Abstract
The tumor microenvironment can profoundly affect tumor cell survival as well as alter antitumor drug activity. However, conventional anticancer drug screening typically is performed in the absence of stromal cells. Here, we analyzed survival of myeloma cells co-cultured with bone marrow stromal cells (BMSC) using an automated fluorescence microscope platform, ScanR. By staining the cell nuclei with DRAQ5, we could distinguish between BMSC and myeloma cells, based on their staining intensity and nuclear shape. Using the apoptotic marker YO-PRO-1, the effects of drug treatment on the viability of the myeloma cells in the presence of stromal cells could be measured. The method does not require cell staining before incubation with drugs, and less than 5000 cells are required per condition. The method can be used for large-scale screening of anticancer drugs on primary myeloma cells. This study shows the importance of stromal cell support for primary myeloma cell survival in vitro, as half of the cell samples had a marked increase in their viability when cultured in the presence of BMSC. Stromal cell–induced protection against common myeloma drugs is also observed with this method.
Introduction
Multiple myeloma is a hematological malignancy of plasma cells (for a recent review of multiple myeloma, see Palumbo and Anderson 1 ). Multiple myeloma is still considered an incurable disease despite new treatment regimens. 2 One characteristic of multiple myeloma is that the malignant plasma cells, except during the very last stages of disease, reside within the bone marrow. It is generally believed that growth and survival of myeloma cells are critically dependent on the bone marrow microenvironment. Bone marrow stromal cells (BMSC) have also been shown to protect myeloma cells from common cytostatic or cytotoxic drugs in vitro.3–7 Conveniently, bone marrow aspirates taken of multiple myeloma patients for routine diagnostic purposes can be used to isolate relatively easily both malignant and stromal cells.
Multiple myeloma is a heterogeneous disease, with some patients having aggressive disease and other patients having disease responding well to treatment. Furthermore, there is an urgent need to identify patients who may benefit from a particular treatment as well as patients who may be resistant toward a specific drug. Recently, extensive attempts have been made to correlate sensitivity toward drugs in cancer cell lines to the presence of genetic aberrations in the same cells.8,9
The aim of this work was to establish a robust and simple method for the measurement of drug sensitivity in myeloma cells under conditions mimicking aspects of the bone marrow microenvironment (i.e., in the presence of stromal cells). Common in vitro assays used for high-throughput drug screening, such as those based on the MTT (3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide) assay, cannot discriminate between stromal and tumor cell responses in co-cultures. A few recent studies have overcome this problem, such as the fluorescence-based model described by Ramasamy et al. 10 and the compartment-specific bioluminescence imaging platform described by McMillin et al. 11 However, the application of stable transfection for labeling of cells makes these methods complicated. This limits the practical application of these co-culture studies to cell lines and excludes primary myeloma cells that may be hard to transduce even by retroviral vectors. We have recently shown that survival of myeloma cell lines in co-cultures with stromal cells can be analyzed by flow cytometry. 12 However, this and similar methods,4,5,7,13 may require labeling of cells before co- culture, as well as enzymatic removal of cells from the tissue culture trays before analysis.
To address these limitations, we here describe a method for the analysis of live-cell co-cultures of multiple myeloma cells and BMSC by the automated fluorescence microscope platform, ScanR. We found that BMSC can be distinguished from myeloma cells, and thus BMSC and myeloma cells can be gated into different populations, based on their nuclear area and staining intensities. This allows for specific quantification of the myeloma cell viability in the presence of BMSC. The main advantages of this method are the nonnecessity of cell manipulation before co-culture and the low number of myeloma cells that are needed per measurement, which makes the method ideal for experiments with primary myeloma cells. This method should be well suited for large-scale screening of primary myeloma cells, as the cells are stained in situ with no washing, centrifugation, or fixation steps before analysis.
By the use of this novel method, it was possible to show the importance of stromal cell support for survival of primary myeloma cells, thus providing a powerful tool to investigate drug responses ex vivo.
Materials and Methods
Cell Culture and Reagents
The multiple myeloma cell lines used in this study were INA-6 (a kind gift from Dr. Martin Gramatzki, Erlangen, Germany), RPMI-8226 (from American Type Culture Collection, Rockville, MD), and ANBL-6 (a kind gift from Diane Jelinek, Mayo Clinic, Rochester, MN). INA-6 and ANBL-6 cells were grown in RPMI 1640 (Sigma-Aldrich, Schnelldorf, Germany) supplemented with L-glutamine (100 µg/mL) and gentamicin (20 µg/mL; referred to as RPMI) and supplemented with 10% heat-inactivated fetal calf serum (FCS) and recombinant human interleukin (IL)–6 (1 ng/mL; Biosource, Camarillo, CA). RPMI-8226 cells were maintained in 20% FCS in RPMI. All cells were cultured at 37 °C in a humidified atmosphere containing 5% CO2. The media were replenished twice weekly. Stock solutions of the MYC inhibitor 10058-F4 (40 mM; Sigma-Aldrich) were made in DMSO. Melphalan (Sigma-Aldrich) and bortezomib (LC Laboratories, Woburn, MA) were dissolved in ethanol. Cyclophosphamide monohydrate (Sigma-Aldrich) was dissolved in distilled water. Fluorescein isothiocyanate (FITC)–conjugated antibodies (CD90, CD44, CD45, CD209, HLA-ABC, and HLA-DR) and PE-conjugated antibodies (CD14, CD34, CD29, CD54, CD73, and CD105; BD Biosciences, San Jose, CA) were used to immunophenotype the BMSC by flow cytometry.
