Abstract
Despite advances toward understanding the prevention and treatment of many cancers, patients who suffer from oral squamous cell carcinoma (OSCC) confront a survival rate that has remained unimproved for more than 2 decades, indicating our ability to treat them pharmacologically has reached a plateau. In an ongoing effort to improve the clinical outlook for this disease, we previously reported that an essential component of the mechanism by which the proteasome inhibitor bortezomib (PS-341, Velcade) induced apoptosis in OSCC required the activation of a terminal unfolded protein response (UPR). Predicated on these studies, the authors hypothesized that high-throughput screening (HTS) of large diverse chemical libraries might identify more potent or selective small-molecule activators of the apoptotic arm of the UPR to control or kill OSCC. They have developed complementary cell-based assays using stably transfected CHO-K1 cell lines that individually assess the PERK/eIF2α/CHOP (apoptotic) or the IRE1/XBP1 (adaptive) UPR subpathways. An ˜66 K compound collection was screened at the University of Michigan Center for Chemical Genomics that included a unique library of prefractionated natural product extracts. The mycotoxin methoxycitrinin was isolated from a natural extract and found to selectively activate the CHOP-luciferase reporter at 80 µM. A series of citrinin derivatives was isolated from these extracts, including a unique congener that has not been previously described. In an effort to identify more potent compounds, the authors examined the ability of citrinin and the structurally related mycotoxins ochratoxin A and patulin to activate the UPR. Strikingly, it was found that patulin at 2.5 to 10 µM induced a terminal UPR in a panel of OSCC cells that was characterized by an increase in CHOP, GADD34, and ATF3 gene expression and XBP1 splicing. A luminescent caspase assay and the induction of several BH3-only genes indicated that patulin could induce apoptosis in OSCC cells. These data support the use of this complementary HTS strategy to identify novel modulators of UPR signaling and tumor cell death.
Keywords
Introduction
O
The UPR is orchestrated through activation of the protein kinases IRE1 and PERK and cleavage of the transcription factor ATF6.4,5 These three endoplasmic reticulum (ER) resident transmembrane stress sensors are maintained in an inactive state through interaction with the ER lumen chaperone BiP/GRP78. Upon accumulation of misfolded protein, BiP dissociates from the sensors in an attempt to facilitate the restoration of homeostatic polypeptide folding. BiP release permits IRE1 and PERK homodimerization, autophosphorylation, and activation. IRE1 and PERK activation initiates parallel yet genetically distinct signaling cascades that culminate in the transcriptional activation of genes encoding ER chaperones and the ER-associated degradation (ERAD) machinery and the attenuation of global protein synthesis. 6 IRE1 activation elicits an endoribonuclease function that mediates a nonconventional splicing reaction on XBP1 mRNA to activate genes encoding ER protein chaperones, lipid biosynthetic enzymes, and multiple factors that function in ERAD.7,8 Activated PERK phosphorylates eIF2α at Ser 51 to inhibit the initial step of polypeptide biosynthesis, leading to general translation attenuation, which has become a hallmark of ER stress.9,10 Recent studies indicate that induction of CHOP, a C/EBP homologous transcription factor that forms heterodimers with other C/EBP family members, following eIF2α phosphorylation is essential for the apoptotic response that occurs when defects with ER protein folding are not resolved.11–13 Although many mechanistic details of CHOP-induced apoptosis remain unknown, it is required for induction of the proapoptotic factors DR5 and TRB3 and can repress the transcription of antiapoptotic BCL2.14,15 Small molecules that could selectively induce proapoptotic CHOP but not adaptive XBP1 signaling are therefore extremely attractive as chemotherapeutics. By enhancing stress, such small molecules might directly induce apoptosis in highly secretory malignant cells or sensitize them to conventional chemotherapies.
