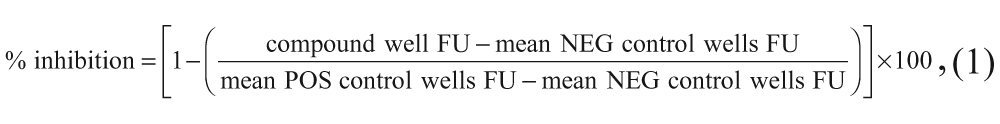Abstract
DNA methylation is an important epigenetic regulator of gene expression. Abnormalities in DNA methylation patterns have been associated with various developmental and proliferative diseases, particularly cancer. Targeting DNA methyltransferases (DNMTs) represents a promising strategy for the treatment of such diseases. Current DNMT inhibitors suffer important drawbacks with respect to their efficacy, specificity, and toxicity. In this study, we have set up a robust in vitro bacterial M.SssI DNMT activity assay to systematically screen a collection of 26 240 compounds that were predicted to compete with the S-adenosyl-L-methionine (SAM) substrate of DNMT. This resulted in the identification of a novel set of structurally distinct inhibitors of M.SssI DNMT activity. Although molecular docking studies using an M.SssI homology model suggest that these compounds might compete with SAM binding, mode of activity (MoA) assays are still needed to confirm this hypothesis. Our set of novel M.SssI DNMT inhibitors, once confirmed in an orthogonal DNMT assay, may thus serve as a starting point to identify and characterize suitable lead candidates for further drug optimization.
Introduction
DNA methylation represents a crucial mechanism for the epigenetic control of gene expression and plays a pivotal role in establishing and maintaining gene expression patterns in both normal and malignant cells. 1 In mammals, DNA methylation is found at the C5 position of cytosine residues, predominantly in the context of CpG dinucleotide sequences. Hypermethylation of CpG islands, defined as discrete ~1-kb regions with high CpG density that occur in approximately 70% of mammalian gene promoters, leads to strong gene silencing that is inherited by daughter cells upon division. 2 DNA methylation is involved in various cellular processes, including parental imprinting, X-chromosome inactivation, suppression of retrotransposon and repetitive elements, and silencing of pluripotency and tissue-specific genes. In addition, aberrant DNA methylation patterns have been associated with various developmental and proliferative diseases. 1 For example, erroneous promoter hypermethylation, and thereby silencing, of tumor suppressor genes is considered one of the hallmarks of cancer. By targeting genes involved in cell cycle regulation (e.g., CDKN2a, CDKN2b, and RB1), DNA repair (e.g., BRCA1, MGMT), and apoptosis (e.g., DAPK), aberrant DNA hypermethylation provides a growth advantage either by increased proliferation or increased resistance to apoptotic factors and thereby plays an important role in cancer development and maintenance. 3
DNA methylation patterns need to be actively maintained upon cell division and are therefore inherently reversible. 2 Thus, pharmacological targeting of the DNA methylation process represents a promising strategy to reactivate (aberrantly) hypermethylated genes in, among others, the treatment of cancer. 3 In addition, such an approach may help to further enhance our understanding of the biological functions of DNA methylation and may support, for example, somatic cell reprogramming. 4
DNA methylation patterns are established and maintained by the coordinated action of DNA methyltransferases (DNMTs), which use S-adenosyl-L-methionine (SAM) as their methyl donor. 2 At present, four active mammalian DNMTs have been identified, including the two de novo methyltransferases DNMT3a and DNMT3b and the maintenance methyltransferase DNMT1. 2 Although structurally similar to the other DNMTs, the role of the fourth DNMT, DNMT2, is less well understood, and it appears to function as a tRNA methyltransferase rather than a DNA methyltransferase. 5 Among the mammalian DNMT family, DNMT1 is the most abundant and is considered the major contributor to cellular DNMT activity. It shows a preference for hemi-methylated DNA substrates and associates itself with the replication machinery, consistent with a primary function in the duplication, and thus maintenance, of DNA methylation patterns during replication. 2 DNMT3a and DNMT3B can associate with specific transcription factors and mediate de novo methylation during early embryogenesis, whereas their activity is reduced upon embryonic stem cell differentiation. 2 DNMT knockout studies suggest a considerable degree of cooperation between these individual family members, indicating an additional level of complexity to their function. 2
