Abstract
P2Y14 is a member of the pyrimidinergic GPCR family. UDP-Glc has been previously shown to activate human P2Y14, whereas UDP was unable to activate the receptor. In this study, the authors used conventional and nonconventional methods to further characterize P2Y14 and its ligands. Conventional calcium mobilization and nonconventional cellular impedance functional assays revealed that UMP and UDP selectively activated HEK cells coexpressing P2Y14 and Gαqi5. In the impedance assays, the presence of exogenous Gαqi5 resulted in agonist-induced Gq signaling, whereas in the absence of exogenous Gαqi5, the signal was indicative of Gi. The authors established the first P2Y14 membrane filtration binding assay using a novel optimized expression vector and [3H]UDP as radioligand. UDP-Glc, UMP, and UDP dose dependently inhibited [3H]UDP binding in the binding assay, and saturation analysis revealed that UDP bound P2Y14 with a KD = 10 nM and a Bmax = 110 pmol/mg. The authors screened a phosphonate library and identified compound A, which inhibited UDP-Glc–mediated calcium signaling in the fluorometric imaging plate reader assay (IC50 = 2.3 µM) and competed for [3H]UDP binding in the novel binding assay with a Ki = 1280 nM.
Introduction
Chambers et al. 2 first reported that nucleotide sugars activate P2Y14 exogenously expressed in both yeast and mammalian cells. Other sugar nucleotides such as ADP-glucose, naturally occurring nucleotides (UMP, UDP, and UTP), and nucleosides (uridine) were unable to activate P2Y14. In 2008, Fricks et al. 7 reported that UDP was a competitive antagonist of human P2Y14 but a potent agonist of the rat receptor, and recently, Carter et al. 5 illustrated that UDP was a potent agonist of human P2Y14.
In this study, we further characterize the P2Y14 receptor using conventional Ca2+ mobilization and nonconventional cell impedance technologies. We develop functional assays and demonstrate that nucleotides uncoupled to glucose specifically activate P2Y14 in HEK cells stably expressing human, chimpanzee, or mouse P2Y14 in combination with Gαqi5. Given that nucleotides containing a specific number of phosphate groups differentially activate P2Y14 in our functional assays, we screened a phosphonate library in the calcium mobilization assay. Finally, the successful establishment, to our knowledge, of the first P2Y14 filtration binding assay allowed us to determine the affinity of the ligands and provide data for the first identified nonnucleotide, small-molecule antagonist of P2Y14.
Materials and Methods
Construction of expression vectors
Human P2Y14 was PCR amplified from the I.M.A.G.E. clone obtained from Invitrogen (Carlsbad, CA) using primer hGPR105_5′_KpnI AAAGGTACCGCCACCAT GATCAATT CAACCTCCAC and PRIMER hGPR105_3′_NotI AAAGCGG CCGCTCACAAAGTATCTGTGCTTTCAAG. The codon encoding glutamate at position 54 in the wild-type human P2Y14 was changed to code for lysine. The PCR conditions were 50 ng of the I.M.A.G.E. clone #5265592; 1X final concentration of cloned Pfu DNA polymerase reaction buffer; 300 µM each of dATP, dCTP, dTTP, and dGTP; 300 nM of each primer; dH2O to a final volume of 50 µL; and 2.5 units PfuTurbo hotstart DNA polymerase (Stratagene, La Jolla, CA). The PCR reaction was 5 cycles at 94 °C for 30 s, 65 °C for 30 s, and 72 °C for 90 s and then 25 cycles at 94 °C for 30 s, 70 °C for 30 s, and 72 °C for 90 s. The last cycle was followed by 7 min at 72 °C. The PCR-amplified human P2Y14 was digested with KpnI and NotI and cloned into a pcDNA3.1 (+) vector previously digested with KpnI and NotI to place the DNA fragment in the proper orientation downstream of the CMV promoter.
Chimpanzee P2Y14 nucleic acid fragment was synthesized based on the Ensembl Gene ID ENSPTRG00000030093 by Bio S&T (Montreal, Canada) to be flanked with KpnI and NotI restriction enzyme sites. The DNA fragment was digested with KpnI and NotI and then cloned into a pcDEF3 vector previously digested with KpnI and NotI to place the DNA fragment in the proper orientation downstream of the human EF-1a promoter.
Mouse P2Y14 was subcloned unidirectionally from the I.M.A.G.E. clone from Invitrogen (Cat. No. 6314145; in vector pCMV-SPORT 6.1) by digesting the clone with HindIII and NotI and then cloning the DNA fragment containing the P2Y14 into a pcDNA3.1(+) vector digested with HindIII and NotI to place the DNA fragment in the proper orientation downstream of the CMV promoter.
