Abstract
Flap endonuclease-1 (FEN1) is a highly conserved metallonuclease and is the main human flap endonuclease involved in the recognition and cleavage of single-stranded 5′ overhangs from DNA flap structures. The involvement of FEN1 in multiple DNA metabolism pathways and the identification of FEN1 overexpression in a variety of cancers has led to interest in FEN1 as an oncology target. In this article, we describe the development of a 1536-well high-throughput screening assay based on the change in fluorescence polarization of a FEN1 DNA substrate labeled with Atto495 dye. The assay was subsequently used to screen 850 000 compounds from the AstraZeneca compound collection, with a Z′ factor of 0.66 ± 0.06. Hits were followed up by IC50 determination in both a concentration-response assay and a technology artifact assay.
Introduction
Flap endonuclease-1 (FEN1) is a member of the Rad2 family of metallonucleases and is involved in several DNA repair and replication pathways, including the processing of Okazaki fragments during DNA replication, long-patch DNA base excision repair pathway, and rescue of stalled DNA replication forks that encounter DNA. 1 These activities are important particularly at G-rich regions of telomeric DNA that are prone to form G-quadruplexes, which can stall replication forks at the ends of chromosomes. 2 FEN1 is highly conserved and has been demonstrated to play a critical role in both prokaryotes and eukaryotes. Deletion of the yeast FEN1 homologue Rad27 results in severe growth defects and cell cycle arrest, whereas homozygous mouse knockouts of FEN1 are embryonic lethal.3,4 FEN1 overexpression has been found in a number of cancers, including testis, lung, bladder, and brain tumors. The mutation or downregulation of FEN1 has been shown to sensitize cells to chemotherapeutic agents such as cisplatin, temozolomide, and MMS. 5 This has led to interest in developing FEN1 inhibitors as stand-alone agents for treating cancers that rely on its activity and as a therapy in combination with chemotherapeutic agents that cause DNA damage.
A small number of putative FEN1 inhibitors have appeared in the literature, but none of these were deemed leadlike enough to pursue for development of a drug.6,7 A high-throughput screening (HTS) approach was therefore considered to be a key strategy in finding novel leads. Various technologies have been exploited in studies of FEN1 activity in the past. Assays based on the separation of radio-labeled or fluorescently labeled DNA substrate and product by gel electrophoresis or high-performance liquid chromatography (HPLC) have been used extensively to determine the kinetics and substrate specificity of FEN1.8,9 These methods are too low-throughput to be used for a screening campaign, and this has led to the development of several high-throughput assays including fluorogenic donor/quencher assays and a chemiluminescence-based AlphaScreen assay.6,7
