Abstract
Phosphorus flame retardants are a group of chemicals that are used to slow or prevent the spread of fire. These compounds have been detected in different environments including human organism. In the present study, we have investigated DNA-damaging potential and effect on DNA methylation of tris(2-chloroethyl) phosphate (TCEP) and tris(1-chloro-2-propyl) phosphate (TCPP) in human peripheral blood mononuclear cells (PBMCs). In order to determine DNA damage and repair, the alkaline and neutral versions of the comet assay were used. The level of DNA methylation was determined with specific antibodies against methylated DNA. PBMCs were exposed to TCEP and TCPP at the concentrations in the range of 1–1000 µM for 24 h. We have observed that TCEP and TCPP induced DNA damage—DNA breaks and alkali-labile sites. All DNA damages were effectively repaired during 120-min repair incubation. The results have also shown that TCEP and TCPP decreased the level of DNA methylation in PBMCs. In the case of TCEP, this effect was observed at a very low concentration of 1 µM.
Introduction
Flame retardants (FRs) are a group of chemicals that are designed to prevent the initiation and spread of fire. Moreover, several FRs are also serving as plasticizers. FRs are extensively used in furniture, textiles, mattresses, and electronics and also in plastic equipment used in homes and offices. Some FR compounds can be released from materials to the outside. So, they are found in various environments: sediments and soils, water, air, and indoor dust. These compounds are detected in marine and freshwater animal, avian, insect, and human samples such as human breast milk, blood, hair, adipose tissue, and various organs. 1,2 Currently, one of the important groups of FR compounds are organophosphate FRs (OPFRs) that replace brominated FRs (BFRs) such as polybrominated diphenyl ethers and hexabromocyclododecane. OPFRs are synthetic phosphoric acid derivatives and can be divided into three groups: chlorinated OPFRs, alkyl-OPFRs, and aryl-OPFRs. 1
World consumption of FRs will reach 2.8 million ton in 2018. 3 OPFRs production accounts for 20% of the world production of FR used in plastics and textiles. 4 A recent study in Germany showed that the total concentration of OPFRs in indoor environments is about eight times higher than in outdoor environments, suggesting the presence of major sources of OPFRs in indoor environments. 5 Tris(2-chloroisopropyl) phosphate (TCPP), tris(isobutyl)phosphate, and tributyl phosphate were the dominating compounds both in indoor and in outdoor air. 5 There are several ways of absorbing these compounds by the human body by inhalation, ingestion, and skin absorption. 6 –8
Toxicological studies have shown that OPFRs have the potential to induce adverse reproductive, endocrine, and neurological effects in animals as a result of long-term exposure. 9 –12 Meeker and Stapleton 13 observed that concentrations of OPFRs in house dust may be associated with altered hormone levels and decreased sperm concentration in men. Some of OPFRs such as tris(2-chloroethyl) phosphate (TCEP), tris (1-chloro-2-propyl) phosphate (TCIPP), and Tris(1,3-dichloro-2-propyl) Phosphate (TDCIPP) have been proven to be carcinogenic. 1 They can accumulate in the liver, kidneys, and testis and induce tumors. 1,9
TCEP and TCPP are the most widely used chlorinated OPFRs. On April 5, 2018, European Chemicals Agency published screening report about an assessment of whether the use of TCEP, TCPP, and tris (1,3-dichloro-2-propyl) phosphate (TDCP) in materials should be restricted. 14 We can read in this report that TCEP is toxic for reproduction and TCEP have the carcinogenic properties. TCPP is identified as substance with similar properties and uses to TCEP. 14 The results point to a relevant carcinogenic potential of especially TCEP, but the mechanism of is not known. Carcinogenic action of the chemical compound may arise from induction of DNA damage. Unrepaired DNA damage can make mutations, and these may transform normal cells to cancer cells. On the other hand, epigenetic changes including DNA methylation may also affect carcinogenesis. Reducing the level of DNA methylation can lead to overexpression of oncogenes, whereas DNA hypermethylation can cause down-expression of tumor suppressor genes.
Given the above, we decided to compare the DNA-damaging potential of TCEP and TCPP in human peripheral blood mononuclear cells (PBMCs) using the single-cell gel electrophoresis (comet assay). The kinetics of repair of DNA damage induced by TCEP and TCPP was also measured. Moreover, we examined the effects of TCPP and TCEP on global DNA methylation in PBMCs.
Materials and methods
Chemicals
TCPP and TCEP were purchased from Sigma-Aldrich (St Louis, Missouri, USA). Medium used for PBMCs isolation (lymphocyte separation medium (LSM)) as well as Roswell Park Memorial Institute medium (RPMI) 1640 medium and fetal bovine serum was obtained from Cytogen (Germany). Low melting point (LMP) and normal melting point (NMP) agarose, phosphate-buffered saline (PBS), 4′,6-diamidino-2-phenylindole (DAPI), dimethyl sulfoxide (DMSO), and hydrogen peroxide (H2O2) were purchased from Sigma-Aldrich. All other chemicals were of the highest commercial grade available.