Primary Cells
CD138-positive myeloma cells were isolated from bone marrow samples obtained from the Norwegian Myeloma Biobank using RoboSep automated cell separator and the Human CD138 Positive Selection Kit (StemCell Technologies, Grenoble, France). The purity of plasma cells after isolation was >90%.
BMSC were prepared by seeding the remaining mononuclear cells after CD138-positive selection. The cells were seeded in culture flasks in RPMI with 10% heat-inactivated human serum (HS; Department of Immunology and Transfusion Medicine, St. Olav’s University Hospital, Trondheim, Norway). After 3 d of culture, nonadherent cells were removed, and the remaining cells were expanded and split after about 10 d. After 3 wk of culture, stromal cells from 10 different patients were mixed to obtain a batch of standardized BMSC for use in co-culture experiments with myeloma cells. The cells in different batches of BMSC were characterized by immunophenotyping, analyzed using an LSRII flow cytometer (BD Biosciences). The patients had given informed consent to give bone marrow cells to research purposes (REK 2011-2029).
ScanR Cell Viability Assay
Treatment and cultivation of cells
BMSC (2500 cells/well) were seeded in 96-well plates and allowed to adhere for at least 3 h before addition of primary myeloma cells (5000 cells/well) or myeloma cell lines (2000 cells/well). The cells were cultivated in RPMI medium supplemented with 2% HS in a total volume of 200 µL/well, and drugs were added at the concentrations indicated. Experiments were also performed with myeloma cells or BMSC alone. All samples were run in duplicate.
Staining of cells
After 3 d of incubation, YO-PRO-1 (1 µM; Invitrogen, Carlsbad, CA) was added to the wells, and the plates were incubated for 30 min at 37 °C. DRAQ5 (2.5 µM; eBioscience, San Diego, CA) was added to the wells 15 min before image acquisition without further processing. The plates were allowed to settle to room temperature (RT) for 15 min before starting the scan.
Automated image acquisition
ScanR (Olympus, Hamburg, Germany) is a modular epifluorescence microscope-based imaging platform designed for fully automated image acquisition and was used to acquire images of the co- cultures. 14 Images were taken with a UPlanSApo 10× objective (Olympus) with a numerical aperture of 0.4 and a working distance of 3.1 mm. Excitation filters (wavelength [nm]/width [nm]) used were YO-PRO-1 (485/20) and DRAQ5 (650/13). A combination emission filter was used (440/521/607/700) for all fluorophors (Chroma Technology Corp, Bellows Falls, VT). For each well, 21 different fields of view were analyzed. The area analyzed corresponded to approximately 40% of the well area, and using these settings, the number of analyzed myeloma cells per well varied from 1000 to 1800. We used a combination of hardware autofocus (50% of the positions) and software autofocus to ensure that all imaged wells were in focus. With these specifications, it took approximately 2.5 h to screen 60 wells.
Image analysis and assay parameters
The images were analyzed by the ScanR Analysis software (Olympus). 15 With this software, the cells could be classified and gated according to their size, circularity, and fluorescence intensity. For every gate or classified group of objects, a reference gallery of objects of the selected area can be viewed and manually reviewed for accuracy of the gate settings. Here, nuclei stained with DRAQ5 were defined as the main objects using edge detection, and for these objects, the area, circularity, and mean intensity in all color channels were measured. After the gates were defined, randomly selected images of objects belonging to the respective gates were manually controlled for quality of the gate settings and to identify potential errors.
Gating of viable cells was performed on objects that were gated and classified as myeloma cells. YO-PRO-1 is a DNA-binding dye that stains early apoptotic cells. 16 Myeloma cells that showed low YO-PRO-1 staining and high-intensity DRAQ5 staining were characterized as viable. In addition, all analyzed objects were gated in a separate dot plot, with circularity factor versus nucleus staining intensity, to exclude potential noncellular particles.
CellVue Staining
The myeloma cells were stained with the far red staining CellVue Claret (Sigma-Aldrich) according to the manufacturer’s protocol. In short, a final concentration of 2 µM CellVue Claret in 250 µL staining solution was used for 4 to 8 × 105 myeloma cells. Dye was added to the cells, followed by incubation for 3 min and three washes in 10% FCS in RPMI. After staining, the cells were cultured either in the presence or absence of BMSC and analyzed on the ScanR microscope as described above, except that 0.2 µg/mL Hoechst 33342 (Invitrogen) was used as nuclear stain instead of DRAQ5. The excitation filter for Hoechst 33342 was (377 nm/50 nm).
