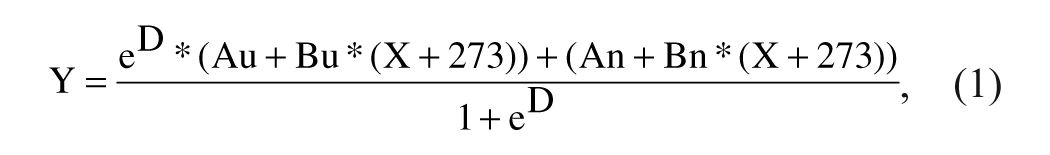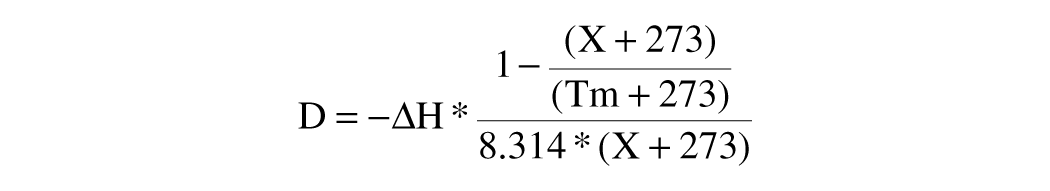Abstract
The authors describe the discovery of anti-mycobacterial compounds through identifying mechanistically diverse inhibitors of the essential Mycobacterium tuberculosis (Mtb) enzyme, pantothenate kinase (CoaA). Target-driven drug discovery technologies often work with purified enzymes, and inhibitors thus discovered may not optimally inhibit the form of the target enzyme predominant in the bacterial cell or may not be available at the desired concentration. Therefore, in addition to addressing entry or efflux issues, inhibitors with diverse mechanisms of inhibition (MoI) could be prioritized before hit-to-lead optimization. The authors describe a high-throughput assay based on protein thermal melting to screen large numbers of compounds for hits with diverse MoI. Following high-throughput screening for Mtb CoaA enzyme inhibitors, a concentration-dependent increase in protein thermal stability was used to identify true binders, and the degree of enhancement or reduction in thermal stability in the presence of substrate was used to classify inhibitors as competitive or non/uncompetitive. The thermal shift–based MoI assay could be adapted to screen hundreds of compounds in a single experiment as compared to traditional biochemical approaches for MoI determination. This MoI was confirmed through mechanistic studies that estimated Kie and Kies for representative compounds and through nuclear magnetic resonance–based ligand displacement assays.
Keywords
Introduction
The World Health Organization estimated in 2010 that ~11.1 million people across the globe are infected with Mycobacterium tuberculosis in a given year, with an associated mortality of 1.3 million. 1 Furthermore, 4% of all cases harbored resistance to at least one frontline drug in the current anti–tuberculosis (TB) regimen. 2 This, coupled with the extended duration of treatment, makes it imperative to discover new drugs to combat tuberculosis.
Cofactor synthesis pathways are rich sources of potential drug targets, as a single cofactor is used by many proteins in a network. Coenzyme A (CoA), one such cofactor, is an essential and ubiquitous part of the metabolism of many organisms. A recent review by Spry et al. 3 estimates that 9% of 3500 identified activities in the database BRENDA use CoA or a derivative as a cofactor.
In most bacteria, CoA is synthesized from pantothenic acid in five steps. 3 The first committed step is the phosphorylation of pantothenate by the enzyme pantothenate kinase (PanK or CoaA), with adenosine triphosphate (ATP) serving as the phosphate donor. The PanK reaction is also the rate-limiting step in CoA biosynthesis and, in many bacteria and eukaryotes, feedback regulated by the product of the pathway, CoA, and its thioesters. 4 Three distinct types of PanK have been characterized thus far. The type I PanK, exemplified by the Escherichia coli CoaA protein, was originally thought to represent the majority of PanK enzymes from prokaryotic sources.5,6 Type II PanKs predominantly occur in eukaryotic organisms.7,8 Recently, a third type of PanK was identified in bacteria (encoded by the gene coaX, to be distinguished from coaA). Type III PanKs have a much wider phylogenetic distribution than the better known type I PanKs and are present in 12 of the 13 major bacterial groups, as well as in many pathogenic bacteria. 9
The Mtb genome has a single copy of the gene encoding the type I PanK, coaA. CoaA has been shown to be essential in M. tuberculosis by transposon mutagenesis 10 and gene knockout experiments. 11 The Mtb genome also contains a type III PanK, coaX, but this gene has recently been shown to be nonessential. 11 Furthermore, there is low sequence homology between prokaryotic (type I) and eukaryotic (type II) pantothenate kinases. 12 Hence, an opportunity exists for the discovery of Mtb CoaA inhibitors that are specific to bacteria and have minimal interaction with the human enzyme.