Methods
Cells
The human PBMCs were isolated from the leukocyte buffy coat collected from blood of healthy, nonsmoking men (volunteers; aged 18–55). The cells were isolated by centrifugation in a density gradient of LSM (30 min, 600 ×
DNA damage
The final concentrations of each studied retardants were in the range from 1 µM to 1000 µM, which correspond to the level of these compounds determined in the environment. 4,9 We used the same concentration as in our previous article on TCPP and TCEP. 16 Then, we observed that TCEP at 1 mM and TCPP at 0.5 mM decreased viability of PBMCs. At 1 mM, TCPP decreased PBMCs viability up to 47.7%, while TCEP up to 76.9% (vs. control 83.5%). 16 Because of the limited water solubility in water, tested compounds were dissolved in DMSO with final concentration of this solvent in sample equal 0.4%. In the alkaline version of the comet assay, PBMCs were incubated with TCEP and TCPP at the concentrations of 1, 10, 100, 500, and 1000 µM. In the neutral version of the comet assay, PBMCs were incubated with TCEP and TCPP at the concentrations of 10, 100, 250, 500, and 1000 µM. The incubation was performed at 37°C in 5% CO2 atmosphere in the darkness for 24 h. Each sample was analyzed in triplicate (PBMCs from three blood donors were taken). The cells after treatment with TCEP and TCPP were washed and resuspended in RPMI 1640 medium. A freshly prepared suspension of the cells in LMP agarose dissolved in PBS was spread onto microscope slides. The slides were processed as describe below in the section “Comet assay.”
DNA repair
TCEP and TCPP were added to the suspension of the cells to give the final concentrations of 100, 500, and 1000 µM in the alkaline version of the comet assay. In the neutral version of the comet assay, TCEP and TCPP were added to the suspension of the cells to give the final concentrations of 500 and 1000 µM. PBMCs were incubated for 24 h at 37°C in 5% CO2 atmosphere in the darkness. Each sample was analyzed in triplicate (PBMCs from three blood donors were taken). The cells after the treatment with TCEP and TCPP were washed and resuspended in a fresh, RPMI 1640 medium preheated to 37°C. Aliquots of the cell suspension were taken immediately (time “0”) and at 60 and 120 min later. Placing the samples in an ice bath stopped DNA repair. The slides were processed as described below in the section “Comet assay.”
Comet assay
The comet assay was performed at pH > 13 essentially according to the procedure of Singh et al. 17 as described previously. 18 We also performed the comet assay at pH 9 to determine the double-strand breaks (DSBs) level only. 19 A freshly prepared suspension of the cells (1 × 105 cells/ml) in 0.75% LMP agarose dissolved in PBS was spread onto microscope slides precoated with 0.5% NMP agarose. The cells were then lysed for 1 h at 4 °C in a buffer consisting of 2.5-M NaCl, 100-mM EDTA, 1% Triton X-100, 10-mM Tris, and pH 10. After the lysis, the slides were placed in an electrophoresis unit, DNA was allowed to unwind for 20 min in the solution consisting of 300-mM NaOH and 1-mM EDTA, pH > 13. Electrophoresis was conducted in the solution consisting of 30-mM NaOH and 1-mM EDTA, pH > 13 at ambient temperature of 4°C (the temperature of the running buffer did not exceed 12 °C) for 20 min at an electric field strength of 0.73 V/cm (28 mA). The slides were then washed in water, drained and stained with 2-μg/ml DAPI, and covered with cover slips. To prevent additional DNA damage, all the steps described above were conducted under dimmed light or in the dark. Each experiment in the alkaline version of the comet assay included a positive control, which were PBMCs incubated with H2O2 at 20 μM for 10 min at 4°C. Negative control was cells incubated with DMSO at 0.4% for 24 h at 37°C.
The comet assay at pH 9 was performed essentially according to the same procedure as the alkaline version except the pH value. In the version at pH 9, electrophoresis was run in a buffer consisting of 100-mM tris and 300-mM sodium acetate at pH adjusted to 9.0 by glacial acetic acid. 19 Electrophoresis was conducted for 60 min, after a 20-min equilibrium period, at 12 V (100 mA) at 4°C. Each experiment in the neutral version of the comet assay included a positive control, which were PBMCs exposed to irradiation in a dose of 9 Gy at 37°C. Negative control were cells incubated with DMSO at 0.4% for 24 h at 37°C.