The mechanisms by which microenvironmental stresses modulate growth, vascularization, and metastasis in solid tumors remain an avid area of research interest. Recent in vivo studies using luciferase reporters have revealed that ATF4 and XBP1 levels are elevated in the hypoxic regions of xenografts with human fibrosarcoma and primary mammary tumors, respectively, and may predict tumor growth.16,17 Several intriguing reports have together examined approximately 50 OSCC cell lines and observed increased rates of protein synthesis that the authors independently attributed to the overexpression of the translation factors eIF4E and eIF2α.18–21 Considered together, and predicated on our discovery that bortezomib-induced death of OSCC cell lines was dependent on the induction of a terminal UPR, these studies strongly support our hypothesis that small molecules that induce ER stress and overwhelm the hyperactive secretory pathway of OSCC cells will be an effective therapeutic approach to reduce tumor burden. Importantly, targeting the translation machinery of OSCC cells and not a specific signaling cascade or intermediate provides a unique approach whereby the events leading to cell death cannot be short circuited by any redundant signaling mechanism.
Bioactive natural products isolated from plants and microorganisms such as bacteria and fungi have been a rich source of pharmaceuticals, with an estimated 60% of drugs currently on the market deriving from natural sources. 22 In terms of cancer therapies, rapamycin, the vinca alkaloids vincristine and vinblastine, anthracyclines such as daunorubicin, doxorubicin, the quinoline alkaloid camptothecin, and the cyclic diterpene paclitaxel are all naturally derived. Although historically underrepresented in terms of therapeutic use, many studies are currently under way examining the benefit of marine-derived natural products for diverse clinical applications ranging from analgesics to cancer and inflammation. Importantly, the number of compounds isolated from living organisms undergoing preclinical and clinical evaluation continues to expand with many promising candidates, 23 and the recent Food and Drug Administration granting of “orphan drug” status to ET-743 (trabectedin) for soft-tissue sarcomas and ovarian cancer. 24
A collection of 5036 bacterial and fungal-derived prefractionated natural extracts was screened for the ability to activate the CHOP and XBP1 luciferase reporters. Using bioassay-guided fractionation, we identified and isolated methoxycitrinin from an extract that preferentially activated the CHOP reporter. Purified methoxycitrinin, its analog citrinin, and patulin and ochratoxin A, two compounds with similar structural features, were examined for their ability to induce UPR signaling in reporter and gene expression assays. Patulin was found to be a robust UPR activator, whereas citrinin demonstrated activity similar to methoxycitrinin and ochratoxin A was unable to activate either UPR reporter. Patulin is a secondary metabolite produced by Aspergillus and Penicillium species and has been demonstrated to have a cytotoxic effect in human promyelocytic leukemia and hepatoma cell lines via a mechanism that involves perturbation of redox status and inhibition of amino acid uptake, respectively.25,26 Further analyses with a panel of OSCC cell lines revealed activation of caspases and induction of BH3-only proapoptotic genes following low-dose patulin exposure. This study demonstrates the value of HTS of natural product extracts to identify novel modulators of the unfolded protein response and cell death in human OSCC cell lines.
Materials and Methods
Cell lines, reporters, and reagents
For screening, single high-expressing clones that provided Z′ values >0.5 were selected from stably transfected CHO-K1 cells that individually report on either the PERK/eIF2α/CHOP or the IRE1/XBP1 pathway. The CHO-CHOP-luciferase reporter cell line was screened previously. 27 In this cell line, approximately 8 kb of the murine CHOP promoter is upstream of the luciferase coding region and can be induced by ER stress. The CHO-XBP1-luciferase cell line was made by transfection and G-418 selection of CHO-K1 cells with pcDNA3.1-XBP1-luc. pcDNA3.1-XBP1-luc contains a 26 base intron from XBP1 placed into the 5′ end of the luciferase mRNA; luciferase is not translated unless the intron is removed via IRE1-mediated splicing that occurs upon activation of the UPR. 28 The human floor-of-mouth squamous cell carcinoma lines UMSCC1 and UMSCC14A and laryngeal squamous cell carcinoma cell line UMSCC23 were kindly provided by Dr. Thomas Carey at the University of Michigan. The tongue carcinoma cell line CAL27 (CRL-2095), the pharyngeal carcinoma cell line FaDu (HTB-43), and the epidermoid carcinoma cell line A-253 (HTB-41) were from ATCC (Manassas, VA). OSCC cell lines were grown in DMEM supplemented with penicillin and streptomycin and 10% fetal bovine serum. Normal human epidermal keratinocytes (nHEK; ScienCell Research Laboratories, Carlsbad, CA) were grown in keratinocyte medium (#2101) with keratinocyte growth supplement (#2152; ScienCell Research Laboratories). Citrinin (ALX-380-058), patulin (ALX-270-111), and ochratoxin A (ALX-630-089) were purchased from Enzo Life Sciences (San Diego, CA).
