Abstract
Chronic hepatitis B virus (HBV) infection is an independent risk factor for the development of hepatocellular carcinoma (HCC). The HBV HBx gene is frequently identified as an integrant in the chromosomal DNA of patients with HCC. HBx encodes the X protein (HBx), a putative viral oncoprotein that affects transcriptional regulation of several cellular genes. Therefore, HBx may be an ideal target to impede the progression of HBV infection–related HCC. In this study, integrated HBx was transcriptionally downregulated using an artificial transcription factor (ATF). Two three-fingered Cys2-His2 zinc finger (ZF) motifs that specifically recognized two 9-bp DNA sequences regulating HBx expression were identified from a phage-display library. The ZF domains were linked into a six-fingered protein that specified an 18-bp DNA target in the Enhancer I region upstream of HBx. This DNA-binding domain was fused with a Krüppel-associated box (KRAB) transcriptional repression domain to produce an ATF designed to downregulate HBx integrated into the Hep3B HCC cell line. The ATF significantly repressed HBx in a luciferase reporter assay. Stably expressing the ATF in Hep3B cells resulted in significant growth arrest, whereas stably expressing the ATF in an HCC cell line lacking integrated HBx (HepG2) had virtually no effect. The targeted downregulation of integrated HBx is a promising novel approach to inhibiting the progression of HBV infection–related HCC.
Introduction
Hepatocellular carcinoma (HCC) is one of the most common cancers 1 and is the third most common cause of death from cancer worldwide. 2 Chronic hepatitis B virus (HBV) infection is a strong independent risk factor for HCC, 3 increasing the likelihood of HCC 5- to 15-fold compared with the general population. 4 Approximately 85% to 90% of HBV-positive HCC cases involve integration of HBV DNA into chromosomes of infected hepatocytes, the initial sites of natural acute infections.5,6 Specifically, the HBV HBx gene is the most frequently detected integrant in the chromosomal DNA of patients with HCC. 2 A study using an HBx transgenic CD1 mouse model reported that 80% to 91% of males and 60% to 67% of females developed HCC, with most males dying from clear cell HCC at 11 to 15 months of age and most females dying from clear cell HCC at 17 to 21 months of age. 7 In contrast, wild-type CD1 mice did not develop spontaneous liver tumors. The HBx protein contributes to viral genome replication and transcriptionally regulates host genes involved in oncogenesis, cell proliferation, cell cycle regulation, protein degradation, and DNA repair.3,5,7 During late stages of tumor progression, HBx drives cellular genes associated with metastasis and angiogenesis. 8 As a “viral oncoprotein” that directs HBV-mediated hepatocarcinogenesis, HBx is an ideal target for blocking the progression of HBV infection–related HCC. Previous studies have evaluated antisense agents such as hammerhead ribozymes, 9 antisense oligodeoxynucleotides, 10 and short interference RNA (siRNA)8,11 to repress HBx gene expression in HCC cell lines and in HBx transgenic mice models. Although these antisense agents demonstrated significant HBx inhibition, further study and clinical application are restricted by their short half-lives and potent “off-target” effects. Potential small molecular inhibitors of HBx have been screened virtually for active site targeting using molecular docking methodologies, 12 highlighting HBx as a promising drug target.
Cys2-His2 (C2H2) zinc finger (ZF)–based artificial transcription factors (ATFs) have been used successfully to alter the expression of specific endogenous genes.13,14 Native transcription factors (TFs) typically consist of a DNA-binding domain and an effector domain. The DNA-binding domain functions to localize the TF to a specific site on chromosomal DNA, whereas the effector domain recruits other factors to the site, resulting in transcriptional activation or repression. 13 Using this basic structural blueprint, ATFs can be constructed by combining an effector domain with a specifically targeted DNA-binding domain selected from a ZF module library.13,14 In this study, a novel ATF composed of a six-fingered ZF DNA-binding domain and a Krüppel-associated box (KRAB) repression domain was engineered to specifically downregulate integrated HBx in the human HCC cell line, Hep3B.
Materials and Methods
Construction and Screening of ZF Libraries
Two ZF phage-display libraries, Lib12 and Lib23, each of which contained half of the three-fingered C2H2 motif of Zif268, were constructed according to the “bipartite complementary” strategy.15–17 Briefly, ZF genes were randomized at their α-helical DNA-contacting residues across each library’s portion of the Zif268 scaffold by synthesizing and annealing complementary oligonucleotides (Invitrogen, Carlsbad, CA). The synthesized fragments were inserted into the modified vector, pHB-1HScFv, and were expressed as fusions to the phage minor coat protein, pIII, as described previously.
17
A continuous 20–base pair (bp) double-stranded DNA fragment (5-
Scanning Mutagenesis Enzyme-Linked Immunosorbent Binding Assay
Each site in the U target and D target was sequentially mutated into its transitional base (A↔G or C↔T) to generate a series of oligonucleotide targets. Both strands of the DNA targets were synthesized; for each target, one strand was biotinylated at the 5′-end. The binding abilities of single clones against their wild-type targets and the series of mutated targets were evaluated by an enzyme-liked immunosorbent assay (ELISA), as previously described. 16 Briefly, double-stranded, biotin-labeled DNA targets generated by annealing two synthesized fragments were added to 96-well plates coated with streptavidin (Roche Applied Science), and phage displaying ZF proteins to be tested then were added. Binding was detected using a horseradish peroxidase (HRP)–conjugated anti-M13 IgG antibody (GE Healthcare, Buckinghamshire, UK).