The discovery of compounds with exquisite inhibitory effect against purified enzymes does not always translate into antibacterial activity. In addition to permeability and efflux issues, the target enzyme could also exist in different forms inside the cell, with potency optimization happening on the incorrect enzyme form. Kinetic analysis of the E. coli type I PanK indicates that substrates bind in an ordered fashion, with ATP binding first and pantothenate binding to the ATP–PanK complex. 6 In-house nuclear magnetic resonance (NMR) experiments studying the order of addition of substrates using a nonhydrolysable analog of ATP (ATPγS) and pantothenate corroborate this observation for Mtb CoaA, arguing that Mtb CoaA could exist as free and ATP-bound forms inside the cell. We therefore explored the possibility of screening for mechanistically diverse inhibitors that could bind either the free enzyme (biochemically competitive with ATP), the ATP–CoaA complex (uncompetitive with ATP), or both (noncompetitive inhibitors).
Information on inhibitor mechanism is routinely obtained by detailed biochemical analysis, but this technique is limited in the numbers of compounds that can be characterized. Obtaining mechanistic information for the large numbers of hits produced by high-throughput screening (HTS) necessitates a technique with adequate throughput. In this article, we explore the modulation of inhibitor-enhanced protein thermal stability by the presence of enzyme substrates as a tool to screen for desired inhibitor mechanisms without compromising throughput. We further demonstrate the validity of this method by characterizing the modes of inhibition (MoI) of inhibitors thus discovered, using a variety of techniques.
Materials and Methods
Protein expression and purification
The gene coding Mtb pantothenate kinase (coaA) was cloned, expressed, and purified in E. coli as reported previously. 13 Briefly, coaA was cloned in a pET15b vector under a T7 promoter and expressed as a Histidine6-tagged protein in BL21(DE3) cells (Novagen, Gibbstown, NJ) at 37 °C. Protein was purified using metal affinity chromatography (Ni-NTA; Qiagen, Valenica, CA), followed by gel permeation chromatography (GPC) in buffer containing 50 mM Tris-HCl (pH 7.5), 300 mM NaCl, 10% glycerol, 2 mM dithiothreitol (DTT), and 0.1 mM EDTA. Protein aliquots were stored at −70 °C. Only once-thawed aliquots were used for each experiment.
Thermal stability measurements
Thermal stability measurements were recorded in 96-well PCR plates covered with optical film (Bio-Rad, Hercules, CA), using a real-time PCR thermal cycler (initially Mx3000 from Stratagene [La Jolla, CA], subsequently iQ5 from Bio-Rad). Protein solution at a concentration of 0.15 to 0.2 mg/mL in buffer containing 50 mM HEPES-Na (pH 7.5) and 100 mM NaCl in a 50-µL volume was mixed with compound at appropriate concentrations or with equivalent amounts of DMSO. Final DMSO concentration was maintained at 4% (v/v). Protein–DMSO or protein–compound mixes also contained 0.5 mM or 1 mM ATP as required.
The environmentally sensitive dye, SYPRO Orange (Invitrogen, Carlsbad, CA), was used to monitor protein unfolding at a final arbitrary concentration of “6.25X” (the dye is sold as a DMSO stock marked “5000X”). Thermal unfolding was monitored at 0.5 °C intervals from 25 °C to 85 °C. Thermal melting profiles were analyzed using a six-parametric equation (1) in GraphPad Prism version 3.0 (GraphPad Software, La Jolla, CA):
where
where Tm is the midpoint of the unfolding transition, ΔH is the enthalpy change, Au is the maximum value of fluorescence observed, An is the minimum value of fluorescence observed, Bu is slope of the unfolded baseline, and Bn is the slope of the folded baseline.
Enzyme assay for Mtb CoaA characterization
Mtb CoaA activity was measured using a pyruvate kinase–lactate dehydrogenase (PK-LDH) coupled assay. A previously reported assay 14 was modified and adapted to the Mtb enzyme in the 96-well format. This assay has also been optimized in the 384-well format for the HTS campaign (unpublished data). All reactions were performed at 25 °C. Kinetic parameters for pantothenate and ATP using Mtb CoaA were determined in a 100-µL reaction mixture containing 50 mM PIPES–NaOH (pH 7.0), 25 mM KCl, 20 mM MgCl2, 0.1 mM EDTA, 2 mM DTT, 0.002% Brij, 0.5 mM phosphoenol pyruvate (PEP), 0.24 mM NADH, 10 U/mL PK-LDH, and 20 nM Mtb CoaA (0.75 µg/mL). The reaction was started with the addition of Mtb CoaA to the reaction mix. Change in absorbance at 340 nm (NADH consumption) was monitored for 60 min in a SpectraMax (Molecular Devices, Sunnyvale, CA) spectrophotometer. Km for ATP was determined by varying the concentration of ATP from 20 µM to 1 mM while maintaining D-pantothenate at a saturating concentration of 3 mM. Km for D-pantothenate was determined by varying its concentration from 50 µM to 3 mM while maintaining ATP at a saturating concentration of 1 mM.