Human GPRA was cloned from a cDNA library and subcloned into a pcDNA3.1 (+) vector.
Chimpanzee P2Y14 was PCR amplified from the pcDEF3-chimpanzee P2Y14 construct using primers containing a 5′ XbaI restriction site and a 3′ NotI restriction site. The following primers were used: human P2Y14_5′_XbaI: 5′-AAATCTAGA GCCACCATGATCAATTCAACCTCCACACAG-3′ and human P2Y14_3′_NotI: 5′-AAAGCGGCCGCTCACAAAGTATCTG TGCTTTCAAG. Human P2Y14 was PCR amplified from the pcDNA3.1(+)-human P2Y14 construct using the same primers as for chimpanzee. The pYD7 vector 8 and PCR products were cut using XbaI and NotI enzymes and then ligated to create the chimpanzee and human pYD7 constructs.
Construction of stable clonal cell lines
For construction of the HEK cell line expressing P2Y14 and Gαqi5, plasmid encoding human, chimpanzee, or mouse P2Y14 and pCEP4 encoding the chimeric Gαqi5 were transfected into HEK EBNA cells using FUGENE 6 (Roche, Mannheim, Germany), according to the manufacturer’s instructions. For construction of the HEK cell line expressing GPRA and Gαqi5, conditions were similar to that for P2Y14. Cells were grown under selection with 100 to 200 µg/mL hygromycin B and 250 µg/mL geneticin to generate a bulk stable population. Single cells were seeded onto 96-well Nunc plates, and subsequent to the formation of a cell monolayer, receptor-expressing clones were identified based on agonist-induced stimulation of Ca2+ in the Ca2+ mobilization assay (described below).
Calcium mobilization functional assay
P2Y14- and GPRA-expressing clonal cells were selected for by UDP-Glc- or GPRA peptide–induced Ca2+ release using a fluorometric imaging plate reader (FLIPR), as follows. HEK cells expressing Gαqi5 and P2Y14 or GPRA were plated onto 384-well, poly-D-lysine–coated plates (Corning, Corning, NY) in DMEM growth medium (see above) containing 10% fetal bovine serum. Fluorescent no-wash dye (MDS, Sunnyvale, CA) was added to the cell monolayer without removing medium, and the plate was incubated for 60 min at 37 °C and 5% CO2. Agonist in HBSS was added, and Ca2+ signaling was monitored using a FLIPR.
For antagonist assays, compound was added to cell/dye mixture; following a 20 min incubation, EC80 concentration of agonist solution was added to cells, and Ca2+ signaling was monitored by FLIPR.
Quantitation of the percentage stimulation (% stimulation) or inhibition (% inhibition) of Ca2+ signaling by agonist or antagonist, respectively, was calculated using the Max-Min fluorescent signal detected.
Cellular impedance assays using CellKey
HEK293 cells expressing P2Y14 in the absence or presence of Gαqi5 or mock-transfected control HEK cells were plated in normal growth medium onto CellKey (MDS) standard 96W cell plates. For transient transfections, P2Y14 or vector control plasmid DNA were transfected into parental HEK cells using Fugene 6 (Roche) according to the manufacturer’s instructions. Twenty-four hours posttransfection, cells were transferred to standard Cellkey 96W cell plates and incubated overnight. Immediately prior to assay, medium was removed from the cell monolayer and replaced with assay buffer (Hanks’ Balanced Salt, supplemented with 20 mM Hepes + 0.01% bovine serum albumin). Cells were allowed to equilibrate in assay buffer for 30 min. After a 5 min baseline measurement, cells were treated with P2Y14 agonists dose response. To confirm Gi signaling, assays were identical to those described above except that cells were pretreated overnight in the presence of 40 ng/mL pertussis toxin. Maximum change in impedance (dZiec) was used to generate dose-response curves using Graphpad Prism. The unique pattern of the change of impedance over time was used to identify the signaling pathway.
Nucleotide stability analysis in functional and binding assays
Uridine nucleotides in the binding assay mixture were extracted with EDTA (pH 8) and acetonitrile or in the calcium mobilization assay with acetonitrile/water (2:1, v:v). Samples were centrifuged at 4 °C. For anion exchange chromatography, supernatant was injected onto a Waters 2695 system (Waters, Milford, MA) with a Zorbax SAX, anion-exchange column (Agilent, Santa Clara, CA). The analyses were performed at 4 °C with a gradient elution mode using 50 mM KH2PO4 as buffer A and 750 mM KH2PO4 as buffer B, both at pH 3.0. After 0.5 min of 50% buffer B, a linear gradient was from 50% to 100% buffer B over 4.5 min, maintaining 100% buffer B for 6 additional minutes, returning to initial conditions (50% buffer B) over 0.5 min, and then held for 2.5 min for a total run time of 14.0 min. Flow rate was 1 mL/min until 11 min, after which the flow rate was 1.5 to prepare for the next injection. Radioactivity was monitored online using a flow scintillation analyzer (Packard 150TR radiometric, Perkin Elmer, Waltham, MA), and the nucleotides were identified by their co-elution with known [3H]nucleotide standards.