As FEN1 is a structure-specific DNA endonuclease, physiologically relevant substrates should contain certain key features including a 5′ flap and a single-nucleotide 3′ flap. All substrates used in previously published high-throughput assays lack a single nucleotide 3′ flap, which has been shown to be critical for specificity of the FEN1 cleavage site. 10 The presence of an unpaired single-nucleotide 3′ flap results in an eightfold increase in FEN1 nuclease activity and a fourfold reduction in KM compared with an identical substrate that lacks the 3′ flap. 9 The recent crystal structure of human FEN1 in complex with a DNA substrate containing a single-nucleotide 3′ flap showed the presence of a 3′ flap-binding pocket in which the 3′ hydroxyl group of the DNA substrate forms a bifurcated hydrogen bond with the backbone carbonyl of Lys314 and the hydroxyl group of Thr61. 11
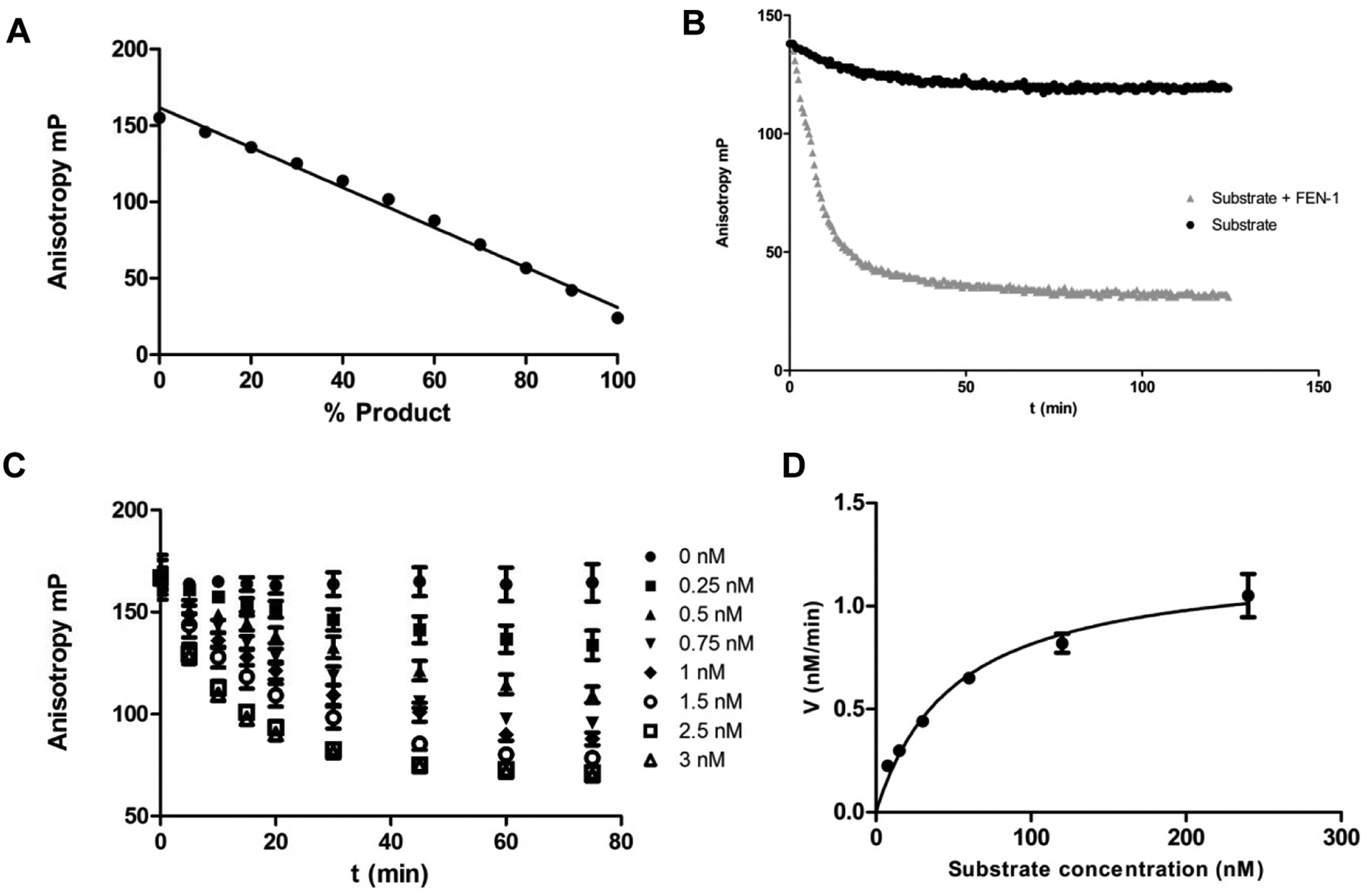
We developed an alternative, miniaturized, homogenous assay, using an FEN1 DNA substrate containing both a 5′ and 3′ flap. The assay uses fluorescence polarization (FP) to monitor the catalytic activity of FEN1 as it cleaves the FEN1 substrate. The FEN1 substrate contains a 5′ flap labeled with Atto495. Cleavage of the 5′ flap results in a labeled product with reduced size and a concomitant increase in depolarization of the fluorescence signal ( Fig. 1B ). The assay has been used to screen a large screening set of 850 000 compounds from the AstraZeneca compound collection.
(
Materials and Methods
Unless otherwise stated, all chemicals were purchased from Sigma-Aldrich (Gillingham, UK).
Expression and Purification of FEN1
DNA encoding full-length FEN1 (NP_004102) was codon-optimized by Life Technologies (Paisley, UK) using the Geneart proprietary GeneOptimizer technology and cloned into a proprietary AstraZeneca pET expression vector containing a C-terminal 6-His tag.
Freshly transformed BL21 Gold Escherichia coli cells containing the FEN1-6His plasmid were used to seed 75 mL flasks of LB broth containing 50 µg/mL kanamycin and 12.5 µg/mL tetracycline. The flasks were incubated overnight (o/n) at 37 °C, shaking at 180 rpm. The OD600s of the 75 mL o/n seeders were measured, and the appropriate volume was added to 600 mL flasks of LB broth containing 50 µg/mL kanamycin and 12.5 µg/mL tetracycline to give a starting OD600 of 0.05. The flasks were incubated at 37 °C, 180 rpm, until OD600 reached ~0.3, then cooled to 18 °C and induced with IPTG (final concentration 0.1 mM) when the ODs reached 0.4 to 0.5. The flasks were incubated o/n at 18 °C, 180 rpm. Cells were harvested and stored at −80 °C.
All steps in the purification were undertaken at 4 °C. Cells were lysed in buffer (50 mM NaH2PO4, pH 7.5, 300 mM NaCl, 5 mM imidazole, 1 mM TCEP, 1 mM PMSF, 1 µM aprotinin, pepstatin, leupeptin, 1:10 000 dilution of benzonase) by two passages through an Emulsiflex C-5 homogeniser (Avestin Inc, Ontario, Canada) set at 10 000 psi. The clarified lysate was applied to a 2 mL Ni-sepharose (GE Healthcare) column, freshly equilibrated in lysis buffer. The column was washed to baseline with buffer A (50 mM NaH2PO4, pH 7.5, 300 mM NaCl, 5 mM imidazole, 1 mM TCEP) followed by buffer A plus 35 mM imidazole, and finally the remaining bound protein was eluted with elution buffer (50 mM NaH2PO4, pH 7.5, 300 mM NaCl, 500 mM imidazole, 1 mM TCEP). Fractions containing FEN1-6His were pooled and buffer exchanged into storage buffer (50 mM HEPES, pH 7.5, 50 mM NaCl, 10 % v/v glycerol, 1 mM TCEP, 1 mM MgCl2) by passage through two in-series HiPrep 26/10 desalting columns (GE Healthcare). The FEN1-6His was stored at −80 °C until required. Protein concentration was determined by absorbance measurement at A280 using a molar extinction coefficient of 22340 M−1 cm−1.
