–regulated ABCA1 and ABCG1 expressions were dependent on LXRα activation. In addition, the LXRα activation effect of IMB-151 was clearly inhibited when coincubated with GGPP in the LXRα-GAL4 chimera reporter assay ( Fig. 5C ).
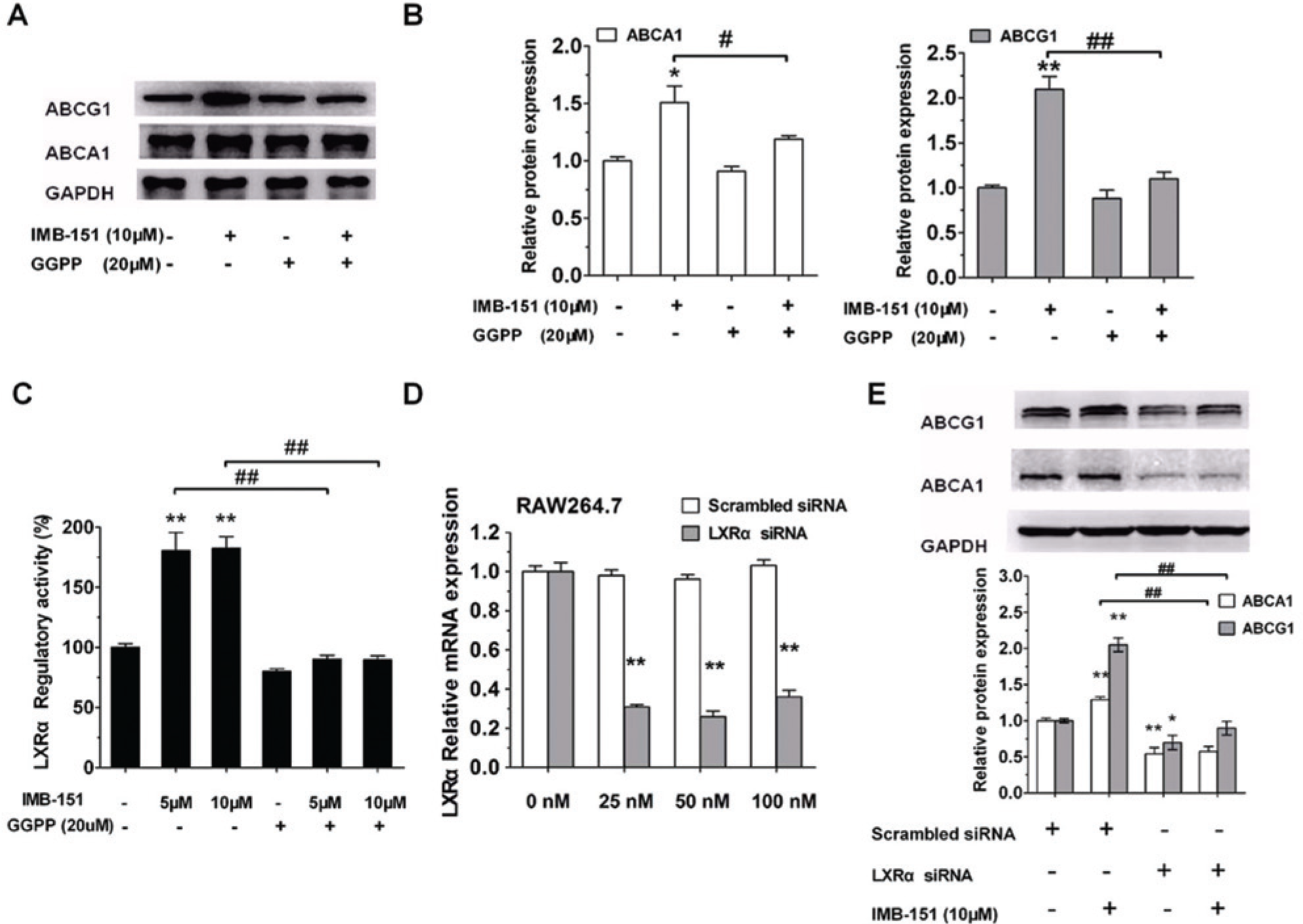
IMB-151 activates liver X receptor α (LXRα) via the LXRα-ABCA1/ABCG1 pathway. (
To further confirm whether the upregulating effects of IMB-151 on ABCA1 and ABCG1 were dependent on LXRα, siRNA knockdown experiment on LXRα in macrophages was also performed ( Fig. 5D ). RAW264.7 macrophages were treated as described in the Materials and Methods. Western blotting analysis showed that the upregulating effects of IMB-151 on ABCA1 and ABCG1 markedly decreased when LXRα was silenced ( Fig. 5E ), which also indicated that IMB-151–regulated ABCA1 and ABCG1 expressions depended on LXRα activation.