For C18 reverse-phase chromatography, the samples were injected onto an Agilent 1100 system (Agilent, Santa Clara, CA) equipped with a diode array detector and an Eclipse XDB-C18 reverse-phase column. The mobile phase consisted of 2% MeOH/98% aqueous with a flow rate of 1.5 mL/min. The aqueous phase was acetic acid, 0.025% (v/v) dimethylhexylamine (DMHA) adjusted to pH 7.5 using ammonium hydroxide, to pH 8.3 using DMHA. The absorbance was monitored at 260 nm and the nucleotides identified by their co-elution with standards.
Preparation of membranes for binding assay
HEK 293 EBNA cell pellets, transiently transfected with human or chimpanzee P2Y14 cDNA in pdy7 vector, or pdy7 vector control, 9 were obtained from Biotechnology Research Institute from the National Research Council of Canada according to Loignon et al. 8
Membranes were prepared by N2 cavitation using a Parr unit (Parr Instrument Company, Moline, IL). Cells were thawed and resuspended in 10 mM Hepes/1 mM EDTA, pH 7.4, containing complete protease inhibitor (Roche Biochemicals, Mannheim, Germany) at a maximum concentration of 108 cells per milliliter. All subsequent steps were performed either on ice or at 4 °C. Cells were disrupted using a Dounce homogenizer, and homogenates were added to the cavitation unit. The pressure was adjusted to 800 psi, and samples were left under pressure for 30 min. The homogenates were released drop by drop and centrifuged at 1000g for 10 min. The supernatent was subjected to a final centrifugation at 160 000g (rotor type 60 Ti, 40 000 rpm) for 30 min. Resultant pellets were resuspended in 10 mM HEPES/KOH pH 7.4, 1 mM EDTA, in 1/10 the original volume by Dounce homogenization, aliquoted, and stored frozen at −80 °C.
Membrane filtration binding assay
Binding assays were performed in 10 mM Hepes pH 7.4 (KOH) containing 5 mM MnCl2 (added fresh for each experiment) and 5 mM MgCl2. For binding and competition experiments, membranes were incubated with [3H]UDP, [3H]UMP, or [3H]UDP-Glc in the presence of DMSO or compounds diluted in DMSO (2% final in assay). Incubations were conducted for 50 min (human P2Y14) or 60 min (chimpanzee P2Y14) at room temperature with shaking to reach equilibrium. Binding in wells containing 2 µL DMSO only (no compound) represents total binding, whereas binding measured in the presence of 1000-fold excess unlabeled nucleotide in DMSO corresponds to nonspecific binding. Incubation mixtures were rapidly filtered on wash buffer preequilibrated GF/C filters (Perkin Elmer), and filters were washed with ice-cold 10 mM Hepes/KOH buffer, pH 7.4, using a Tomtek Mach III harvester (Tomtek, Hamden, CT) to separate bound from free [3H]UDP. GF/C filters were dried at 55 °C in a vacuum oven, and plate bottoms were sealed before addition of 25 µL/well Ultima Gold F scintillation fluid (Perkin Elmer). Filter-bound cpm were detected using a Wallac Microbeta counter (Perkin Elmer) normalized for 3H. Counts (ccpms) generated for nonspecific binding were subtracted from total counts to give specific binding, which was expressed as percentage of total binding. KDs for [3H]UDP in the human and chimpanzee assays were determined by saturation binding. Binding of [3H]UDP to HEK 293 EBNA cell membranes from cells mock transfected with the pdy7 vector was used to demonstrate [3H]UDP binding specificity to P2Y14.
Data analysis
Affinity (KD) and maximum binding (Bmax) of [3H]UDP to membranes expressing P2Y14 were calculated using nonlinear curve fitting for one site binding with GraphPad Prism, version 4 (Graphpad Software, San Diego, CA). Kis were calculated using a four-parameter curve-fitting program and the inflection point of the sigmoidal dose-response curve generated for each compound. Agonist dose-response curves were generated and EC50 values were calculated using experimental data fit to a four-parameter logistic equation using GraphPad Prism.