Identification of Myeloma Cells Using Fluorescently Labeled Antibodies
To validate the gating of myeloma cells, wells with either co-cultures of myeloma and stromal cells or with myeloma cells alone were stained with anti-human CD138-FITC (BD Biosciences, Cat. No. 552723). In short, media were removed from the wells so that 50 µL remained, 7 µL CD138-FITC was added, and the wells were incubated for 30 min at RT. DRAQ5 (2.5 µM) was added to the wells 15 min before cells were analyzed. Immediately before analysis on the ScanR microscope, unbound antibodies were diluted by addition of 150 to 200 µL RPMI/well.
Detection of Apoptotic Myeloma Cells by Flow Cytometry
Co-culture experiments to be analyzed by flow cytometry were performed essentially as previously described. 12 Briefly, BMSC were allowed to adhere in wells, media were removed, and the cells were stained by 1 µM CFDA-SE (100 µL/well), followed by incubation for 15 min at 37 °C. The cells were then washed once with phosphate-buffered saline. Next, unstained myeloma cells and drugs were added. To be able to analyze all cells, BMSC and adhering myeloma cells were detached from culture plates using Accutase (Sigma-Aldrich) at day 3 for further processing. The extent of apoptosis was measured using annexin V Alexa Fluor 647 conjugate (Invitrogen). In short, cells were incubated with annexin V in binding buffer for 1 h and analyzed using an LSRII flow cytometer (BD Biosciences). Cells negative for annexin V Alexa Fluor 647 and CFDA-SE were considered viable myeloma cells. Data were analyzed with FlowJo 7.6 software (Tree Star Inc, Ashland, OR).
Results
The aim of this work was to establish a robust and simple method for the measurement of drug sensitivity of myeloma cells co-cultured with BMSC. BMSC were characterized by immunophenotyping using flow cytometry. More than 90% of the BMSC were positive for CD90, CD44, CD73, CD105, CD29, and HLA-ABC, whereas antibodies toward CD209, CD45, CD14, and CD34 also stained a small fraction of the cells (
