Abstract
A high-throughput prioritization method was developed for use with a validated confirmatory method detecting organophosphorus nerve agent exposure by immunomagnetic separation high-performance liquid chromatography tandem mass spectrometry. A ballistic gradient was incorporated into this analytical method to profile unadducted butyrylcholinesterase (BChE) in clinical samples. With Zhang et al.’s Z′ factor of 0.88 ± 0.01 (SD) of control analytes and Z factor of 0.25 ± 0.06 (SD) of serum samples, the assay is rated an “excellent assay” for the synthetic peptide controls used and a “double assay” when used to prioritize clinical samples. Hits, defined as samples containing BChE Ser-198 adducts or no BChE present, were analyzed in a confirmatory method for identification and quantitation of the BChE adduct, if present. The ability to prioritize samples by highest exposure for confirmatory analysis is of particular importance in an exposure to cholinesterase inhibitors such as organophosphorus nerve agents, in which a large number of clinical samples may be collected. In an initial blind screen, 67 of 70 samples were accurately identified, giving an assay accuracy of 96%, and it yielded no false-negatives. The method is the first to provide a high-throughput prioritization assay for profiling adduction of Ser-198 BChE in clinical samples.
Keywords
Introduction
Cholinesterases are a family of enzymes composed of acetylcholinesterase and pseudocholinesterase, also known as butyrylcholinesterase (BChE).1,2 These enzymes are responsible for triggering hydrolysis of the neurotransmitter acetylcholine.3,4 Hydrolysis of acetylcholine follows nerve stimulation and allows the activated neuron to return to its resting state. Inhibitory compounds such as organophosphorous nerve agents can block this process by binding to the cholinesterase serine active site, causing a buildup of acetylcholine.4,5 This in turn causes the neuron to remain stimulated, sending repeated nerve impulses and leading to constant neuromuscular contraction. In this state, the nervous system is suppressed and patients experience a variety of symptoms related to cholinergic inhibition. 5
When patients exhibit symptoms of cholinergic inhibition, their cholinesterase activity levels are quickly screened so that treatment, if needed, can be administered immediately. Initial screens of activity are effective for fast diagnoses but lack the specificity needed to identify and confirm the chemical agent of exposure. 4 Confirmatory methods undergo formal validation and are reviewed annually within a quality assessment system. The methods to detect organophosphate exposure range from gas chromatography–mass spectrometry 6 and matrix-assisted laser desorption/ionization time-of-flight mass spectrometry 7 to high-performance liquid chromatography tandem mass spectrometry (HPLC-MS/MS).7–11 For each of these methods, the time to first result can vary due to availability of reference materials, the number of analytes monitored, and the complexity of the sample matrix. These limiting factors can be further affected by the number of samples received for analysis.
Although confirmatory methods ensure high-quality analyses for all samples, little discussion on prioritizing those samples has been available until recently. 12 By profiling a validated method analyte known to be affected by exposure, analysts can organize samples so that those suspected of highest exposure are analyzed first. The following HPLC-MS/MS analysis using a ballistic gradient is designed to complement the validated confirmatory method currently in use. This high-throughput assay is used to profile BChE Ser-198 adduction8,10 in serum samples and consequently provide an effective first-tier tool for prioritizing samples for further analysis.
Materials and Methods
Materials
Synthetic unlabeled and stable isotopically labeled BChE peptides (FGESAGAAS and 13C9-FGESAGAAS, respectively) of ≥90% purity were obtained from TNO (the Netherlands). BChE monoclonal antibodies from clone 3E8 were commercially available from ThermoFisher Affinity Bioreagents (Rockford, IL). HPLC-grade solvents acetonitrile and deionized water were purchased from Tedia (Fairfield, OH). Formic acid (98%), phosphate-buffered saline with Tween 20 dry powder, dimethyl pimelimidate dihydrochloride, 0.2 M Tris buffered saline 10× concentrate, triethanolamine buffer solution, and pepsin from porcine gastric mucosa were all ordered from Sigma Aldrich (St. Louis, MO). Dynabeads protein G were purchased from Life Technologies (Carlsbad, CA). KingFisher 96 Flex microplates (200 µL), KingFisher 96-tip combs for deep-well magnets, KingFisher Flex microtiter deep-well 96 plates (v-bottom), protein precipitation plates, 96-well PCR plates (ABgene product), PCR foil, and easy-pierce 20 µm heat-sealing foil were all obtained from Fisher Scientific (Rockford, IL).
