Abstract
In evaluating kinase inhibitors, kinetic parameters such as association/dissociation rate constants are valuable information, as are equilibrium parameters KD and IC50 values. Surface plasmon resonance (SPR) is a powerful technique to investigate these parameters. However, results are often complicated because of impaired conformations by inappropriate conditions required for protein immobilization and/or heterogeneity of the orientation of immobilization. In addition, conventional SPR experiments are generally time-consuming. Here we introduce the use of single-site specifically biotinylated kinases combined with a multichannel SPR device to improve such problems. Kinetic parameters of four compounds—staurosporine, dasatinib, sunitinib, and lapatinib—against six kinases were determined by the ProteOn XPR36 system. The very slow off-rate of lapatinib from the epidermal growth factor receptor and dasatinib from Bruton’s tyrosine kinase and colony stimulating factor 1 receptor (CSF1R) were confirmed. Furthermore, IC50 values were determined by an activity-based assay. Evaluating both physicochemical and biochemical properties would help to understand the detailed character of the compound.
Introduction
Recently, kinases have become eagerly investigated molecular targets, because aberrant kinase activation is implicated in the etiology of a wide variety of diseases including cancer, inflammation, and autoimmunity. 1 There are now 25 low-molecular weight kinase inhibitors that have been approved by the Food and Drug Administration for treatment of cancer and immune diseases, and numerous kinase inhibitors are currently in clinical development. 2 Under such circumstances, there is a growing need for more efficient technologies to evaluate kinase inhibitors.
In evaluation of kinase inhibitors, biophysical assays are increasingly of value. Of these, surface plasmon resonance (SPR) technology is a quite powerful technique because it can measure parameters such as rates of association (ka) and dissociation (kd) as well as the equilibrium dissociation constant (KD). Moreover, the required protein amount for SPR is relatively small compared with other biophysical assays, such as calorimetry. Although KD and the half-maximal inhibitory concentration (IC50) have been used as indicators of potency of molecularly targeted drugs, the dissociation rate constant is increasingly recognized as a crucial metric for compound optimization. 3
To use the SPR technology for evaluation of kinase inhibitors, the target kinase needs to be immobilized onto a sensor chip surface. This step has been a bottleneck because of the heretofore required harsh conditions and/or heterogeneity of the orientation of immobilized proteins. The immobilization step is generally conducted through covalent amine coupling or specific interactions such as the antibody-antigen reaction and biotin-avidin interaction. In the case of amine coupling, the low pH required for ligand electrostatic preconcentration on the sensor chip surface often causes irreversible conformational change of immobilized ligand. Although a substantial amount of immobilization can be most likely achieved by using a low pH, the orientation of immobilization is heterogeneous because random lysine residues on kinases are used for the coupling. The molecular weight of the antibody is considerably large and makes it difficult to obtain sensitive results, particularly in the case of protein–small molecule interaction.
Although biotin is often used as an affinity tag for immobilization on surfaces of binding experiments, chemical reagents that conjugate biotin to proteins usually modify multiple sites heterogeneously. This random biotinylation often results in inactivation of target proteins, either directly by the modification of active sites or indirectly because of immobilization in sterically undesirable orientations. On the other hand, single-site–specific biotinylation can be achieved by in vivo enzymatic biotinylation. 4 Such proteins can be immobilized onto the device surface in a favorable orientation and free from direct inactivation. In this study, single-site specifically biotinylated kinases and a multichannel SPR device were combined to circumvent many problems related to SPR-based binding experiments.
Whereas biotin carboxyl carrier protein (BCCP) is known to be a unique physiological substrate of biotinylation enzyme, biotin holoenzyme synthetase (BirA) in Escherichia coli, the artificial peptide substrate of BirA, named as biotin acceptor peptide (BAP), was isolated by screening a random library. 5 Although at least 87 residues are required for the biotinylation of BCCP by BirA, BAP sequence functions as a biotinylation substrate with only 20 amino acids, facilitating its application in protein tagging. Enzymatic and single-site specifically biotinylated kinases can be obtained by fusion of BAP sequence to kinase genes.
Here we present results of using single biotinylated kinases with the ProteOn XPR36 system to determine physicochemical parameters in order to aid assessment of kinase inhibitors.
Materials and Methods
Strains, Media, and Chemicals
Spodoptera frugiperda (Sf9) insect cells were used for the propagation of recombinant baculoviruses. Sf21 cells were used for the expression of proteins. The cells were cultivated in Grace’s insect media (Life Technologies, Carlsbad, CA) supplemented with 10% fetal bovine serum (FBS) and 50 µM biotin. Staurosporine was obtained from Calbiochem (San Diego, CA). Dasatinib 6 and sunitinib 7 were synthesized at Carna Biosciences, Inc. (Kobe Japan). Lapatinib was extracted from the tablet preparation.