DNA Substrate Preparation
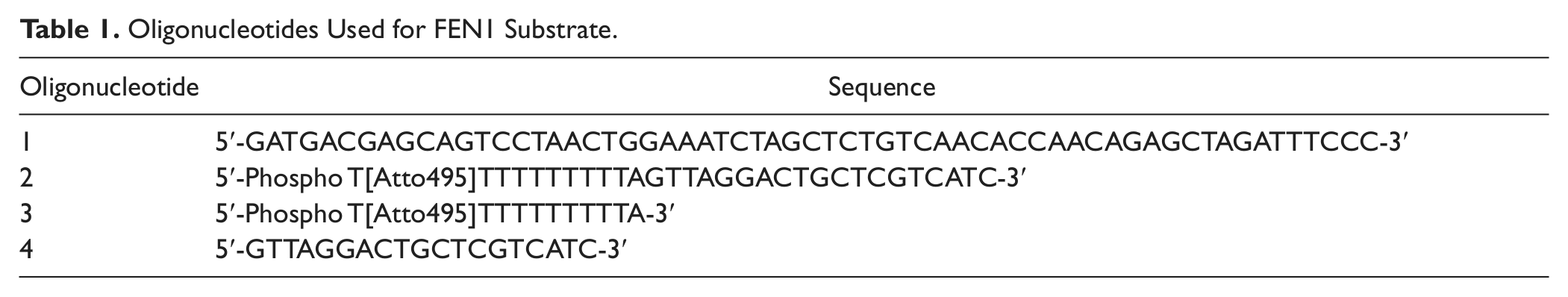
All oligonucleotides were purchased from Eurogentech (Seraing, Belgium) and made up to a concentration of 100 µM in nuclease-free water. FEN1 substrate was assembled using 1000 pM of oligonucleotide 1 and 1000 pM oligonucleotide 2 ( Table 1 ) in buffer (10 mM Tris-HCl pH 7.4, 50 mM NaCl, 1 mM EDTA). FEN1 product was assembled using 1000 pM of oligonucleotide 1, 1000 pM oligonucleotide 4, and 1000 pM of oligonucleotide 3 ( Table 1 ) in buffer (10 mM Tris-HCl pH 7.4, 50 mM NaCl, 1 mM EDTA). Both substrate and product mixtures were heated in the dark at 99 °C for 10 min followed by slow cooling to room temperature. This yielded a 10 µM (10 pM/µL) stock.
Oligonucleotides Used for FEN1 Substrate.
Assay Buffer
All reactions were performed at room temperature in freshly prepared assay buffer containing 20 mM Tris pH 7.5, 10 mM MgCl2, 10 mM KCl, 0.01 % v/v Tween20, and 1 mM dithiothreitol.
Compound Preparation
Compounds were dispensed into black 1536-well medium-binding microplates (Greiner Bio-One, UK, No. 782076) using an Echo 555 (Labcyte Inc., Sunnyvale, CA) as previously described. 12 For assay development, 40 nL of 10 mM compounds solubilized in DMSO were dispensed acoustically into the wells. This gave a final concentration of 1 % v/v DMSO and a final in-well compound concentration of 100 µM after addition of all reagents. For single-shot screening, 10 nL of 4 mM compounds solubilized in DMSO were dispensed acoustically into the wells. This gave a final concentration of 0.25% v/v DMSO and a final in-well compound concentration of 10 µM after addition of all reagents. For concentration-response screening, varying amounts of compound were dispensed acoustically to give a half-log dilution range. Wells were backfilled with the required volume of DMSO to ensure a final concentration of 1% v/v after addition of all reagents.
FP Measurements
The FP measurements were performed on a BMG PheraStar plate reader (BMG Labtech GmbH, Ortenberg, Germany) using an FP optic module containing an excitation-polarization filter (Ex = 485 nm) and emission filter (Em = 520 nm; parallel and perpendicular) with a positioning delay of 0 seconds and 50 flashes per well. The reader was set to give an FP reading of 200 mP using uncleaved substrate. All FP values are expressed as anisotropy (R). Anisotropy is calculated using the following formula:
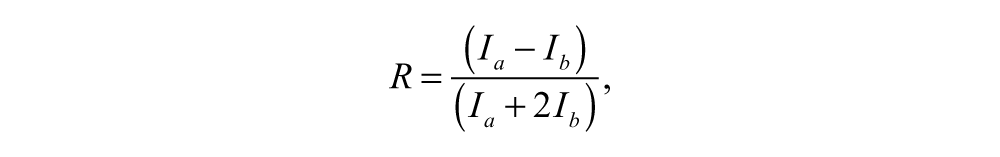
where Ia is the parallel emission intensity and Ib is the perpendicular emission intensity.