HTS
Reporter cells were plated onto white 384-well tissue culture plates (5000 cells/well) using a Multidrop (Thermo Fisher Scientific, Waltham, MA). Plates were incubated for 16 hours at 37 °C, 5% CO2, prior to compound addition. Compounds were added using a high-density replication tool on a Biomek FX liquid handler (Beckman Coulter, Brea, CA) in a 0.2-µL volume (DMSO concentration = 0.4% in all wells). All compounds were tested as singletons. A total of 2.5 µg/mL tunicamycin (Tm) or an equal volume of DMSO was each added to one row of wells on each plate for assay controls. Six to 8 h later, the medium was aspirated to 10 µL with an Elx 405 plate washer (Bio Tek U.S., Winooski, VT), and 10 µL of Steady-Glo (Promega Corp., Madison, WI) was added to each well. Luminescence was measured on a Pherastar multimode plate reader (BMG Labtech, Cary, NC). A hit was defined as a compound able to induce luciferase to a level ≥50% of the tunicamycin control. For the CHOP assay, this definition produced 1064 hits for an overall hit rate of 1.6%. Within the natural products collection, this definition produced 39 hits (0.8%). The 1025 hits (excluding the natural products) were further scrutinized by the Vahlteich Medicinal Chemistry Core synthesis laboratory (VMCC) at the University of Michigan. Compounds were triaged based on their ability to activate reporters in other cell-based assays (including our XBP1 reporter) or if they had molecular weights, total polar surface areas, AlogP values, or known structural alerts that might preclude them from being reasonably considered for therapeutic development. Compounds surviving this level of scrutiny were subjected to confirmatory dose-response assays performed in duplicate with both the CHOP and XBP1 reporter cell lines. The initial triage and subsequent dose-response assays identified 368 bona fide CHOP-selective hits, reducing the overall hit rate for the screen to 0.6%. The 368 compounds were clustered using SARNavigator software at 65%+ similarity, which produced 278 clusters. Signposts for the 278 clusters were provided to the VMCC, where the compounds were prioritized as high (222), medium (75), or low (71) for further study based on their predicted potential for therapeutic development.
Fermentation and extraction
Fungal strain 7792-2H was isolated from marine sediment, collected in Las Baulas Marine National Park in Costa Rica in 2006. The strain was identified based on a phylogenetic sequence analysis of 18S rRNA using the primers ITS1 and ITS4. The rRNA gene sequence data (553 bp) was submitted to a match search using BLAST (http://www.ncbi.nlm.nih.gov/blast/), showing 99% similarity with Penicillium sp. The fungus was cultured for 3 wk at 28 °C in two 6.0 L Fernbach flasks containing 2.5 L of ISP2 medium (1 L of deionized water, 10 g malt extract, 4 g yeast extract, 4 g dextrose, and 30 g NaCl) with shaking at 160 rpm. Cultures were centrifuged at 7000 rpm, and the fermentation medium was extracted with amberlite XAD-16 resin. After shaking overnight, the resin was washed with 1 L of deionized H2O and sequentially extracted with MeOH, acetone, and EtOAc. The resulting extracts were concentrated by rotary evaporation and redissolved in DMSO at 15 mg/mL for storage at −28 °C.