Sample Preparation
Samples were processed as previously described and recently modified.9,10 Briefly, 125 µL of each serum sample was pipetted into a multiscreen HTS HV 0.45 µm opaque filter plate, sealed with adhesive PCR foil, and centrifuged at 3000g for 5 min at 20 °C to remove any fibrous tissue. A 75 µL aliquot of the filtered serum was transferred to a 96-well KingFisher deep-well plate. A 50 µL aliquot of prepared BChE antibody-coated magnetic beads was placed into all wells of a 96-well KingFisher shallow-well plate that corresponded to serum-containing wells. A ThermoScientific KingFisher Flex magnetic particle processor was used to transfer the BChE antibody-coated magnetic beads to the wells containing serum. BChE protein was allowed to bind to the antibody beads by shaking with an Eppendorf MixMate at 1400 rpm for 2 h at room temperature. The protein-bound beads were transferred using the KingFisher to a plate containing pepsin for protein digestion. The pepsin plate contained 10 µL of 850 ppb internal standard (isotopically labeled synthetic unadducted BChE peptide, 13C9-FGESAGAAS) and 75 µL 0.6% formic acid. The digestion plate was heated at 37 °C for 30 min on an Eppendorf Thermoshaker mixing at 1000 rpm for 10 s/min. The beads were removed from the sample following digestion using the KingFisher Flex. A 285 µL aliquot of acetonitrile was added to a Pierce 2 mL protein precipitation plate, 0.2 µm filter. The samples from the digestion plate were then transferred by manual pipetting. The samples were filtered through the protein precipitation plate using a vacuum manifold, eluting into a 2 mL 96-well Nunc plate. The samples were dried to completeness under nitrogen pressure using a TurboVap at 70 °C. The samples were resuspended in 75 µL of 0.6% formic acid and transferred to a 96-well PCR plate. The plate was heat sealed with foil for HPLC-MS/MS analysis.
Preparation of Stock Solutions
Quality control (QC) pooled human serum was obtained from Battelle Memorial Institute (Columbus, OH) and characterized for use.9,10 QCs were used to set the threshold for the prioritization method. The QC materials consisted of unexposed serum and VX exposed serum.
HPLC-MS/MS
BChE levels in human serum were determined on an AB Sciex 6500 triple quadrupole mass spectrometer (AB Sciex, Framingham, MA). The instrument was tuned and calibrated monthly over a mass range of m/z 118–2720 using an Agilent ESI tuning mixture (P/N G2421A) and manufacturer autotune procedure. HPLC elution was performed using an Agilent 1290 Infinity series HPLC system (Agilent, Santa Clara, CA). Samples (2 µL) were injected onto a Waters Acquity UPLC BEH C18 VanGuard PreColumn (2.1 × 5 mm, 1.7 µm; Waters, Milford, MA). PreColumn and autosampler temperatures were 25 °C and 10 °C, respectively. Mobile phases were 0.1% formic acid in (A) water/acetonitrile (99.9:0.1) and in (B) water/acetonitrile (0.1:99.9). The following ballistic gradient conditions were used at a constant 250 µL flow rate and average back pressure of 143 bar: 0 min 0%B, 0 to 0.01 min linear gradient 100%B, 0.01 to 0.35 min 100%B, 0.35 to 0.36 min linear gradient 0%B, and 0.36 to 0.50 min 0%B. The following optimized instrument parameters were applied for the detection of the unlabeled analyte and the isotopically labeled internal standard: collision gas = 12 psig, curtain gas = 10 psig, ion source gas 1 = 40 psig, ion source gas 2 = 60 psig, ion spray voltage = 5500 V, temperature = 500 °C, collision exit potential = 20 V, declustering potential = 136 V, entrance potential = 10 V, dwell time = 35 ms, 5.0 mm vertical probe height, and a unit resolution of 0.7 amu at full-width half max. Detection of the BChE pepsin digest product peptide FGESAGAAS was determined by selected reaction monitoring (BChE detection ion m/z 796.3→691.3, collision energy = 34 V; confirmation ion m/z 796.3→620.3, collision energy = 36 V, and 13C9-BChE internal standard m/z 805.4→700.3, collision energy = 34 V) in ESI positive ion mode.