Inhibitor screening and kinetic parameters
Routine screening and IC50 estimations for the Mtb CoaA inhibitors were carried out by keeping both substrate concentrations at their respective Km under the buffer conditions mentioned above. For selected inhibitors, competition with ATP was checked by measuring IC50 at a higher ATP concentration (50 × Km = 6 mM). Compounds listed in Table 1 were further characterized for their kinetic MoI with respect to the ATP substrate. The inhibitory effect was monitored using an appropriate range of compound concentrations in the presence of varying amounts of ATP (1.5 mM to 11.7 µM) and a constant amount of D-pantothenate (3 mM).
Mtb CoaA Inhibitors Representative Compounds from Each Mechanistically Distinct Class Are Listed along with Their Inhibitory Properties
Initial rates in all the reactions were estimated using SoftMaxPro data analysis software (Molecular Devices). Inhibition constants were determined by fitting the data to four different equations, equations (2) to (5), for the competitive, pure noncompetitive, mixed noncompetitive, and uncompetitive modes of inhibition where Kie and Kies indicate the dissociation constants for free enzyme–inhibitor complex and substrate bound enzyme–inhibitor complex, respectively.
Competitive inhibition:
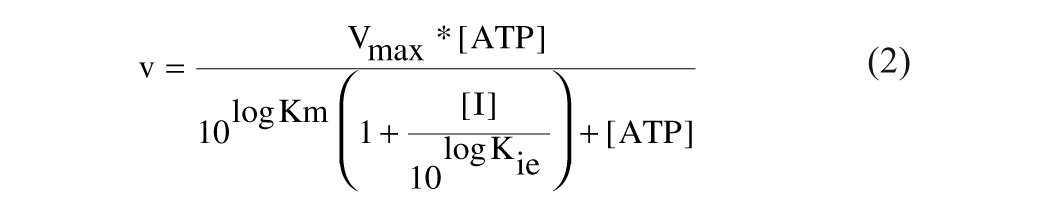
Pure noncompetitive:
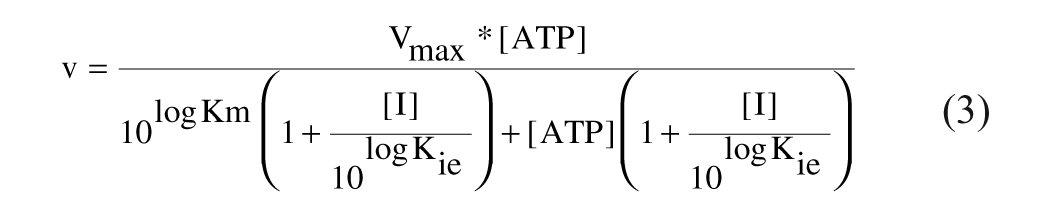
Mixed noncompetitive:
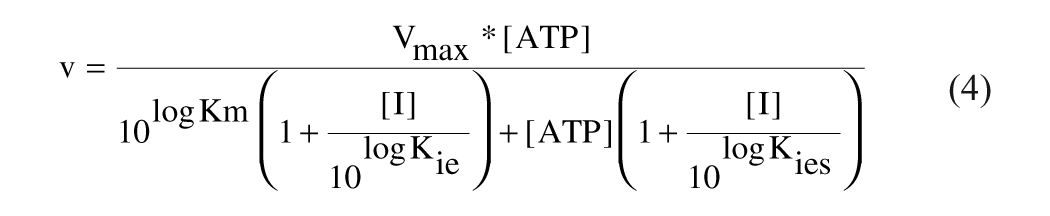
Uncompetitive:
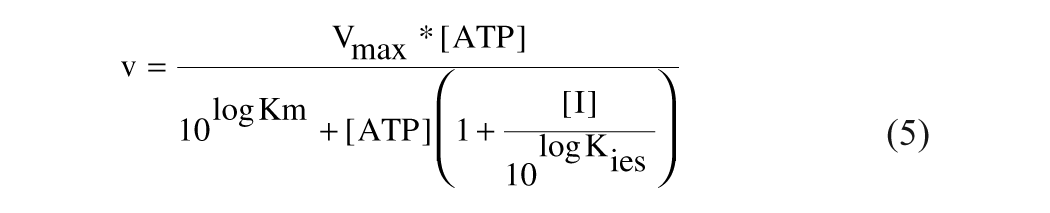
GraFit 5.0.11 was used for nonlinear regression analysis. Eadie-Scatchard plots (v/[ATP] vs v) using the initial velocity data were used to discriminate between various modes of inhibition.