Construction of Three-fingered and Six-fingered ATFs
The six-fingered protein, U5D13, was constructed by linking the three-fingered proteins D13 and U5 with the conserved TGEKP linker peptide. 18 ZF motifs U5, D13, and U5D13, functioning as putative DNA-binding domains, were fused with KRAB repression domains 30 to generate three-fingered and six-fingered ATFs. Each ATF then was cloned into pcDNA3.1(+) (Invitrogen) at NheI/XhoI sites in frame with a C-terminal FLAG tag and an N-terminal nuclear localization signal as reported previously. 19
Cell Culture and Transfection
293T cells were maintained in minimum essential medium (MEM; Invitrogen) supplemented with 10% (v/v) fetal bovine serum (FBS). Hep3B cells were maintained in MEM supplemented with 10% FBS, 2 mM L-glutamine (L-Gln), 1 mM sodium pyruvate, and 0.1 mM nonessential amino acids. HepG2 cells were cultured in RPMI 1640 medium supplemented with 10% FBS. All culture media were supplemented with penicillin (100 U/mL) and streptomycin (100 µg/mL) in a humidified incubator at 37 °C and 5% CO2. Transient transfections were performed using the standard calcium phosphate–DNA co-precipitation procedure. 20 Hep3B cells were plated in six-well plates at a density of 1 million cells per well. Increasing amounts (0, 1, 2 and 3 µg) of ATF composed of ZF U5D13 and the KRAB transcription repression domain were transfected into each well using the calcium-phosphate precipitation protocol. Stable transfectants were selected with 0.7 (for HepG2) or 0.5 (for Hep3B) mg/mL G418 48 h after transfection. Following selection, both cell lines were maintained in 0.3 mg/mL G418.
Luciferase Assay
Luciferase reporter plasmid pUDtarget-SV40-Luc was constructed by cloning fragment 5′-GCCAAGTGTT-TGCTGACGCA-3′ (HBV subtype adw, 1175–1194) into the pGL3-Control vector (Promega, Madison, WI) at MluI/SmaI sites. Another luciferase reporter plasmid, pHBX-promoter-Luc, was constructed by inserting the upstream Enhancer I (EnhI)/HBx promoter region (HBV subtype adw, 1155–1355) into the pGL3-basic vector (Promega) at XhoI/HindIII sites. 21 The Renilla luciferase-expressing vector, pRL-TK (Promega), was used as an internal control to normalize transfection efficiency. ATF expression vectors, luciferase reporter vectors, and pRL-TK were cotransfected into 293T cells seeded into 24-well plates at a density of 50 000 cells per well. Cell lysates were harvested 42 h after transfection, and dual-luciferase reporter assays were performed using a GloMax 96 Microplate Luminometer (Promega).
Real-time PCR
Total RNA was extracted from harvested cells using an RNeasy kit (Qiagen, Mainz, Germany), and first-strand cDNA was synthesized using the Superscript III First-Strand Synthesis system (Invitrogen). Reverse transcriptase (RT)–PCR was performed as previously described, 22 using HBx primers (forward, 5′-gctgctaggctgtgctgc-3′; reverse, 5′-atgcctcaaggtcggtcgt-3′). β-Actin primers (forward, 5′-caaccgcgagaagatgac-3′; reverse, 5′-agggtacatggtggtgcc-3′) were used as an internal control. Real-time quantitative PCR (qPCR) was performed using SYBR Green Master Mix (Applied Biosystems, Foster City, CA) on an ABI PRISM 7000 Sequence Detection System (Applied Biosystems) with HBx primers (forward, 5′-accgtgtgcacttcgcttc-3′; reverse, 5′-ttcacggtggtctccatgc-3′). GAPDH (forward, 5′-ccatgttcgtcatgggtgtga-3′; reverse, 5′-catggactgtggtcatgagt-3′) was amplified as an internal control. Each sample was analyzed in triplicate.
MTT Assay
Cells were seeded into 96-well plates at densities of 5 × 103 cells per well. Fresh culture medium (100 µL) containing 20% methyl thiazol tetrazolium (MTT) stock (5 mg/mL) was added to each well at indicated time points. Cells then were incubated for 4 h, and medium was replaced with 100 µL DMSO. Plates were agitated for 10 min at room temperature, and absorbances were measured at 570 nm. All samples were assayed in triplicate.
Statistical Analysis
Student t test or one-way analysis of variance (one-way ANOVA) and post hoc multiple comparisons were performed using SPSS v.10.0 statistical software (SPSS, Inc., an IBM Company, Chicago, IL). All data were presented as mean ± standard deviation (SD). Statistical significance was assigned at p < 0.05.
Results
Design of ATFs Targeted to HBx Gene Integrated in Hep3B Cell Genome
Phage-display ZF libraries, Lib12 and Lib23, were constructed based on the backbone of the three-fingered C2H2 ZF domain of Zif268. C2H2 ZFs are one of the most abundant classes of DNA-binding domains in human TFs, 14 and they provide a flexible and versatile tool for targeting ATFs to desired promoter regions. Each C2H2 ZF comprises approximately 30 amino acids and recognizes a 3-bp DNA sequence with its α-helix via specific base–side chain interactions. 23 ZF libraries may be generated by randomizing amino acids in the α-helix of a seed ZF motif to obtain novel ZFs that bind specifically to desired DNA sequences. To recognize longer DNA sequences, C2H2 ZFs may be joined in tandem arrays by consensus linkers, such that a three-fingered protein could bind a 9-bp DNA target.