Assay Optimization
The linearity of response with increasing FP signal was confirmed by mixing varying ratios of substrate and product DNA oligomers. Each reaction was made up to a final volume of 8 µL using assay buffer. The final DNA concentration was 30 nM.
Cleavage of the FEN1 substrate was confirmed by incubating 30 nM DNA substrate in assay buffer with 2.5 nM FEN1 in a final volume of 8 µL. The FP signal was read every minute for 120 min.
Kinetic parameters for the FEN1-catalyzed hydrolysis of the FEN1 substrate were determined by incubating varying concentrations of FEN1 substrate with 2.5 nM FEN1. The FP signal was read every minute for 120 min. Initial rate data were fitted to the Michaelis-Menten equation using GraphPad Prism V.5 (GraphPad Software Inc.).
Inhibition of FEN1 by Tool Compounds
Two microliters of 60 nM FEN1 substrate in assay buffer and 2 µL of 3 nM FEN1 were added separately to plates that had been predispensed with compound. The plates were covered and left to incubate for 1 hour at room temperature, before the enzymatic reaction was quenched by the addition of 4 µL 0.5 M EDTA. The IC50 data were fitted using Origin 7.5 (OriginLab Corporation, Northampton, MA).
High-Throughput Assay Format Development
To determine the effect of DMSO on the FP assay, the time course was repeated using 30 nM FEN1 substrate and 1.5 nM FEN1 in buffer containing 0 to 10% v/v DMSO.
Reagent stability was assessed at room temperature for up to 2.5 h prior to initiation of the reaction. Five milliliters 5 mL of 3 nM FEN1 in assay buffer and 5 mL of 60 nM FEN1 substrate in assay buffer were stored at room temperature on the bench for up to 2.5 h prior to the start of the reaction. Two microliters of both substrate and FEN1 were then added to black 1536-well medium-binding plates. The plates were covered and left to incubate for 1 h at room temperature before the enzymatic reaction was quenched by the addition of 4 µL 0.5 M EDTA.
High-Throughput Compound Library Screening
Test compounds from the AstraZeneca compound collection were dissolved in DMSO to form stock solutions (4 mM). A final concentration of 0.25% v/v of DMSO and 10 µM test compound was achieved after reagent additions. A Multidrop Combi (Thermo Scientific, Waltham, MA) was used to dispense 2 µL of 60 nM FEN1 substrate and 2 µL of 3 nM FEN1 to all wells. The plates were incubated for 1 h at room temperature before the enzymatic reaction was quenched by the addition of 4 µL 0.5 M EDTA and the plates were read. Data were analyzed using HBase, proprietary AstraZeneca software.
Concentration Response Follow-up Screening
Seven-point, half-log concentration-response curves, with a top concentration of 100 µM were constructed. The assay was performed using the process described for the high-throughput compound library screening.
Artifact Assay Development
To identify compounds that interfere with the FP signal rather than inhibit FEN1 activity, an artifact assay was developed. Substrate and product oligonucleotides were titrated to give a signal that matched that seen in the activity assay (~60% conversion). Final concentrations of 12 nM FEN1 substrate oligo and 18 nM of the DNA product oligo were used in the artifact assay. Two microliters of 60 nM FEN1 substrate/product mixture and 2 µL of assay buffer (in place of FEN1) were added to all wells. The plates were incubated for 1 h at room temperature before 4 µL 0.5 M EDTA was added to each well and the FP was read.
Results
Design of Oligonucleotide Substrate
The synthetic FEN1 substrate used for the development of the FP assay was carefully designed to ensure it incorporated the necessary features for biologically relevant catalysis by FEN1.
The FEN1 substrate consists of 2 oligonucleotides, which are annealed to form a duplex. The oligonucleotides required for the FEN1 substrate are shown in Table 1 (oligonucleotides 1 and 2). Oligonucleotide 1 forms the backbone of the substrate. This oligonucleotide has a region of self-complementation that allows it to form a hairpin loop structure during annealing. The 3′ nucleotide is mismatched and forms a 3′ flap, which has been shown to be essential for optimal cleavage of the substrate by FEN1. 9 Oligonucleotide 2 consists of a 19-base complementary region and a short poly-T noncomplementary region. Upon annealing with oligonucleotide 1, the noncomplementary region forms the 5′ flap structure. Oligonucleotide 2 is further modified to contain an Atto495 fluorophore at the 5′ terminus. A schematic of the assay and the substrate oligonucleotide can be found in Figure 2 .