Isolation of compounds 1–6
The methanol and acetone extracts (3.2 g and 2.2 g) were first fractionated by C18 LOBAR B (Ø = 25 × 310 mm) LiChroprep column eluting with MeOH/H2O 7:3. The active fractions were further subjected to high-performance liquid chromatography (HPLC) purification on a XBridge Prep C18 column (Ø = 10 × 250 mm) with MeOH/H2O gradient to afford citrinin 29 together with five more nonactive compounds,30,31 including one that has not been previously described. These structures were determined by a combination of spectroscopic and spectrometric experiments. IR spectra were recorded with a PerkinElmer BX FT-IR infrared spectrometer using NaCl plates. Mass spectra were carried out with a Micromass AutoSpec Yltima Magnetica mass spectrometer. 1H, 13C, and 2D NMR spectra were recorded in CD3OD on a Varian INOVA 600 MHz NMR spectrometer at 600 MHz for 1H NMR and 150 MHz for 13C, using TMS as internal standard.
PCR
Total mRNA was harvested with Trizol (Invitrogen Corp., Carlsbad, CA), and cDNA was made with iScript cDNA kits (BioRad, Hercules, CA). Semiquantitative reverse transcription PCR (RT-PCR) analysis of spliced and unspliced XBP1 was performed with a single human-specific primer pair ACA CGC TTG GGA ATG GAC AC (forward) and CCA TGG GAA GAT GTT CTG GG (reverse); amplicons were visualized with a Qiagen Qiaxcel automated nucleic acid fragment analyzer using a high-resolution cartridge on the M500 setting using a 15 bp to 1 kb alignment marker and a 50 bp to 800 bp size marker. Quantitative real-time RT-PCR (qRT-PCR) was performed following Life Technologies Taqman primer/probe sets: CHOP/DDIT3 (Hs01090850_m1), 18S (Hs99999901_s1), GADD34 (Hs00169585_m1), ATF3 (Hs00910173_m1), ATF4 (Hs00909569 _g1), BiP/GRP78 (Hs99999174_m1), Bax (Hs99999001_m1), Bak (Hs00832876_g1), Bim (Hs00708019_s1), Bid (Hs00609632 _m1), NOXA (Hs00560402_m1), PUMA (Hs00248075_m1), and NBK (Hs00154189_m1), as described. 32
Viability and caspase assays
Cell Titer-Glo Luminescent Cell Viability Assay (G7570) and Caspase-Glo 3/7 Assay (G8090; Promega) were performed with 12 500 cells (in 50 µL) per well in 96-well white opaque cell culture plates, as described. 32
Results
Construction of reporter cell lines
To identify small molecules that modulate specific subpathways of the UPR, we developed two (complementary) cell-based assays using stably transfected CHO-K1 cells that individually report on the PERK/eIF2α/CHOP or the IRE1/XBP1 subpathways of the UPR. Counterscreening with these two cell lines has allowed us to identify compounds that specifically perturb only one pathway and has allowed us to rule out many compounds that generally perturb properties of protein synthesis or folding such as ER lumenal Ca2+ concentration or cellular redox status. The CHO-CHOP-luciferase reporter cell line was previously described and used successfully for a chemical genomic screen. 27 The CHOP reporter construct consists of approximately 8 kb of the murine CHOP promoter cloned immediately upstream of the luciferase coding region ( Fig. 1A ). The CHO-XBP1-luciferase cell line was created by the stable transfection of a reporter that contains a 26 base intron from XBP1 placed into the 5′ end of the luciferase mRNA; luciferase is not translated unless the intron is removed via IRE1-mediated splicing that occurs upon activation of the UPR ( Fig. 1B ). 28 Stable high-expressing clones from each cell line were chosen that provided Z′ values >0.5 for HTS ( Fig. 1C ). 33 The assay was optimized for a 384-well format and screened with a 66 000 compound library that included a collection of 5036 prefractionated natural product extracts. Hit criteria, rates, and chemoinformatic analysis are described in the Materials and Methods section.