The comets were observed at 200× magnification in an Eclipse fluorescence microscope (Nikon, Tokyo, Japan) attached to COHU 4910 video camera (Cohu, San Diego, California, USA) equipped with a UV-1 filter block (an excitation filter of 359 nm and a barrier filter of 461 nm) and connected to a personal computer-based image analysis system Lucia-Comet v. 6.0 (Laboratory Imaging, Praha, Czech Republic). Fifty images were randomly selected from each sample.
DNA extraction and purification
Genomic DNA was extracted and purified with QIAamp DNA Blood Mini Kit (QIAGEN, Germany) according to the manufacturer’s instructions. RNA digestion was performed at 37 °C for 1 h using RNase (QIAGEN). The concentration and quality of the DNA samples were determined by spectrophotometry (Nanodrop 2000; Thermo Scientific, Waltham, Massachusetts, USA) and 1% agarose gel electrophoresis.
DNA methylation
The DNA methylation test was performed according to the manufacturer’s instruction with Methylated DNA Quantification Kit (ABCAM, ab117128). This kit is a complete set of optimized buffers and reagents to quantify global DNA methylation by specifically measuring levels of 5-methylcytosine (5-mC) in a microplate-based format. Methylation levels were calculated relatively to the methylated control DNA (positive control included in the kit) and expressed as a percentage of methylated DNA (5-mC). The calculation of 5-mC amount was done with the use of standard curve created by using defined dilutions of methylated DNA. PBMCs were incubated with TCEP and TCPP at the concentrations of 1, 10, 100, and 250 µM for 24 h at 37°C. Negative control were cells incubated with DMSO at 0.4% for 24 h at 37°C. Each sample was analyzed in triplicate (PBMCs from three blood donors were taken). The assay was performed according to the manufacturer’s instructions, with quantitation at absorbance 450 nm on a microplate reader Synergy HT (BioTek Instruments, Winooski, VT, USA).
Data analysis
The values of the comet assay in this study were expressed as mean ± standard error of the mean from three experiments, that is, data from three experiments were pooled, and the statistical parameters were calculated. The data were analyzed using Statistica software (StatSoft, Tulsa, Oklahoma, USA). The values of the DNA methylation experiments were presented as mean ± standard deviation from three experiments. Statistical analysis was conducted using the Mann–Whitney test (samples with distributions departing from normality) and the Student’s