Data Acquisition and Processing
Crystal structures were produced using PyMOL v. 1.5 from Schrödinger LLC (Portland, OR). Data acquisition and spectral analysis were carried out using AB Sciex Analyst v.1.6 build 6536. The capacity factor for the separation was calculated by k′ = (tr – tm)/tm, where tr is the analyte retention time and tm is the system void volume. Zhang et al. 1999’s Z factor of the 96-well plate assay was calculated as Z = 1 – ((3σ s + 3σ c )/│µ s – µ c │), where σ is the standard deviation, µ is the mean, c is the control, s is the sample, and abs is the absolute value. 13 Similarly, the Z′ factor was calculated. 13 The percentage relative standard deviation %RSD = (SD/Cavg) × 100 was calculated as a measure of assay precision, where Cavg is the average concentration calculated and SD is the standard deviation of Cavg. Intraplate precision was determined with an analyte n ≥ 48, and interplate accuracy was determined with a plate n = 3. Assay accuracy was calculated as %Accuracy = (ncorr/ntotal) × 100, where ncorr is the number of samples prioritized correctly and ntotal is the number of samples total. The assay’s confirmation rate was determined as a percentage by %CR = (hits confirmed/hits selected) × 100, where hits are the samples found to have low unadducted BChE levels and thereby high priority for further analysis.
Convenience Sample Set
A set of 100 individual human sera samples was commercially obtained from Tennessee Blood Services (Memphis, TN).
Results and Discussion
A novel validated method for extracting human BChE from serum or plasma by immunomagnetic separation coupled with isotope-dilution HPLC-MS/MS was developed by Sporty and colleagues in 2010.8,10 This combination of analytical techniques enabled analysts to obtain and quantitatively measure a direct protein biomarker of organophosphate exposure from clinical samples. Because BChE is functional in ≥99.9% of the general population, 14 profiling this biomarker provides a meaningful target for prioritizing samples received during a potential exposure to cholinesterase inhibitors. Detection of organophosphate adducts to the BChE target are projected up to 55 days postexposure, expanding response time by 95% from the urinary metabolite method’s 3 day response time.6,11 Although further automation of the adduct method led to more efficient sample processing times, sample reporting remained dependent on a 10 h/96-well plate HPLC-MS/MS analysis and the subsequent data review of the multianalyte system. 9 If samples were prioritized before they were analyzed with the more complex confirmatory methods, samples with the highest exposure profiles would have a faster time to report. The benefits of this prioritization design ensure that investigators obtain information about a chemical emergency in the order of highest exposure first.
The key to adding a prioritization method before the confirmatory method was to not prolong the original time to first result. The confirmatory method analysis time for initial calibration and QCs was 1.75 h. A high-throughput prioritization method was designed to profile at least 96 unknown clinical samples in less than 1.75 h. The desired method was achieved using a ballistic HPLC gradient. First termed in 2000, ballistic gradients are rapid gradients at high flow rates on short and narrow-bore columns. 15 A 0.5-min ballistic gradient HPLC-MS/MS analysis was developed to target unadducted BChE peptide at a fifth of the confirmatory method’s sample volume ( Fig. 1 ). A BChE peak at 0.24 min was observed with a measured k′ = 3.8. Because the BChE peptide used was the same as the biomarker peptide extracted for the confirmatory method, the method had the additional benefit of not altering the original sample-processing protocol. 10 The method enabled BChE profiles to be collected from a 96-well plate in 1.6 h and queued into the confirmatory analysis method in order of suspected exposure.