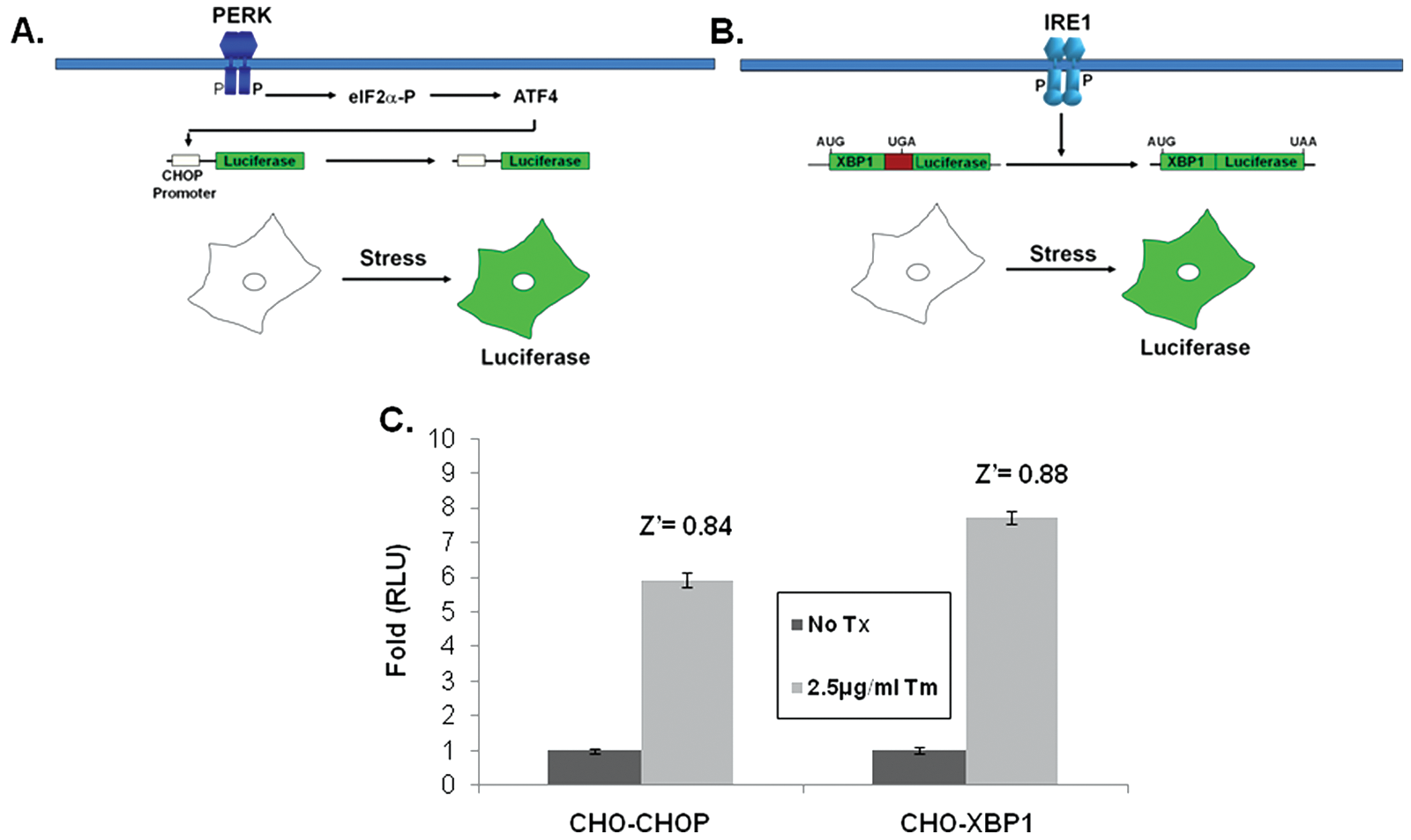
Reporter construct and cell lines. Illustration of the CHOP-luciferase (
Complementary screens identify active natural product extracts
To support our effort to identify bioactive compounds derived from natural products, we focused our attention toward the prefractionated natural products that demonstrated selectivity for the CHOP reporter. The methanol (SID-10794) and acetone (SID-10795) extracts from strain 7792-2 potently activated the CHOP-luciferase but not the XBP1-luciferase reporter, suggesting that active metabolites able to selectively induce the apoptotic arm of the UPR might be present ( Fig. 2A ). Subsequent dose-response assays performed with aliquots of all three extracts used in the primary screen confirmed this activity (data not shown). In an attempt to isolate and identify the active metabolite(s) in SIDs 10794 and 10795, archived spore stocks from the original fungal strain were recultured in 5 L of ISP2 medium, and methanol and acetone extracts were prepared as described. The extracts were separated using a medium-pressure column into eight fractions, and each was retested with triplicate samples in a dose-response fashion (3.75 µg/mL–300 µg/mL) for the ability to activate the CHOP-luciferase reporter. Active fractions were further purified by HPLC and a MeOH adduct of citrinin, which was present as an isomer in