The structure of DNMT1, DNMT3A, and DNMT3B consists of an N-terminal regulatory domain and a C-terminal catalytic domain harboring the active center of the enzyme. 2 The catalytic domain is structurally conserved between eukaryotic and prokaryotic DNA cytosine C5 methyltransferases. More particularly, the catalytic domains of eukaryotic and prokaryotic DNMTs share 10 conserved amino acid motifs and have a common core structure, known as the “SAM-dependent methyltransferase fold.” 2 This domain is involved in both cofactor binding (motifs I, II, and X) and catalysis (motifs IV, VI, and VIII). A variable region located between motifs VIII and IX, known as the target recognition domain, is involved in DNA recognition and specificity. 2
Along with the structural conservation of their catalytic domain, the mechanism of cytosine C5 methylation is also considered to be similar for all C5 DNA methyltransferases. 2 First, the methyltransferase forms a complex with DNA and SAM, whereby the target cytosine is flipped out from the DNA base-pairing position. The thiol group of the catalytic cysteine residue, located in a proline-cysteine-glutamine (PCQ) motif (motif IV), then performs a nucleophilic attack on the 6-position of the target cytosine, resulting in the generation of a covalent DNA-enzyme intermediate with nucleophilic properties at the cytosine 5-position. This activated intermediate accepts a methyl group from SAM, forming S-adenosyl-L-homocysteine (SAH) and a 5-methyl covalent adduct, which is subsequently resolved by deprotonation at the 5-position to generate the methylated cytosine and the free enzyme.
DNMTs can be inhibited effectively by nucleoside analogues, such as 5-azacytidine (5AC), 5-aza-2′-deoxycytidine (5AdC), 5-fluoro-2′-deoxycytidine, and the cytidine analog zebularine, which all act as mechanism-based inhibitors that are incorporated into the DNA, where they cause covalent trapping and subsequent depletion or degradation of active DNMT enzymes. 6 However, this inhibitory mechanism is inherently cytotoxic due to the formation of DNMT-DNA adducts. 6 It is therefore important to identify nonnucleoside compounds that are able to directly block DNMTs in their activity. To date, only a small number of compounds, including hydralazine, procainamide, (–)-epigallocathechin-3-gallate (EGCG), curcumin, and RG108, have been shown to directly target DNMTs and to be able to reactivate hypermethylated genes, although in most cases far less effectively than 5AdC.7–9
Thus, currently only a limited number of chemical scaffolds are available that could be used for the development of DNMT inhibitors with increased efficacy and specificity and reduced toxicity. Therefore, expanding this set of chemical scaffolds will improve the probability of developing a more successful drug. On the basis of these considerations, we set out in this study to systematically screen for novel DNMT inhibitors. To this end, we set up a robust in vitro assay for DNMT activity, using the bacterial CpG DNA methyltransferase from Spiroplasma sp. (strain MQ-1; M.SssI) as a model system. This enzyme has the same specificity for CpG dinucleotides as the mammalian DNMTs and has previously been used as an experimental tool for studying various aspects of DNA methylation in higher eukaryotes. 10 Given the structural similarity between the cofactors SAM and adenosine triphosphate (ATP), we reasoned that screening a library of compounds predicted to bind in the ATP binding pocket of protein kinases should give us a good chance of identifying compounds that could compete with SAM binding to DNMTs. Our screen resulted in the identification of a novel set of small-molecule M.SssI CpG methyltransferase inhibitors.