Results and Discussion
Agonist-induced signaling in HEK cells
We were unable to initially demonstrate UDP-sugar nucleotide signaling in HEK cells transiently expressing P2Y14 only. Signaling was observed upon co-expression of P2Y14 and the chimeric G-protein Gαqi5. Consequently, we generated stable HEK clonal cell lines coexpressing P2Y14 and Gαqi5 and monitored calcium signaling using a calcium-binding fluorescent dye and a FLIPR. The cell lines were used to confirm dose-dependent sugar-nucleotide signaling in cells expressing human as well as chimpanzee and mouse P2Y14 (
Fig. 1A
;
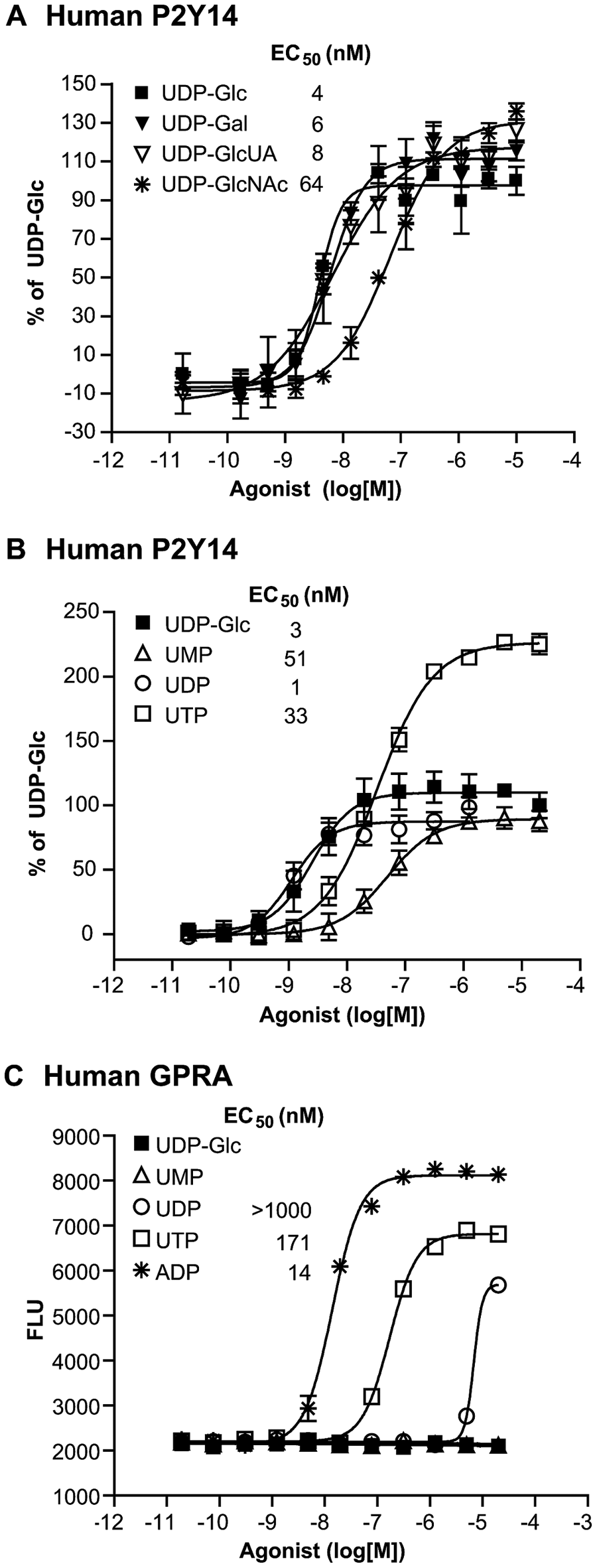
Agonist dose-response profiles in HEK cells stably expressing human P2Y14 or a control GPCR, GPRA, and Gαqi5 in a calcium mobilization assay. HEK cells expressing human P2Y14 and Gαqi5 were incubated in the presence of dose responses of UDP-Glc (■),UDP-Gal (▼), UDP-GlcUA (▽), and UDP-GlcNAc (*) (
Fricks et al. 7 demonstrated that UDP behaves as a competitive antagonist of human but a full agonist of rat P2YR14. Our data illustrating that UDP behaves as a full agonist of chimpanzee and mouse P2Y14 are consistent with Fricks’ rat P2Y14 data; however, we have been unable to demonstrate UDP-mediated antagonism of human P2Y14. Recently, Carter et al. 5 showed that UDP is an agonist of human P2Y14 expressed in HEK 293, C6 glioma, and Chinese hamster ovary cells and that receptor activation results in inhibition of forskolin-stimulated cAMP. We have also shown, using FLIPR and impedance assays, that UDP is an agonist of P2Y14 (expressed in HEK cells concomitantly with Gαqi5), with the expected receptor activation resulting in Ca2+ mobilization. We have never been able to directly detect UDP-Glc– or UDP-mediated inhibition of forskolin-stimulated cAMP using standard biochemical assays, either in the absence or presence of Gαqi5. (Please see the next section for discussion.)