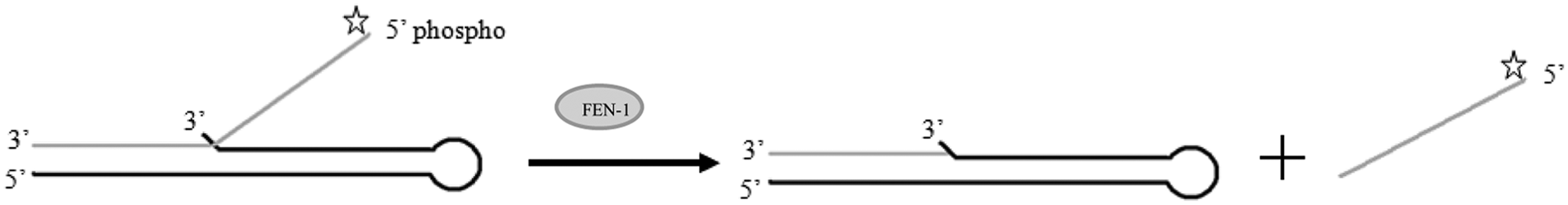
Assay schematic: The substrate for FEN1 consists of oligonucleotide 1 (black) annealed with oligonucleotide 2 (gray), which is labeled at the 5′ end with Atto495. Incubation of the substrate with FEN1 results in cleavage of the single-stranded 5′ flap oligonucleotide. The reduction in mass of the labeled oligonuclotide allows the detection of FEN1 activity by FP.
Design, Specificity, and Optimization of the FEN1 FP Assay
To ensure proportionality of the FP signal with the ratio of substrate and product in the reaction mixture, the annealed substrate was titrated with oligonucleotide product with the overall concentration of DNA oligonucleotide kept constant at 30 nM ( Fig. 1A ).
The ability of FEN1 to cleave the Atto495-labeled FEN1 substrate was determined by incubating 30 nM FEN1 substrate in the presence and absence of FEN1. A decrease in FP signal was seen only when the substrate is incubated with FEN1 ( Fig. 1B ), and the rate of reaction was proportional to the concentration of FEN1 present in the reaction ( Fig. 1C ). An enzyme concentration of 1.5 nM was chosen for subsequent experiments as the reaction showed linear behavior at this concentration for more than 1 h.
The Michaelis-Menten parameters were then determined by monitoring the rate of reaction at several different concentrations of substrate ( Fig. 1D ). A KM value of 50.5 ± 8.6 nM was determined. A substrate concentration of 30 nM (~KM) was chosen for subsequent experiments.
As DMSO was used to solubilize the compounds used for screening the FEN1 FP assay needed to be tolerant of the presence of low concentrations of DMSO (1% [v/v] DMSO in each well). The effect of DMSO concentration on the HTS assay was therefore checked, and no change in signal was found in the presence of up to 10% (v/v) DMSO ( Fig. 3A ).
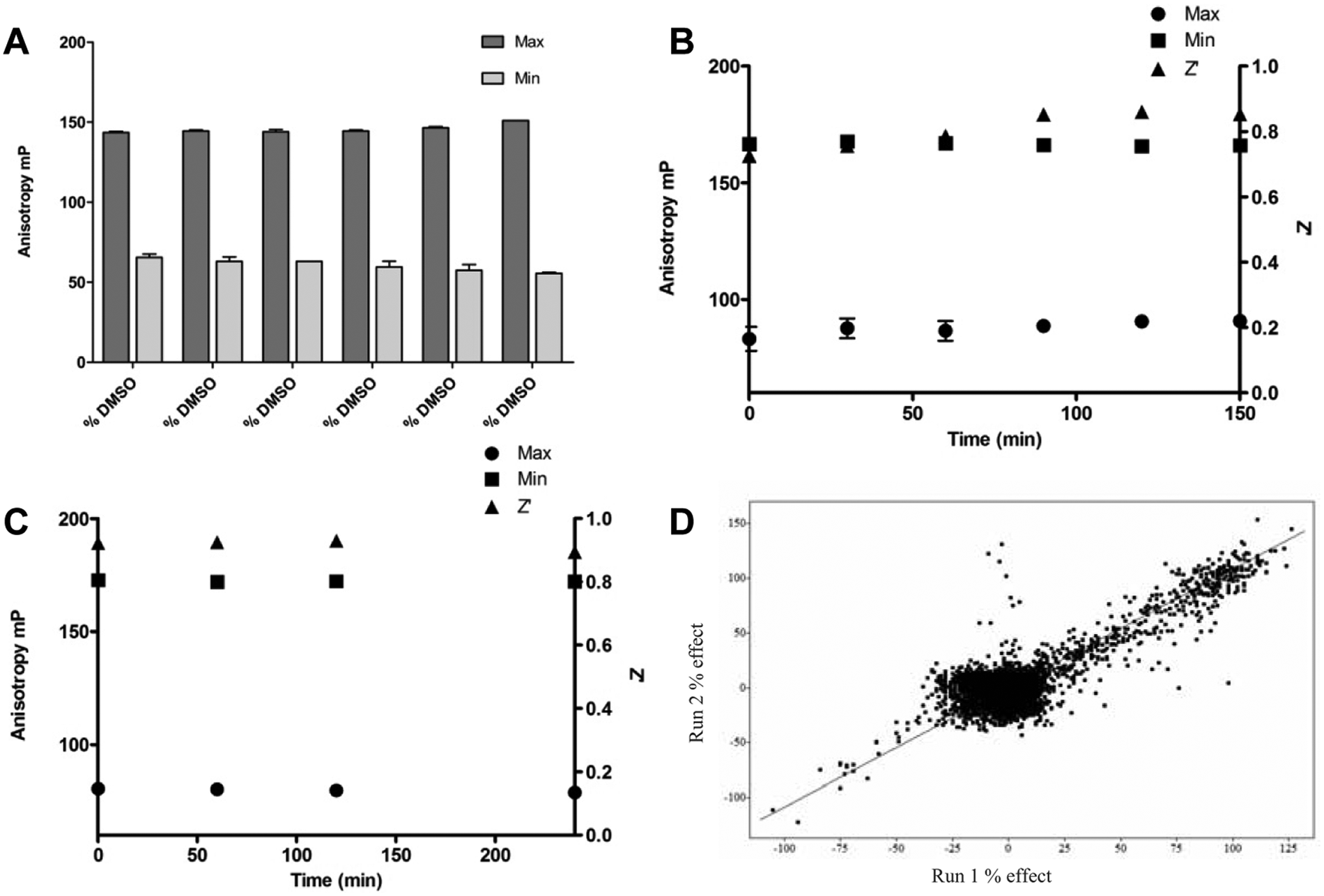
(A) DMSO tolerance: 30 nM FEN1 substrate was incubated with 1.5 nM FEN1 in the presence of varying concentrations of DMSO. The data shown are representative data from a single experiment. The data points are the mean of the replicates from that experiment, and the error bars are the standard deviation of the replicates within the experiment. The assay was found to be tolerant of DMSO concentrations of up to 10 % v/v. (