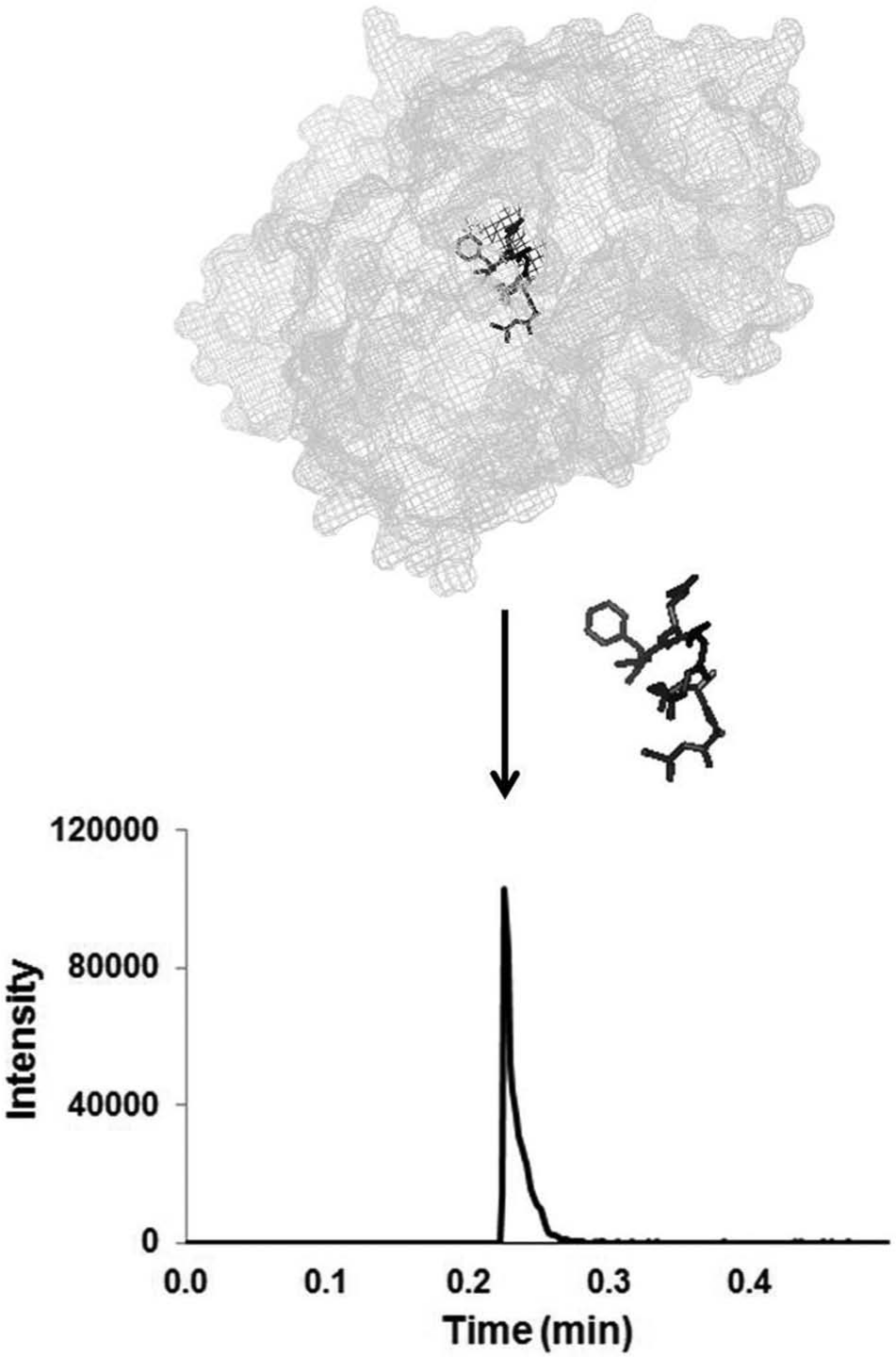
Human butyrylcholinesterase (BChE) was extracted by immunomagnetic separation, digested by pepsin, and analyzed by high-performance liquid chromatography tandem mass spectrometry analysis. The x-ray crystal structure of human BChE with waters removed for clarity was created using PyMOL (PDB identifier 1P0M). The Ser-198 containing BChE peptide (FGESAGAAS) is highlighted in black. A representative extracted ion chromatogram of the peptide transition m/z 796.3→691.3 is shown at retention time 0.24 min (k′ = 3.8).