To estimate the effects of drugs or small-molecule inhibitors on myeloma cells, either primary myeloma cells or cell lines were seeded in wells with adherent BMSC in 96-well plates, before drugs were added. Then, after 72 h, the cells were stained with DRAQ5 for visualization of cell nuclei and with YO-PRO-1 for identification of apoptotic cells, before analysis on a ScanR automated fluorescence microscope. Even though the dyes were used without washing steps, both stains showed a strong and stable fluorescence with low background noise. As shown in Figure 1A and C , the myeloma cells had a round nuclear shape with a relatively high-intensity staining, whereas the nuclei of the BMSC ( Fig. 1B, C ) had a characteristic oval shape and lower staining intensity. The nuclei stained with DRAQ5 were used as the primary objects for discrimination between cell types. Thus, on the basis of the fluorescence intensities and the area of the stained nuclei, the BMSC and the myeloma cells could be gated as different populations ( Fig. 1A–C ). A main advantage to using ScanR analysis software is that the objects in every gate or region of interest can be observed in a reference gallery. In each image, the objects that are located to the different regions/gates can be marked and thus visually controlled for accurate gate settings and correct object recognition, as indicated by the arrows in Figure 1C .
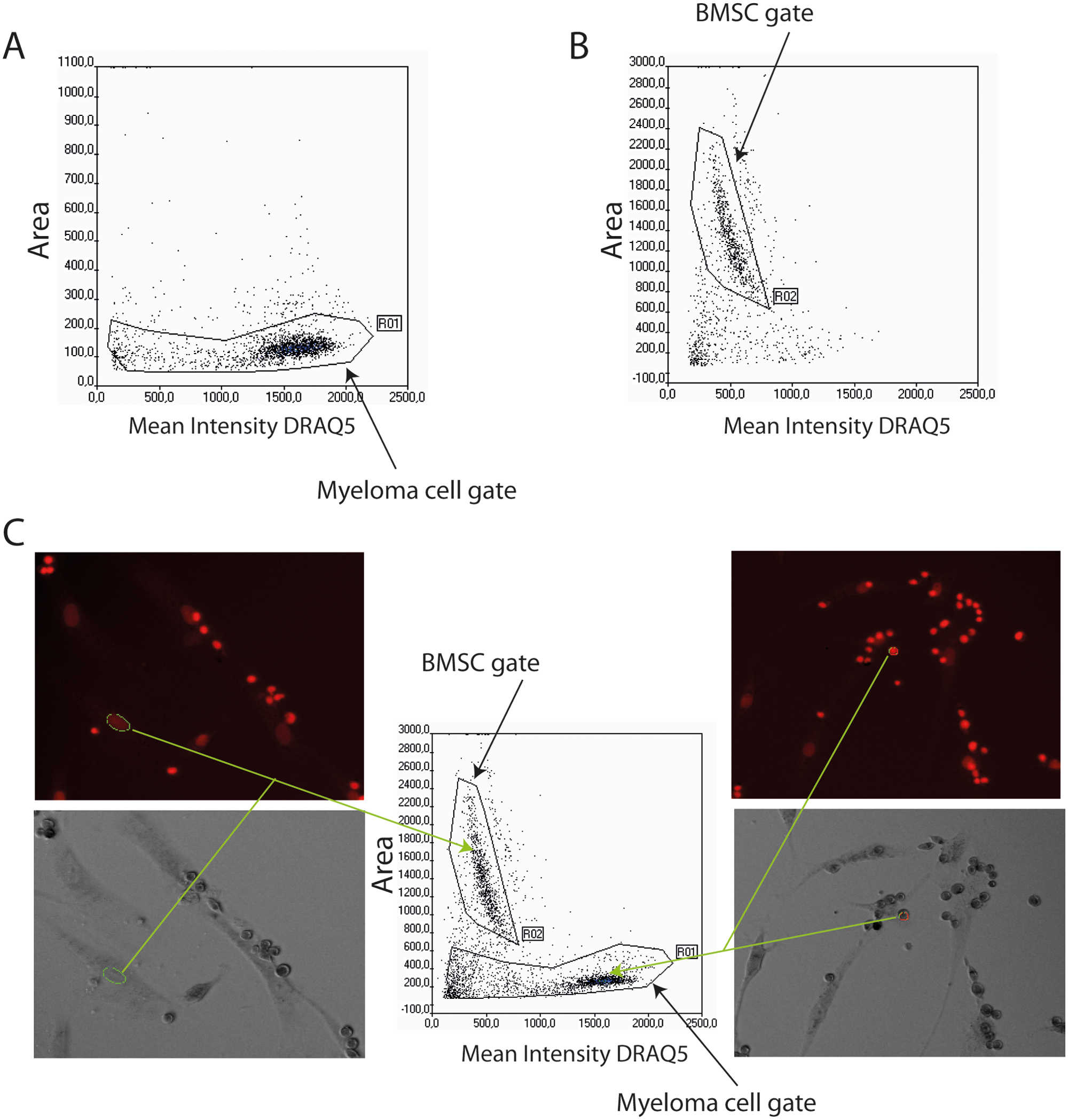
Discrimination between cell types by nuclear area and staining intensity. A typical example of the location of primary myeloma cells and bone marrow stromal cells (BMSC) in the area versus nucleus staining intensity plot in the ScanR analysis program is shown. (
In co-culture experiments, primary myeloma cells from the myeloma cell gate (as shown in
Fig. 1
) were analyzed by plotting YO-PRO-1 versus DRAQ5 staining intensity. The viable cells were gated by a high-intensity DRAQ5 nuclear staining and low YO-PRO-1 staining (