Construction of Baculovirus Vectors
To generate the recombinant baculovirus constructs, Escherichia coli BirA and each kinase-encoding gene N-terminally tagged with the BAP sequence and DYKDDDDK (FLAG) sequence were inserted into the multiple cloning sites under the p10 promoter (Pp10) and polyhedron promoter (PPH) of pFastBac Dual (Life Technologies), respectively. As kinase genes, the following six fragments were used: anaplastic lymphoma kinase (ALK), cytoplasmic domain (1058–1620 amino acids of accession number BAG10812.1), Bruton’s tyrosine kinase (BTK), full-length (2–659 amino acids of accession number NP_000052.1), epidermal growth factor receptor kinase (EGFR), cytoplasmic domain (669–1210 amino acids of accession number NP_005219.2), colony stimulating factor 1 receptor (CSF1R), cytoplasmic domain (538–972 amino acids of accession number NP_005202.2), insulin-like growth factor-1 receptor (IGF1R), cytoplasmic domain (959–1367 amino acids of accession number NP_000866.1), polo-like kinase 1 (PLK1), and full-length (1–603 amino acids of accession number NP_005021.2). GST-tagged RAF1 and His(×6) tagged MAP2K1 kinases were constructed as described before. 8
Purification of Kinases
To express single-site specifically biotinylated kinases, Sf21 cells were infected with baculovirus, which contains designed construct in Grace’s insect media supplemented with 10% FBS and 50 µM biotin. After 48 to 72 h of incubation, the cells were harvested, washed with cold phosphate-buffered saline (PBS) buffer, and stored at −80 °C until purification. The frozen cells were thawed and lysed in lysis buffer (50 mM Tris-HCl, pH 7.5, 150 mM NaCl, 0.5 mM EDTA, 0.5 mM EGTA, 1% Nonidet P-40, 1 mM DTT, 2 µg/mL leupeptin, 2 µg/mL aprotinin, 1 mM NaF, 100 µM sodium orthovanadate, 1 µM cantharidin) on ice. All purification procedures thereafter were carried out at 4 °C. The cell lysate was applied to ultrasonication for 1 min and centrifuged at 9000 g for 20 min. The supernatant was recovered and mixed with 1 mL slurry of anti-FLAG beads. After 60 min of incubation at 4 °C, the mixture was centrifuged at 3000 rpm for 2 min, and the supernatant was removed. The slurry was packed in a column and washed three times with 10 mL of wash buffer (50 mM Tris-HCl, pH 7.5, 500 mM NaCl, 10% glycerol, 0.5 mM EDTA, 0.5 mM EGTA, 1 mM DTT). Proteins were eluted with elution buffer (50 mM Tris-HCl, pH 7.5, 150 mM NaCl, 10% glycerol, 100 µg/mL FLAG peptide, 0.05% Brij35), and protein-containing fractions were pooled. GST-tagged RAF1 and His(x6)-tagged MAP2K1 kinases were purified as described before. 8
Protein Identification
The purified proteins were applied to sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) followed by Coomassie brilliant blue (CBB) staining. Each corresponding band was excised and trypsin digested. The peptide mass was analyzed by Autoflex III matrix-assisted laser desorption/ionization reflection time-of-flight mass spectrometry as previously described. 9 Peptide identification was performed using Mascot Peptide Mass Fingerprinting.
Determination of Protein Purity and Biotinylation Ratio
To determine the purity, purified proteins were applied to SDS-PAGE followed by CBB staining and scanned with a densitometer. To estimate the biotinylation ratio, purified proteins were incubated with streptavidin-sepharose HP (GE Healthcare, UK) in storage buffer (50 mM Tris-HCl, 150 mM NaCl, 0.05% Brij35, 1 mM DTT, pH 7.5) followed by centrifugation. Supernatant was applied to SDS-PAGE alongside untreated purified protein and quantified bovine serum albumin protein as standards; rates of protein unbound to streptavidin-sepharose were calculated.
SPR Experiments by Multichannel Sensor Chip
Interactions between biotinylated kinases and kinase inhibitors were investigated by SPR using ProteOn XPR36 system (Bio-Rad Laboratories, Tokyo, Japan). Immobilization of high-density NeutrAvidin (Thermo Fisher Scientific, Waltham, MA) onto the ProteOn GLH sensor chip and subsequent capture of biotinylated kinases were performed as follows. NeutrAvidin is a nonglycosylated avidin mutant known to show much less nonspecific binding than avidin and streptavidin. 10 PBS with 0.005% Tween20 was used as a running buffer during immobilization of NeutrAvidin. After preactivation with a mixture of 0.2 M 1-ethyl-3-(3-dimethylaminopropyl) carbodiimide hydrochloride and 0.05 M N-hydroxysulfosuccinimide, 50 µg/mL of NeutrAvidin was injected for 5 min at a flow rate of 30 µL/min. After the injection of NeutrAvidin, the surface was blocked with 1 M ethanolamine HCl. The final immobilization level for NeutrAvidin was approximately 16,000 RU. Thereafter, 50 µg/mL of each biotinylated kinase was run into each vertical ligand channel for 5 min or more in TBS running buffer composed of 10 mM Tris-HCl, pH 7.5, 0.15 M NaCl, 10 mM MgCl2, 0.05% Tween20, giving immobilization levels ranging from 4000 to 7000 RU. Five concentrations of compounds were analyzed, and the association/dissociation rate constants were calculated by global fitting of the data to the 1:1 binding interaction model.