Materials and Methods
DNMT Activity Assay
The DNMT activity assay described in this study was based on the commercially available Methyltransferase HT activity kit (Enzo Life Sciences, Farmingdale, NY), which detects the SAM-dependent methyltransferase reaction product SAH through a coupled enzyme reaction that converts SAH to a fluorescent product. This assay consists of two parts: an initial methyltransferase reaction, in which M.SssI uses SAM as a methyl donor to methylate a DNA substrate, thereby producing SAH, followed by a detection reaction, in which SAH is enzymatically processed into a fluorescent product. We selected the commonly used synthetic oligonucleotide poly(2′-deoxyinosinic-2′-deoxycytidylic acid) (poly(dIdC)) as a DNA substrate since it has been reported to be a good substrate for DNMTs. 11
The DNMT activity assay was performed in small-volume 384-well black FLUOTRAC 200 HiBase microplates (Greiner Bio-One, Monroe, NC). The initial methyltransferase reaction contained 1 U M.SssI (New England Biolabs, Ipswich, MA), 5 µM SAM (Enzo Life Sciences), and 0.004 U poly(dIdC) (Sigma-Aldrich, St. Louis, MO) in a total volume of 10 µL assay buffer (10 mM Tris-HCl [pH 7.9], 50 mM NaCl, and 10 mM MgCl2) with 0.5% DMSO and was carried out for 3 h at 37 °C. For the subsequent detection reaction, a total of 5 µL assay buffer containing 50× diluted “detection mix” and 20× diluted “methyltransferase reaction mix” components from the Methyltransferase HT activity kit (Enzo Life Sciences) was added, followed by another 3-h incubation at room temperature. Finally, fluorescent emission at 520 nm (355 nm excitation) was measured using an EnVision 2100 multilabel plate reader (PerkinElmer, Waltham, MA). Resulting fluorescence data were analyzed using Prism 5.01 (GraphPad Software, La Jolla, CA).
Assay Optimization
We used an M.SssI titration series in the activity assay described above to determine the optimal M.SssI concentration for the assay. In addition, we varied the concentration of SAM and the incubation time of the initial methyltransferase reaction to estimate the apparent Km for SAM under our assay conditions.
SAH standard curves were generated by adding SAH titration series (ranging from 0–50 µM) directly to the reaction part of the assay, and the resulting fluorescence values were used to determine the amount of SAH produced by M.SssI in the complete assay.
High-Throughput Screening
A subset of 26 240 compounds that previously have been predicted to have characteristic features of ATP-competitive protein kinase inhibitors and, in addition, fulfilled drug-likeness criteria was selected from the full Merck library for the high-throughput screen (HTS).
Compounds were tested at a 10-µM concentration in the initial methyltransferase reaction. Each microplate was set up with 16 wells each, for both negative (“NEG,” without poly(dIdC) or compound) and positive (“POS,” with poly(dIdC), without compound) controls, as well as 320 wells for compound testing.
Assay performance was assessed by calculating Z′ values, a commonly used parameter to describe assay quality, using the NEG and POS controls on each plate. 12
The percent inhibition of enzyme activity by each compound was calculated using equation (1), based on the NEG and POS control wells of the plate the compound was in.
whereby FU (fluorescence units) refers to the measured fluorescence intensity values.
Plate maps containing % inhibition results for each well were visually inspected, and aberrant results (i.e., whereby a complete row or column showed outlier values, likely due to a technical error) were removed (referred to as “quality control”).
Deselection Assay
To deselect hits that inhibited the secondary detection reaction instead of the primary methyltransferase reaction in the DNMT activity assay, we performed only the detection part of the assay in the presence of 10 µM compound and 2.5 µM SAH in a total volume of 15 µL. Similar to the HTS screen, POS (with SAH, without compound) and NEG (without SAH and compound) control wells were used to determine Z′ values and to calculate the % inhibition of SAH processing by each compound.