equilibrium with the parent compound ( Fig. 2B ) 34 identified by NMR and mass spectrometry analysis ( Fig. 2C ). Citrinin (compound 1) was able to activate the CHOP luciferase reporter only at 15 µg/mL (˜60 µM) and then appeared to be toxic (120 µM+) as evidenced by lower than baseline luminescence. Additional citrinin derivatives that failed to activate the CHOP-luciferase reporter (at the same concentrations) were purified in adjacent fractions, including one that had not been previously described (structure 6; Fig. 2D ). The structure of this new citrinin congener was determined by a combination of HRMS and 1D and 2D NMR experiments including 1H NMR, 13C NMR, COSY, HSQC, HMBC, and NOESY (see supporting information). Purified methoxycitrinin was found to induce the CHOP reporter at 80 µM, and experiments performed with the human OSCC cell line UMSCC1 revealed that methoxycitrinin could induce the expression of the UPR genes CHOP, GADD34, ATF3, ATF4, and BiP after 6 h ( Fig. 2E ). Conventional PCR analysis of the same cDNA failed to detect XBP1 splicing (data not shown), suggesting methoxycitrinin might preferentially activate the apoptotic arm of the UPR.
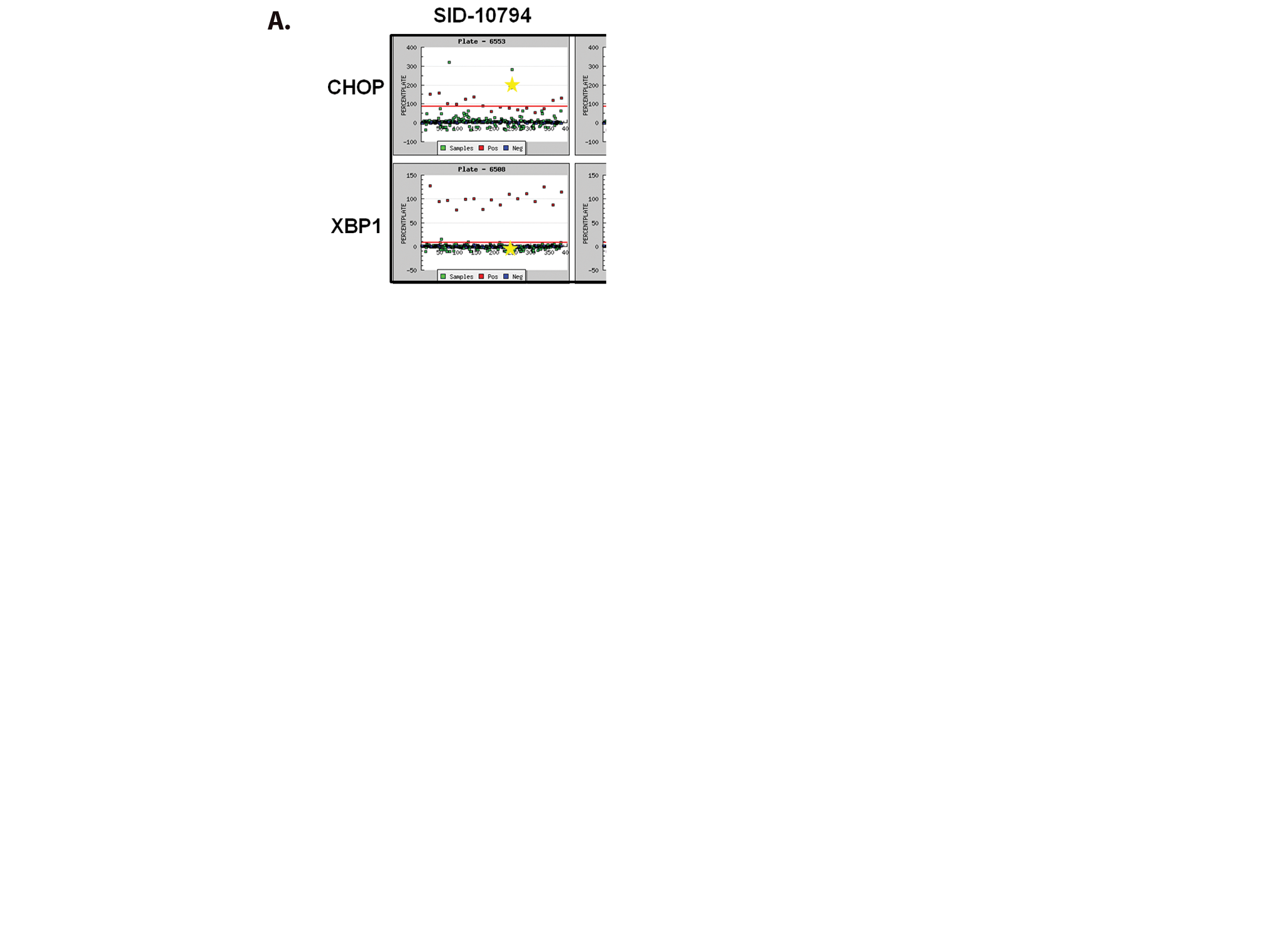
Complementary screens identify CHOP-selective natural extracts. (
Structure-based analog selection identifies more potent compounds