Fig. 2A
) and could thus be separated from apoptotic and dead cells obtained by treatment with bortezomib (10 nM;
Fig. 2B
). For most of the myeloma cell lines and primary cells we have tested, the same viability gating could be used with only minor modifications for all conditions (i.e., in the absence or presence of BMSC and with different drug treatments;
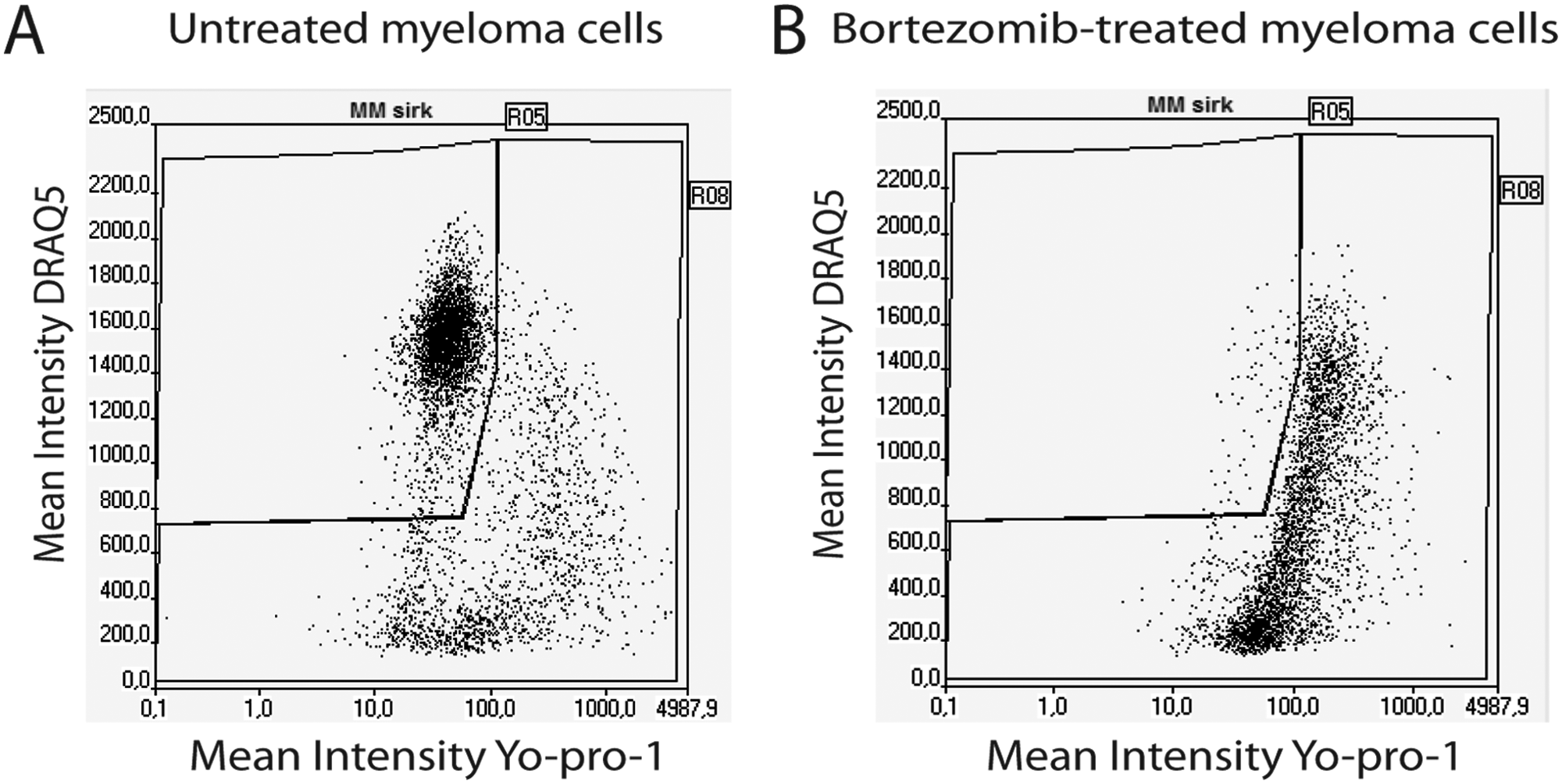
Measurement of myeloma cell apoptosis in co-cultures. Cells gated as myeloma cells in
Figure 1
are either shown without treatment (
As shown in
Figure 1B
, some BMSC appear in the area of the myeloma cell gate. These are BMSC that have shrunken to a smaller and round shape. However, by using doses nontoxic for the BMSC, this number of stromal cells in the myeloma gate will be low and constant and will not affect the estimation of myeloma cell viability, as usually 1000 to 1800 myeloma cells versus ~100 BMSC are counted in the myeloma cell gate in each well. Furthermore, by performing experiments with wells containing BMSC alone and increasing doses of drugs, an indication of the toxicity of common drugs toward stromal cells can be obtained (
To verify that this method of estimating myeloma cell death in co-culture with BMSC was valid, we performed experiments in which the myeloma cells were identified by labeling with a membrane dye before addition to the BMSC. Shown in Figure 3A is a typical experiment in which the RPMI-8226 cells were labeled with the fluorescent dye CellVue Claret, before addition to BMSC and treatment with increasing concentrations of the MYC inhibitor 10058-F4. Analysis of surviving cells was done after 3 d by YO-PRO-1 staining. The effect of MYC inhibitor on RPMI-8226 cells prelabeled with CellVue Claret (right-hand panel) was compared with the effect of MYC inhibitor on RPMI-8226 cells stained with DRAQ5/YO-PRO-1 (left-hand panel). As shown in Figure 3A , the results obtained were in general very similar.