Enzyme Assay
A kinase activity-based assay was performed by an off-chip mobility shift assay using LabChip 3000 as previously described. 8 Briefly, to determine the IC50 value, each compound was diluted in DMSO in a 10-point half-log series. Each concentration of compound was mixed with kinase and FITC-labeled peptide substrate in assay buffer (20 mM HEPES, pH 7.5, 0.01% Triton X-100, 2 mM DTT, 5 mM MgCl2, and 1 mM ATP) to start the kinase reaction. After 1 or 5 h of incubation at 25 °C, the kinase reaction was stopped by the addition of three times its volume of termination buffer (127 mM HEPES, pH 7.5, 0.01% Triton X-100, 26.7 mM disodium EDTA, 1% DMSO, 0.13% Coating-3 Reagent). Thereafter, the peak height of phosphorylated and nonphosphorylated peptide substrates was measured by LabChip3000 (Perkin-Elmer, Alameda, CA), and the phosphorylation rate of the substrate was defined by P/(P + S). The inhibition percentage of each compound against kinase activity was determined from the phosphorylation percentage of the substrate; IC50 was calculated by interpolation on a log-concentration-response curve fitting for four-parameter logistic equation.
Results and Discussion
Construction and Expression of Single-Site Specifically Biotinylated Kinases
Kinases with N-terminal specific biotinylation are free from active site modification and can be immobilized onto the SPR sensor chip via a biotin-avidin interaction in the mechanically directed orientation with mild condition. To produce such kinases, the BAP sequence was fused at the N-terminus of kinase genes. We cloned a set of kinase genes that are N-terminally fused with BAP and FLAG sequences for ease of purification alongside BirA into the dual expression vector pFastBac Dual, which can express both genes generously in insect cells ( Fig. 1A ). The FLAG sequence also works as the linker, which makes it easier for small molecules to access its binding site in kinases. Kinase genes were the following six species: ALK, BTK, EGFR, CSF1R, IGF1R, and PLK1. BTK and PLK1 were cloned into the vector as full-length genes. For the four transmembrane receptor-type tyrosine kinases, ALK, EGFR, CSF1R, and IGF1R, only the cytoplasmic domains were cloned into the vector (see the “Materials and Methods” section). From insect cells infected with these recombinant baculoviruses, BAP-FLAG–fused kinase proteins were purified using anti-FLAG affinity gel. Purified kinases were applied to SDS-PAGE followed by CBB staining; densitometric analysis revealed that all six kinases were more than 80% pure ( Fig. 1B ). To determine the biotinylation ratio, these kinases were incubated with streptavidin-sepharose beads, and the proportion of absorbed proteins was investigated. The supernatant fraction of streptavidin-sepharose biotinylated kinase incubation mixture and the untreated kinase solution were applied together to SDS-PAGE. HIS-tagged MAP2K1 protein and GST-tagged RAF1 protein without the BAP sequence were treated likewise as negative controls. Although the protein bands were seen on the lanes of the supernatant of HIS-MAP2K1 and GST-RAF1, the bands on the lanes of the supernatant of BAP-fused six kinases were almost not seen, indicating that BAP-fused proteins were nearly completely biotinylated ( Fig. 1C ). Compared with the standard curve generated from threefold dilution series of bovine serum albumin bands, all six species of BAP-fused kinases were estimated to be more than 95% biotinylated ( Fig. 1C ).