The analysis of the unadducted BChE in each sample included monitoring the ESI-MS/MS target ion transition m/z 796.3→691.3, confirmatory target ion transition m/z 796.3→620.3, and the isotopically labeled internal standard ion transition m/z 805.4→700.3. The peak area ratio of the target to the internal standard was used to establish assay parameters. Assay quality was determined using Zhang et al.’s high-throughput Z-factor parameter. 13 A checkerboard 96-well plate of alternating unadducted and adducted synthetic BChE peptide standards was used to assess the developed method’s ability to discriminate between unadducted BChE target and adducted BChE ( Fig. 2A ). The peptide standards were prepared at 37 ng/mL, equivalent to the QC material concentration measured in the confirmatory method. Only baseline detection of adducted BChE synthetic standards was observed due to the high specificity of MS/MS analysis. The calculated Z′ factor was 0.88 ± 0.01 (SD) and described an “excellent assay” capable of distinguishing the target with confidence. 13 The high precision of the peptide standard was 4%RSD intraplate (n = 48) and 7%RSD interplate (n = 3). A naturally lower Z factor was predicted and observed for matrix serum targets from clinical samples ( Fig. 2B ). The calculated Z factor for serum QC reference material was found to be 0.25 ± 0.06 (SD) and termed a double assay. 13 This classification was as intended because the profiling method was designed as a means of prioritization before the validated confirmatory method. Profiles of unadducted BChE extracted from the unexposed QC reference material were found to be less precise than the synthetic standards but still within a high working range of 13%RSD intraplate (n = 48) and 15%RSD interplate (n = 3).
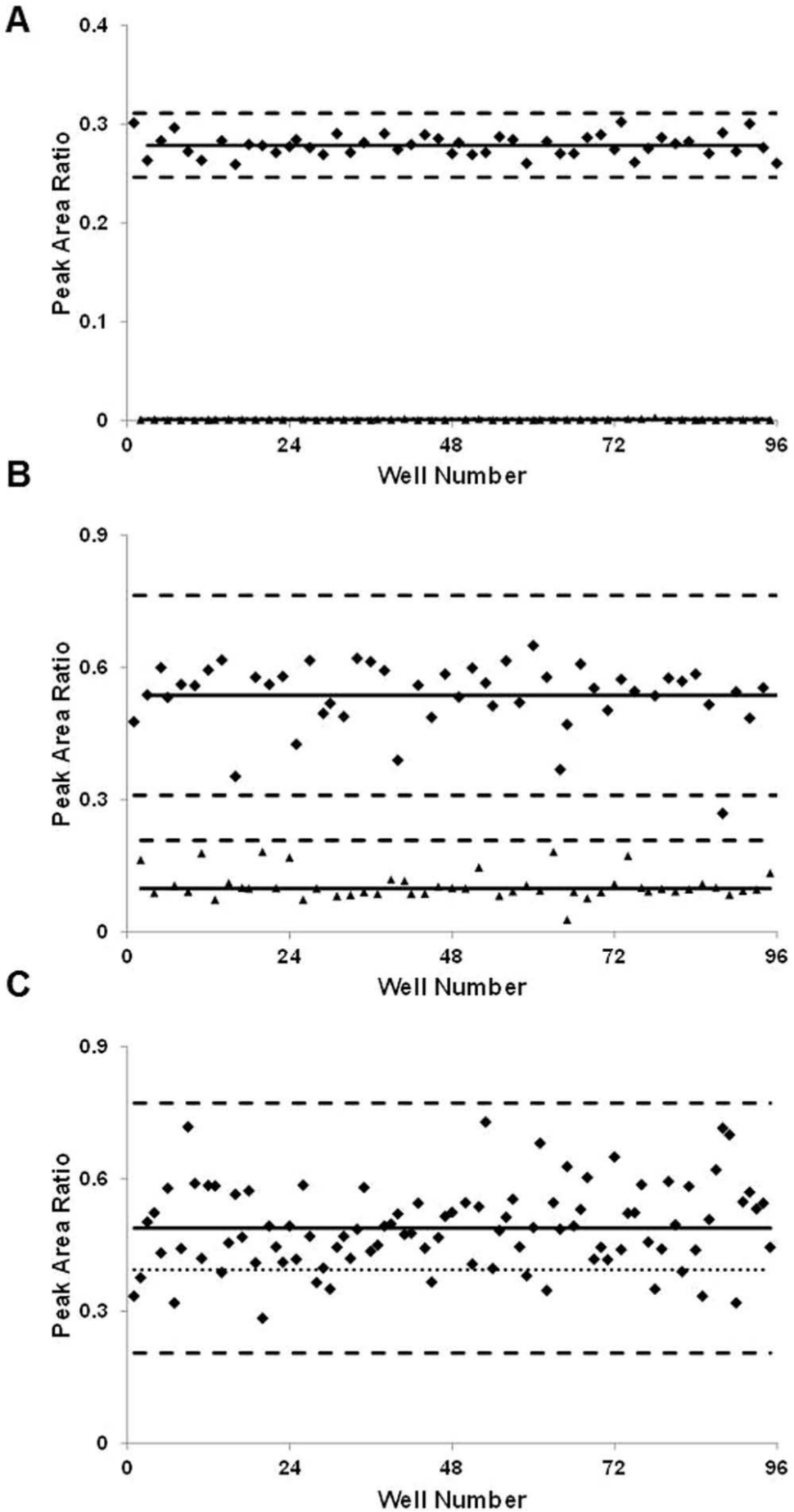
Zhang et al.’s Z factor
13
was used to evaluate assay quality across each 96-well plate. The Z factor parameter was calculated using a checkerboard 96-well plate analysis for the system’s (