In an attempt to identify more potent UPR-inducing natural products (more potent than methoxycitrinin), we obtained citrinin (Penicillium citrinum), patulin (Penicillium expansum), and ochratoxin A (Aspergillus ochraceus; Fig. 3A ), three mycotoxins that have previously been studied together for their ability to inhibit lymphocyte growth. 35 Similar to purified methoxycitrinin, citrinin could activate the CHOP-luciferase reporter at 80 µM, and ochratoxin A was unable to induce luciferase expression at any concentration tested ( Fig. 3B ). Patulin at 2.5 µM induced a 2-fold increase in luciferase expression and then appeared to be toxic, as shown by lower than baseline luminescence, suggesting patulin might be a more potent UPR activator than any citrinin congener ( Fig. 3B ). To determine the relative ability of citrinin and patulin to modulate the expression of CHOP and other UPR-associated genes, dose-response assays were performed with OSCC cells. UMSCC14A cells were treated with citrinin (20–80 µM) or patulin (1.25–5 µM) for 6 h. Quantitative real-time gene expression analysis revealed that patulin treatment led to a more robust accumulation of mRNA transcripts of CHOP, GADD34, and ATF3 than citrinin ( Fig. 3C and D ). In addition, comparatively low doses of patulin led to an accumulation of the spliced form of XBP1, indicating that this natural product, unlike any citrinin congener, could activate the UPR ( Fig. 3E and F ).