Results and Discussion
Assay Design and Optimization
To identify new DNMT inhibitors, we set up a homogeneous assay to allow for robust HTS of bacterial M.SssI DNMT activity based on the enzymatic conversion of the SAM-dependent methyltransferase reaction product SAH into a fluorescent signal (
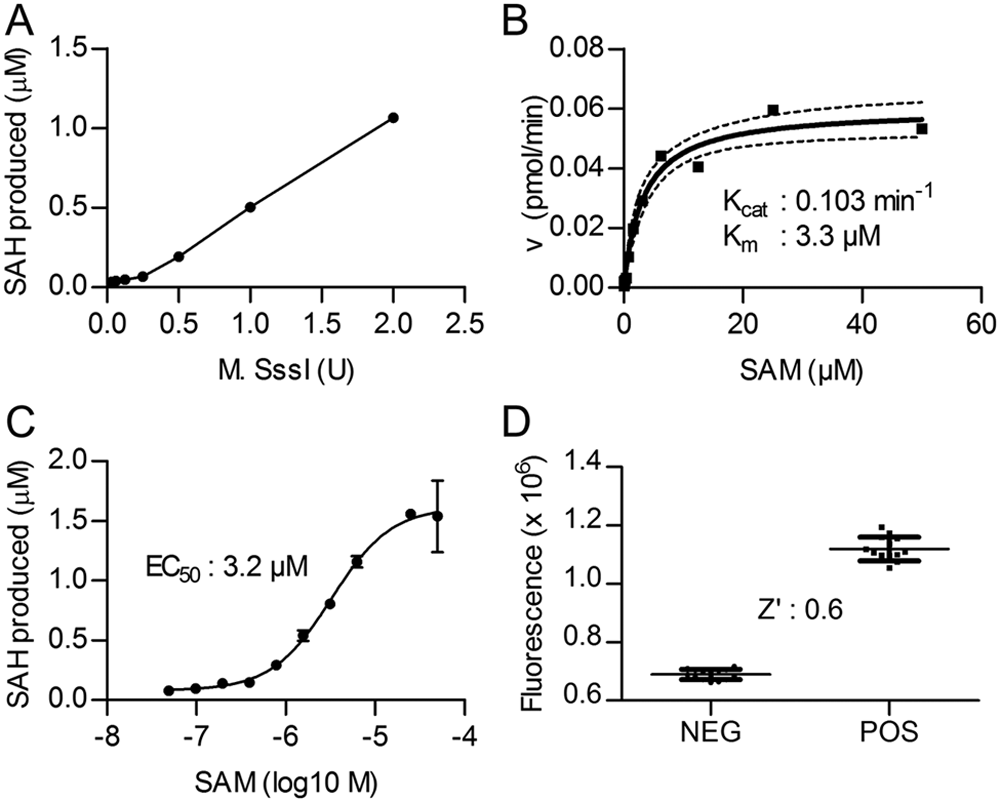
DNA methyltransferase (DNMT) activity assay: optimization of the DNA methyltransferase reaction. (
To enable the identification of SAM-competitive compounds, we decided to design our assay to be particularly sensitive to changes in SAM availability by using a SAM concentration near its apparent Km value. Poly(dIdC) substrate, on the other hand, was used in an excess concentration (0.0004 U/µL, which corresponds to ~30 µM methylation sites). To estimate the apparent Km value of SAM for our assay, initial reaction rates were measured using a constant amount (0.004 U) of poly(dIdC) and (1 U) M.SssI, in combination with variable SAM concentrations. The plot of these reaction rates against SAM concentration was fitted to the Michaelis-Menten equation ( Fig. 1B ), resulting in an apparent Km for SAM of 3.3 µM (95% confidence interval, 1.8–4.7 µM), which agrees well with previously reported values.11,13 In addition, a SAM dose-response curve (DRC) of the relative fluorescence intensity values obtained for the 3-h M.SssI incubation time point ( Fig. 1C ) demonstrated an EC50 value of 3.2 µM. On the basis of these results, we selected 5 µM as the SAM concentration for the high-throughput screen since this value approximates the estimated Km, thereby providing a “balanced” condition with an equal population of free enzyme and SAM-enzyme complex available for inhibitor interactions and within the area of the SAM DRC with the highest dynamic range.
Using these final assay conditions, we subsequently established the assay signal in the presence and absence of poly(dIdC), representing the POS and NEG controls, respectively, that would be present on each 384-well plate during the high-throughput screen. The data presented in Figure 1D show that the final assay has an acceptable Z′ value of 0.6.