It is not surprising that different expression systems reveal altered ligand efficacy or signaling pattern. It has been shown that one GPCR can couple to a number of intracellular signaling cascades to modulate downstream effectors. In recombinant systems, the stoichiometry of the receptor, G-protein, adapter and accessory proteins, as well as receptor reserve/density and localization can all modulate ligand efficacy. Coupling of a receptor to an alternative G-protein may result in a distinct receptor conformation and signaling, and the expression of a heterologous receptor at high concentrations may alter the signaling pathway through alternative receptor dimerization or by altering the levels and/or localization of the endogenous receptors. Therefore, although UDP behaves as a full agonist in our high-expressing cell line, UDP may show partial agonist activity in cells expressing more physiologically relevant receptor expression levels or in tissues.
Agonist activity in HEK cells expressing P2Y14 and Gαqi5 using CellKey
The Cellkey System, a novel, live-cell, label-free, real-time technology, was used to further corroborate P2Y14 activation by UDP and UMP. This functional assay is based on cellular dielectric spectroscopy (CDS) and monitors changes in impedance of the applied electrical current through and around the cell monolayer upon receptor activation.
10
Sugar nucleotides, UMP, UDP, and UTP, induced signaling in the cellular impedance assay (data not shown;
Several groups have shown that activation of P2Y14 results in signaling through Gi with a consequent inhibition of cAMP and also through Gq with a consequent increase in intracellular calcium.3,5,6,7,11 Cellular dielectric impedance measurements can generate unique response profiles that are characteristic of the specific G-protein coupling of the activated receptor. 10 We have used this technology to illustrate that UDP-Glc, UMP, and UDP stimulation of HEK cells expressing P2Y14 and Gαqi5 resulted in the expected characteristic early, small, and rapid decrease followed by an increase in Ziec, typical of activation of Gαq-coupled receptors ( Fig. 2A ; data not shown). We have also observed a similar Gαq profile in UDP-activated HEK cells expressing P2Y14 without recombinant Gαq protein. In contrast, UDP-Glc– and UMP-induced activation of HEK cells expressing P2Y14 without recombinant G-protein resulted in an increase without the early, small dip in Ziec, characteristic of activation of Gαi-coupled receptors ( Fig. 2B ). Incubation of cells with 100 ng/mL pertussis toxin resulted in the reduction or near abolishment of the UDP-Glc– and UMP-mediated Gi signal, further confirming that P2Y14 signals via inhibition of cAMP in these cells ( Fig. 2B ). We were unable to detect agonist-induced suppression of forskolin-induced cAMP accumulation or ERK signaling in HEK cells expressing P2Y14 using conventional biochemical assays. Perhaps the ability of GPCRs to signal through multiple pathways (Gi, Gq, Gs, Go, arrestin dependent and arrestin independent) and activate multiple downstream effectors is better captured using sensitive cellular real-time kinetic impedance assays that capture the aggregate effect of multiple signaling pathways.