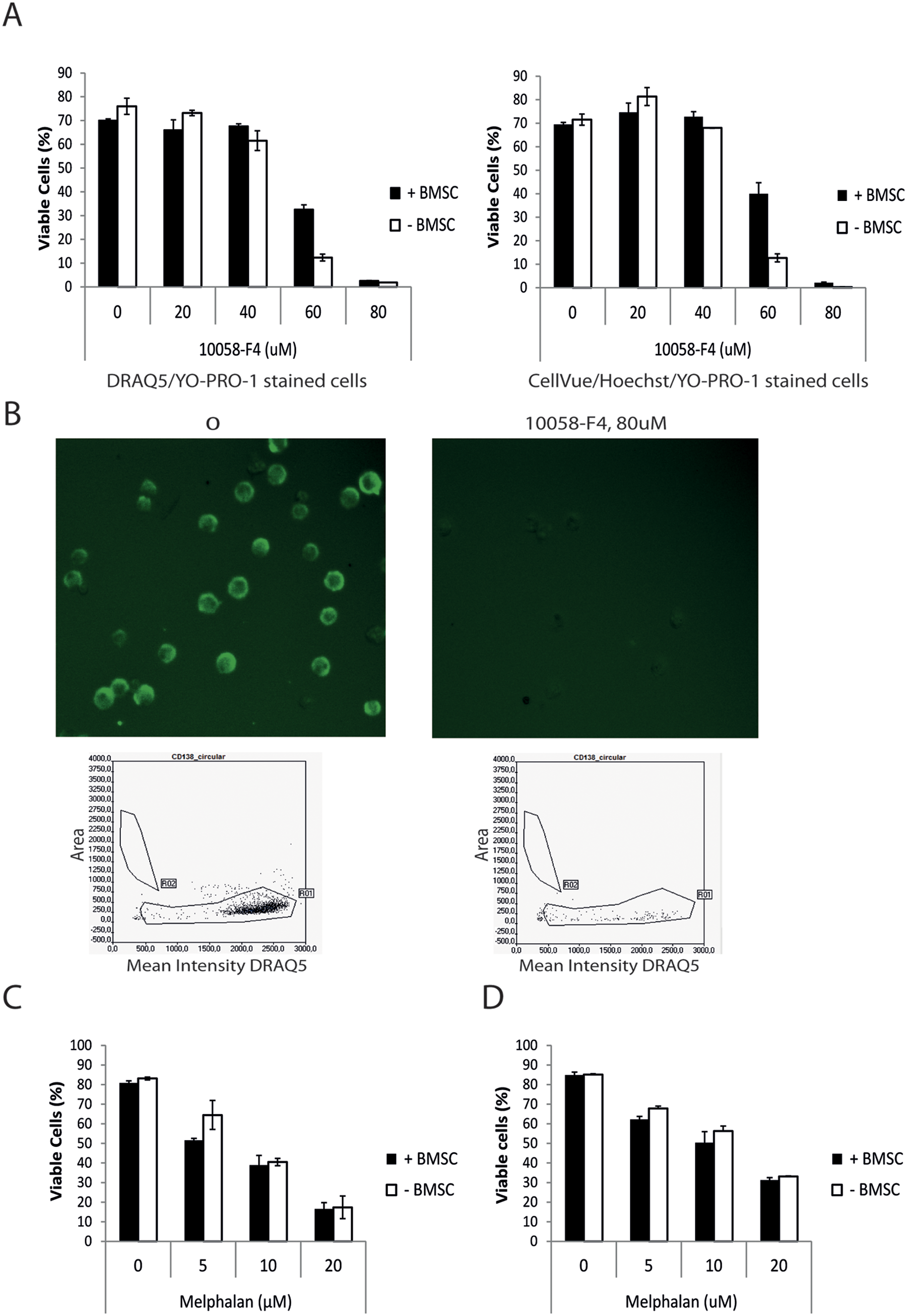
Comparison with other methods for identification of myeloma cells and viability measurements with and without bone marrow stromal cells (BMSC). (
To further ensure that the myeloma cells were correctly gated, we also performed an experiment in which ANBL-6 cells were identified by labeling with an FITC-conjugated antibody against CD138. The result showed that the CD138-positive myeloma cells were placed correctly in the myeloma cell gate ( Fig. 3B , lower left panel). However, in accordance with earlier observations, 17 ANBL-6 cells that had become apoptotic, in this case after treatment with the MYC inhibitor 10058-F4, lost their expression of CD138 ( Fig. 3B , right-hand picture and panel). Thus, identification of myeloma cells by their cell surface marker CD138 is not useful when analyzing apoptotic cells.
Comparison of Measurements of Cell Viability by ScanR Fluorescence Microscopy and by Flow Cytometry
To examine if the viability assays performed on the ScanR fluorescence microscope were comparable with conventional methods for detecting cell viability, parallel experiments applying earlier published methods were performed. Thus, the effects of melphalan on cell viability of the ANBL-6 cell line were measured by ScanR using YO-PRO-1 and in parallel by flow cytometry, using annexin V labeling to detect apoptotic cells, as described earlier. 12 The dose-response curves obtained for ANBL-6 cells treated with melphalan were similar in both cases and showed the same trends ( Fig. 3C, D ). However, the effects of melphalan treatment were somewhat more evident when analyzed by the ScanR system ( Fig. 3C ) than by flow cytometry ( Fig. 3D ; EC50YO-PRO-1 = 11 µM vs. EC50Annexin V = 15 µM).
BMSC Are Important for Myeloma Cell Viability
To show that the BMSC preparations as applied here were important for myeloma cell survival in co-culture experiments, we took advantage of earlier reports indicating that stromal cells may replace the requirement of IL-6 for myeloma cells. 18 INA-6 is an IL-6–dependent myeloma cell line, and we plated INA-6 cells in the presence or absence of BMSC, and in media with or without IL-6, before estimation of INA-6 cell survival after 3 d. The results showed that when INA-6 cells were cultivated in the presence of BMSC, the cells survived without addition of IL-6 ( Fig. 4A ). Thus, in this respect, the BMSC apparently could replace the growth factor requirement of a myeloma cell line.