The design of ATFs targeted to the HBx integrant in the Hep3B genome is illustrated in Figure 1 . Hep3B is a well-differentiated human hepatoma cell line established from a liver tumor biopsy. 24 This cell line is known to contain integrated HBV (subtype adw) sequences, including intact HBx promoter and upstream EnhI regions, which are regarded as the complete HBx regulatory region. 25 Hep3B cells synthesize detectable HBx transcripts, providing an ideal experimental model to evaluate strategies of HBx downregulation. Using this model, Cheng et al. 8 reported that siRNA directed against HBx suppresses Hep3B cell growth and induces apoptosis. In our study, two 9-bp sequences in EnhI (HBV subtype adw, 1175–1194) were used to screen three-fingered phage-display libraries for ATF-mediated HBx downregulation. The target DNA region consisted of a U target (5-GCCAAGTGT-3) and a D target (5-GCTGACGCA-3), with an intervening dinucleotide (TT). The U and D targets were used to screen Lib12 and Lib23, respectively ( Fig. 1 ). The three-fingered ZFs were joined by a linker to create a six-fingered motif that could potentially bind both the U and D targets. The DNA-binding region was fused with a KRAB transcriptional repression domain to produce an ATF designed to bind and downregulate HBx expression.
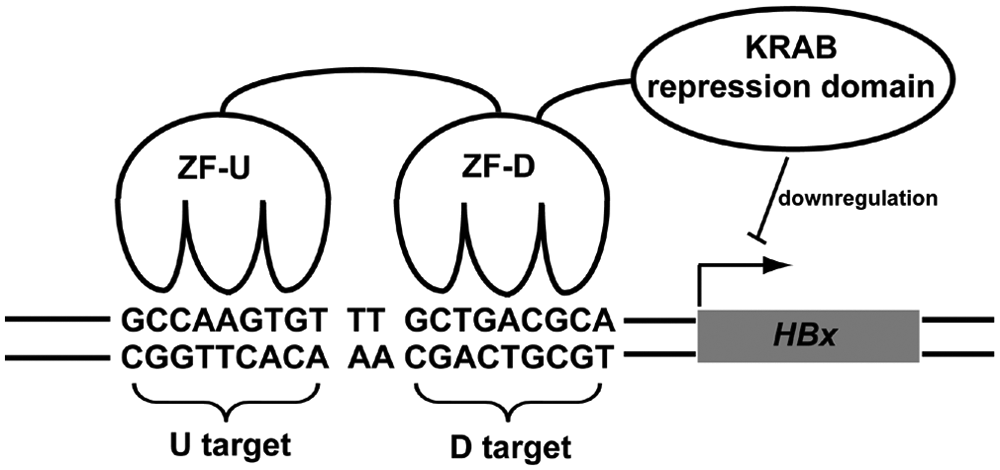
Artificial transcription factor (ATF) targeting to HBx integrant of Hep3B cells. ATFs are composed of two domains: the DNA-binding domain consists of six Cys2-His2 zinc fingers and was generated by linking two three-fingered zinc finger proteins (ZF-D and ZF-U) acquired from a screen of phage-display libraries; the effector domain is a natural transcriptional repressor called the Krüppel-associated box (KRAB).
ZF Motifs Specifically Bind the HBx Promoter Region
According to the bipartite strategy, ZF libraries Lib12 and Lib23 are subject to selection in parallel. Four rounds of selection were carried out for each biotinylated target. The affinities of phage supernatants rescued from the eluate in each round of selection were validated by ELISA. We observed substantial enrichment of phage clones displaying high-affinity ZF motifs after the second round of selection at the U target ( Fig. 2a ) and the D target (data not shown). Phage pools from the parallel selection processes then were cloned and recombined to produce a small library for further selection at the 9-bp target of interest. Phage clones U5 and U37 demonstrated specificity to the U target, and clones D13 and D46 were specific to the D target ( Fig. 2b ). The binding specificity of clones U5 and D13 was examined further using a scanning mutagenesis ELISA binding assay against the 9-bp target fragments containing a series of transition mutations. Clones U5 and D13 exhibited dramatically higher affinities for their own target sites than for the mutated targets, even though only single transitional mutations were made ( Fig. 2c , d ). Table 1 details the amino acid compositions of the randomized DNA-binding positions on the α-helices of wild-type three-fingered Zif268, U5, and D13. The backbones of phage-display libraries Lib12 and Lib23 are also given, with X indicating the randomization sites ( Table 1 ).