WaterLOGSY
NMR spectra were recorded at 298K on a Bruker AvanceII 500-MHz spectrometer equipped with a 5-mm triple resonance inverse probe. Reference spectra were recorded for all samples prior to each 1D WaterLOGSY experiment. Pulse program was from Dalvit et al.15,16 The first water-selective pulse was 6 ms long, whereas the two water-selective 180° square pulses of the double spin echo scheme were 3 ms long. A weak rectangular pulsed field gradient was applied for the duration of the mixing time (1.8 s), which was followed by a gradient recovery time of 2 ms before the detection pulse. The experimental sweep width was 12 379 Hz, with an acquisition time of 1.3 s. Data were multiplied with an exponential function (line broadening 2 Hz) and then processed.
Results
Using changes in protein thermal stability to determine mode of inhibition
The thermal stability of an enzyme can be enhanced by the binding of a ligand, and the extent of stabilization is guided by ligand binding affinity and the number of binding sites. This enhancement in stability can be measured as a positive shift in the midpoint of the melting curve (Tm) in the presence of ligand. The magnitude of observed thermal shifts can be modulated by the addition of a second ligand. If the first ligand were an enzyme inhibitor and the second ligand a substrate, three scenarios are possible ( Figure 1 ; assumption: inhibitor and substrate do not have similar contributions to protein thermal stability on binding. Ligands bind preferentially to the native form of the protein.).
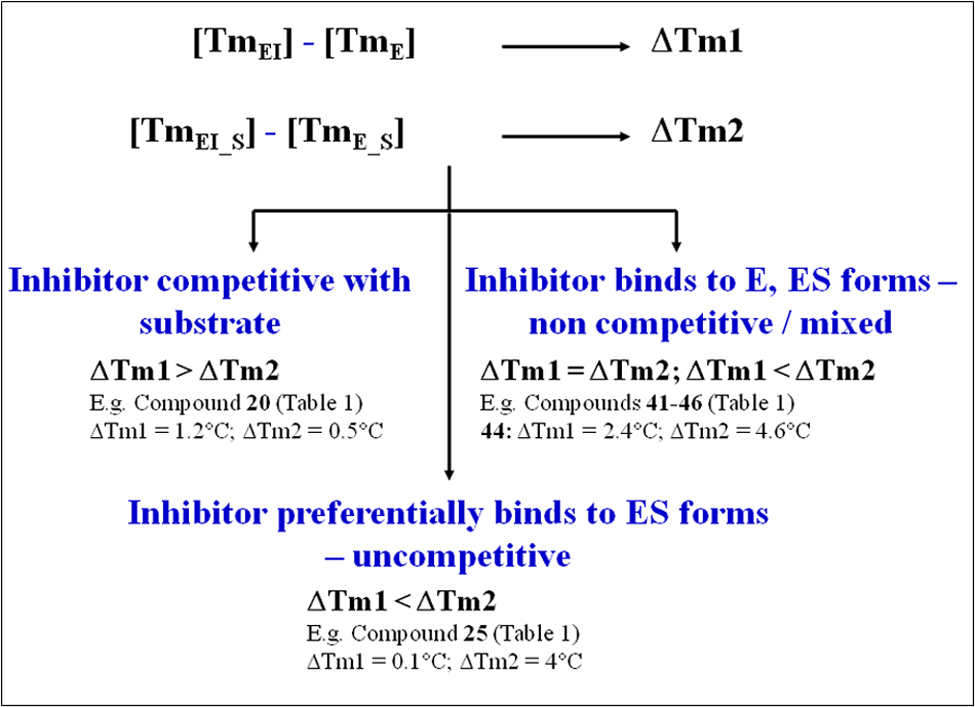

Deriving mode of inhibition (MoI) using changes in the magnitude of thermal melting points upon binding of inhibitor in the presence and absence of substrate. TmEI, Tm observed on inhibitor binding to protein in the absence of substrate; TmE, Tm of protein alone; TmEI_S, Tm observed on inhibitor binding in presence of substrate; TmE_S, Tm of protein in presence of substrate; ΔTm, shift in Tm on inhibitor binding.
Scenario 1: Inhibitor with higher affinity is partially displaced by a substrate molecule competing to bind at the same site as inhibitor (competitive inhibitor). In such a scenario, the magnitude of an inhibitor-induced thermal shift in the presence of substrate is lowered as compared to the shift observed in its absence.
Scenario 2: Inhibitor preferentially binds to the substrate-bound form of the enzyme (uncompetitive inhibitor). In such a scenario, the magnitude of shift in the presence of substrate is higher as compared to the shift observed in its absence. For purely uncompetitive inhibitors, no Tm shift may be observed in the absence of substrate.
Scenario 3: Inhibitor binds equally to the free and substrate-bound forms of the protein (noncompetitive inhibitor). In such cases, the magnitude of shift in the presence of substrate is either higher as compared to the shift observed in its absence, or little difference is observed.
