Abstract
The macromolecular synthesis assay was optimized in both S. aureus and E. coli imp and used to define patterns of inhibition of DNA, RNA, protein, and cell wall biosynthesis of several drug classes. The concentration of drug required to elicit pathway inhibition differed among the antimicrobial agents tested, with inhibition detected at concentrations significantly below the minimum inhibitory concentration (MIC) for tedizolid; within 4-fold of the MIC for ciprofloxacin, cefepime, vancomycin, tetracycline, and chloramphenicol; and significantly above the MIC for rifampicin and kanamycin. In a DNA gyrase/topoisomerase IV structure-based drug design optimization program, the assay rapidly identified undesirable off-target activity within certain chemotypes, altering the course of the program to focus on the series that maintained on-target activity.
Introduction
Bacterial resistance to existing drugs has increased dramatically over the past two decades, and many pathogenic strains are multiply resistant to nearly all major classes of antibacterial agents. This poses a serious and growing threat to public health. 1 Since 2000, the Food and Drug Administration (FDA) has approved only two antibiotics that act via novel mechanisms. However, both agents (daptomycin and linezolid) are inactive against infections caused by Gram-negative pathogens. Consequently, there is an urgent medical need for new antibacterial agents active against these organisms, where clinical resistance rates are high. A new agent, acting via a novel mechanism of action, would be particularly valuable. Despite the acute needs, the pharmaceutical industry has struggled to introduce new antibiotic drugs, especially those of novel classes. 2
A vital part in the discovery and development of antibacterial drugs is an accurate evaluation of mechanism of action (MOA). This is particularly important for compounds acting via novel mechanisms. The effect of test compounds on the synthesis of macromolecules has been measured for more than 40 years by monitoring inhibition of incorporation of radiolabeled precursors into major biosynthetic pathways, including DNA, RNA, protein, and cell wall biosynthesis.3–5 These assays were originally designed in large culture formats with 1-mL aliquots removed at the end of incubation periods that were then precipitated with trichloroacetic acid (TCA), filtered, washed, dried, and counted. 3 Incorporation was sometimes measured over long incubation times (1–2 h), where secondary effects of the antimicrobial agent could confound the results.6,7 Microplate formats were later developed to aid in assay throughput. 7 The Escherichia coli imp (increased membrane permeability) strain, which contains a mutation in an outer membrane protein, has been used in these assays since the strain is more susceptible to a wide range of antimicrobial compounds, enabling assessment of compounds that typically do not permeate Gram-negative bacteria. 7
Many of these methodologies have been used to corroborate a particular mechanism late in a research program, rather than identify potential off-target activities early in optimization. Hilliard et al. 8 examined compounds from their histidine protein kinase (HPK) inhibitor program, as well as compounds from other published HPK programs, some of which had progressed to acute in vivo toxicity experiments. Using a combination of macromolecular synthesis and membrane integrity assessments, they demonstrated that bacterial killing was likely due to membrane disruption rather than HPK inhibition. 8
As part of our structure-based drug design (SBDD) efforts, we required a rapid, high-throughput assay capable of evaluating the MOA of novel antibacterial agents and identifying unwanted, off-target liabilities. To this end, a modified protocol was established for high-throughput macromolecular labeling of both a Gram-positive and Gram-negative organism: Staphylococcus aureus and E. coli imp. As part of the validation/characterization process, we evaluated a variety of known antibacterial agents that target different cellular processes to obtain mechanistic fingerprints for different cellular targets. Here we report both the mechanistic fingerprints and the use of these assays to identify off-target liabilities during an SBDD program targeting DNA gyrase and topoisomerase IV.
Materials and Methods
Reagents
All reagents were obtained from Sigma-Aldrich (St. Louis, MO) except ciprofloxacin (Cellgro, Manassas, VA), daptomycin (Biotang, Waltham, MA), linezolid (ChemPacific, Baltimore, MD), and Mueller Hinton cation-adjusted (MH-CA) medium (Remel, Lenexa, KS). Tedizolid and preclinical inhibitors of DNA gyrase/topoisomerase IV were synthesized by Trius Therapeutics, Inc. (San Diego, CA).
Radioisotopes
[3H]-thymidine (80 Ci/mmol) and [3H]-leucine (108 Ci/mmol) were obtained from PerkinElmer (Waltham, MA). [3H]-uridine (30 Ci/mmol) and [3H]-N-acetylglucosamine (30 Ci/mmol) were purchased from American Radiolabeled Chemicals (St. Louis, MO).