Results
DNA damage
Figure 1 demonstrates the level of DNA damage, including DNA single-strand break and DSBs as well as alkali-labile sites, induced by TCPP and TCEP in PBMCs during 24 h incubation at 37°C. Figure 2 shows the example images of comets from this experiment. We detected statistically significant increase in DNA damage after incubation of PBMCs with TCPP at the concentrations of 100, 500, and 1000 µM. TCEP induced significant DNA damage only after incubation at the concentration of 1000 µM. We did not observe an increase in the level of DNA damage in cells incubated with DMSO at 0.4% (negative control). In PBMCs incubated with H2O2 at 20 µM for 10 min on ice (positive control), we noted DNA damage at the level 9.31
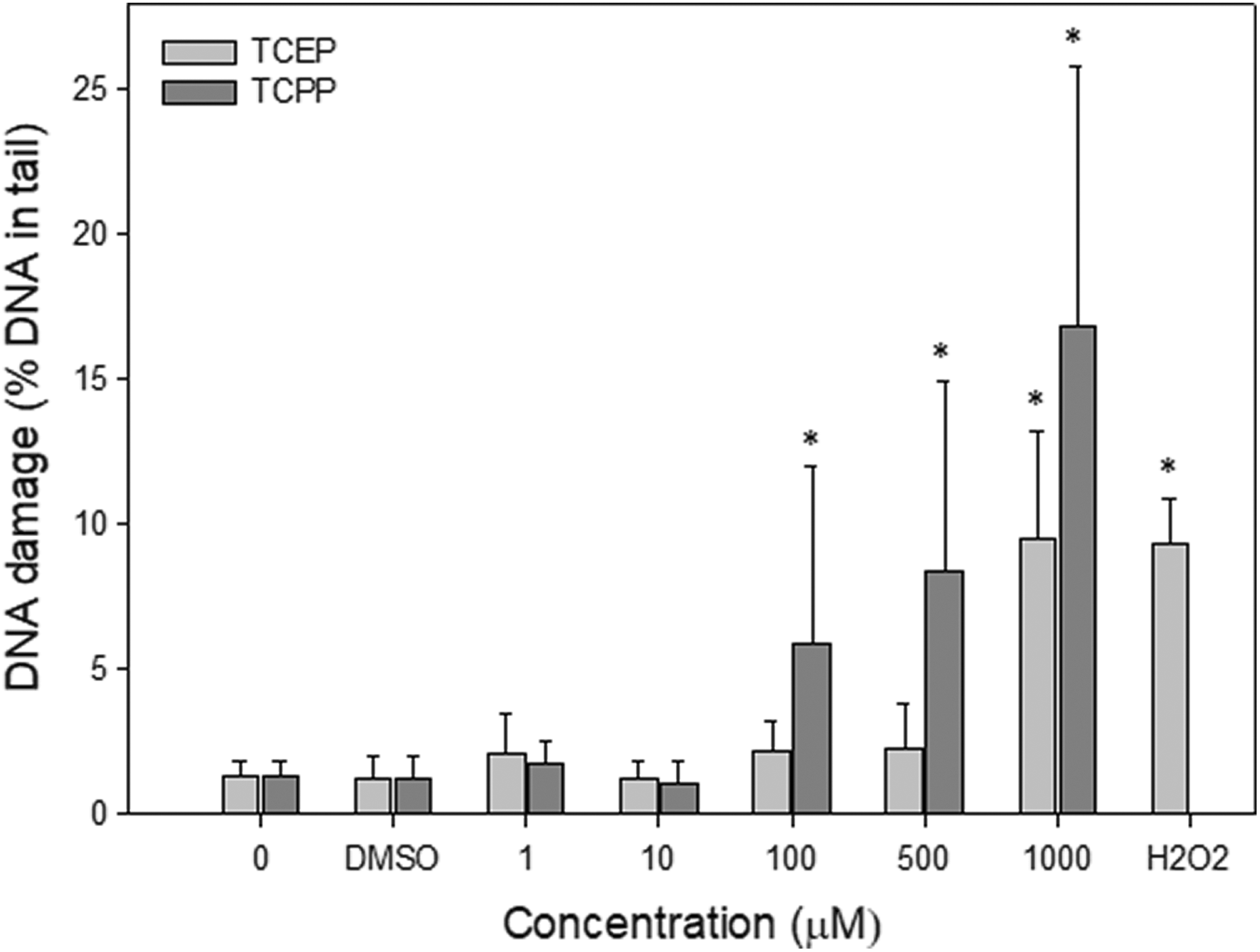
DNA damage (SSBs and DSBs) in human PBMCs incubated with TCEP or TCPP for 24 h at 37°C. DNA damage was measured as percentage of DNA in comet tail using the alkaline version of the comet assay. The mean value represents three independent experiments (donors); error bars represent SD, *
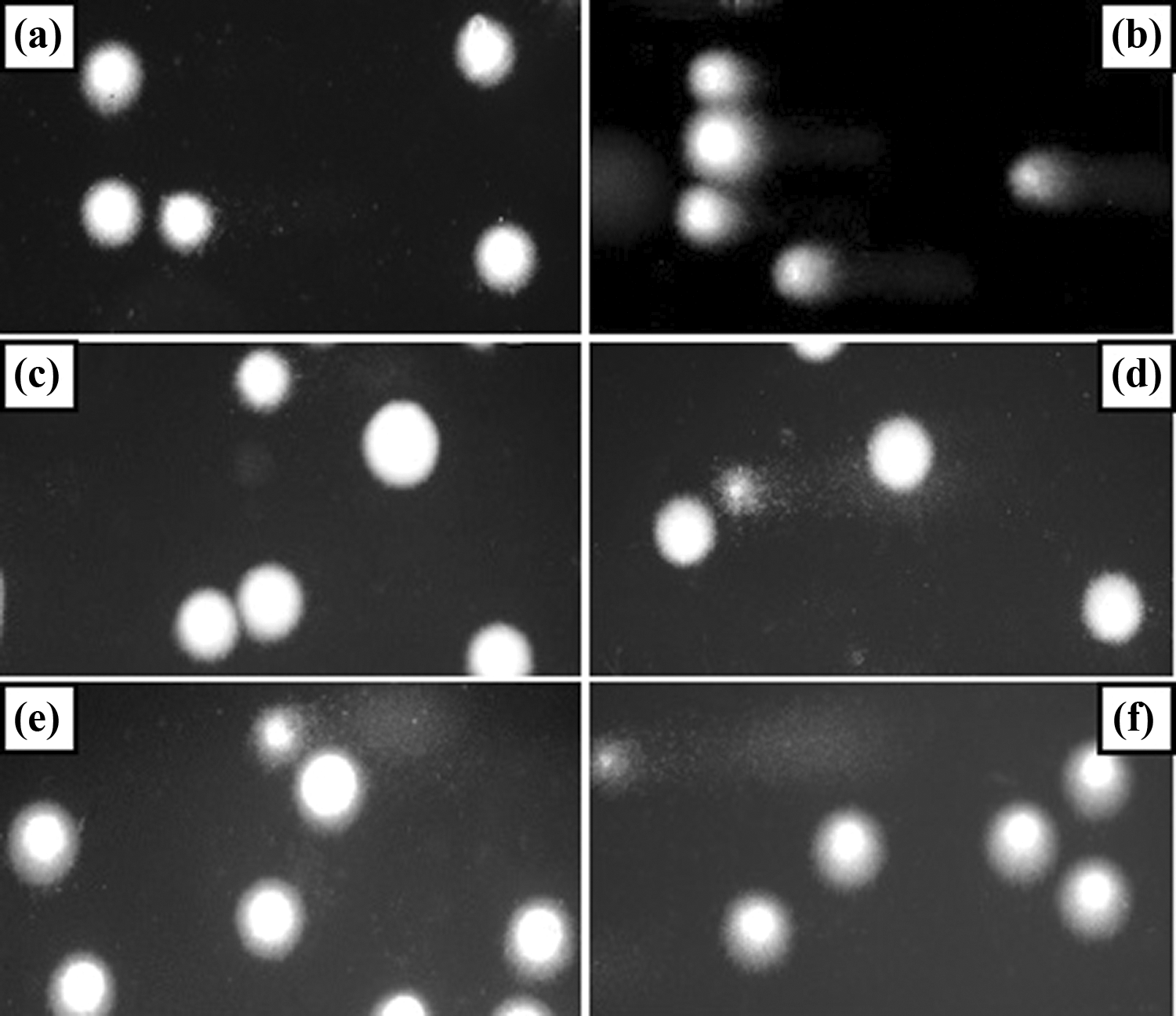
Representative photos of comets obtained in the alkaline version of the comet assay after incubation of PBMCs with TCEP and TCPP for 24 h at 37°C. (a) Negative control (cells incubated with DMSO at 0.4% for 24 h at 37°C), (b) positive control (cells incubated with H2O2 at 20 µM for 10 min at 4°C), (c) 100-µM TCEP, (d) 100-µM TCPP, (e) 1000-µM TCEP, and (f) 1000-µM TCPP. PBMC: peripheral blood mononuclear cell; TCEP: tris(2-chloroethyl) phosphate; TCPP: tris(1-chloro-2-propyl) phosphate; DMSO: dimethyl sulfoxide; H2O2: hydrogen peroxide.
Figure 3 shows the level of DNA DSBs induced in PBMCs by TCPP and TCEP during 24 h incubation at 37°C. In this experiment, cells exposed to irradiation in a dose of 9 Gy were positive control. We observed statistically significant increase in DNA DSBs in cells incubated with TCPP at the concentrations in the range of 100–1000 µM. TCEP induced DNA damage only at the concentration of 1000 µM. As in the case of DNA damage identified by the comet assay in the alkaline version, we did not observe a significant increase in DNA damage in cells incubated with DMSO at the concentration of 0.4% (negative control).