To screen 850 000 compounds in an HTS, it was important to ensure that prepared reagents were stable over the time required to process up to fifty 1536-well assay plates (approximately 2.5 h). Reagents were prepared ready for screening and then stored at room temperature for up to 2.5 h before being used in the assay (reagent stability test results shown in Fig. 3B ). A Z′ factor >0.6 was achieved and remained unaltered over the measured time, indicating that assay reagents were sufficiently stable to allow processing of appropriate-sized batches of plates for an HTS campaign. The FP signal was measured up to 4 h after the addition of EDTA to determine the stability of the assay signal and found not to change significantly over this time period ( Fig. 3C ).
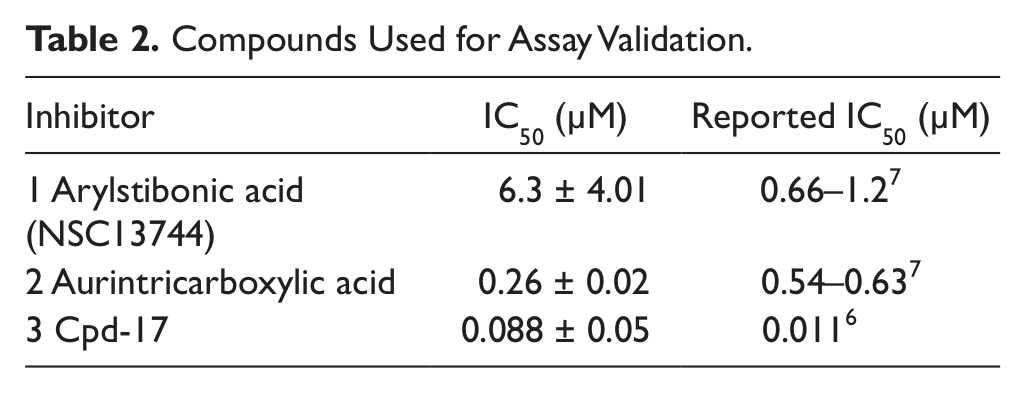
To validate the assay’s performance, we tested several reported FEN1 inhibitors, all of which showed concentration-dependent inhibition of FEN1, with IC50 values consistent with those previously reported ( Table 2 ).6,7
Compounds Used for Assay Validation.
Assay Validation
The suitability of the assay for HTS was assessed by testing a validation set of approximately 12 000 compounds. This validation set was screened at a single compound concentration of 10 µM on two separate occasions with a randomized plate/well assignment, allowing us to estimate screening performance and assay reproducibility under fully automated conditions ( Fig. 3D ). The observed correlation between these two experiments was in good agreement, as evidenced by a linear fit with an R2 value of 0.79.
In addition, control plates containing Min (a concentration of 10 × IC50 of Cpd-17), Max (DMSO), and reference (a concentration of IC50 of Cpd-17) wells were included in the runs to establish the Z′ factor.7,13 An overall screening Z′ factor of ~0.6 was achieved, demonstrating that the assay’s performance is robust and suitable for application in an HTS campaign.