Bacterial Strains
Studies were performed using S. aureus ATCC 29213 (American Type Culture Collection, Manassas, VA) and E. coli imp (BAS849) (Spencer A. Benson, University of Maryland, College Park, MD). 9
Antibacterial Activity
Minimum inhibitory concentration (MIC) values were determined by Clinical and Laboratory Standards Institute broth microdilution method M7-A7 (CLSI, 2006), modified by dilution of the test compound in DMSO or water instead of test media, and interpreted using alamarBlue (Invitrogen, Carlsbad, CA) to visualize cell viability. Assays were conducted in MH-CA medium except for those with daptomycin, which were conducted in Mueller Hinton with the addition of magnesium chloride and calcium chloride to final concentrations of 12.5 µg/mL magnesium and 50 µg/mL calcium. Compound stocks were prepared at 3.2 mg/mL in either water or 100% DMSO. Antibacterial activity was assessed using 16-point, 2-fold serial dilutions covering a range from 64 to 0.002 µg/mL.
Defined Growth Media
Defined growth media (DM) used for macromolecular synthesis comprised the following: 1× M9 salts; 0.5% glucose (w/v); 1 mM MgCl2; 20 µg/mL CaCl2; 2 µg/mL thiamine; 2 µg/mL nicotinic acid; 100 µM L-alanine, L-asparagine, L-aspartate, L-glutamate, L-glycine, L-proline, and L-serine; and 200 µM L-arginine, L-cysteine, L-histidine, L-isoleucine, L-lysine, L-methionine, L-phenylalanine, L-threonine, L-tryptophan, L-tyrosine, and L-valine. Media was supplemented with 1 µM L-leucine for growth of E. coli imp.
Macromolecular Synthesis Assay
Unless indicated, an identical protocol was used for both S. aureus and E. coli imp.
Cell Growth
Cells grown overnight in MH-CA at 37 °C with shaking were diluted 1:200 in prewarmed DM and grown at 37 °C with shaking (300 rpm) until mid-log phase was attained (A600 nm = 0.3).
Sample Preparation
Radioisotope precursors (10× final concentration, in 2× DM) and test compounds (10× final concentration, in 20% DMSO/80% water) were prepared. Typically, for test compounds, an 8-point 2-fold serial dilution was made in 20% DMSO/80% water. Wider dilution series were used as needed. For each [3H]-precursor, a separate 96-well, V-bottomed polypropylene plate (Corning 3357; Corning, Tewksbury, MA) was prepared containing 10 µL of the 10× stock per well. To columns 1 and 2, 10 µL of 20% DMSO/80% water was added. These columns provided background (column 1) and maximal incorporation (column 2) controls. Test compounds were added to columns 3 to 12, with 10 µL of the 10× concentrations per well. Once prepared, plates were prewarmed to 37 °C.
Macromolecular Synthesis
In columns 2 to 12, 80-µL aliquots of mid-log stage cells (A600 nm = 0.3) were added to wells. Media only was added to column 1 (cell-free blank). Plates were incubated at 37 °C with continued shaking for 15 min (E. coli imp) or 25 min (S. aureus). At this point, incorporation was terminated by the addition of 50 µL 30% TCA/70% ethanol to each well. Plates were incubated at room temperature for 60 min with shaking at 300 rpm to enable precipitation of macromolecular material. During this time, 96-well filter plates (MSHVN4B50; Millipore, Billerica, MA) were prepared by washing with 150 µL 70% ethanol according to the manufacturer’s instructions. Test aliquots of 130 µL (of a total 150 µL) were transferred to the filter plates, filtered, and washed with 4 × 150 µL 5% TCA followed by 1 × 150 µL 95% ethanol. Plates were dried, the bottoms were sealed (Millipore MATAH0P00), and 50 µL OptiPhase Supermix (Perkin Elmer, Waltham, MA) was added to each well and the top sealed with transparent film (Millipore MATAHCL00). Plates were counted using a Wallac MicroBeta (PerkinElmer) scintillation counter.
Enzyme Assays and Inhibition Studies
GyrB and ParE activities were evaluated using a coupled spectrophotometric assay in which the enzyme-dependent release of inorganic phosphate from adenosine triphosphate (ATP) hydrolysis was measured. 10 The assay contained between 20 and 100 nM GyrB or ParE (active site concentrations) in 50 mM Tris-HCl buffer (pH 7.6), 2 mM MgCl2, 125 mM NaCl, 0.2 mM 7-methyl-6-thioguanosine, and 1U/mL purine nucleoside phosphorylase. The reaction was initiated by addition of 3 mM ATP and monitored at 360 nm for 30 min at 27 °C. Inhibitor potency was determined by incubating the target enzyme in the presence of various concentrations of inhibitor ranging between 1.5 nM and 50 µM for 10 min prior to addition of ATP substrate. The final concentration of DMSO was kept constant at 2.5% (v/v). Enzyme activity in the presence of inhibitor was expressed relative to the no-inhibitor control and Ki values determined using the Morrison tight-binding equation to account for ligand depletion. All analysis was carried out using GraphPad Prism 4.0 (GraphPad Software, La Jolla, CA).