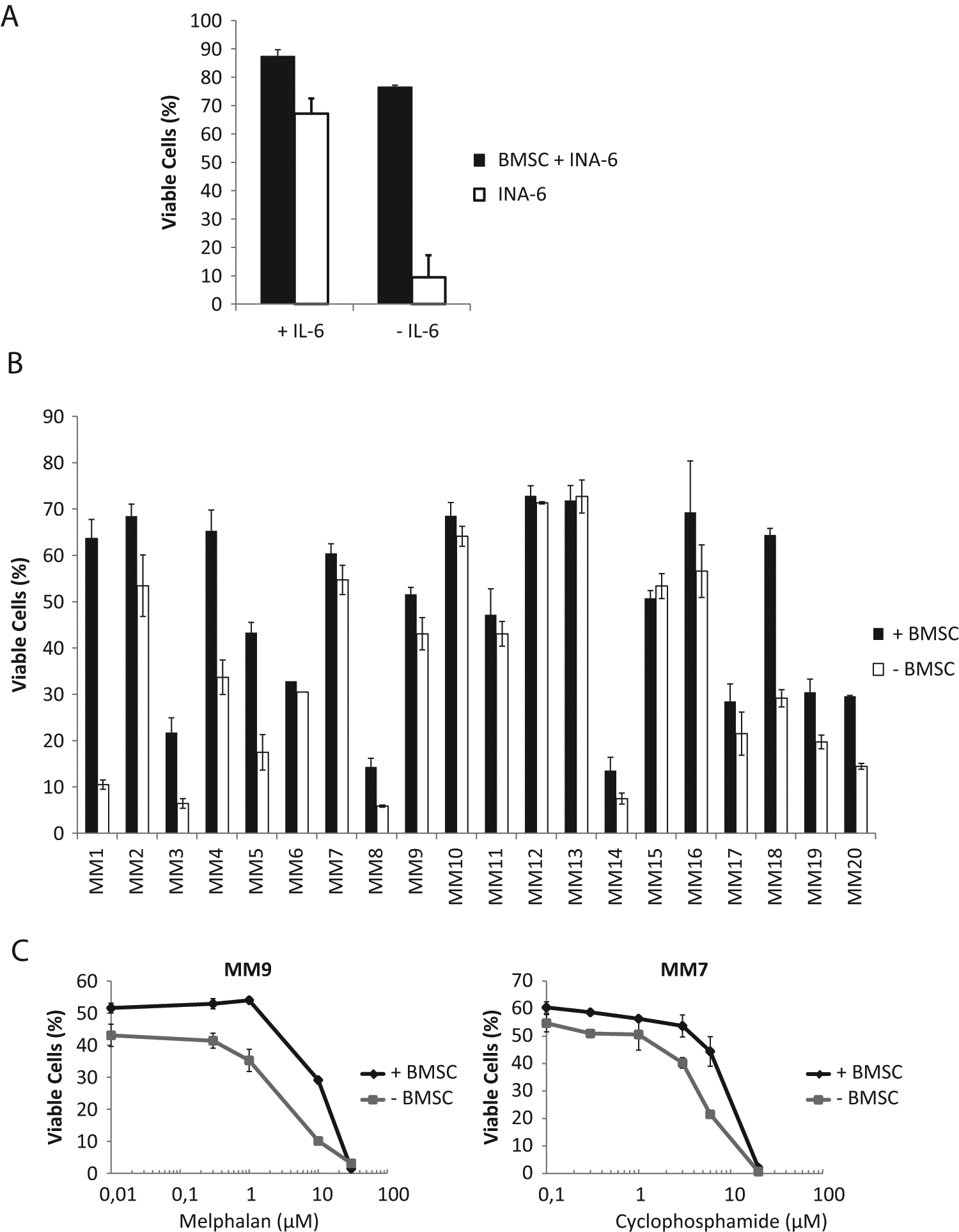
Effects of bone marrow stromal cells (BMSC) on myeloma cell viability and protection against drugs. (
More important, when primary myeloma cells from different patients were cultivated in the presence of BMSC, there was in many cases a marked increase in myeloma cell survival compared with primary myeloma cells cultured alone. Thus, 8 of 20 patient samples had at least twice as high viability after 3 d of cultivation in the presence of BMSC ( Fig. 4B ).
It has been reported that BMSC may protect myeloma cells from the inhibitory action of some drugs.3–7 Thus, the ability of our preparations of BMSC to protect primary myeloma cells from drugs was also observed with some of the compounds tested. Examples of the protective effect of BMSC are shown in Figure 4C , where primary myeloma cells (patients MM7 and MM9) were treated with the common myeloma drugs melphalan and cyclophosphamide in the absence or presence of BMSC. In the presence of BMSC, the EC50 for melphalan increased from 4 µM to 12 µM, whereas the EC50 for cyclophosphamide increased from 5 µM to approximately 10 µM ( Fig. 4C ).