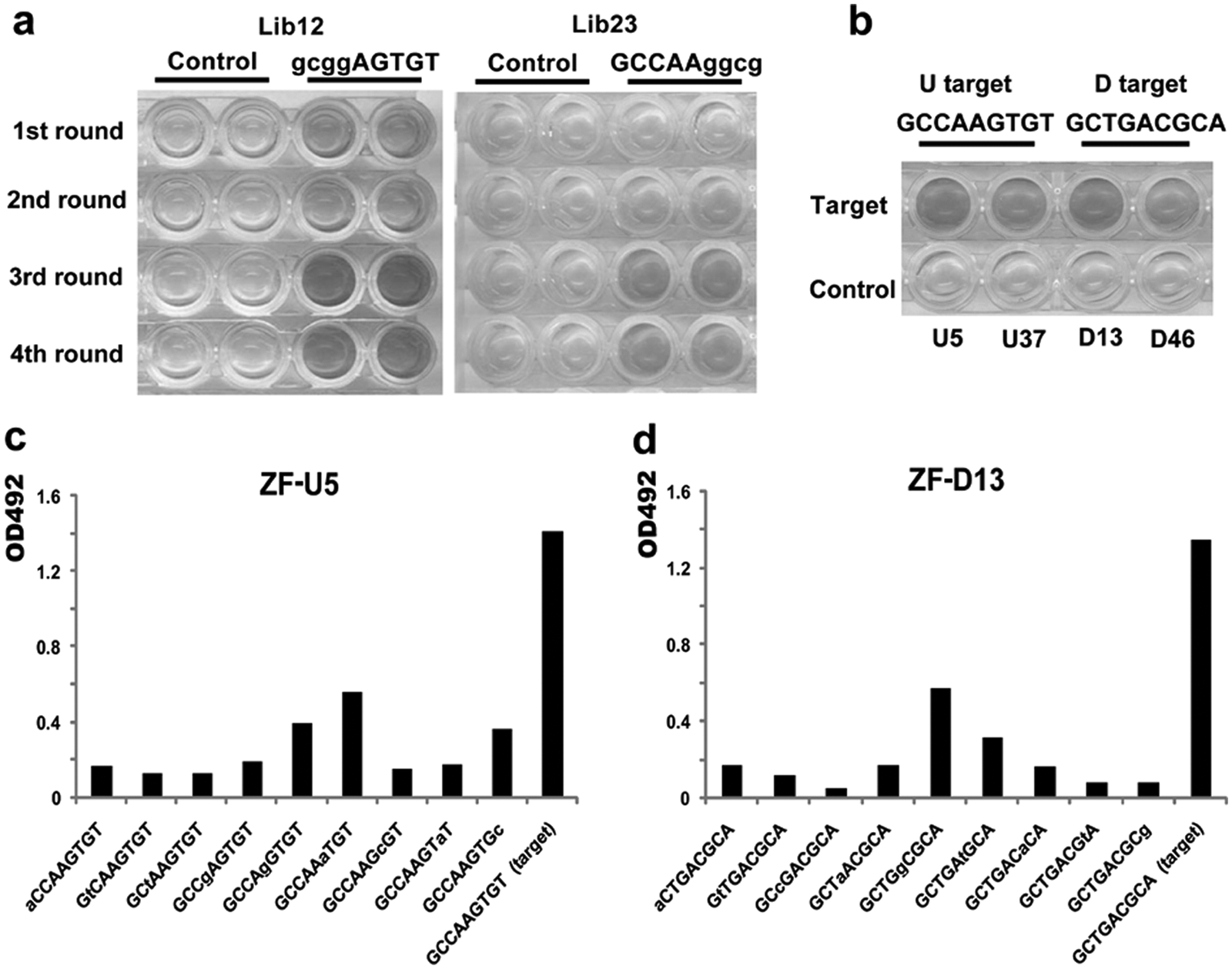
Selection of three-fingered proteins recognizing the Enhancer I region upstream of the HBx promoter. (
Amino Acid Sequences of α-Helical Positions −1 to 6 in Zinc Fingers U5 and D13.
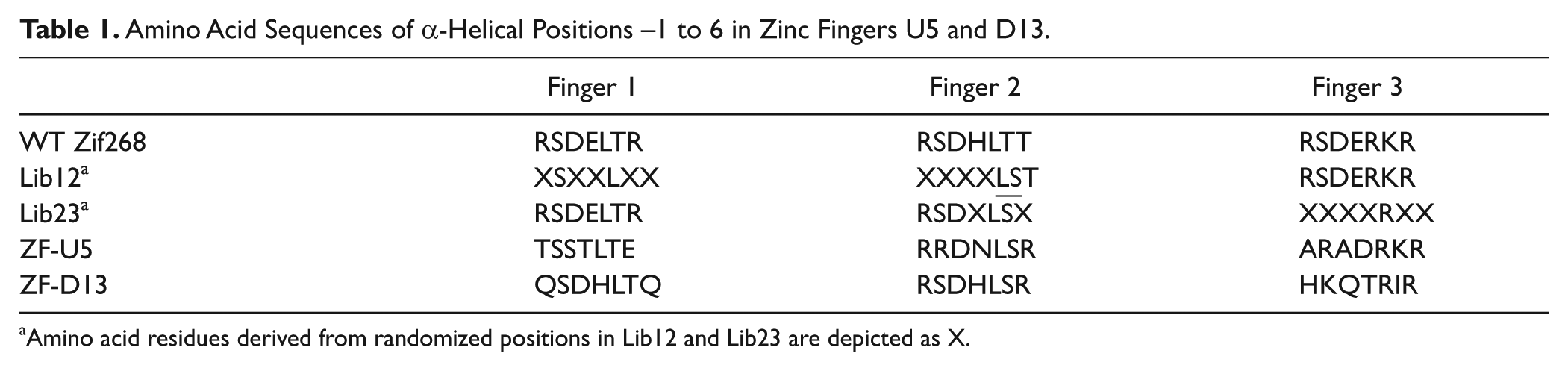
Amino acid residues derived from randomized positions in Lib12 and Lib23 are depicted as X.
ATFs Targeting to the HBx Promoter Downregulate the Reporter Gene
The three-fingered proteins, D13 and U5, were joined with a TGEKP linker peptide to generate the six-fingered protein, U5D13, which recognized the 18-bp combined target. ATFs were engineered by fusing U5, D13, or U5D13 with the KRAB effector domain. ZF(SV40)-KRAB, an ATF targeting the DNA sequence 5-GCAGAGGCC-3 and specifically repressing the SV40 promoter, 17 was used as a control. ATF expression vectors were cotransfected with the luciferase reporter vector and pRL-TK into 293T cells ( Fig. 3a ), and the relative luciferase activities were measured to infer the transcriptional repression effects of the candidate ATFs. The protein expression of each ATF was validated by Western blot, and protein levels were compared to ensure that the ATFs were expressed in similar quantities (data not shown).