Bacterial Growth and Radioisotope Incorporation
To maximize incorporation of radiolabeled precursors into newly synthesized macromolecules in as short a period as possible, we wanted robust and reproducible growth of bacterial cells in defined media containing limited amounts of unlabeled precursors. Preliminary experiments led to the identification of minimal nutrient conditions capable of supporting robust growth (data not shown). No additional sources of thymidine, uridine, leucine, or N-acetylglucosamine were required to support growth of S. aureus, whereas E. coli imp demonstrated more reproducible growth by the inclusion of a low level of leucine (1 µM). For both bacterial species, a short lag phase was noted, followed by exponential growth until the stationary phase was reached, with optical density A600 nm >1. Under these conditions, the doubling time of S. aureus was considerably longer than for E. coli imp, approximately 60 and 30 min, respectively. Growth of both S. aureus and E. coli was highly reproducible under these conditions, with cells reaching mid-logarithmic growth typically within a 15-min window. For subsequent studies, an optical density A600 nm = 0.3 was chosen, representing logarithmically growing cells with a suitably high cellular density, adequate for isotope labeling.
Radioisotope incorporation was measured for approximately half the bacterial generation time (i.e., 25 min for S. aureus and 15 min for E. coli imp) and was linear for the duration of the experiment. The optimized precursor concentrations and specific activities for S. aureus and E. coli imp are summarized in
Results
Evaluation of Known Antibacterials Using the Macromolecular Synthesis Assay
The MIC values of compounds were determined and are shown in
Table 1
. An 8-point dilution series spanning the MIC value was typically used in the macromolecular synthesis assay. If incomplete or no pathway inhibition was observed, the concentration range for testing was expanded. Using this assay, we evaluated the primary effects of a number of known antibacterial agents of different classes against S. aureus and E. coli imp, including inhibitors of nucleic acid synthesis, cell wall synthesis (supplemental information), and protein synthesis (supplemental information). The MIC value for each compound is indicated by a vertical dotted line in
Figures 1
to
3
and
Antimicrobial Activity of Compounds vs Strains Used in the Macromolecular Synthesis Assay.
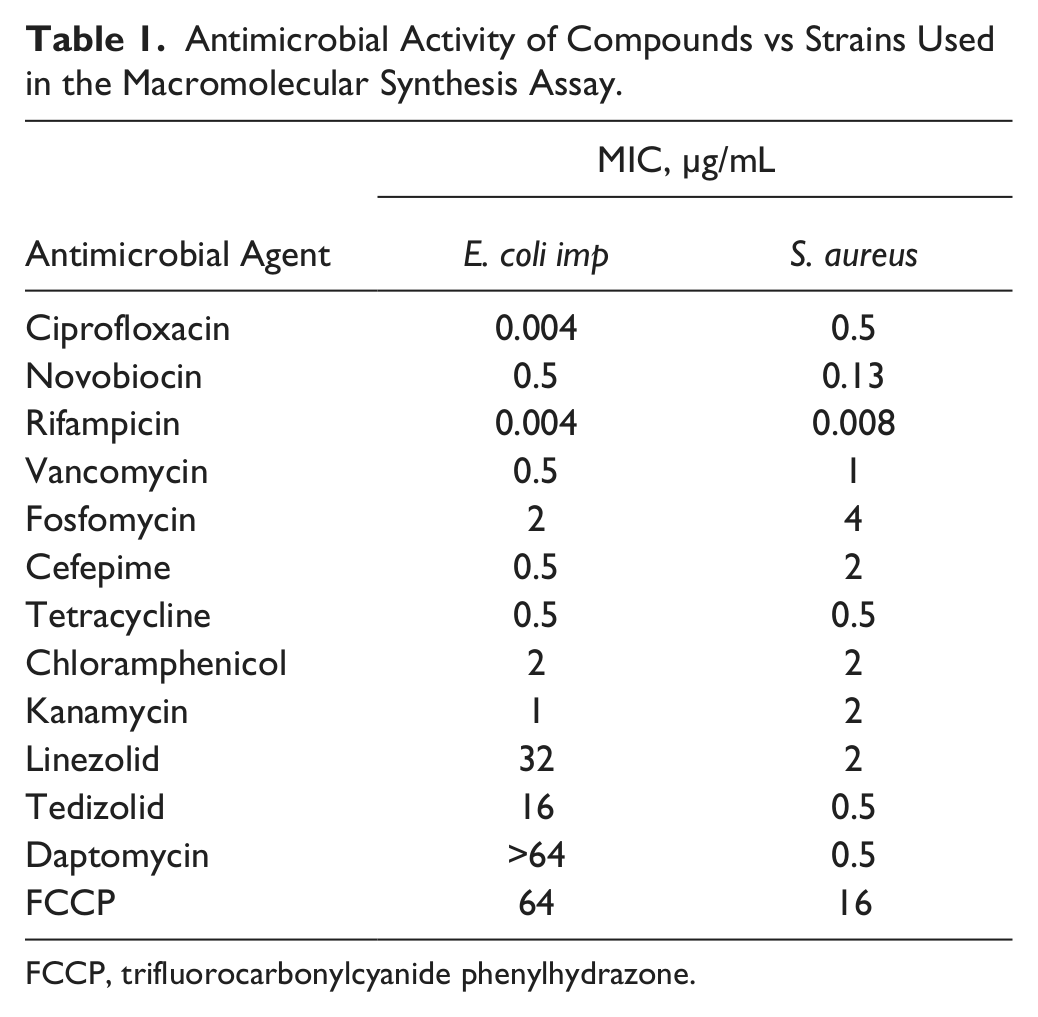
FCCP, trifluorocarbonylcyanide phenylhydrazone.