High-Throughput Compound Library Screening
A high-throughput compound library screen of 850 000 compounds was completed in 1536-well format. Plates were run in batches of 50 with the first and last plate of each batch used for control plates (max/min/reference). The latter were used to calculate the Z′ factor for each batch. In total, 12 batches were completed without issue, and with a Z′ factor across all batches of >0.6, which highlights the consistent performance of the assay during the HTS campaign. Data analysis was completed using an in-house wavelet-based pattern correction algorithm. This was applied to correct for any potential spatial patterns on plates and temporal patterns over the time of the run. 14 Z′ factors were calculated following application of this pattern correction. The percentage effect for each compound was calculated based on the control data for each batch (% effect = 100 × [1 – (test compound – median min control)/(median max control – median min control)]. Figure 4A shows a histogram of one batch of HTS data. The histogram shows a normal distribution of the data around 0% effect of the max control. Using a percentage-based cutoff of ≥30% effect, we designated 8538 compounds in total as active in the screen; this can be seen as a tail on the normal distribution (insert in Fig. 4A ).
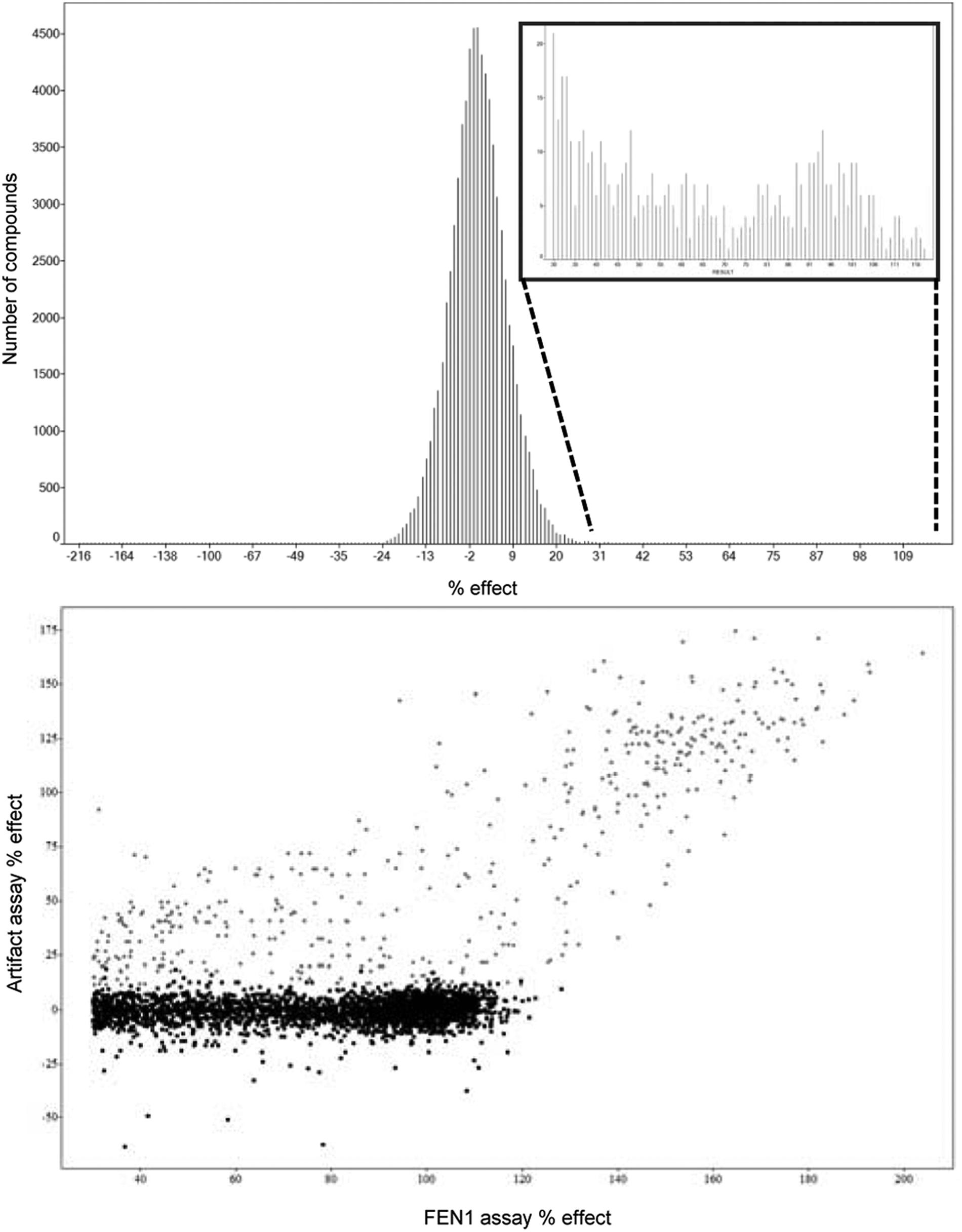
(A) Histogram plot of single-concentration data tested at 10 µM is shown. Percentage effect was calculated, and a percentage cutoff of ≥30% was used to define actives. Insert shows the distribution of the designated actives. (
As FP assays are subject to interference, primarily by light scattering and auto-fluorescent compounds, an orthogonal artifact assay was developed to aid the hit confirmation process. 15 A total of 15% of the actives were identified as artifacts of the technology ( Fig. 4B ). As the artifact assay was run in the absence of FEN1 enzyme, any increase in signal would be independent of FEN1 activity, and compounds exhibiting such behavior were removed as technology artifacts. To correct for such artifactual activity, we used an in-house algorithm, which compared enzyme assay data with artifact data on a well-by-well basis and normalized the data, either by multiplying by a factor if quenching of signal was detected or by subtraction if an increase in signal was detected during the artifact assay. 16
The compounds identified in the single-concentration assay were then screened in a concentration-response assay and IC50 values determined for each compound. The concentration-response assay was run using the same conditions as the single-shot screen. Concentration-response plates contained control wells (max/min), which were used for Z′ factor calculation. In total, 56 compound plates were run with an average Z′ factor of >0.6. Artifact data were used during the analysis of the concentration-response data, and 1280 of the 8538 hits from the primary screen were identified as artifacts of the technology. With the artifacts removed from the data set, a total of 6261 compounds were confirmed as active in the assay with an IC50 of less than 100 µM (73% confirmation rate).