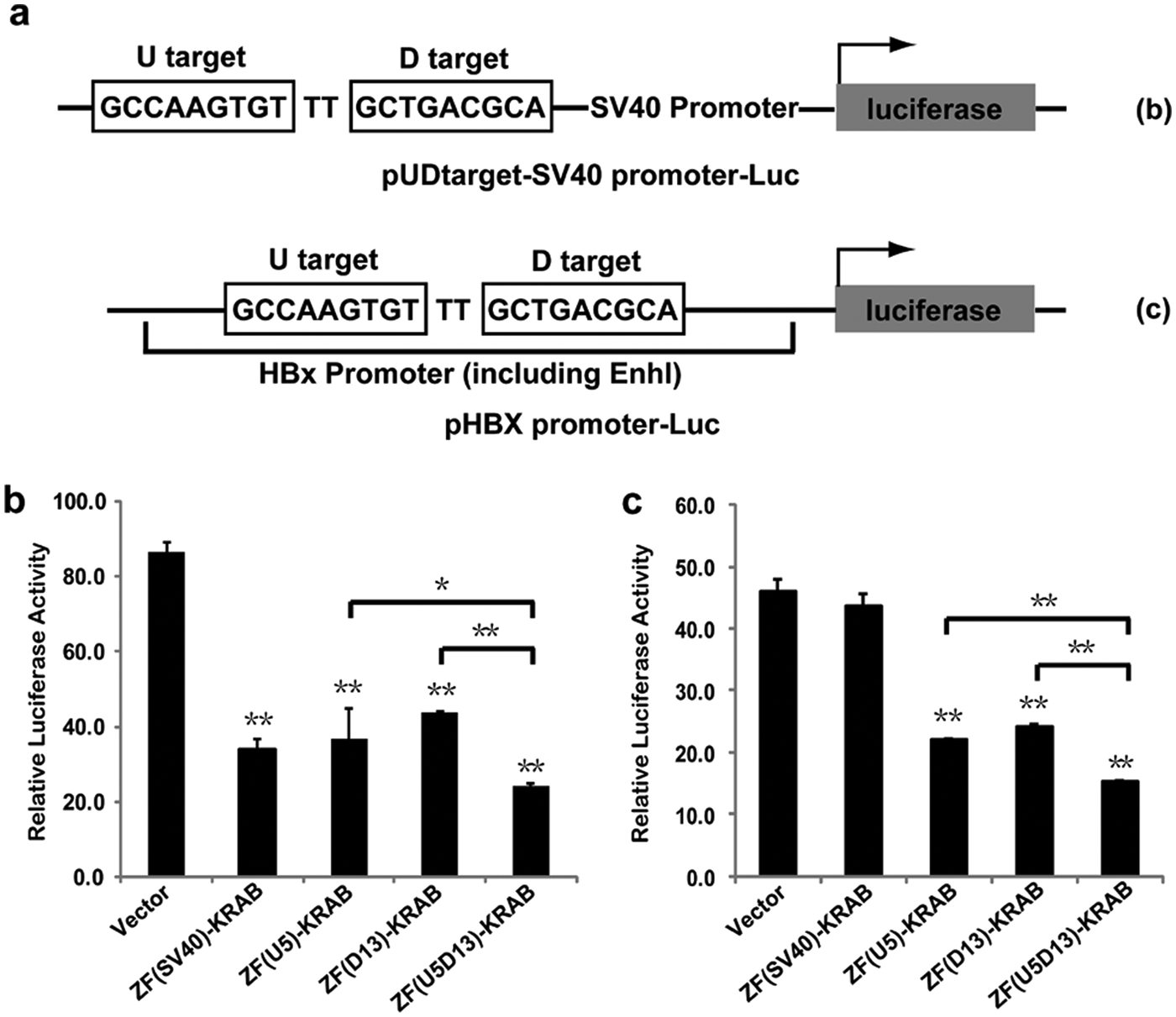
Downregulation of luciferase reporter gene by three- and six-fingered artificial transcription factors (ATFs). (
All assayed ATFs significantly decreased the expression of the luciferase reporter gene compared with the pcDNA3.1(+) vector control (p < 0.05). This was expected as the pUDtarget-SV40 promoter-Luc promoter region contains all the DNA recognition targets of these ATFs. The luciferase reporter gene was driven by EnhI and the HBx promoter in the absence of the SV40 promoter and ZF(SV40)-KRAB’s target ( Fig. 3c ). The ATFs ZF(U5)-KRAB, ZF(D13)-KRAB, and ZF(U5D13)-KRAB significantly downregulated luciferase compared with the control vector (p < 0.05). In contrast, there was no significant difference between the ZF(SV40)-KRAB and control vector activities. This was expected as there are no SV40 promoter sequences in the reporter vector. In both tests, the repressor function of the six-fingered ATF ZF(U5D13)-KRAB was superior to any of the three-fingered ATFs (pUDtarget-SV40 promoter-Luc: 71.8% repression vs. 57.4% with ZF(U5)-KRAB [p < 0.01] and 49.2% with ZF(D13)-KRAB [p < 0.05]; pHBX-promoter-Luc: 66.6% repression vs. 51.9% with ZF(U5)-KRAB [p < 0.01] and 47.2% with ZF(D13)-KRAB [p < 0.01]). These results were in accordance with previous reports that six-fingered ATFs display higher affinities to their targets than three-fingered ATFs.26,27
Six-fingered ATF Downregulates Expression of the HBx Integrant in Hep3B
To evaluate transcriptional repression by the candidate ATFs on HBx expression in Hep3B cells, we transiently transfected Hep3B cells with ATF expression vectors and measured HBx mRNA expression using real-time qPCR 72 h posttransfection. None of the ATF constructs induced significant repression of HBx, presumably because of poor transfection efficiency (~20% as evaluated by pEGFP-C1; data not shown). We then transiently transfected increasing amounts of the ZF(U5D13)-KRAB expression vector into Hep3B cells to determine whether a larger dose of expression vector could repress HBx to a greater extent. The HBx repression rate was somewhat enhanced when the ATF vector input was increased ( Fig. 4a ). However, even at the cells’ maximum tolerable input level (3 µg), the relative HBx mRNA level still was not significantly repressed compared with the vector control.