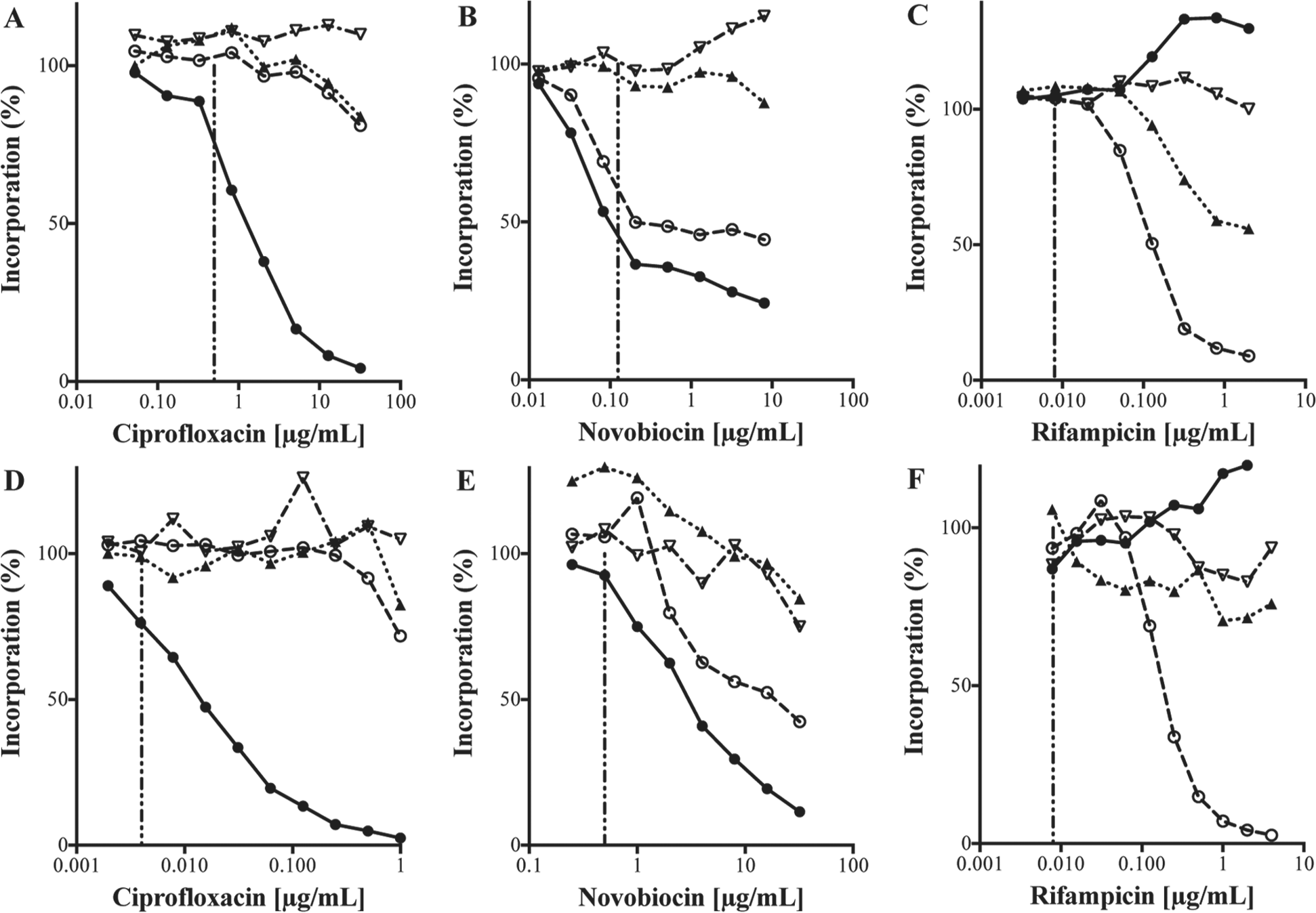
The effect of antibacterial agents targeting nucleic acid pathways on macromolecular synthesis in S. aureus and E. coli imp. Incorporation of [3H]-precursors of DNA (●), RNA (○), protein (▲), and cell wall (▽) was examined as described in the Materials and Methods. The MIC value for each compound is indicated by a vertical dashed line. S. aureus plus ciprofloxacin (
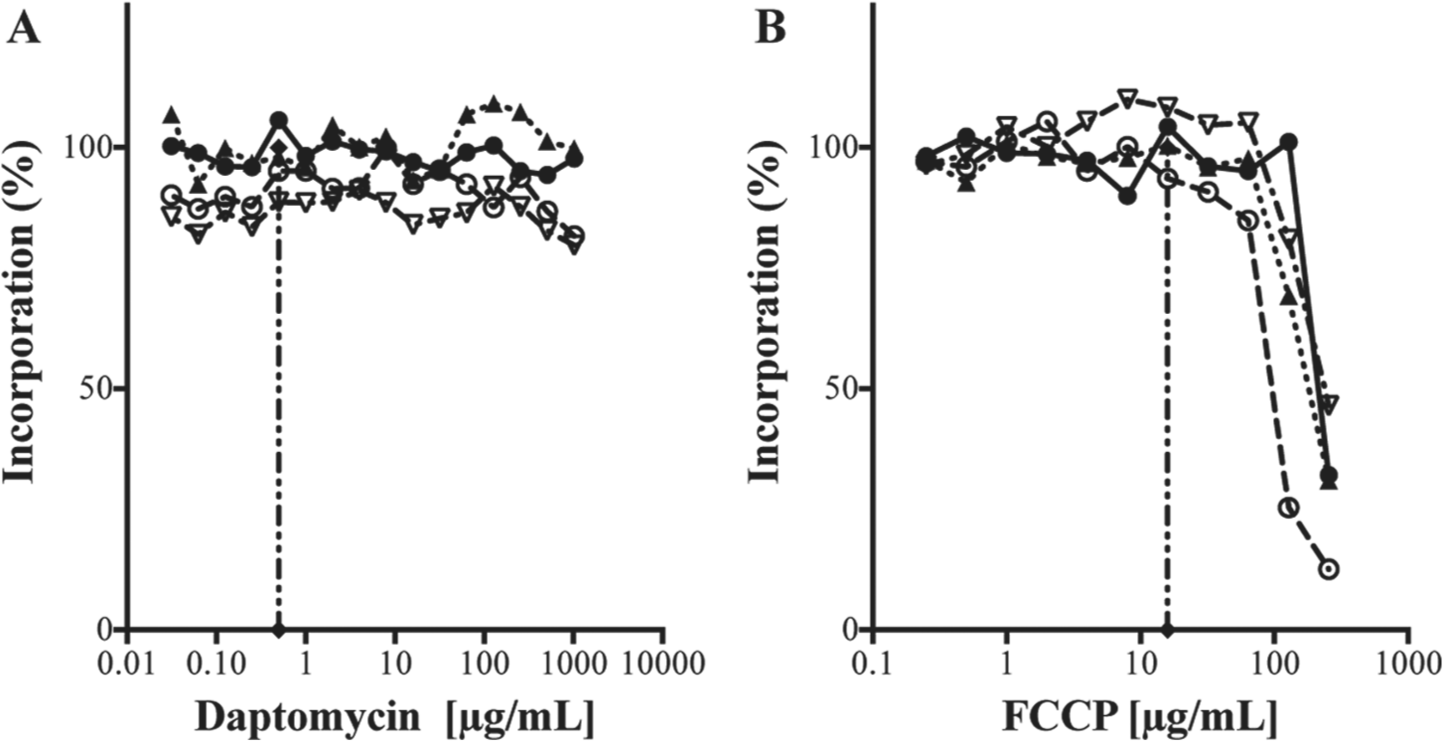
The effects of daptomycin (
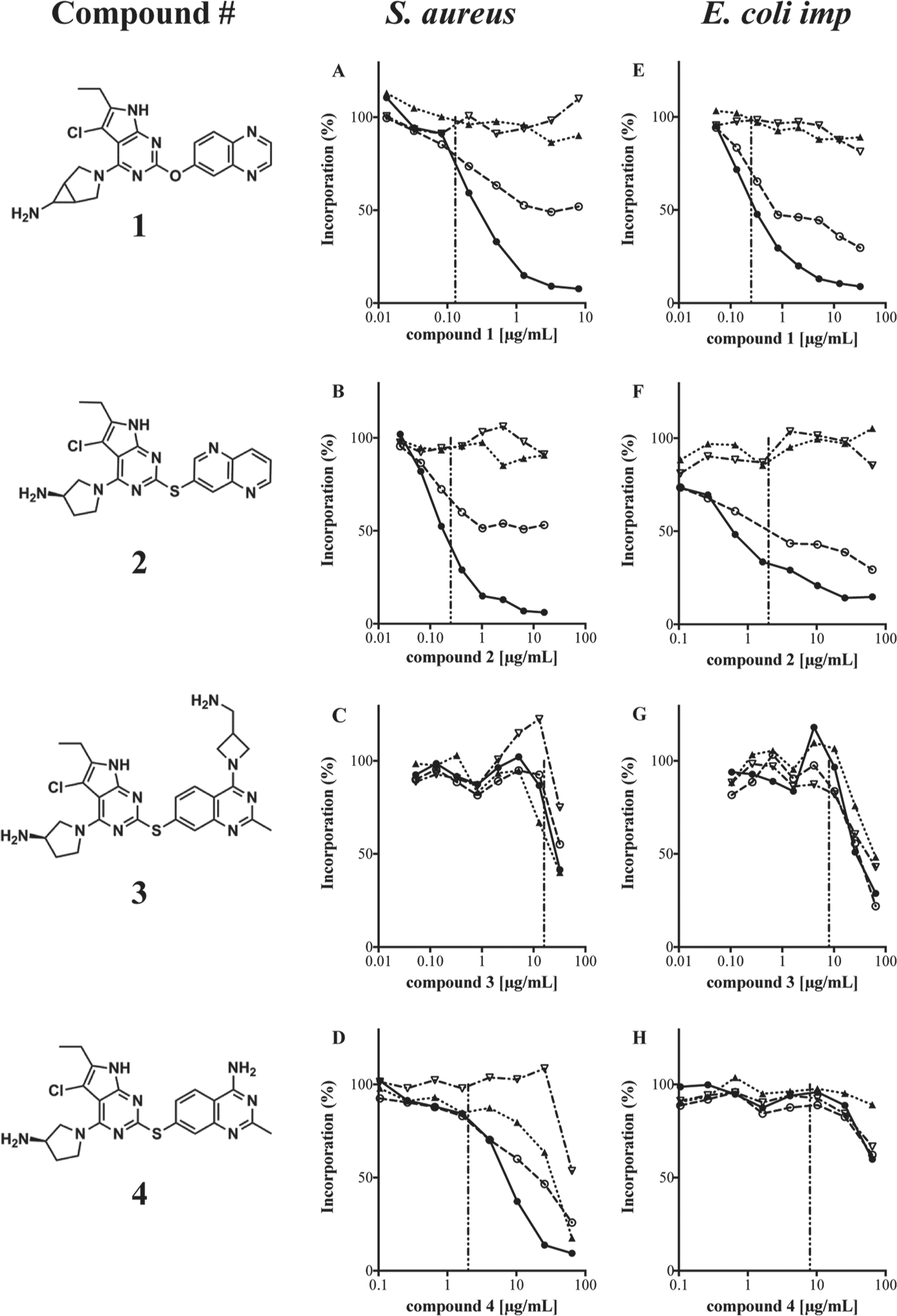
The effects of DNA GyrB/topoisomerase IV ParE inhibitors on metabolic labeling in S. aureus and E. coli imp. Incorporation of [3H]-precursors of DNA (●), RNA (○), protein (▲), and cell wall (▽) was examined as described in the Materials and Methods. The MIC value for each compound is indicated by a vertical dashed line. Compounds
Nucleic Acid Synthesis