Effect of Mtb CoaA inhibitors on protein thermal stability
We conducted two HTS campaigns for inhibitors against the Mtb CoaA enzyme. The first screen (campaign 1) challenged a diverse library of ~70 000 compounds, whereas the second screen (campaign 2) challenged a larger library of ~1 000 000 compounds. Nonpromiscuous inhibitors (IC50s independent of protein and detergent concentration in the enzyme assay and Hill slope ~1) were used for MoI studies described below.
Campaign 1 hits
Campaign 1 chemistry focused on triazole and quinolone scaffolds (see Table 1 for representative molecules). Molecules from these two chemical classes increased the thermal stability of Mtb CoaA ( Fig. 2a ), suggesting specific binding without enzyme denaturation. The increased thermal stability also correlated broadly with inhibitor potency (IC50).
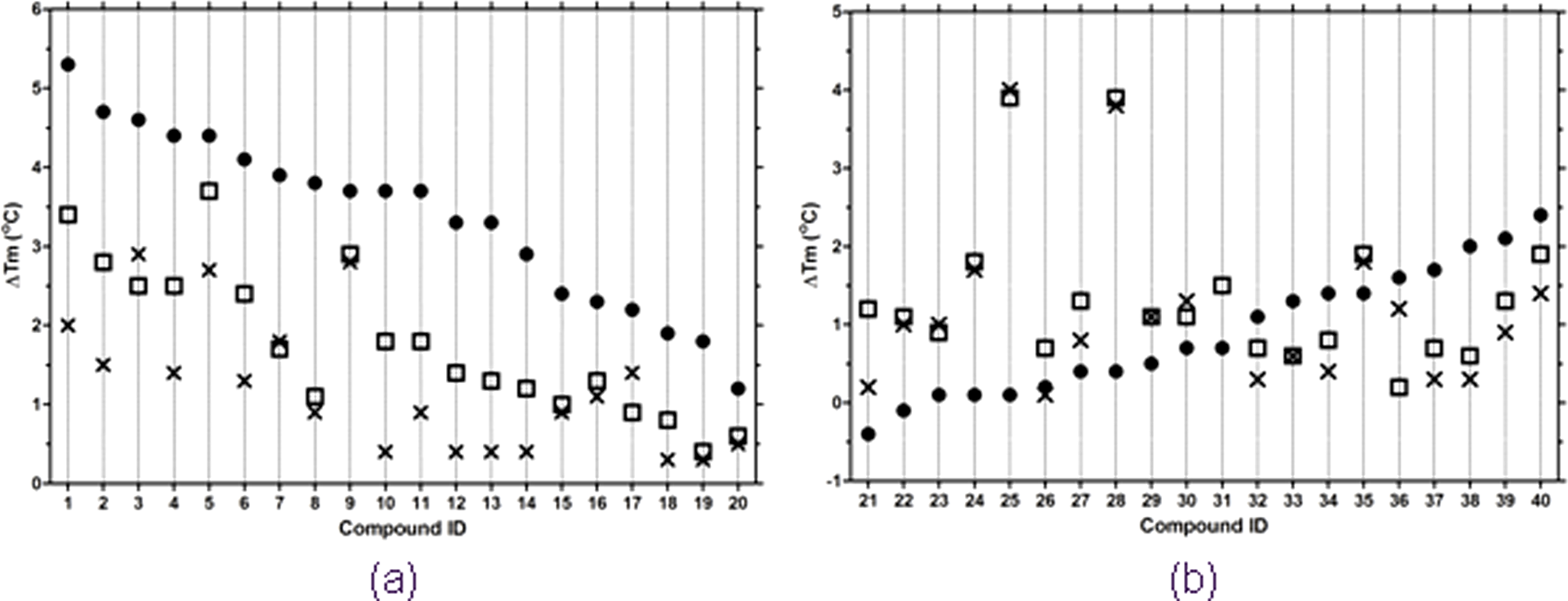
Modulation of inhibitor-induced thermal shift in the presence of adenosine triphosphate (ATP). Compounds were assayed at 25 µM and at three ATP concentrations: 0 (•), 0.5 (□), and 1 (×) mM. (
In the presence of ATP, this stability enhancement was reduced, as evidenced by a decreased Tm shift when compared to the “no-ATP” condition.
Figure 2a
shows the progressive decrease in Tm shift in the presence of increasing concentrations of ATP. For example, the triazole
Campaign 2 hits
Campaign 2 involved the screening of a larger, more diverse library and resulted in a greater number of potential chemistry start points. To identify hits with diverse mechanisms of inhibition, the high-throughput thermal stability-based MoI assay was used to classify inhibitors as competitive or un/noncompetitive with respect to ATP.
Figure 2b shows the behavior of a representative set of compounds from campaign 2 in the presence and absence of ATP. Unlike molecules from campaign 1, all inhibitors tested did not bind and enhance the thermal stability of CoaA in the absence of ATP. In the presence of ATP, CoaA inhibitors derived from campaign 2 displayed a variety of responses.