DNA damage (DSBs) in human PBMCs incubated with TCEP or TCPP for 24 h at 37°C. DNA damage was measured as percentage of DNA in comet tail using the neutral version of the comet assay. The mean value represents three independent experiments (donors); error bars represent SD, *
DNA repair
Figure 4 shows DNA damage (single-strand break and DSBs and as well as alkali-labile sites) in PBMCs incubated with TCEP at 100, 500, and 1000 µM immediately after 24-h incubation and after 60 and 120 min later. We noticed that PBMCs after 120-min incubation repaired all DNA damage induced by TCEP. Similarly, the cells were able to completely repair DNA damage induced by TCPP during 120-min postincubation (Figure 5).

Time course of the repair of DNA damage (SSBs and DSBs) in human PBMCs after 24-h incubation with TCEP. The repair was assessed as a decrease in the extent of DNA damage measured after 60 and 120 min as the percentage of the DNA in comet tail using the alkaline version of the comet assay. The results are mean of three independent experiments (donors). Error bars denote SEM, *

Time course of the repair of DNA damage (SSBs and DSBs) in human PBMCs after 24-h incubation with TCPP. The repair was assessed as a decrease in the extent of DNA damage measured after 60 and 120 min as the percentage of the DNA in comet tail using the alkaline version of the comet assay. The results are mean of three independent experiments (donors). Error bars denote SEM, *
The cells exposed to 20 µM H2O2 for 10 min at 4°C (positive control) were able to total repair DNA damage within the repair incubation time of 120 min (Figures 4 and 5). We did not observe any change in the level of DNA damage during repair incubation of cells exposed to DMSO (negative control).
Figures 6 and 7 show DNA damage, only DSBs, in PBMCs incubated with TCEP and TCPP (respectively) at 500 and 1000 µM immediately after 24-h incubation and after 60- and 120-min postincubation. We observed that DNA DSBs induced by both TCEP and TCPP were effectively repaired by PBMCs during 120-min postincubation. The cells exposed to irradiation in a dose of 9 Gy (positive control) were also able to total repair DNA double-strand breaks within the repair incubation time of 120 min (Figures 6 and 7).