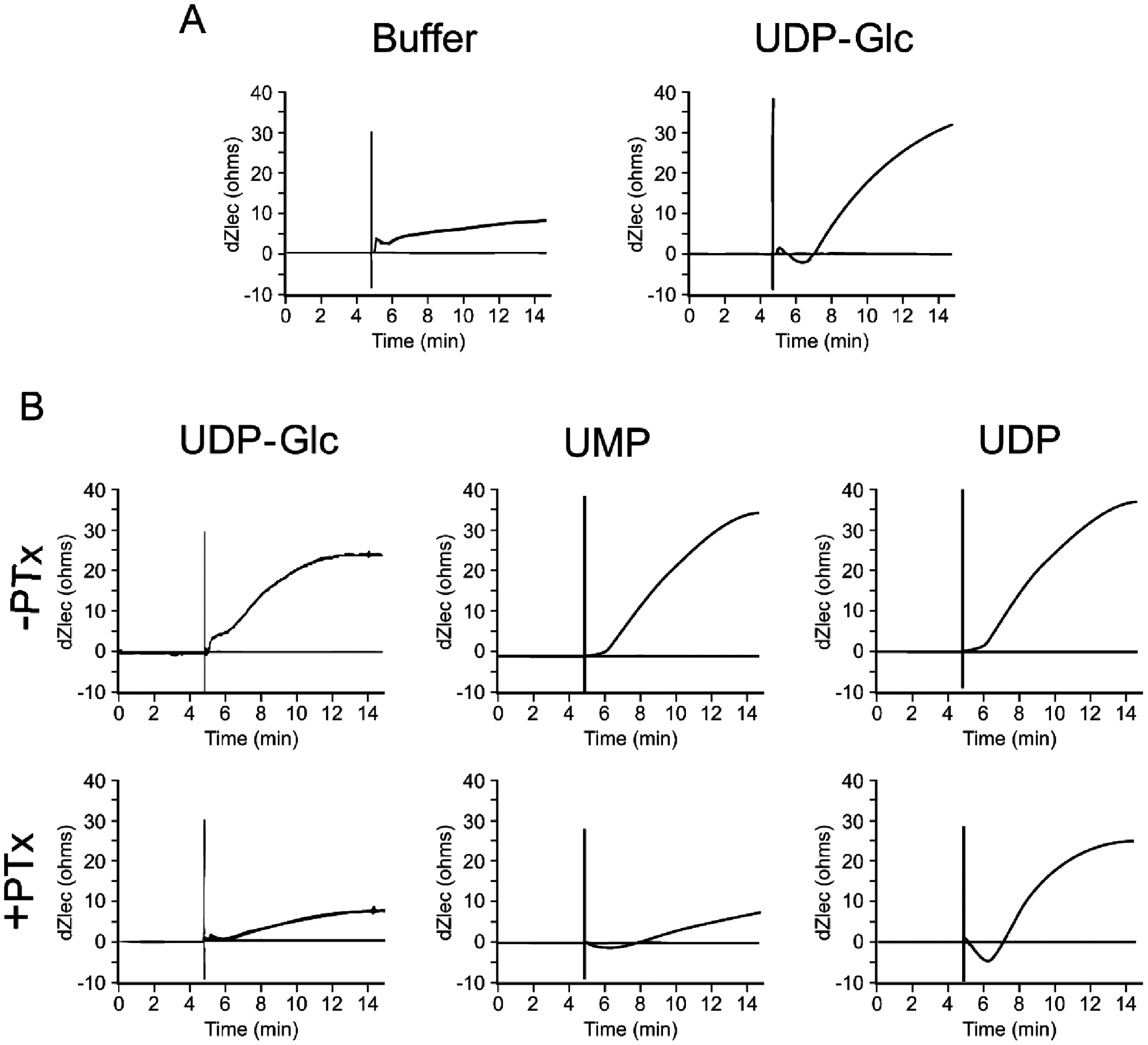
Agonist-induced signaling in the CellKey cellular impedance assay in HEK cells expressing P2Y14 in the absence or presence of Gαqi5. HEK cells stably expressing P2Y14 and Gαqi5 (
Activity of nucleotides, nucleosides, and sugar nucleotides in HEK cells expressing P2Y14 and Gαqi5
Our observation that UDP can activate P2Y14 led us to further investigate ligand specificity for this receptor. Structure activity relationships of purine and pyrimidine nucleotides, nucleosides, and sugar nucleotides in HEK cells expressing either P2Y14 or GPRA and Gαqi5 are shown in
Determination of affinity of P2Y14 ligands
We attempted to establish a membrane filtration binding assay in HEK cells transiently expressing P2Y14 using [3H]UDP-Glc as radioligand; however, we were unable to demonstrate a reasonable specific binding window over mock-transfected cells (data not shown). Based on the observation that the sugar moiety was not essential to activate P2Y14 in the functional assay, we sought to develop a membrane filtration binding assay using [3H]UMP or [3H]UDP as radioligand. We did observe a specific binding window over mock-transfected cells using [3H]UDP and membranes from HEK cells transiently expressing chimpanzee or human P2Y14 (data not shown).
Saturation binding experiments were performed in membranes from HEK cells transiently expressing P2Y14 to assess [3H]UDP affinity and maximum binding to P2Y14. The results of a typical binding isotherm are shown in Figure 3A . A single high-affinity binding site for [3H]UDP was detected in membrane preparations. Results from replicate experiments yielded a mean [3H]UDP KD value of 16.8 nM and Bmax of 112 pmol/mg of membrane protein.