Set 1: The binding of compounds such as
Set 2: Compounds such as
Set 3: Compounds such as
Chemically desirable hits that were also not competitive with ATP in the thermal shift–based MoI assay were prioritized for chemistry expansion. Specifically, biaryl acetic acid and quinoline carboxamide scaffolds (represented in
Fig. 2b
by compounds
To demonstrate that the thermal shift–based MoI assay reliably picks compounds with desirable MoI, inhibition kinetics was studied in greater detail for selected compounds synthesized during the structure–activity relationship (SAR) exploration of the biaryl acetic acid and quinoline carboxamide series ( Table 1 ). An uncompetitive inhibitor (thiazole class) and a competitive inhibitor (triazole class) were also included in the study for comparison.
Confirming mode of inhibition in enzyme kinetic assays
Mode of Mtb CoaA inhibition was elucidated using the PK-LDH coupled assay adapted to the 96-well plate format. Estimates of Kcat, Km for ATP, and D-pantothenate were confirmed in four replicates during characterization of the Mtb enzyme and were reconfirmed during eight independent studies on mode of inhibition for the enzyme inhibitors using varying concentrations of ATP. Estimated values of Kcat, Km for ATP, and D-pantothenate were 5 ± 2 s–1, 122 ± 25 µM, and 395 ± 9 µM, respectively.
Enzyme kinetics data obtained in the presence of eight inhibitors (
Table 1
) were modeled using equations (1) to (4). Inhibition constants, Kie (indicative of the affinity for the free form of enzyme) and Kies (indicative of the affinity for the substrate-bound form of enzyme) (
Table 1
), were used along with Eadie-Scatchard plot analysis (
Fig. 3
) to elucidate the MoI. The plot of v/[S] versus v for compound
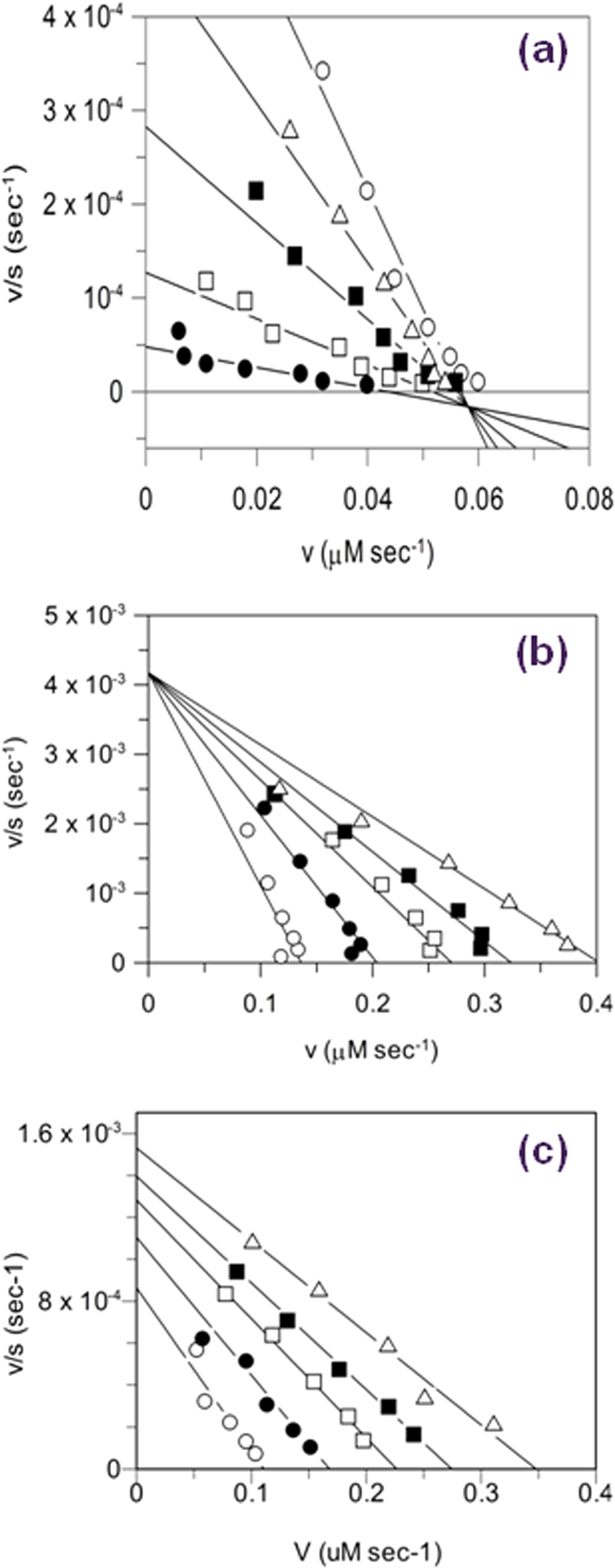
Eadie-Scatchard plots for Mtb CoaA inhibitors with various modes of inhibition. Plots are shown for three compounds from
Table 1
and exemplify the three different modes of inhibition (MoIs) observed with respect to adenosine triphosphate (ATP) substrate. (
Similarly, the thiazole
Kinetic data for the quinoline carboxamide
Kie/Kies estimates for the representatives of the biaryl acetic acid class (compounds
The kinetic MoI data obtained for the above inhibitors thus confirmed the MoI indicated by the high-throughput thermal melting–based assay.