Ciprofloxacin is a broad-spectrum fluoroquinolone that acts via inhibition of DNA replication, specifically binding to DNA gyrase (GyrA subunit) and topoisomerase IV (ParC subunit). The mechanism involves the formation of a ternary complex between the drug, protein, and DNA. 11 For both S. aureus and E. coli imp, the only significant effect was on DNA synthesis, and in both organisms, this disruption was evident at the MIC concentration ( Fig. 1A,D ). Complete cessation of [3H]-thymidine incorporation was seen at the highest concentrations tested, whereas little or no effect was observed on the other three pathways. Novobiocin, a natural product aminocoumarin, also acts via inhibition of DNA replication; however, the mechanism involves the competitive inhibition of the ATPase activity of DNA gyrase (GyrB subunit) and topoisomerase IV (ParE subunit). 12 Again, a dose-dependent disruption of DNA synthesis was observed for both organisms ( Fig. 1B,E ). However, in contrast to ciprofloxacin, a secondary effect was also observed, with a clear dose-dependent disruption of [3H]-uridine incorporation indicating decreased RNA synthesis.
Rifampicin, a semi-synthetic derivative of the naturally occurring rifamycin class of antibacterials, is a specific inhibitor of DNA-dependent RNA polymerase. 13 The primary effect on macromolecular synthesis was on RNA synthesis in both S. aureus ( Fig. 1C ) and E. coli imp ( Fig. 1F ), whereas a secondary effect on protein synthesis was also seen in S. aureus. RNA inhibition was evident only at concentrations >8-fold above the MIC. In addition, an apparent increase in [3H]-thymidine incorporation was noted. As part of a marine natural product screening campaign, a large number of rifamycin-like molecules were identified, which invariably gave this characteristic fingerprint (unpublished observations).