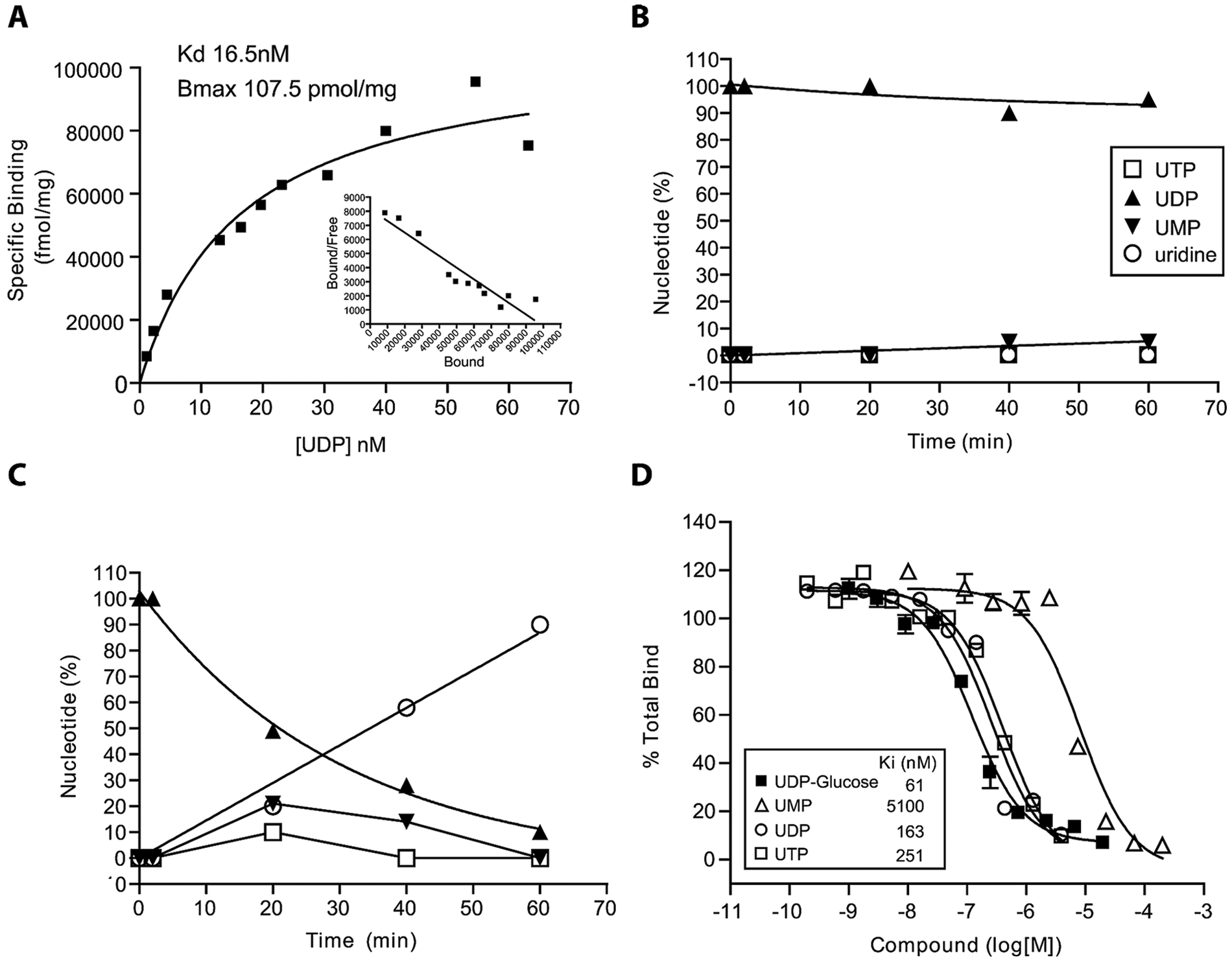
Development of a P2Y14 radioligand binding assay. (
UDP-Glc, UMP, and UDP successfully competed for [3H]UDP binding to chimpanzee P2Y14 in a competition binding assay with similar Ki values for UDP-Glc and UDP ( Fig. 3D ). UMP was significantly less potent in competing for P2Y14 binding with a Ki of approximately 5000 nM ( Fig. 3 ). Similar results were seen using membranes expressing human P2Y14 (data not shown). [3H]UTP was shown to rapidly degrade to [3H]UDP and then remain stable as [3H]UDP for the duration of the binding assay (data not shown), and as a result, the Kis obtained in the binding assay for UTP and UDP were similar ( Fig. 3D ).