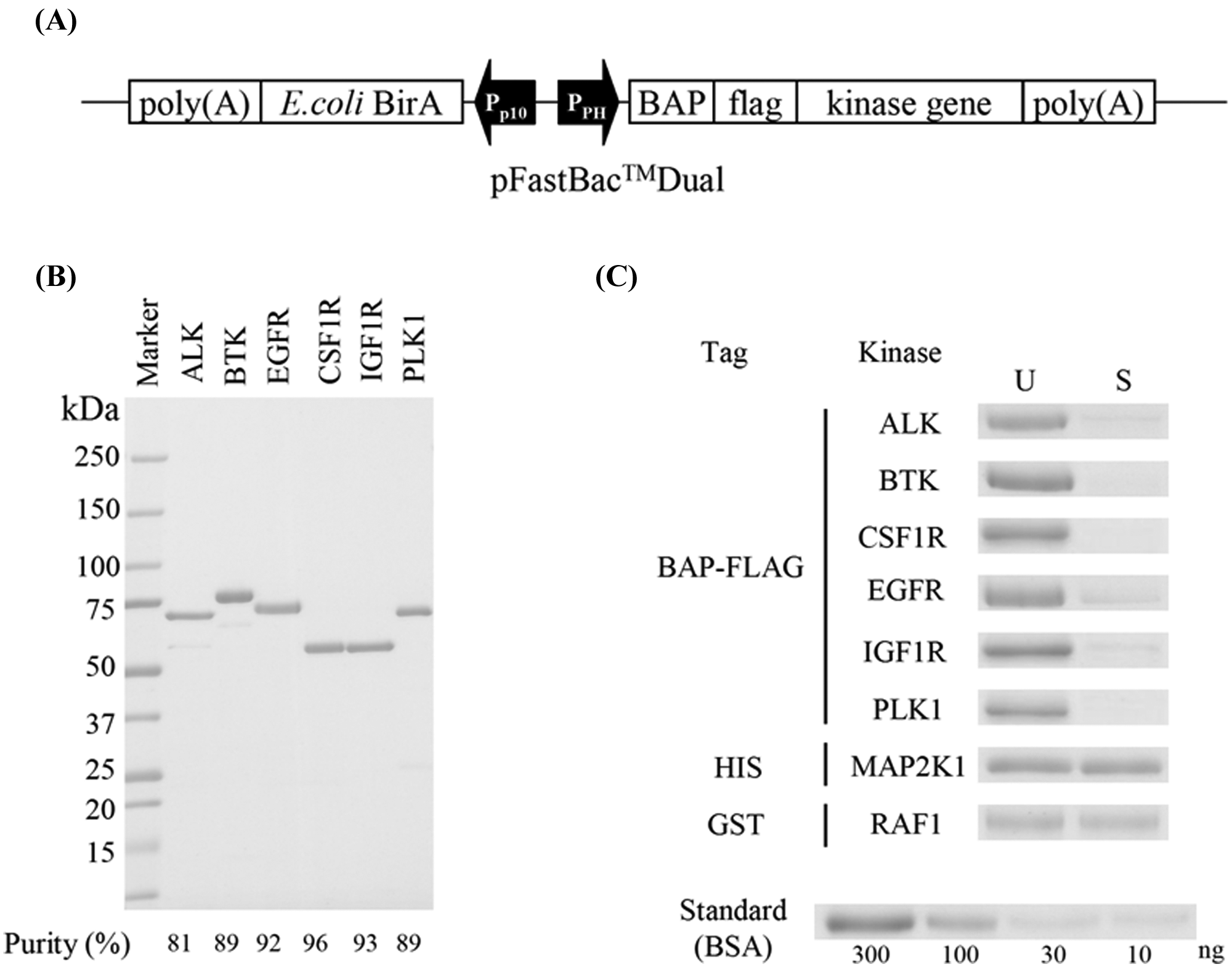
Single-site specifically biotinylated kinases. (
Evaluation of Kinase Inhibitors Using Multichannel SPR
To determine both the association/dissociation rate constants and equilibrium-binding affinities of kinase inhibitors against their targets, SPR-based direct binding assays were conducted using these in vivo enzymatically biotinylated kinases. The ProteOn XPR36 system was adopted, by which it is feasible to measure at once up to 36 interactions, the combination of six immobilized ligands and six analytes. At first, all six vertical channels were coated with NeutrAvidin by amine coupling. The six kinases (i.e., ALK, BTK, EGFR, FMS, IGF1R, and PLK1) were then flowed onto each vertical channel and captured on NeutrAvidin-coated sensor chips. Across all six surfaces, five concentrations of compounds and blank control sample were flowed into six horizontal channels. The analyzed kinase inhibitors were a pan-kinase inhibitor staurosporine and three approved kinase inhibitors (i.e., dasatinib, lapatinib, and sunitinib). All sensorgrams are shown in Figure 2 ; determined kinetic parameters are shown in Table 1 . All six kinases showed some responses to the pan-kinase inhibitor staurosporine, as expected; KD values ranged from 2.1 × 10−8 M to 4.5 ×10−7 M. Dasatinib showed interaction against three kinases and, in each case, showed slow dissociation rates from BTK, FMS, and EGFR in order of decreasing dissociation rate. No interaction was observed between dasatinib and the other three kinases. Lapatinib, which is known to be a highly selective and slowly dissociating EGFR inhibitor, 11 interacted only with EGFR. Sunitinib showed weak interaction against ALK and relatively strong interaction against CSF1R. Among especially strong kinase-inhibitor pairs, such as BTK-dasatinib, CSF1R-dasatinib, and EGFR-lapatinib, association rates were not necessarily high (4.90 × 105 M−1s−1, 1.62 × 105 M−1s−1, and 9.79 × 104 M−1s−1, respectively; Table 1 ). Rather, the association rate of lapatinib-EGFR was relatively slow, but the slow dissociation rate (1.20 × 10−3 s−1) outweighed the slow association rate, giving a low KD value (1.22 × 10−8 M). As for dasatinib, the rank order of equilibrium dissociation constant KD was identical to the dissociation rate constant kd, indicating a key role for kd in the realization of low KD values (BTK 8.50 ×10−10 M, CSF1R 8.64 × 10−9 M, EGFR 1.74 × 10−8 M, respectively).