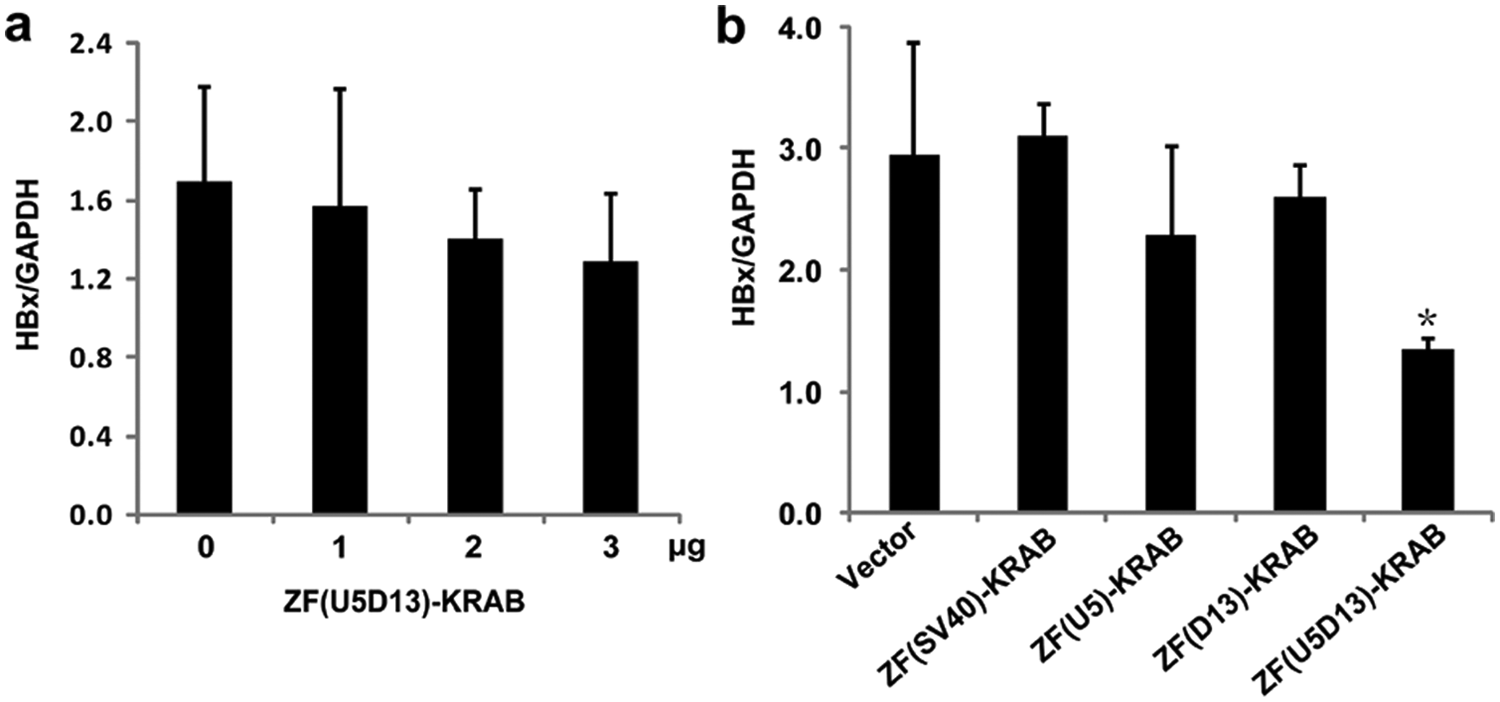
Stable expression of a six-fingered artificial transcription factor (ATF) based on U5D13 represses HBx mRNA. (
Stable Hep3B cell lines expressing the candidate ATFs then were established to avoid the drawbacks of transient transfections. The repressor function of the six-fingered ATF, ZF(U5D13)-KRAB, was superior to all other ATFs tested, inducing a significant decrease in relative HBx mRNA expression ( Fig. 4b , p < 0.05).