Mapping of inhibitor binding site by 1D NMR
A robust campaign to improve inhibitor potency is greatly aided by knowledge of the inhibitor binding pocket. In the absence of co-crystal structures, competition with substrate or an exemplary inhibitor often provides part of this information. During a large part of the two chemistry campaigns, we did not have co-crystal structures of Mtb CoaA with any inhibitor. Also, the substrate binding pocket of CoaA is an extended zone that accommodates both ATP and D-pantothenate or the feedback inhibitor, coenzyme A.13,17,18 To map the binding sites of the various inhibitors in this extended pocket, a series of displacement experiments was planned. CoA or the inhibitors discovered from screening were sequentially used as probes to check if inhibitors bound Mtb CoaA in the extended pocket occupied by CoA and if the various inhibitor classes occupied the same binding site.
Figure 4
shows the binding to Mtb CoaA of a representative quinoline carboxamide,
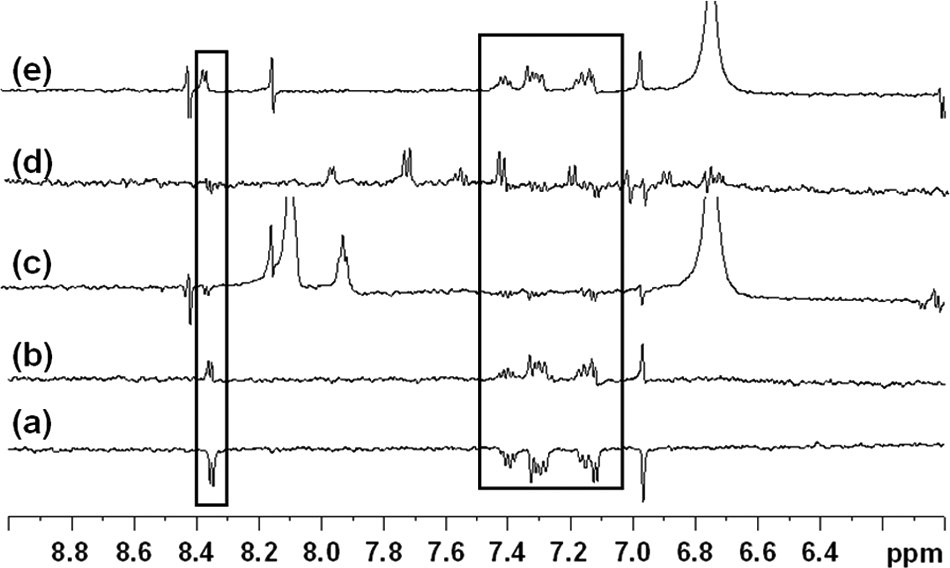
WaterLOGSY experiments showing (
WaterLOGSY data obtained using the uncompetitive thiazole compound
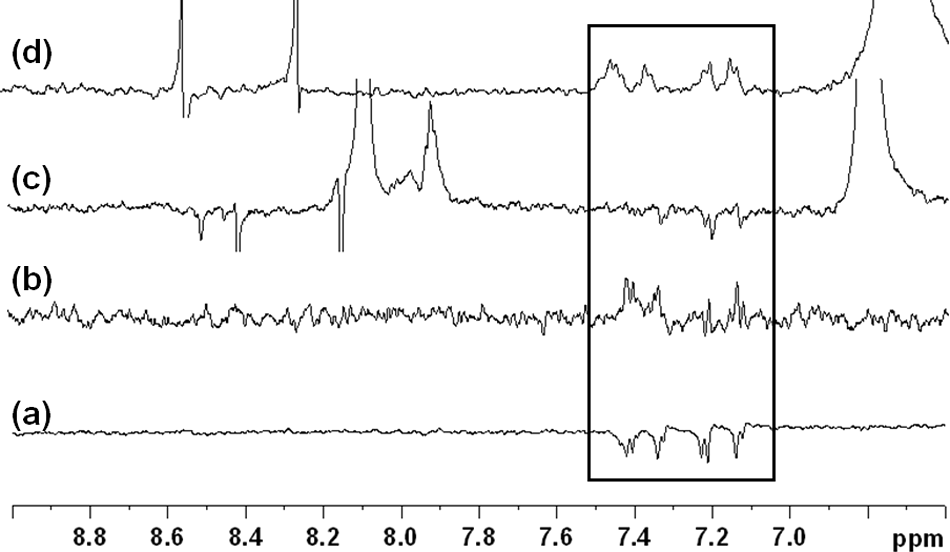
WaterLOGSY experiments showing (
The above NMR data were consistent with the observation that the quinoline carboxamide and thiazole classes of inhibitors were not competitive with ATP but interestingly also suggested that their binding site(s) overlapped with that of known ATP competitors such as coenzyme A and a triazole class of inhibitors. Crystal structures of Mtb CoaA in complex with triazole and biaryl acetic acid inhibitors were obtained as this article was being prepared, and the inhibitors were observed to occupy overlapping positions with coenzyme A and each other, as suggested by the NMR displacement experiments (T. Alwyn Jones, personal communication, 2008).