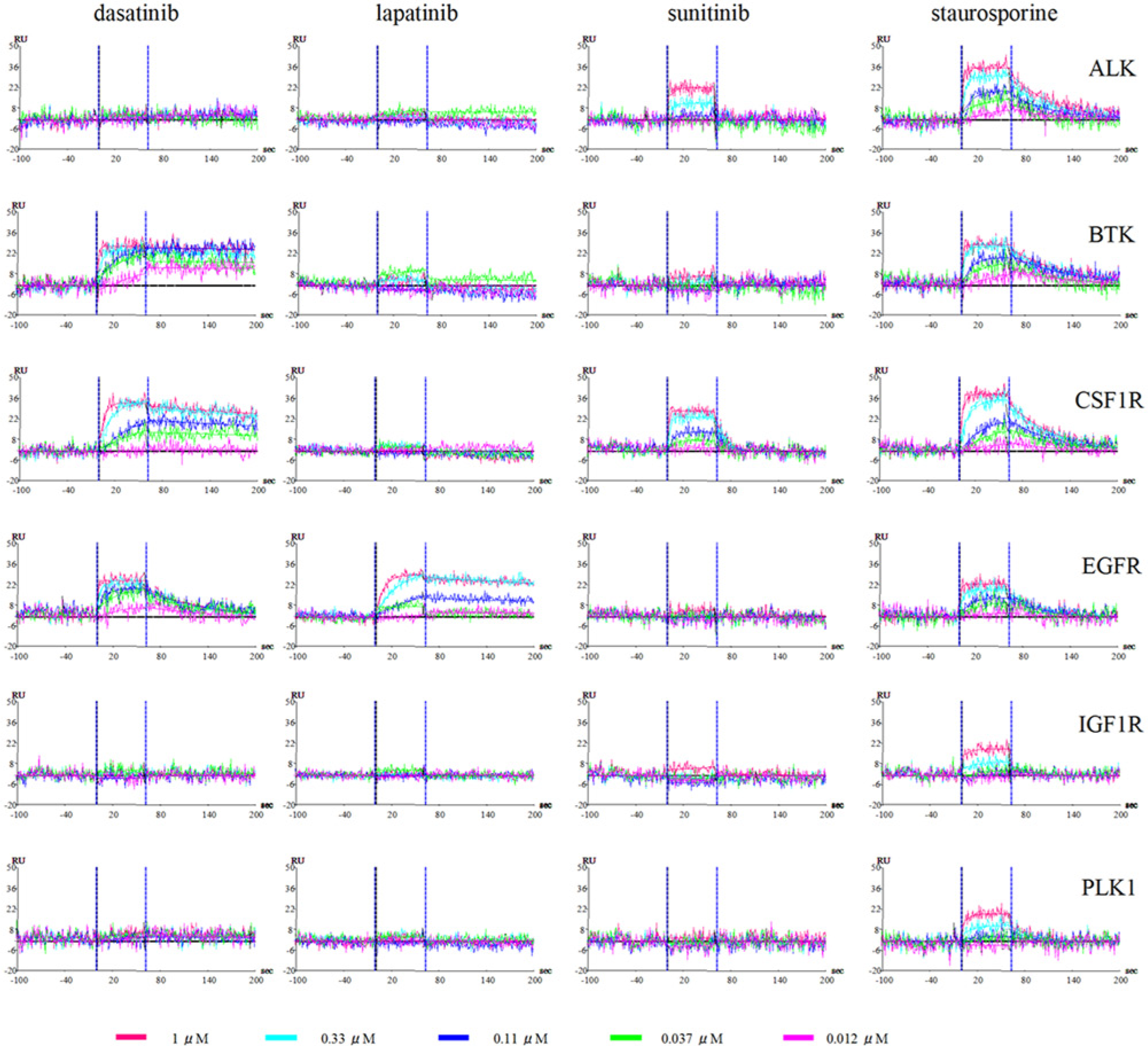
Sensorgrams of four kinase inhibitors binding to immobilized biotinylated kinases. Six species of single-site specifically biotinylated kinases were immobilized onto six vertical channels of a NeutrAvidin-precoated ProteOn GLH sensor chip. Serial threefold dilution of four compounds, including three clinical inhibitors (dasatinib, lapatinib, sunitinib) and a nonspecific kinase inhibitor (staurosporine) were then injected into six horizontal channels of the sensor chip. The association rate constant (ka) and dissociation rate constant (kd) were calculated for each interaction. The concentrations of inhibitors are 1.0 µM (red), 0.33 µM (cyan), 0.11 µM (blue), 0.037 µM (green), and 0.012 µM (magenta).