Six-fingered ATF Specifically Inhibits Growth of Hep3B Cells
HepG2, an HCC cell line known to lack HBV integrants ( Fig. 5a ), was used as a control to determine the specificity of the six-fingered ATF. ZF(U5D13)-KRAB and pcDNA3.1(+) blank vector were transfected into Hep3B and HepG2 cells, and stable transfectants were selected with G418. Transcriptional repression of HBx in a selected Hep3B stable clone that expressed the six-fingered ATF was confirmed by real-time qPCR. As shown in Figure 5b , the six-fingered ATF caused a 57.2% decrease in HBx transcription compared with the Hep3B stable clone that transfected blank vector (p < 0.05). Cell growth of selected stable clones was measured by the MTT assay. Stable expression of ZF(U5D13)-KRAB in Hep3B cells was associated with dramatic cell growth inhibition compared with mock control at the earliest observation time point (24 h) and lasted during the entire experimental process ( Fig. 5c ). In HepG2 cells, stable expression of ZF(U5D13)-KRAB resulted in no significant difference in cell growth rate compared with mock control.
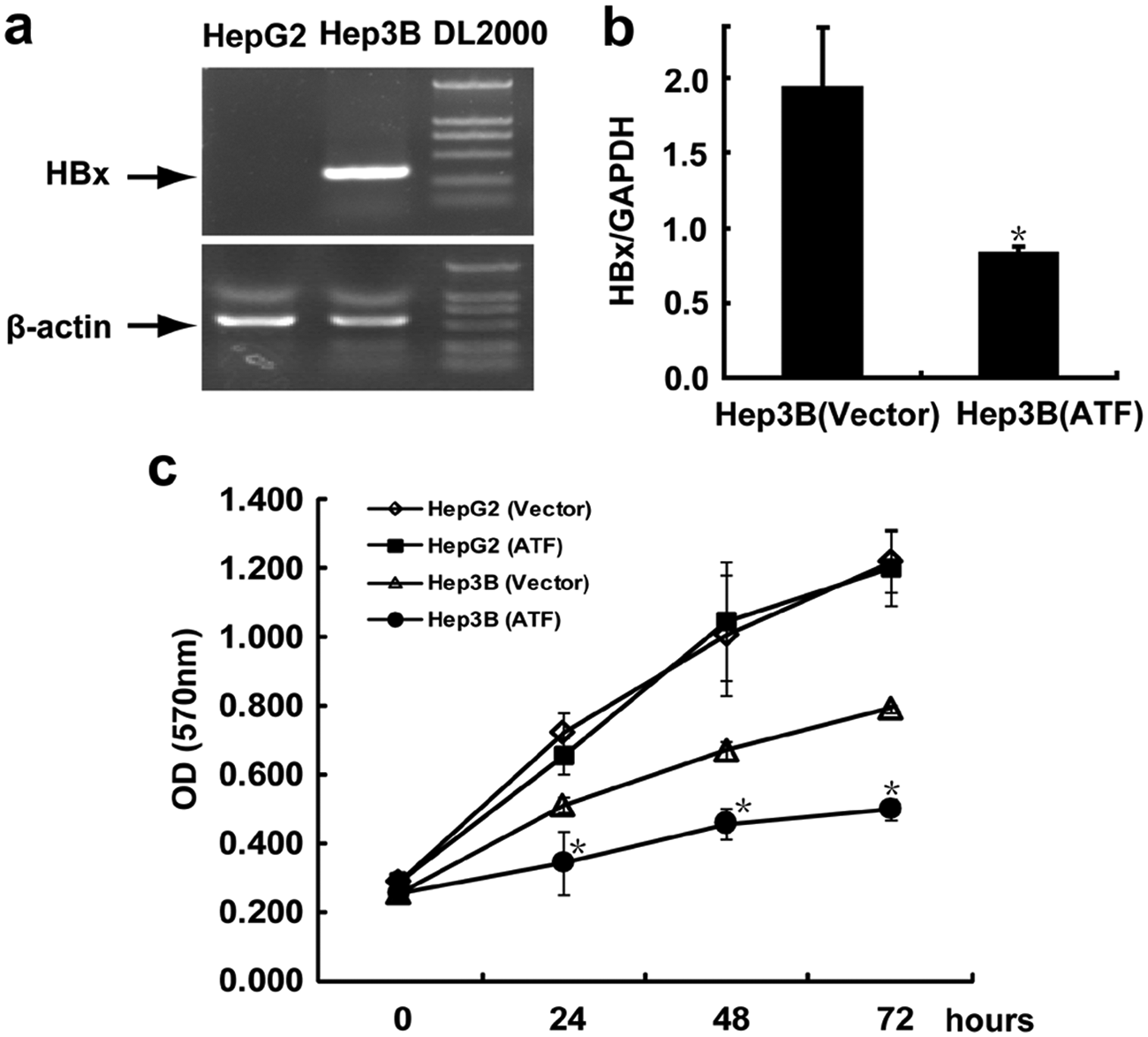
Six-fingered artificial transcription factor (ATF) based on U5D13 binding motif inhibits Hep3B cell growth. (
Discussion
C2H2 ZF-based ATFs have successfully been applied to the regulation of clinically relevant endogenous genes in human cell lines (e.g., VEGF, EPO, erbB-2, erbB-3).26,28 Although engineered three-fingered motifs are capable of regulating endogenous genes, 28 six-fingered ATFs generally demonstrate superior binding affinity and performance.18,27 An optimal ATF should regulate only its target gene. 14 Since the haploid human genome comprises more than 3 billion bp, an address of at least 16 bp is required for specific delivery of a DNA-binding protein to a single DNA target site within the genome. 18 An 18-bp DNA target, recognized by a six-fingered protein, would be unique within 68 billion bp (i.e., 418) and therefore could sufficiently target a single locus in the human genome (avoiding a highly repetitive sequence). 18 Because no native ZF motifs were identified that matched the full DNA sequence and because three-fingered phage-display ZF libraries are widely used to screen ZFs targeted to 9-bp DNA sequences, a six-fingered motif was generated by linking two three-fingered proteins, each recognizing half of the 18-bp region. In the present study, the six-fingered ATF, ZF(U5D13)-KRAB, was engineered to recognize the upstream HBx promoter region. As determined by luciferase reporter assays, the three-fingered ATFs (ZF(U5)-KRAB and ZF(D13)-KRAB) and the six-fingered ATF all substantially repressed HBx expression ( Fig. 3 ), but the repression efficiency of the six-fingered ATF was superior (**p < 0.01, *p < 0.05). However, when the ATFs were transiently transfected into Hep3B cells, the low transfection efficiency failed to produce HBx downregulation ( Fig. 4a ). Therefore, clonal cell lines stably expressing each ATF were established ( Fig. 4b ). Only the stable cell line originally transfected with the six-fingered motif exhibited statistically significant repression of HBx. These data suggest that three-fingered ATFs (specifying 9-bp DNA targets) might be sufficient to control gene expression from an episomal expression vector but might be inadequate to precisely target a single sequence among the 3 billion base pairs of the human genome. Thus, six-fingered ATFs (specifying 18-bp DNA targets) are recommended to ensure specific and efficient gene regulation.