Other Mechanisms of Action
The proposed mechanism of action of the lipopeptide daptomycin is binding to the bacterial membrane, resulting in a loss of membrane potential, which leads to inhibition of protein, DNA, and RNA synthesis. 14 The macromolecular synthesis assay in S. aureus revealed relatively no effect on any of the pathways tested within the 25-min time frame of the assay, even at concentrations 1000× MIC ( Fig. 2A ). These data are consistent with results of a previous study in which the effect of daptomycin on macromolecular biosynthesis was monitored over a 2-h period, but no inhibition was observed during the first 30 min. 15
Trifluorocarbonylcyanide phenylhydrazone (FCCP) is a protonophore that disrupts ATP synthesis by collapsing the proton gradient required for oxidative phosphorylation. 16 This uncoupling agent caused a characteristic disruption of all pathways in the macromolecular synthesis assay at concentrations >4-fold above the MIC ( Fig. 2B ).
Use of the Macromolecular Synthesis Assay in Early Drug Discovery to Guide Optimization
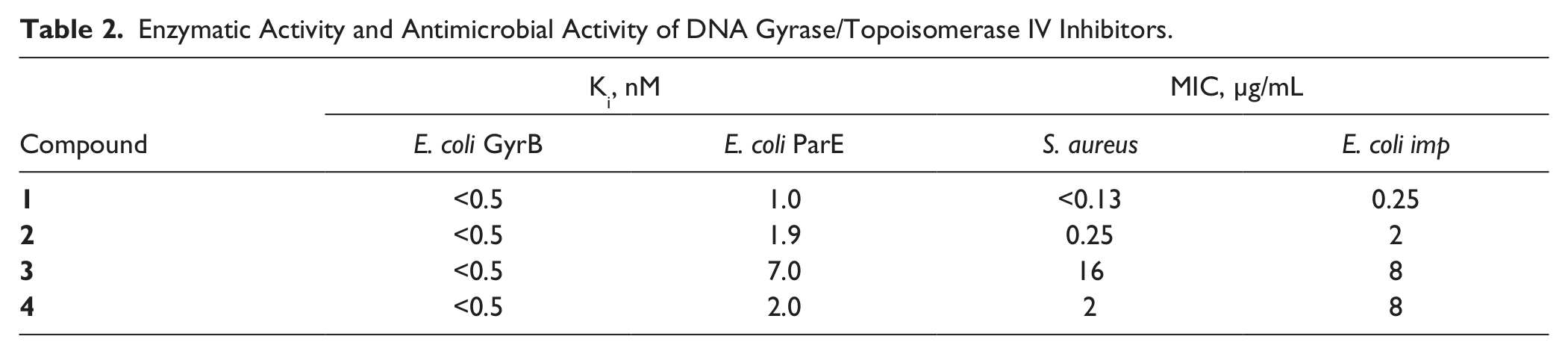
The macromolecular synthesis assay was used in a structure-based drug design program targeting the related bacterial enzyme subunits DNA gyrase B and topoisomerase IV ParE. Data for four representative members of a pyrrolopyrimidine-based class of compounds generated early in this program are shown (
Table 2
and
Fig. 3
). The closely related compounds were all potent, competitive inhibitors of both the DNA gyrase B (GyrB) and topoisomerase IV (ParE) ATPase activities, with similar nanomolar Ki values.
17
However, comparison of the macromolecular synthesis profiles for these compounds revealed striking mechanistic differences. The primary effect of compounds
Enzymatic Activity and Antimicrobial Activity of DNA Gyrase/Topoisomerase IV Inhibitors.
Discussion
Numerous methodologies to monitor the disruption of radioisotope precursor incorporation have been developed. Although highly valuable in determining the MOA of newly identified natural products or as part of antimicrobial drug discovery efforts, some of the protocols used long incubation times, resulting in the observation of secondary effects that are not due to the initial target-dependent events. 7 This is exemplified by the observation that daptomycin inhibits all four molecular pathways at time points greater than 40 min. 15 The use of cells that are not growing logarithmically or adapted to the test media or the use of a single test concentration at large multiples above the MIC can also result in artifacts.
The current assay was tailored for use with S. aureus and E. coli imp, a strain that is highly susceptible to many drug classes. Growth rates, precursor concentrations, and specific activities for the simultaneous determination of DNA, RNA, protein, and cell wall synthesis were optimized to ensure linear incorporation with adequate signal-to-noise ratios. Moreover, our conditions enabled us to expose logarithmically growing cells to test compounds or known drugs for a short duration, thus enabling the primary, compound-induced events within the cell to be identified. The assay lengths were less than half the generation time, which limited changes in cell density. The throughput was such that large numbers of compounds could rapidly be assessed; a single researcher could readily assay 100 compounds a week against all four pathways.