Noteworthy here is that alternative (DNA) methyltransferase activity assays equally suitable for high-throughput applications have been described previously by others, although they have not yet been applied for this purpose. These include variants to the assay described in this study, in which enzymatic conversion of SAH leads to the formation of fluorescent products through alternative pathways.11,14 Another strategy makes use of internally quenched fluorescent hairpin DNA substrates, which, when methylated by DNMT, are cleaved by a methylation-sensitive restriction enzyme, resulting in an increase of fluorescence. 15 The present study describes, to the best of our knowledge, the first application of such an in vitro biochemical DNMT activity assay to perform a systematic high-throughput screen of a compound library for DNA methyltransferase inhibitors.
High-Throughput Screen and Activity Confirmation
High-throughput screening was performed on a library of 26 240 compounds, each at a 10-µM concentration, as described in Materials and Methods (
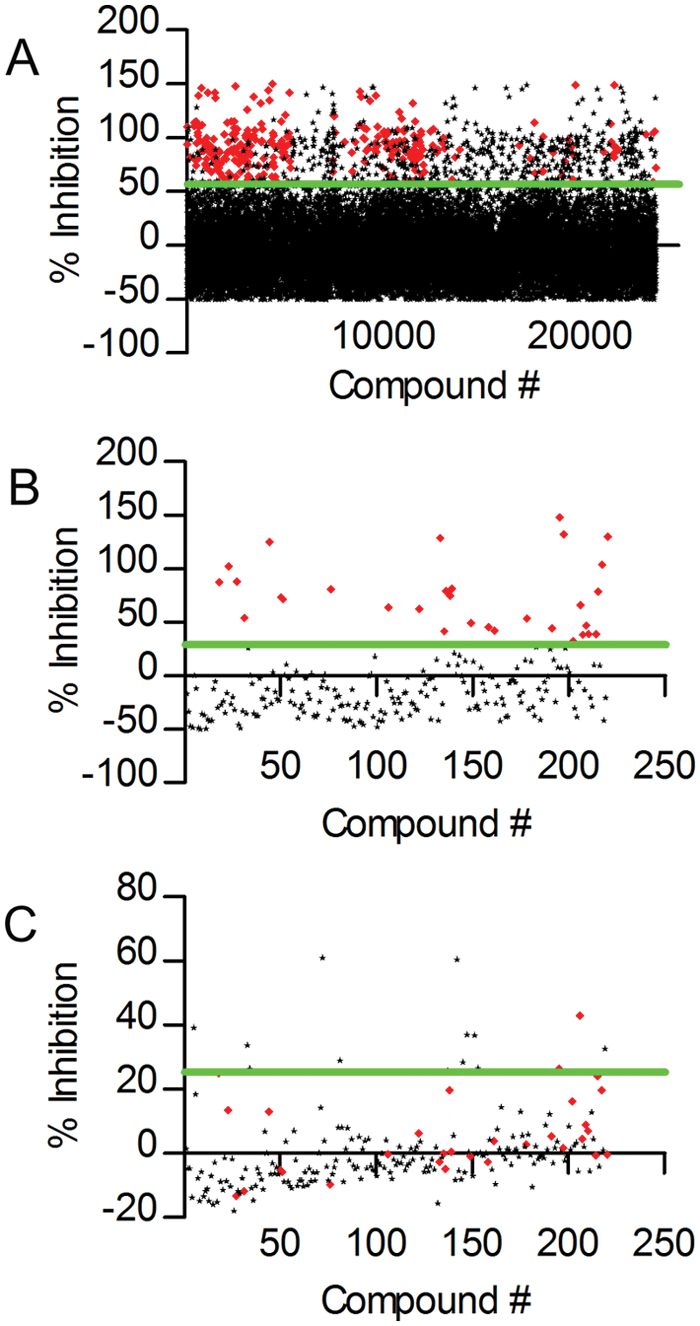
High-throughput screen and hit confirmation. (
Subsequently, this selection of compounds was tested in triplicate in two different assays: a confirmation test, with the same assay conditions and quality control process as the high-throughput screen, and a deselection assay to rule out compounds inhibiting the detection reaction as opposed to the initial methyltransferase reaction. In the deselection assay, SAH was directly added to the detection reaction (so in the absence of an initial M.SssI methyltransferase reaction) in the presence of these compounds to determine their effects specifically on the secondary detection reaction. Figure 2B shows the distribution of activities of the selected compounds in the confirmation test. A threshold for active confirmation was now set at the 10% trimmed mean plus 2 × SD of this distribution, resulting in 31 actives (14%; Fig. 2B , red dots). Figure 2C represents the activities of these compounds in the deselection assay, whereby a threshold for interference with the detection reaction was set at the mean plus 2 × SD. This shows that 14 compounds (6.3% of the screen actives) have an apparent inhibitory effect on the secondary detection reaction and are therefore considered to be nonspecific for the M.SssI methyltransferase reaction. Of the 31 confirmed actives, 29 compounds were inactive in the deselection assay, resulting in 29 actives specific for the initial methyltransferase reaction (13.2% of the screen actives).