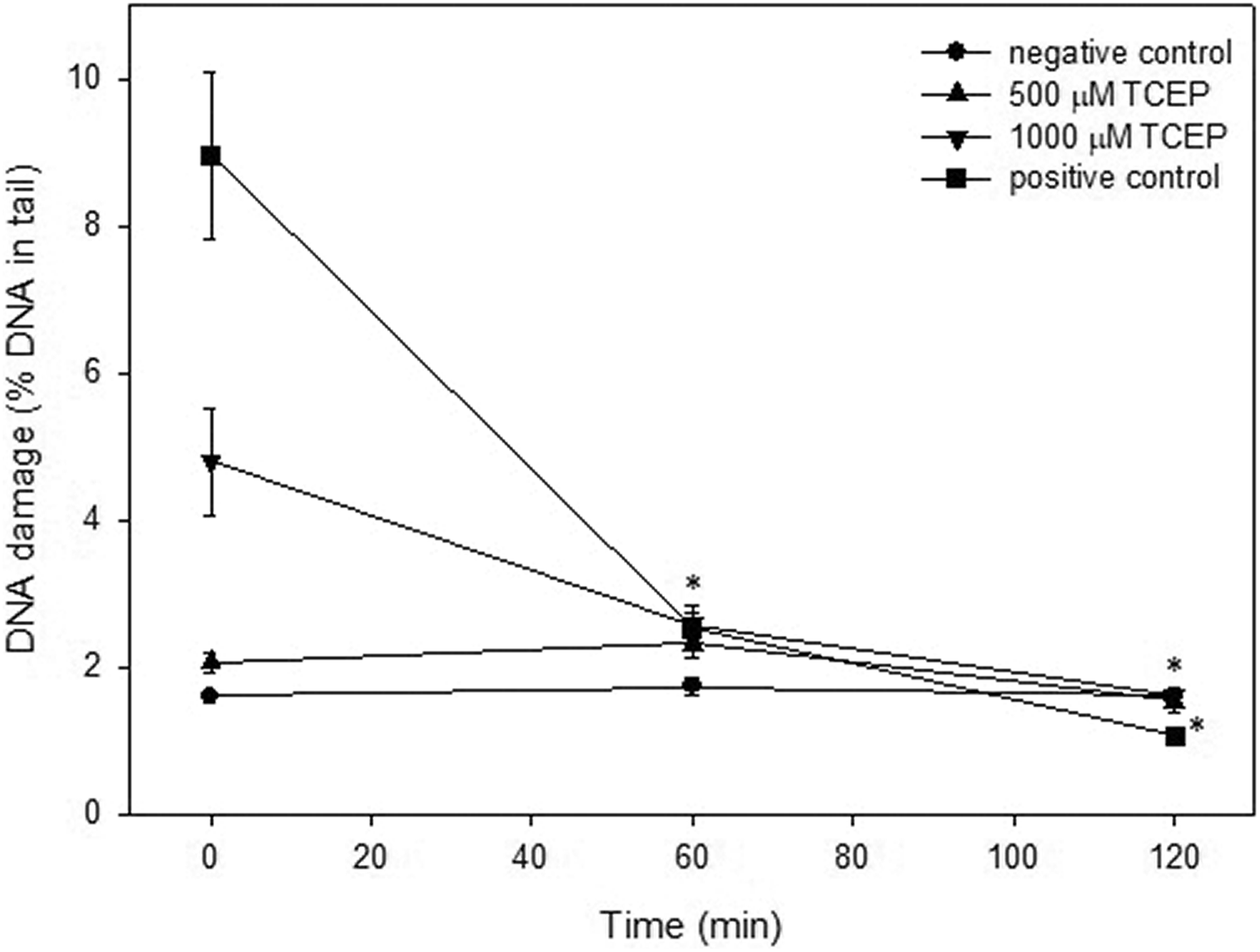
Time course of the repair of DNA damage (DSBs) in human PBMCs after 24 h incubation with TCEP. The repair was assessed as a decrease in the extent of DNA damage measured after 60 and 120 min as the percentage of the DNA in comet tail using the neutral version of the comet assay. The results are mean of three independent experiments (donors). Error bars denote SEM, *

Time course of the repair of DNA damage (DSBs) in human PBMCs after 24-h incubation with TCPP. The repair was assessed as a decrease in the extent of DNA damage measured after 60 and 120 min as the percentage of the DNA in comet tail using the neutral version of the comet assay. The results are mean of three independent experiments (donors). Error bars denote SEM, *
DNA methylation
Statistically significant changes were observed in the level 5-mC after incubation of PBMCs at all used concentrations of TCEP (1, 10, 100, and 250 µM; Figure 8). As compared to control cells, the level of global DNA methylation was decreased. TCPP also reduced the 5-mC level, but these results were not statistically significant with exception of the cell sample incubated for 24 h with TCPP at 100 µM (Figure 8). In this sample, we noticed the highest decrease of the level of global DNA methylation among of all testes performed with TCEP and TCPP.