In general, we used an 8-point dilution series spanning the measured MIC values in the macromolecular synthesis assay. Although the data obtained from MIC determinations and the macromolecular synthesis assay cannot be directly compared because of differences in methodology, including media (MH-CA vs minimal media, necessitated by labeled precursor uptake), size of the inoculum (5 × 105 vs ~5 × 107 cells/mL), and length of the assay (16–20 h vs 15–25 min), we observed that for most compounds, the MIC value was generally within 2- to 4-fold of the concentration in which pathway inhibition is first observed in the macromolecular synthesis assay.
Characterization of a number of known antibacterials reveals considerable diversity within subsets. Ciprofloxacin and novobiocin both target DNA replication, via disruption of DNA gyrase/topoisomerase IV function; however, the underlying mechanisms are different. Ciprofloxacin targets the GyrA (and ParC) subunit, forming a ternary complex with the protein and DNA. The stabilized protein-DNA complex blocks DNA tracking enzymes, such as RNA and DNA polymerase, and movement of the replication fork. 18 Cellular processes acting upon the ternary complexes have been hypothesized to be responsible for the double-stranded DNA breakage that occurs, along with the rapid induction of the SOS response.19,20 It has been suggested that the rapid inhibition of DNA synthesis is specifically associated with gyrase molecules located at the replication fork. 21 Early studies of the addition of nalidixic acid to E. coli showed no inhibition of RNA synthesis for ~2 h along with continued increase in cell mass, consistent with the microscopic observation of cell filamentation. 22 In contrast, the coumarin antibiotics competitively inhibit the ATP hydrolysis activity of the GyrB (and, to a lesser extent, ParE) subunit, thus inhibiting supercoiling, but do not result in the cleavage of DNA. 20 Novobiocin has also been shown to rapidly (within 30 min) inhibit the synthesis of stable ribosomal RNA in vivo, with little effect on messenger RNA synthesis. 23 Thus, within the timeframe of the macromolecular synthesis assay, rapid inhibition of DNA synthesis is observed for ciprofloxacin, whereas first DNA and then RNA synthesis inhibition is observed for novobiocin and the pyrrolopyrimidine class of inhibitors.
Frequent and accurate evaluation of antibacterial mechanism of action is a vital part in the discovery and development of novel antibacterial drugs. There have been several attempts to define the properties consistent with activity against Gram-negative organisms through examination of the property space of antibiotics.
24
The difficulty stems from a requirement for appropriate charge and solubility to enable penetration through outer membrane porins, an uncharged hydrophobic state within the periplasm allowing traversal of the cytoplasmic membrane, and a charged state with low potential for efflux such that sufficient compound remains trapped within the cytosol to reach the target.
25
Compounds such as fluoroquinolones and tetracyclines, with multiple charges, meet the criteria and exemplify a possible path to achieving Gram-negative activity. With those features in mind, molecules within a new pyrrolopyrimidine class of inhibitors were designed to evaluate the antibacterial effect of different charged groups. Although all four of the pyrrolopyrimidine-based compounds were highly potent enzyme inhibitors, only two of the four compounds (compounds
Evaluation of additional analogues in the macromolecular synthesis assays revealed that incorporation of charged groups off a specific analoging vector of the pyrrolopyrimidine scaffold introduced non–target-based components to the observed antibacterial activity. 17 These data suggest that the general use of zwitterions or multiple charges for achieving Gram-negative activity must be approached with caution, to avoid potential issues due to the introduction of membrane disruption or other off-target activity. The results of the macromolecular synthesis assays enabled us to distinguish between otherwise similar compounds, identify and deprioritize those compounds that contained an unwanted nonspecific component, and focus subsequent optimization efforts on compounds with only target-based antibacterial activity. The remarkably different profiles observed across very similar compounds illustrate the need to regularly examine mechanism of action throughout all stages of an optimization project. It is important to note that selecting target-based mutants in resistance studies performed on a novel molecule, although supportive of proof of mechanism of action, would not by itself eliminate the potential for off-target antimicrobial activities associated with the novel molecule.
In summary, we have developed a modified macromolecular synthesis methodology for both Gram-positive and Gram-negative bacteria that enables high numbers of pure compounds or partially purified extracts to be evaluated for on-target mechanism early in the drug discovery process. Proof of mechanism of action in two species, such as S. aureus and E. coli, avoids the discrepancies that can occur in early stage programs, especially where target essentiality or number of orthologs differs in different species. The regular use of these assays has proven to be a key driver in our optimization efforts by identifying non–target-based antimicrobial activity at an early stage of research.
Footnotes
Acknowledgements
We thank J. Finn, J. B. Locke, M. Stidham, and L. W. Tari for critical reading of the manuscript.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article. All authors are employees of Trius Therapeutics, Inc.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Part of this work has been funded with federal funds from the National Institute of Allergy and Infectious Diseases, National Institutes of Health, Department of Health and Human Services, under contract no. HHSN272200800042C.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