Discussion
Mtb CoaA is an essential bacterial enzyme that has been evaluated in this study as a target for the discovery of anti-TB drugs. Target-driven drug discovery is greatly aided by enzyme targets such as Mtb CoaA that are amenable to a range of biochemical and biophysical studies. Although this frequently leads to the discovery of well-characterized, potent inhibitors, such inhibitors may fail to demonstrate antibacterial activity due to a variety of reasons, such as poor permeability into or efficient efflux out of the bacterial cell. The enzyme may also exist in the cell in forms significantly different from that used to screen for inhibitors. In a target-based, enzyme potency–driven drug discovery campaign, it is not easy to predict permeability or efflux issues during screening or potency optimization phases. It is also difficult to accurately predict the biologically relevant form of the protein. One way to mitigate these risks could be to select chemical start points with maximum structural and mechanistic diversity.
In the specific case of Mtb CoaA as a target, ATP-competitive inhibitors may require very high potency to inhibit the enzyme inside the cell as the intracellular ATP concentration (~1 mM; unpublished data)19,20 is in ~10-fold excess of its Km for ATP. As per the Cheng-Prusoff relationship 21 between IC50 and Ki, a competitive inhibitor may be required at a much higher concentration (>10×) to achieve the same degree of inhibition as obtained by a noncompetitive or uncompetitive inhibitor with equivalent Ki values. Similar arguments have been put forth by Duggleby 22 to overcome the phenomenon of “metabolic resistance,” where increasing substrate accumulation on enzyme inhibition by a competitive inhibitor ultimately reverses the effect of the inhibitor. In such cases, inhibitors that are uncompetitive or noncompetitive with accumulating substrate are more effective in the cellular context. Therefore, determination of MoI immediately after an HTS screen could be useful in either prioritizing compounds with desirable MoI or ensuring the inclusion of compounds with mechanistic diversity, thus increasing chances of success of obtaining antibacterial activity.
Thermal melting of proteins has been extensively used to explore protein function23,24 and search for optimal buffer conditions for purification 25 or crystallography, 26 as well as in drug discovery to screen for small-molecule ligands that bind the drug target under consideration.27-29 Changes in the thermal stability of a protein with varying concentrations of ligand have also been used to determine dissociation constants.30,31 For ligands with similar binding thermodynamics, the magnitude of this change at a single ligand concentration can be used to rank molecules in order of affinity for target. 32
The above examples all stress the ease with which the measurement of protein thermal stability can be adapted to high-throughput formats. In addition to screening for protein ligands, as done by Pantoliano et al.,
27
early binding confirmation can be obtained for the large numbers of enzyme inhibitors picked as hits at the end of HTS campaigns by monitoring increases in protein thermal stability. In this article, we also demonstrate that comparing changes in protein thermal stability in the presence of inhibitor with such changes in the presence of both inhibitor and substrate is a quick, effective, and high-throughput tool to classify HTS hits according to their modes of inhibition (i.e., as either competitive or non/uncompetitive with the substrate). In some cases, such as quinoline carboxamides
All four classes of Mtb CoaA inhibitors discussed in this study could be optimized to nanomolar potencies regardless of MoI (IC50 ≤50 nM when both substrates of Mtb CoaA were maintained at Km). To ensure the various mechanistic classes retained their indicated MoIs, thermal stability behaviors of compounds were tracked during the hit-to-lead campaign (
Fig. 2a
—competitive inhibitors;
Thus, mechanistic information on scaffolds at the onset of chemistry optimization can enhance the chances of identifying potent leads with enzyme inhibition and cellular activity. Such mechanistic information can be efficiently and reliably obtained for large numbers of potential scaffolds using the protein thermal stability-based MoI assay.
Footnotes
Acknowledgements
We thank Rajendra Rane for optimizing the Mtb CoaA assay used in inhibitor screening, Pavithra Vishwanath and T. Raga Deepthi for estimating the intracellular concentration of ATP in mycobacterial cells, and Sunita DeSousa for scientific discussions.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.