Discussion
The main result presented here is a description of a simple method for measurement of myeloma cell viability in co-cultures with BMSC. The method can be used to measure drug sensitivity of myeloma cells from individual patients or to study the role of inhibition of particular intracellular signaling pathways in primary myeloma cells under conditions that to some extent may mimic the bone marrow microenvironment. Because of its simplicity and applicability to primary cells, the method has great advantages compared with earlier described methods.
One of the main advantages of the method is that no prestaining of the cells is necessary before incubation with drugs. Thus, the possible negative effects of cell-staining procedures are avoided. This may be particularly important when working with primary myeloma cells, which may tolerate less handling than cell lines. Moreover, because of its simplicity in terms of staining and lack of handling procedures, the method should be useful in large-scale screens of drug activity toward myeloma cells. Another advantage with this method is that it requires relatively few cells per measurement compared with conventional methods. Thus, a number of drugs or inhibitors can be tested on cells from one patient. In fact, the analysis was easier and more robust when using slowly growing cells, that is, by using primary myeloma cells compared with more rapidly proliferating myeloma cell lines, because uneven distribution of the DRAQ5 nuclear stain can be observed when using rapidly growing cell lines, which may complicate the viability gating (data not shown).
For practical reasons, we chose to include standardized preparations of stromal cells in the co-culture experiments. With this approach, a large number of different primary cells can be tested under similar conditions, and upon retesting of the primary cells from a given patient with additional drugs, the results should be comparable with earlier obtained results. Furthermore, the stromal cell populations applied were able to support IL-6–dependent myeloma cell lines without addition of IL-6 and also protected primary cells from the action of common myeloma drugs. More importantly, many of the preparations of primary cells had low viability when cultured for 3 d without BMSC. In several cases, the fraction of primary cells surviving in control wells was as low as 20% on day 3, making it difficult to obtain proper dose-response curves of the tested drugs. However, in the presence of stromal cells, the fraction of viable cells in several cases rose to 60% to 80%.
The classification of cell populations as performed here depends on an even distribution of DRAQ5 staining of the myeloma cells. The analysis also requires the application of drugs at concentrations that do not show toxicity toward BMSC. Thus, control experiments with wells containing only stromal cells should be performed to control for drug toxicity toward these cells.
We also show here that identification of myeloma cell death by antibody staining of CD138-positive myeloma cells is not feasible because apoptotic myeloma cells tend to lose their CD138 expression, in accordance with previously reported results. 17
One obvious application of the method would be to correlate drug sensitivity to the presence of specific genetic aberrations in the malignant plasma cells. Early sequencing experiments suggest that myeloma cells on average may contain approximately 40 mutations or other genetic aberrations19,20; however, the relative importance of each mutation is unclear. By applying inhibitors of intracellular enzymes and signaling pathways, the method can be used to identify some of the mutations that are of critical importance for the survival of myeloma cells. Several attempts have been made to subclassify myeloma patients based on genetic aberrations or gene expression profiling of the malignant plasma cells (reviewed by Morgan et al. 21 ). However, so far such classification has not substantially influenced choice of treatment of multiple myeloma. 22 Another application of the method could be to predict sensitivity or resistance toward new or common myeloma drugs in individual patients.
The relevance of this method in the above-mentioned applications remains to be shown. It is obvious that even though we try to mimic conditions of the bone marrow during incubation of cells with drugs, the conditions in tissue culture wells is very different from the bone marrow in terms of three-dimensional organization, presence of the proper stromal cells or other cell types adjacent to the plasma cells, and so on. Thus it has been suggested that various distinct cell types are important in mediating myeloma cell survival.23-25 Furthermore, for practical reasons, the primary cells from a given patient are not tested in the presence of stromal cells from the patient itself but rather with cells from a kind of average microenvironment.
When interpreting results, it should also be noted that CD138-based purification of cells from bone marrow aspirates, as performed here, will result in purification not only of the malignant cells but also of normal plasma cells. If the fraction of plasma cells in the bone marrow aspirate is low (e.g., approaching 10%) and the patient has normal levels of normal plasma cells in the bone marrow (i.e., 2%), as much as one-fifth of the plasma cells tested could be normal and not malignant.
In conclusion, we here provide a simple method for drug screening on myeloma cells that could be used to try to predict drug sensitivity in myeloma cells from individual patients and to test the activity of novel drugs toward primary myeloma cells in vitro, as well as to investigate the oncogene dependence of primary myeloma cells.
Footnotes
Acknowledgements
We wish to thank Lill Anny Grøseth, Solveig Kvam, and the rest of the staff at the Department of Hematology, St. Olav’s University Hospital, Trondheim, Norway, and the Norwegian Myeloma Biobank for help obtaining primary myeloma samples.
Authors’ Note
The fluorescence imaging and automated analysis part were performed at the Cellular & Molecular Imaging Core Facility, Norwegian University of Science and Technology (NTNU).
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Norwegian Cancer Society, the Research Council of Norway, and the K.G. Jebsen Foundation for Medical Research.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