Discussion
LXRs, which are members of the nuclear receptor superfamily, consist of two subtypes, LXRα and LXRβ.2,25 Oxysterols have been identified as endogenous LXR agonists, and TO901317 and GW3965 are nonsterol synthetic ligands for LXRs.17,26 LXRs act as cholesterol sensors that control the expression of target genes when activated by ligands. LXR activation promotes cholesterol efflux and reduces cellular lipid accumulation, to prevent macrophage foam cell formation. LXRs have recently come to be regarded as potent targets for treating atherosclerosis, and synthetic agonists have been the subject of many studies. In this study, our goal was to find a novel compound with potent antiatherosclerotic activity through screening.
First, we identified IMB-151, a structural analogue of N,N-disubstituted 2,8-diazaspiro[4.5]decane with LXRα agonistic activity, but not for LXRβ. Although LXRα and LXRβ are highly related and share 78% amino acid sequence identity in both DNA and ligand-binding domains, our data indicated that IMB-151 apparently has better selectivity for LXRα.2,25 Later, in the subsequent work, we used HepG2 cells instead of HEK293T cells because LXRα is abundantly expressed in the liver, where it is involved in cholesterol and lipid metabolism.
ABCA1 and ABCG1 are major transporters involved in cholesterol efflux from macrophages and play a vital role in maintaining cellular cholesterol homeostasis. Here we demonstrated that in RAW264.7 macrophages, IMB-151 dose-dependently induced the expression of ABCA1 and ABCG1 proteins and mRNA. Meanwhile, we found that IMB-151 dose-dependently induced LXRα expression at the translational level in HepG2 cells. Our result on LXRα expression induced by IMB-151 in HepG2 cells was in agreement with Laffitte et al., 23 who previously documented that human LXRα has an autoregulation mechanism. In addition, Du et al. 27 found that some derivatives of N,N-disubstituted 2,8-diazaspiro[4.5]decane upregulated apolipoprotein A-I (ApoA-I) expression, which is the major and critical protein component of HDL particles and responsible for RCT. 28 Enhancing the endogenous expression of ApoA-I is also expected to reduce the risk of atherosclerosis. This conclusion is in agreement with the upregulation in expression of the crucial genes involved in the RCT process in our present study. Therefore, we suggest that IMB-151 activated LXRα and increased LXRα, ABCA1, and ABCG1 expressions in vitro.
Here, we found that IMB-151 reduced cellular lipid accumulation in RAW264.7 macrophages, but this effect was weakened by coincubating with GGPP. Meanwhile, IMB-151 also significantly increased cholesterol efflux in RAW264.7 macrophages. LXRα activation promotes cholesterol efflux and stimulates RCT in macrophages. Therefore, we speculated that cholesterol efflux induced by IMB-151 was related to the upregulation of ABCA1 and ABCG1 expression from activating LXRα in RAW264.7 macrophages. This could be of benefit in the prevention of atherosclerosis.
We wondered whether the upregulating effects were related to the LXRα activation. Our data showed that inhibition of LXRα activation by GGPP or LXRα siRNA eliminated the IMB-151–induced upregulation of ABCG1 and ABCA1 expression. At the same time, LXRα regulatory activity in luciferase assays was clearly lower upon treatment with both IMB-151 and GGPP than upon treatment with IMB-151 alone. Together, our data suggested that LXRα played a crucial role in the regulation of the expression of ABCA1 and ABCG1 by IMB-151, and this was potent evidence implying that IMB-151 provided protection from atherosclerosis through the LXRα-ABCA1/ABCG1 pathway. SREBP-1c, regulated by LXRα, is a transcription factor known to regulate the expression of lipogenic genes.29,30 In the present study, we found that IMB-151 slightly increased SREBP-1c protein expression levels compared with TO901317. This indicated that IMB-151 might have a lower lipogenesis side effect in vivo.
In summary, through screening, we identified IMB-151, a derivative of N,N-disubstituted 2,8-diazaspiro[4.5]decane, as a novel potential LXRα agonist lead compound. IMB-151 upregulated the expression of ABCA1 and ABCG1 dependently on LXRα activation. IMB-151 could potentially reduce lipid accumulation in foam cells and increase cholesterol efflux in macrophages. Moreover, IMB-151 slightly increased SREBP-1c protein expression levels compared with TO901317, and this indicated that IMB-151 might have less lipogenesis side effect. Our findings suggest that IMB-151 may be useful for the development of pharmaceutical agents for treating atherosclerosis.
Footnotes
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Key New Drug Creation and Manufacturing Program (2012ZX09301002-003, and 2012ZX09301002-001), the National Natural Science Foundation of China (81102443), National Natural Science Foundation of China General Program (81273515), and the National Natural Science Foundation of China-Guangdong Provincial People’s Government of the Joint Natural Science Fund Projects (U1032007).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