The choice of DNA target sequence is one of the most critical steps in the design and construction of an ATF. Typically, a DNase I hypersensitivity assay is performed to identify TF-accessible chromosomal loci in the regulatory region of the gene to be targeted. We selected the EnhI region of HBV based on prior DNase I experiments indicating that it is accessible and interacts with many host TFs. 25 In addition, we chose an 18-bp target consisting of two 9-bp DNA sequences spaced by a dinucleotide (TT) to improve extension of the ZF domains. Our results ( Fig. 4b ) confirmed the accessibility of ATFs to this DNA target.
Successful applications of ATFs in fruit fly, 29 rat, 30 and mouse 14 model systems have been reported. The C2H2 ZFs provide ideal DNA-binding domains for ATFs because their small and conserved structural framework makes library-based screens possible. Their independent and modular features allow for tandem linkages and thus recognition of asymmetrical DNA sequences of the desired length. Once a DNA-binding domain specifying the unique DNA target is generated, ATFs can be customized by linking the DNA-binding domain to an effector domain, such as KRAB, p65, or VP16, for transcriptional activation; FokI for cleavage; a DNA methyltransferase catalytic domain; or a histone methyltransferase domain.14,28 Compared with other technologies such as antisense and siRNA approaches, downregulating an endogenous gene with zinc finger ATFs has several advantages that make the ATFs suitable for clinical utility. First, the ATFs mimic the natural mode of gene transcriptional regulation, which works before mRNA synthesis. It is a much more economical strategy than the mRNA targeting approaches. Second, ATFs can be used in either DNA (expression plasmids) or protein form. Both forms provide longer half-lives than the RNA molecules used in antisense and siRNA strategies. Third, ZF-DNA interactions are more various than the interactions between nucleic acid molecules, and the library screening strategy will ensure the optimum affinity and specificity of zinc fingers to the target sequences to minimize the “off-target” effect. Finally, ATFs can be used to regulate an endogenous cellular gene and thus provide a workaround solution for genes whose cDNAs are patented. A company in the United States, named Sangamo BioSciences, Inc. (Richmond, CA) has focused on preclinical and clinical studies of zinc finger proteins (including zinc finger artificial transcription factors) for many years.14,28–30 For instance, it is developing an ATF that upregulates the gene for glial cell line–derived neurotrophic factor (GDNF), a potent neurotrophic factor that has shown promise in preclinical testing to slow or stop the progression of Parkinson disease. 30 According to Sangamo’s reports, zinc finger ATFs have been tested in humans and to date have an excellent safety profile. 31
In this study, we constructed ATFs to downregulate the expression of the HBx integrant in the Hep3B genome. The 18-bp DNA target in the HBx EnhI region was divided into two 9-bp targets and was screened in separate phage-display ZF libraries to obtain three-fingered motifs that recognized U and D targets with high specificity. The fused six-fingered protein then was linked to a KRAB repression domain and was validated by a luciferase reporter assay. In a stably expressing Hep3B cell line, the six-fingered ATF repressed HBx expression and arrested cell growth. In contrast, stable expression of this ATF did not alter the growth of HepG2 cells, which lack the integrated HBV sequence. Our results suggest that targeted downregulation of HBx via ATFs might interfere with HBV-associated HCC progression. This study also lends insight to the design and application of C2H2 ZF-based ATFs for the regulation of HBx in liver cells with HBV genome integration. The DNA target of ATFs generated in this study matches one of the most common HBV subtypes in China. But the HBV sequences found in some patients showed variation. In these cases, the target sequences for ATFs can be determined by individual HBV sequence analysis. Zinc fingers with desired target affinity and specificity can be further screened, and KRAB repression domain will be linked with the zinc finger motifs to generate the ATF for optimum HBx transcription repression efficiency. By substituting the KRAB effector domain with other functional domains, such as a methyltransferase enzymatic domain, new avenues become available for the treatment (and “individual treatment”) of HBV-related HCC and for the study of HBx functional mechanisms in hepatocellular carcinoma genesis.
Footnotes
Acknowledgements
We would like to thank BioMed Proofreading, LLC, for advice and manuscript proofreading.
Declaration of Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the National Natural Science Foundation of China (Grant Nos. 81025018 and 81200356), and Central Grade Public Welfare Fundamental Science fund for the Scientific Research Institute of China (Grant No. 2012cj-3).