Kinetic parameters as determined by multichannel SPR assay. a
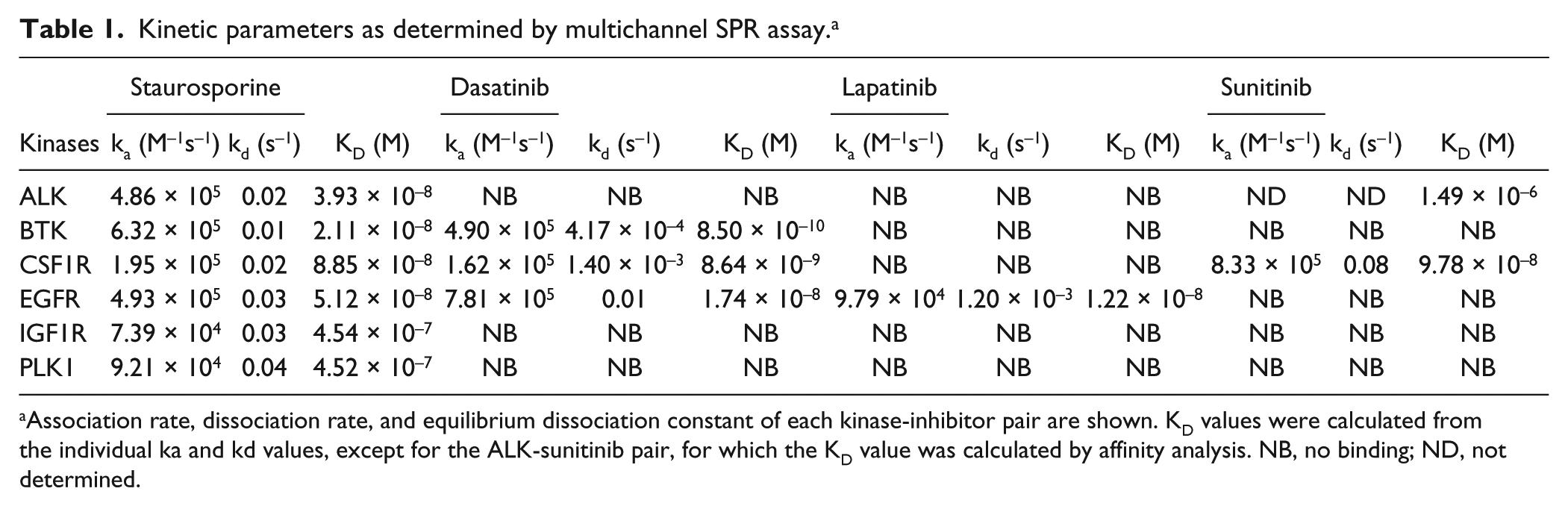
Association rate, dissociation rate, and equilibrium dissociation constant of each kinase-inhibitor pair are shown. KD values were calculated from the individual ka and kd values, except for the ALK-sunitinib pair, for which the KD value was calculated by affinity analysis. NB, no binding; ND, not determined.
Enzymatic Assay Using Biotinylated Kinases
To confirm the availability of enzymatic inhibition data from these six biotinylated protein kinases, IC50 values were determined by activity-based off-chip mobility shift assay. For this purpose, a 10-point half-log dilution series was prepared for each inhibitor, and inhibition rates against the activity of six kinases were determined. Although the activity of EGFR was too low to conduct an activity-based assay, the other five kinases had sufficient activity for IC50 measurement. The determined IC50 values were shown graphically along with KD values ( Fig. 3A ). The IC50 results correlated very well with the KD values. For example, against ALK, staurosporine showed relatively potent interaction, whereas sunitinib showed weak interaction in both the activity-based assay and SPR. These values are very consistent with IC50 values reported elsewhere. 8 Because all four compounds evaluated in this study are ATP-competitive inhibitors, IC50 values are affected by the concentration of ATP used in the assay. To exclude the influence of the concentration of ATP, Ki values were calculated by the Cheng-Prusoff equation using ATP-Km values for each kinase (ALK 240 µM, BTK 89 µM, CSF1R 810 µM, IGF1R 350 µM, PLK1 65 µM, respectively) and plotted to KD values determined by SPR ( Fig. 3B ). As illustrated above, the KD values of these compounds derived from multichannel SPR using single-site specifically biotinylated kinases concurred with data from the activity-based assay. By using these methods, the results can be confirmed mutually.