Dose-Response Curves of Confirmed Actives
Finally, we set out to determine the IC50 values of these resulting 29 actives to evaluate their potencies. To this end, we constructed triplicate DRCs of compound dilution series ranging from 20 µM to 0.6 nM for both our M.SssI activity assay and the deselection assay. On the basis of these assays, we selected nine compounds that showed an IC50 <10 µM on M.SssI activity and an IC50 value of at least 5-fold higher than that for M.SssI in the deselection assay. DRCs for these compounds are shown in Figure 3 , whereby the corresponding IC50, Hill slope, min and max values, and chemical structures are listed in Table 1 . These compounds represent a set of novel, previously unidentified molecular structures with the potential to inhibit M.SssI DNMT activity and thus provide a starting point for the identification and characterization of suitable lead candidates for further drug optimization.
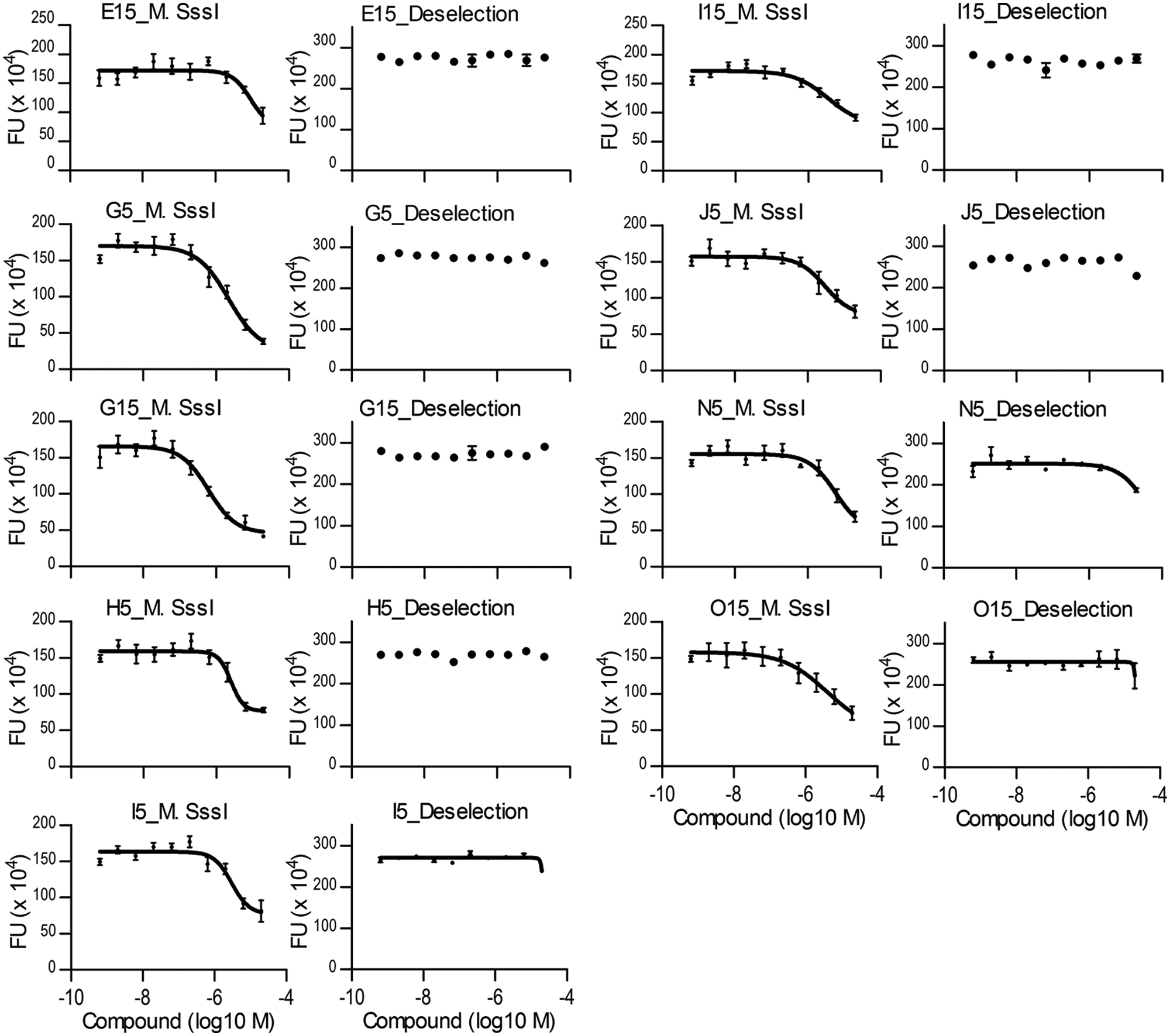
Dose-response curves (DRCs) for inhibition of DNA methyltransferase (DNMT) activity. DRCs of nine compounds with an IC50 <10 µM in the M.SssI activity assay and an IC50 >20 µM in the deselection assay. Chemical structures and IC50 values are indicated in Table 1 . DRCs were generated by measuring fluorescence intensity values (FU; fluorescence units) of 10-point