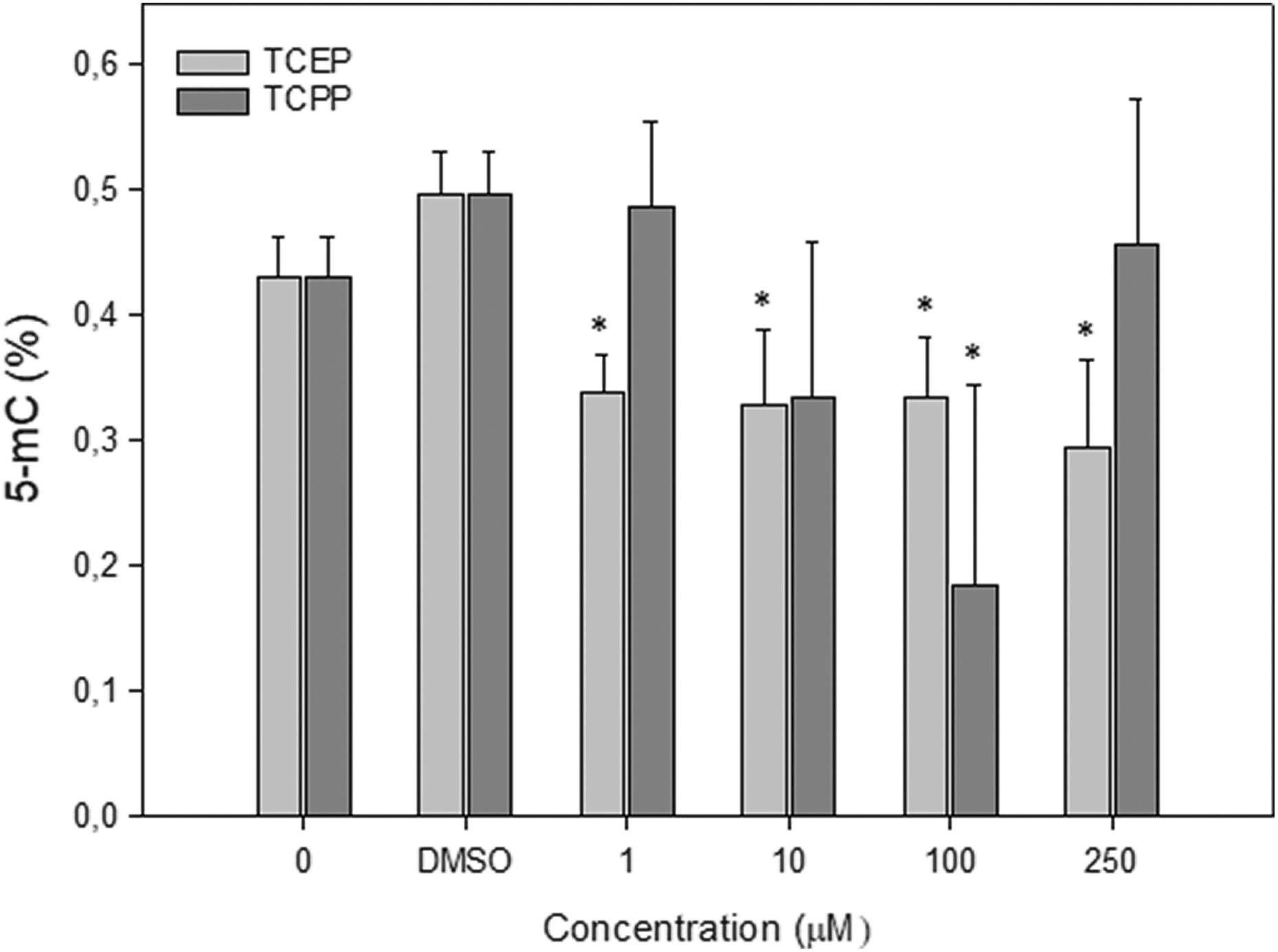
Level of 5-methylcytosine (%) in human PBMCs incubated with TCEP or TCPP for 24 h at 37°C. The mean value represents three independent experiments (donors); error bars represent SD, *
Discussion
OPFRs that are used as additives in polymer industry are not directly attached to the matrix (polymers), and therefore, they can be easily released into the environment. It is currently believed that OPFRs exhibit atmospheric stability, as a result of which they can be transferred at long distances from the emission sources. Despite the widespread occurrence of these compounds in the environment, there are almost no results on their genotoxic potential to human cells. Because of increasing demand for OPRFs, in vitro and in vivo studies should be carried out to exclude or confirm the toxicity of these compounds.