Screening of phosphonate library and identification of nonnucleotide P2Y14 antagonist
The difference in activity between UDP and UMP, and IDP and IMP, in their ability to activate P2Y14 expressed in HEK cells along with Gαqi5 (supplemental
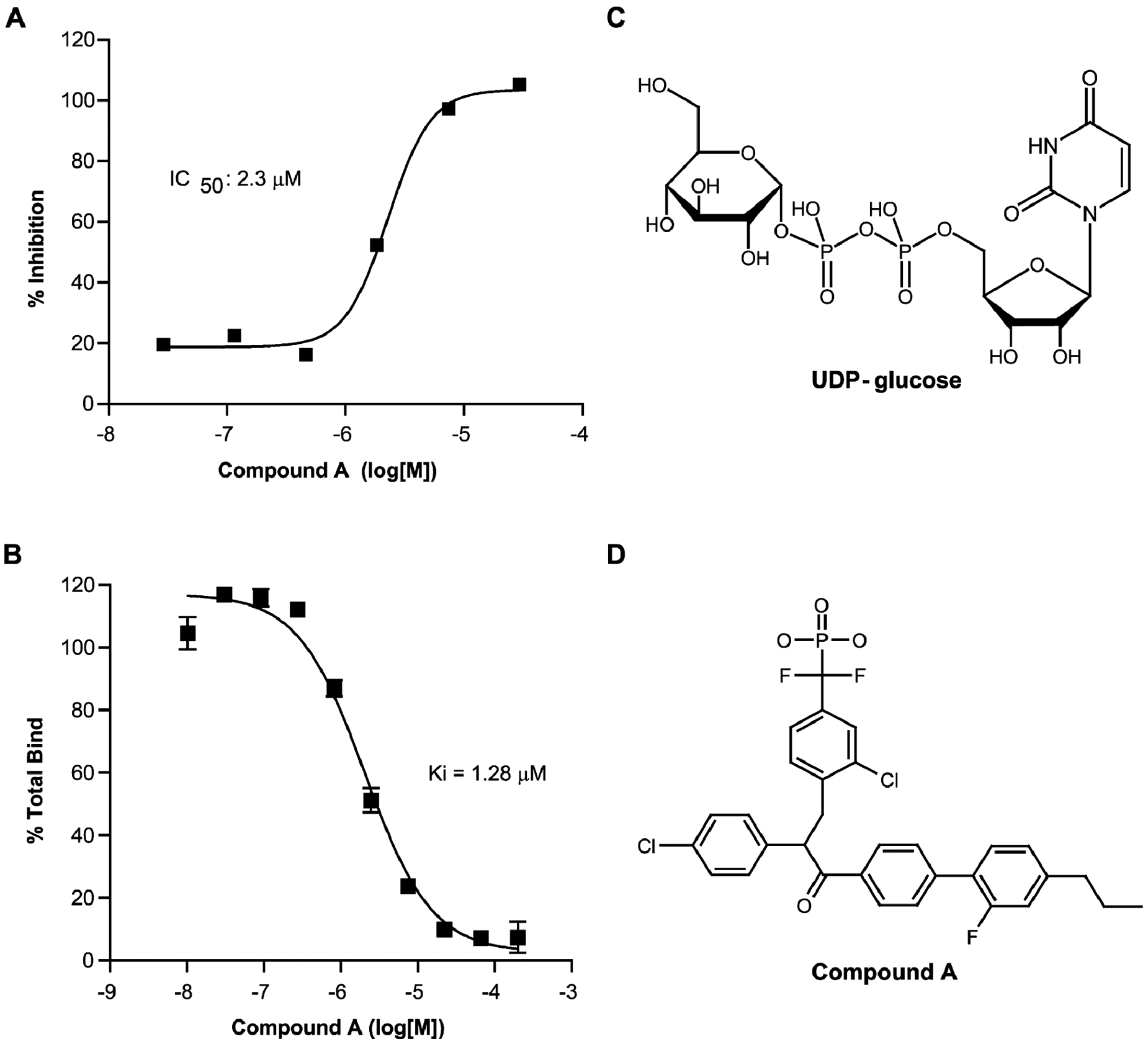
Pharmacological analysis of compound A in HEK cells stably expressing P2Y14 and Gαqi5. (
To our knowledge, this is the first identified nonnucleotide antagonist of P2Y14. It is interesting to compare the antagonist activity of compound A with the agonist activities of the nucleoside monophosphates; AMP, CMP, GMP, and TMP all have IC50 > 20 000 nM, IMP is 5433 nM, and UMP is 54 nM compared with 1280 nM for compound A. Because the diphosphate versions of the nucleotides are more potent, the implication is that the activity of compound A could be improved by altering its structure such that it more closely resembles a diphosphate. This novel binding assay, therefore, is a powerful tool for the identification of potent and selective antagonists of P2Y14, which may help to elucidate the as of yet unknown physiological role of P2Y14.
Footnotes
Acknowledgements
The authors thank Yves Durocher (Biotechnology Research Institute, Montreal, Canada) for the pYD7 expression vector and large-scale preparation of membranes for binding assays and Yael Mamane for the subcloning of human and chimpanzee P2Y14 into pYD7.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