A high confidence in the initial assay quality was established, and a commercial convenience set of individual human serum samples was tested. The BChE target peptide was observed in all of the 96 individual samples with an average response not unlike that observed in the serum QC reference materials ( Fig. 2C ). An intraplate precision of 19%RSD was calculated among the individual samples and was used to set the threshold for the assay. A threshold of 19% of the mean was representative of the reported 22% population variability. 5 Samples with a response lower than one standard deviation of the mean (equivalent to 19%RSD) would be classified as “hits” or high-priority samples in the confirmatory analysis queue. The authors understand a threshold at one standard deviation of the mean will return an abnormally high number of false-positives because of the inherent target BChE variability in the general population, but this low threshold will also reduce, if not eliminate, the number of false-negatives observed. The importance of avoiding false-negatives in a chemical emergency or exposure sample set is of utmost importance for formal investigation purposes.
The final prioritization method parameters were applied to a blind response sample set. A blinded plate containing 70 potential exposure samples was prepared and profiled for priority of suspected organophosphate exposure. Samples were composed of an array of QC materials and individual serum samples. Of the 70 samples analyzed, 67 samples were classified correctly, giving a high assay accuracy of 96%. The confirmation rate of the identifications was also high at 93%, which means that of the 45 hits prioritized, 42 were later confirmed ( Fig. 3 ). The three unconfirmed hits were labeled false-positives; no false-negatives were observed. A tested assay threshold that did not return false-negatives is extremely important when coupled with a confirmatory method for organophosphate exposure.