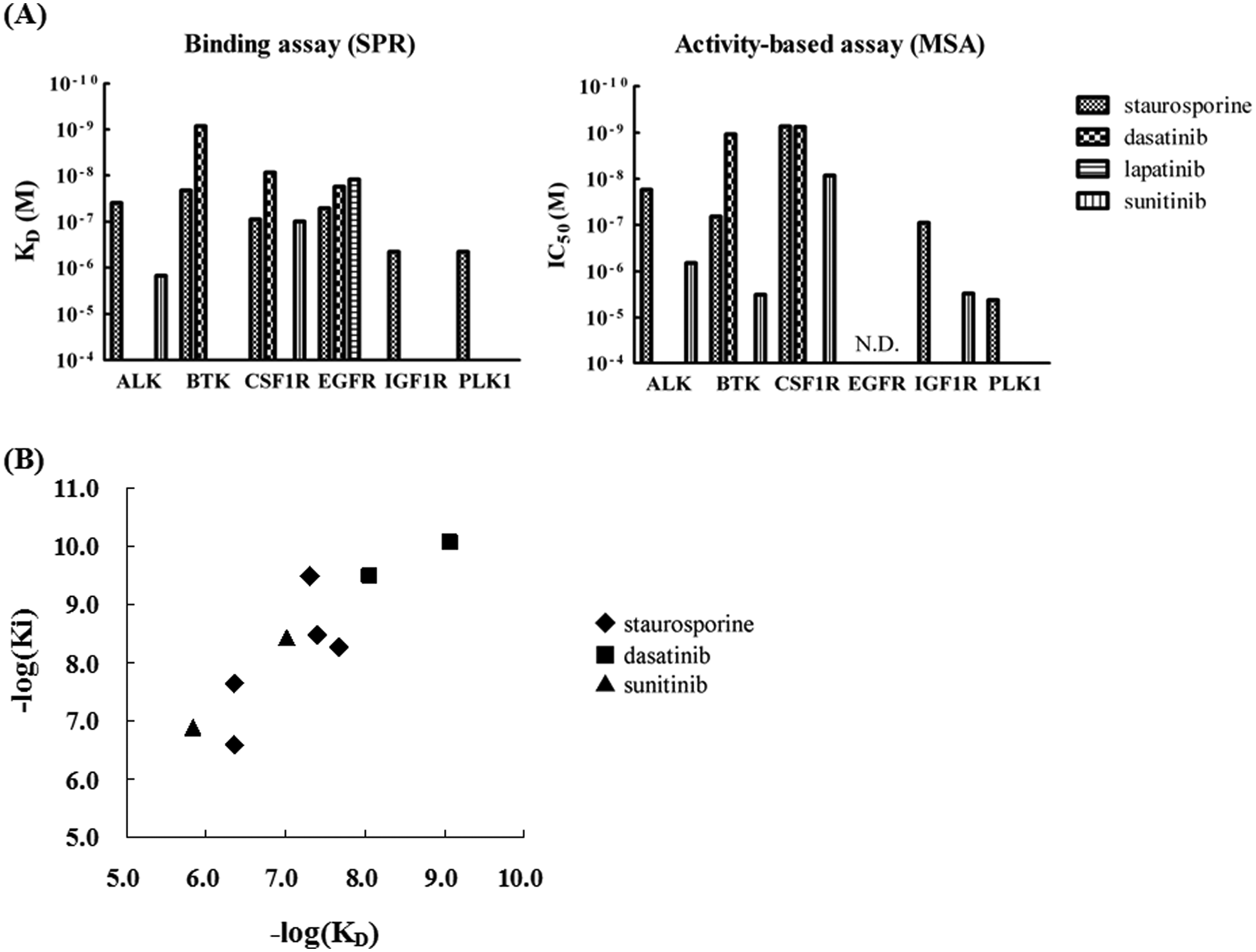
Comparison of results from surface plasmon resonance (SPR) and activity-based off-chip mobility shift assay. (
Implications for Evaluation of Kinase Inhibitors
In kinase inhibitor drug discovery, measuring the IC50 value is relatively easy and convenient because enzyme inhibition of the compound is directly evaluated. However, determining the IC50 against low-activity and inactive kinases is more difficult. Alternatively, by exploiting the affinity assay such as SPR, the equilibrium dissociation constant, KD, can be determined against any kinase state. As shown in this study on EGFR, which has significantly weak kinase activity ( Fig. 3A ), using the present method, KD values were determined easily and quickly.