Discussion
A review of the assay options for FEN1 described in the public domain revealed either low-throughput methods using gel electrophoresis and HPLC, which were unsuitable for HTS, or high-throughput methods, which involved multiple fluorescent labels or the requirement of antibodies or other secondary labels.6–9 None of the methods identified in the literature seemed optimal for delivering a simple, robust assay capable of delivering our HTS campaign. We therefore chose to exploit FP technology to deliver an alternative, miniaturized, homogenous assay to monitor FEN1 activity using a singly-labeled FEN1 DNA substrate.
FP is based on the observation that relatively small, fluorescently-labeled molecules tumble faster in solution than larger molecules. When these fast-tumbling fluorescent molecules are excited with plane-polarized light, the emitted light becomes depolarized with respect to the plane of polarization. Although FP assays are widely used for HTS, most rely on the displacement of a labeled probe from the target of interest. FP assays have also been used to monitor the catalytic activity of enzymes. These assays are generally based on the change in FP signal of substrate or product binding to a larger molecule such as an antibody. 17 There are several examples of FP assays using directly labeled substrates, including an HTS monitoring the cleavage of a DNA/RNA duplex by ribonuclease H, which was used to screen 1.8 million compounds.18–22 The assay we have developed employs FP technology using a singly-labeled DNA substrate consisting of two annealed strands of DNA ( Fig. 2 ).
The DNA substrate adheres closely to literature precedent to ensure suitability for cleavage by the FEN1 enzyme. 9 Initial experiments showed that labeling the flap of the substrate provided a way to monitor DNA flap cleavage through a reduction in mass and therefore a decrease in the FP of the oligonucleotide following cleavage by FEN1 ( Fig. 1B ).
The FEN1 DNA substrate has a KM value of 50.5 ± 8.6 nM, which is in good agreement with the values reported for similar substrates. 9 At a substrate concentration equal to KM, 50% of the enzyme is free in solution, and 50% is in the enzyme-substrate complex. A substrate concentration close to KM was chosen to ensure that the assay was sensitive enough to find both substrate competitive inhibitors, which bind to free enzyme, and uncompetitive inhibitors, which bind only to the enzyme-substrate complex. 23 Noncompetitive and mixed inhibitors are able to bind to both enzyme and enzyme-substrate complex and are therefore less sensitive to the concentration of substrate used in screening.
The FP assay was used to screen a large subset of the AstraZeneca compound collection (850 000 compounds) at a single concentration of 10 µM. From this initial screen, 8538 compounds were flagged as active at 10 µM.
The activity of the hits identified in the single-shot HTS campaign was confirmed in concentration-response studies, and artifact assay data were used to remove compounds that interfered with the technology readout. A final hit confirmation rate of 73% was achieved in the assay, showing the reliability of the assay when screening in single shot or concentration response. Across all of the FEN1 screening runs, a Z′ factor of 0.66 ± 0.06 was maintained, indicating that this assay is suitable for HTS.
Several recent studies have suggested that false-positives in HTS campaigns can account for a substantial proportion of HTS hits. 24 Further annotation of the hits identified by this screen is therefore required to ensure that they are inhibiting FEN1 specifically. False-positive results may arise through several different mechanisms, including aggregation, destabilization of protein structure, or redox activity. Furthermore, many of the hits identified in the FEN1 HTS may act through intercalation of substrate DNA, rather than specific inhibition of FEN1.
FP has shown to be a reliable method for the prosecution of an FEN1 HTS campaign, allowing us to deliver a full HTS campaign with a simple and robust assay readout. We believe that the FP approach could be used in similar ways for other DNA-modifying targets, such as FANCD2/FANCI-associated nuclease 1 (FAN1) and Werner syndrome ATP-dependent helicase (WRN), in which a change in the mass of DNA by either cleavage or annealing of oligonucleotides is seen.
Footnotes
Acknowledgements
We would like to thank Tuseef Chaudhary and Hedley Carr for their input into the design of the FEN1 substrate and Richard Metzger for his help with the HTS campaign and subsequent data processing. We would also like to thank Mark Abbott, Catherine Bardelle, Derek Barratt, Peter Simpson, and Jonathan Wingfield for their comments and scientific input during the preparation of the article.
Declaration of Conflicting Interests
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: All authors work for AstraZeneca.
Funding
All authors are employed by AstraZeneca.