The aim of this study was to assess genotoxic effect of two OPFRs, that is, TCEP and TCPP on human PBMCs.
Firstly, the evaluation of DNA damage using alkaline version of the comet assay in PBMCs treated with OPFRs studied was conducted. The experiments have shown that both TCEP and TCPP induced DNA damage; however, TCPP had a stronger effect on this parameter. Similar differences for these compounds were noticed after analyzing the data obtained by neutral version of the comet assay. At the highest concentration used, TCPP primarily induces DNA DSBs—11.22 ± 1.71% in the neutral version of the comet assay versus 16.86 ± 8.97% in the alkaline version of the comet assay.
Different results concerning genotoxic potential of the OPFRs were obtained previously by Föllmann and Wober
20
who investigated the effect of TCEP and TCPP on DNA breaks formation in hamster fibroblasts (V79 cells). They found that both TCEP and TCPP at the concentrations from 1 µM to 1 mM did not cause statistically significant changes in this parameter neither without an external enzymatic metabolizing system nor in the presence of S9-mix. On the other hand, recently, Yang et al.
21
confirmed genotoxic potential of different concentrations of TCEP on intestinal cells of
In our study, we have also observed kinetics of DNA damage repair induced by OPFRs in PBMCs. It has been shown that PBMCs repaired all DNA damage after 120 min of postincubation of the cells, which were incubated with TCEP or TCPP.
Analysis using the neutral version of the comet assay associated with the detection-only DNA DSBs showed also statistically decrease in the percentage of DNA in the comet tails versus sample—time “0.” Although both tested compounds induced DNA breaks, the results showed effective repair of these lesions suggesting low genotoxic potential of the OPFRs studied. Unfortunately, in the scientific literature, there is lack of information related to the kinetics of DNA single-strand break and DSBs repair induced by OPFRs.
Many studies have shown that toxicity of OPFRs, including TCEP and TCPP, may be associated with generation of reactive oxygen species (ROS) and oxidative stress in cells. 22 –27 Zhang et al. 24 revealed that TCEP increased mitochondrial ROS production, which can cause cell cycle arrest through downregulation of SIRT3 deacetylase. The same group demonstrated that TCEP induced senescence-like growth arrest via the p21Waf1/Cip1-Rb pathway in a p53-independent manner in hepatocytes. 28 We suggest that DNA damage which we observed may be due to the ability of TCEP and TCPP to generate free radicals that can damage biomolecules and affect gene expression.
Studies in recent years have shown that both BFRs and OPFRs can alter DNA methylation and thus affect the expression of genes. It was found that decabromodiphenyl ether (BDE-209) and bisphenol A decreased DNA methylation and reduced DNA methyltransferase (DNMTI and DNMT3A) expression in human embryonic stem cells. 29 Moreover, it was shown that TDCIPP exposure predominantly resulted in hypomethylation of positions outside of CpG islands and within intragenic (exon) regions of the zebrafish genome. 30 Aberrant DNA methylation patterns were also present in human sperm when the exposure to OPFRs was increased. 31 We also observed a significant decrease in the level of DNA methylation in PBMCs incubated with TCEP. This effect was noticeable at a very low concentration of TCEP equal 1 µM. TCPP also significantly reduced the level of DNA methylation in PBMCs but at a higher concentration of 100 µM. It has been suggested that both global hypomethylation and specific hypermethylation of DNA can be linked to oxidative stress. 32 Global DNA hypomethylation and local promoter hypermethylation can be prompted by oxidative stress-induced inhibition of DNA methyltransferases and relocalization of DNA methyltransferases to CpG islands. On the other hand, DNA damage that occurs due to oxidative stress can also affect the expression of genes. It was shown that the presence of DNA damage lesion, 8-oxoguanine, leads to an approximately 300% increase in gene expression. 33
The results obtained in this study showed low genotoxic potential of TCEP and TCPP in PBMCs. This may be due to the effective and rapid repair of DNA damage induced by both compounds in this cell type. However, this genotoxic potential appears to be higher for TCPP, which induced DNA damage at lower concentrations than TCEP. By using the comet assay in the neutral version, it has been shown that TCPP at the maximal used concentration of 1000 µM induces mainly DNA DSBs. We have also shown that TCEP and TCPP reduce the level of DNA methylation in PBMCs. In the case of TCEP, this effect was observed at a very low concentration of 1 µM. Further studies are necessary to determine the mechanism of action of these compounds in human cells, especially to clarify effects of decrease in the level of DNA methylation on the gene expression.
Footnotes
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The research was financed from a student research grant of University of Lodz, which was received by Karol Bukowski. The research was also supported by a statutory research admitted for the Department of Molecular Genetics of University of Lodz and the Department of Biophysics of Environmental Pollution of University of Lodz.