A number of kinase inhibitors, including approved kinase inhibitors imatinib, sorafenib, and lapatinib, target inactive conformation of kinases. 12 Although kinase inhibitors targeting inactive forms of kinases have been found in many cases serendipitously, KD evaluation offers a rational approach to select such compounds. KD is an intrinsic value, which is independent from assay conditions. In fact, there are good correlations between the results of this investigation, which showed KD < 10 µM, and of a previously reported study by another group using different detection technology. 13 For example, the respective KD values of staurosporine against ALK, BTK, CSF1R, EGFR, IGF1R, and PLK1 were 39 nM, 21 nM, 89 nM, 51 nM, 450 nM, and 450 nM in this study, whereas they were 32 nM, 210 nM, 12 nM, 370 nM, 210 nM, and 190 nM in a previous competitive binding assay. 13 Lapatinib interacted only with EGFR among six kinases with a KD value of 12 nM but of 2.4 nM in the previous study. 13
Dissociation rates of several inhibitors against CSF1R were determined with a competitive binding assay by another group. 14 Uidehaag et al. 14 reported that the dissociation rate constants of staurosporine, dasatinib, and sunitinib against CSF1R and all of these values were approximately 0.02 s−1. In the present study, dissociation rate constants of these three compounds were 0.02 s−1, 0.0014 s−1, and 0.08 s−1, respectively. The reason for the differences between the results from the two studies is possibly due to the differences in phosphorylation states of the kinases. The structure of the ATP-binding pocket drastically changes in correlation with kinase activation status. It is reported that the activated CSF1R was used in the competitive binding assay. 14 On the contrary, the ATP Km of CSF1R used in the present study is 810 µM, whereas that of activated CSF1R is approximately 40 µM, indicating the former is less active. Although most kinase inhibitors are rationally designed molecular-targeted agents, there are many obstacles that make the compounds difficult to be optimized to the clinically applicable medicines. One problem in developing kinase inhibitors is poor physiological property and difficulty of predicting in vivo efficacy from in vitro activity. One clue to the proper correlation between in vitro and in vivo drug activity is the dissociation rate, which may better describe cellular efficacy than KD and IC50 among kinase inhibitors and other type of enzyme inhibitors such as proteases.3,15 A good example of this is given in the study of the AIDS drug saquinavir. Although saquinavir is an effective HIV protease inhibitor in treating AIDS, resistant mutants arise through the treatment. In the study of kinetic parameters of saquinavir against these mutants, it was revealed that the dissociation rate constant correlates very well with the in vivo efficacy of the drug. A strong inverse linear correlation was seen between the log of the dissociative half-life of the HIV-1 protease–saquinavir complex and log of the viral replication IC50. Slow dissociation rates contribute to low KD values; moreover, long residence time for drug-target complexes are considered to give drugs appreciable advantages. 3 Long-dwelling drug-target complexes are favorable for drug efficacy because elevation of the local concentration near the target facilitates rebinding the drug to the target. This further enhances the potency and selectivity by reducing the opportunity of binding to off-target kinases. Of course, the dissociation rate constant alone is not sufficient to illustrate the drug efficacy in vivo because various other factors (e.g., pharmacokinetic parameters) are also critical. However, as in the EGFR-specific inhibitor lapatinib, the slow off-rate is considered to be a significant factor for both long-lasting effect and high selectivity. 11
The gatekeeper residue is a key determinant for potency and selectivity of kinase inhibitors. In the case of dasatinib binding, threonine is the most favorable residue. 16 The critical hydrogen bond is formed between the corresponding threonine of BTK and dasatinib. 16 Furthermore, the substitution of the gatekeeper threonine to more bulky residue hampers penetration of dasatinib into the hydrophobic pocket behind the gatekeeper residue, resulting in the complete abrogation of sensitivity. 17 As shown in X-ray crystallography, dasatinib forms several hydrogen bonds with the hinge region of BTK, resulting in very high affinity (KD = 0.85 nM; Table 1 ). Besides BTK, two other kinases (i.e., EGFR and CSF1R) out of six kinases used in this assay have a threonine as gatekeeper residue. EGFR showed relatively moderate affinity to dasatinib (KD = 17.4 nM; Table 1 ). There are at least six substituted residues in EGFR compared with BTK in the dasatinib binding site that are thought to be responsible for lowering the affinity. 17 In the present study, the difference of affinity against dasatinib between BTK and EGFR (8.50 × 10−10 M and 1.74 × 10−8 M, respectively) is well correlated with the difference of dissociation rate (4.17 × 10−4 s−1 and 0.01 s−1, respectively). CSF1R also showed an interaction with dasatinib, whereas kinases that have other residues than threonine (leucine for ALK and PLK1, methionine for IGF1R, respectively) did not show any interaction with dasatinib. Based on above analysis, the gatekeeper is indeed a critical residue for binding of dasatinib, but the other residues in the binding site would also be important for the slow dissociation rate and high affinity.
In addition to obtaining dissociation rate constants, SPR can be used for new approaches in finding kinase inhibitors with characteristic properties. Recently, allosteric site binders have been evaluated by SPR in parallel with ATP competitors; identification of binding sites and site-specific kinetic parameters were performed well using competition assays. 18
In summary, six species of single-site specifically biotinylated kinases were produced by in vivo enzymatic biotinylation. These kinases can be easily immobilized onto avidin-coated surfaces via biotin-avidin interaction under biochemically mild conditions, facilitating application to biophysical binding assays such as SPR. For the four kinase inhibitors examined here, binding parameters such as association/dissociation rate constants and equilibrium dissociation constants against these six kinases could be determined rapidly by the use of a multichannel SPR device. Slow dissociation rates of dasatinib and lapatinib from their targets were confirmed. In addition, by using the same kinases, activity-based assays could be conducted; results were well correlated with those of multichannel SPR. These methods should facilitate kinase inhibitor design by quick delineation of kinetic profiles of inhibitors.
Footnotes
Acknowledgements
We are grateful to Bio-Rad Laboratories for technical assistance with SPR data collection. We thank Dr. Yusuke Kawase for helpful discussion and all of our colleagues at Carna Biosciences for their support and encouragement.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