compound dilution series (ranging from 20 µM to 0.6 nM) in the DNMT activity assay (left) and the deselection assay (right). Data were fitted to a “nonlinear log(agonist) vs. response – variable slope” equation to determine IC50 values.
Chemical Structure, Hill Slope, and Min, Max, and IC50 Values of Confirmed Hits
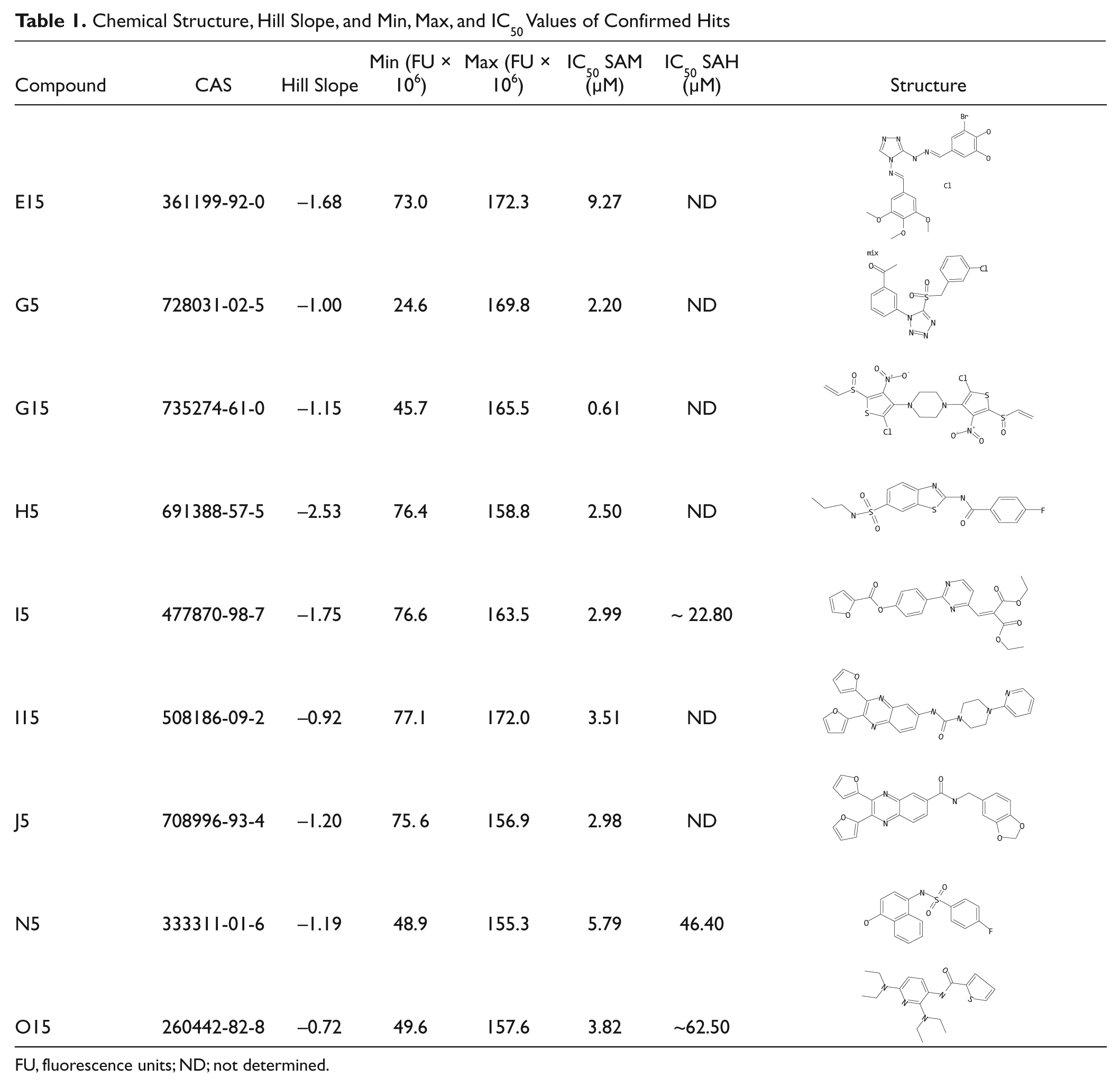
FU, fluorescence units; ND; not determined.
However, we emphasize that these compounds are still in a very early stage of development. Although they demonstrate M.SssI reaction-specific, dose-related activity in our screening assay, this activity will first need to be confirmed in an orthogonal DNMT activity assay. In addition, it will be important to determine the effects of these compounds also on the activity of mammalian DNMTs. Preliminary docking studies of our M.SssI inhibitors into the SAM binding pocket of human DNMT1 and bacterial M.SssI do indicate that they might have similar binding modes in both enzymes (data not shown), although this remains to be experimentally confirmed.
Further in vitro analysis is also required to gain understanding of the mode of action of these compounds. The preliminary molecular modeling studies mentioned above suggest that some of the identified compounds might bind M.SssI in a SAM-competitive mode (
In addition, since we screened a library of compounds that were selected based on their resemblance to ATP-competitive kinase inhibitors, it will be important to assess the selectivity of the identified M.SssI inhibitors against the human kinome.
Thus, future studies are needed to further examine the modes of inhibition, selectivity, potency, and toxicity of the novel set of M.SssI methyltransferase reaction inhibitors identified in this study, and they should focus on improving their inhibitory properties and drug-likeness characteristics. This might eventually result in the development of an improved DNMT inhibitory drug that does not suffer from the important drawbacks of currently available inhibitors, potentially leading to an improved therapy against the aberrant DNA methylation patterning that occurs in various developmental and proliferative diseases, particularly cancer. 1
Footnotes
Acknowledgements
We thank Prof. Gromova for providing us with their homology model of M.SssI CpG methyltransferase. In addition, we thank Bas van de Kar, Els van Doornmalen, Maaike van Hoek, Monique van Amstel, and Simone Ruygrok for their assistance in the HTS.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by a Casimir grant from NWO (project number 018.002.035) and by Merck Sharp & Dohme (Oss, the Netherlands).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.