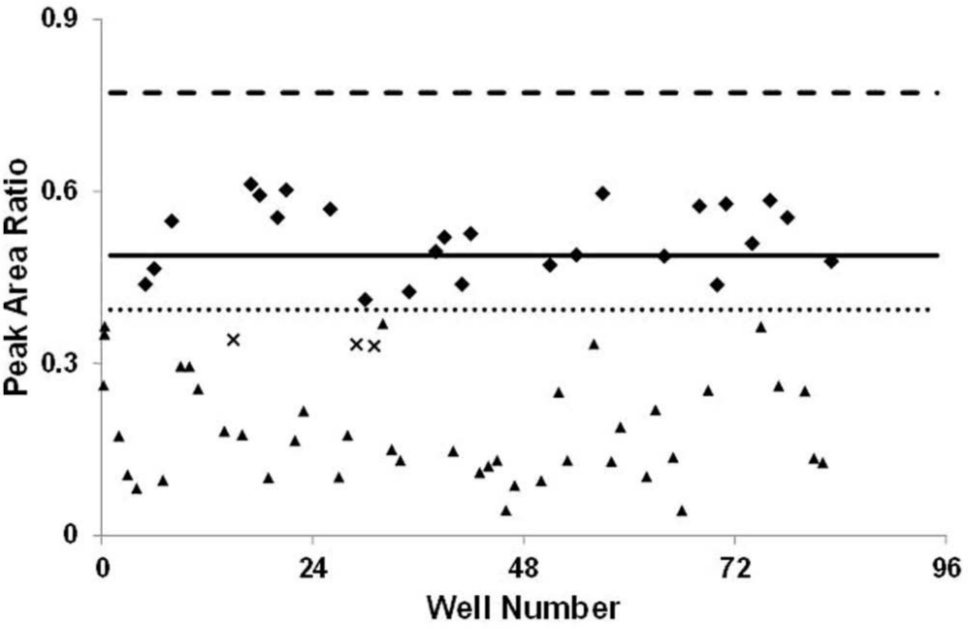
The assay was used to prioritize samples from a blind response exercise. The assay was shown to have an accuracy of 96%, evaluating 67 of the 70 samples as expected. There were 42 known hits of high priority (▲) in the sample set; the 42 hits were confirmed plus three false-positives (x) and no false-negatives, giving a high confirmation rate of 93%. All other samples (♦) are distributed around the mean. The mean (solid line), three standard deviations of the mean (dashed line), and one standard deviation of the mean (dotted line) were the parameters established from the individual serum convenience set.
The described high-throughput prioritization method was developed for use with a validated confirmatory method detecting organophosphate exposure by immunomagnetic separation HPLC-MS/MS. Although all potential exposure samples received would be analyzed with equal care using a validated confirmatory method, sample profiles with suspected organophosphate exposure would be queued for analysis first. A prioritization assay accuracy of 96% and confirmation rate of 93% illustrates the confidence with which this approach can be used. Prioritizing potential exposure samples therefore increases the value of the validated method’s time-to-first-result and will be important in organizing sample analyses in the case of a large-scale exposure event.
Footnotes
Acknowledgements
The authors would like to thank Ms. Chariety Sapp of the CDC’s Incident Response Laboratory (IRL) for dispensing convenience set serum samples prior to analysis.
Disclaimer
The findings and conclusions in this article are those of the authors and do not necessarily represent the views of the Centers for Disease Control and Prevention. Use of trade names is for identification only and does not imply endorsement by the Centers for Disease Control and Prevention, the Public Health Service, or the US Department of Health and Human Services.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by Defense Threat Reduction Agency contract CDC IAA: OT12-010.