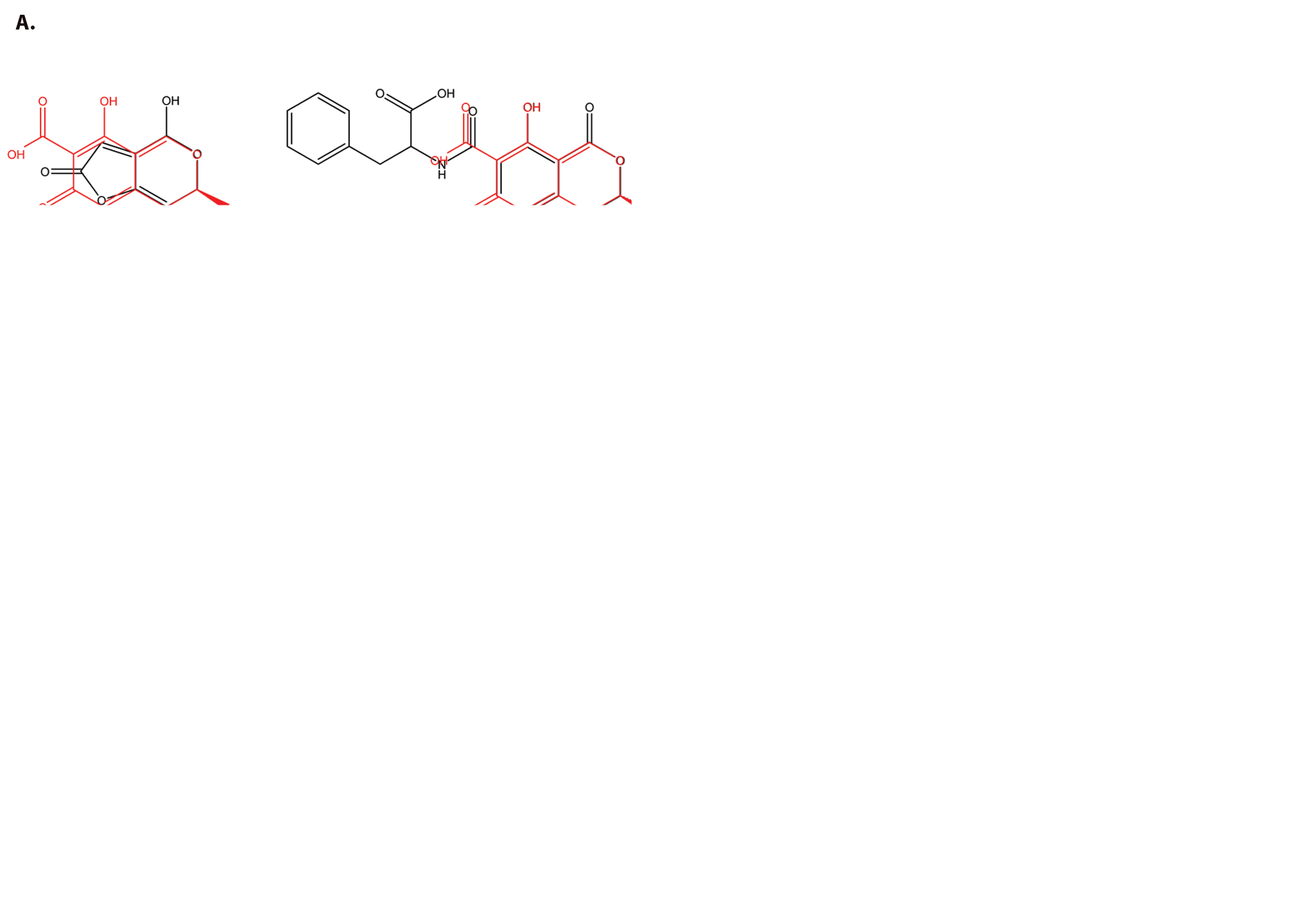
Structure-based analog selection identifies similar mycotoxins. (
Patulin inhibits proliferation of OSCC cells via oxidative stress
Because patulin and citrinin were shown to induce markers of UPR signaling, we next examined their ability to alter the proliferative capacity of a panel of OSCC cells in vitro. CAL 27, A-253, and FaDu cells were cultured with 2.5 to 80 µM citrinin or 0.625 to 20 µM patulin for 24 h. An ATP-based (luciferase) proliferation assay revealed that patulin could substantially inhibit the growth of CAL 27 and A-253 cells but provided only modest inhibition of FaDu ( Fig. 4A ). In addition, UMSCC14A cells were found to be sensitive at a concentration of patulin similar to CAL 27 and A-253 (data not shown). In contrast, these experiments revealed that citrinin could only reduce proliferation in CAL 27 cells at a dose of 80 µM (the same concentration required to activate the CHOP reporter). To determine whether or not patulin was toxic in nonmalignant cells, nHEK were cultured with 1.25 to 10 µM patulin, and proliferation was determined after 8, 16, and 24 h. Modest but significant inhibition of proliferation occurred after 16 and 24 h at 10 µM, a concentration that correlated with CHOP mRNA induction ( Fig. 4B and C ). However, the fact that FaDu cells were able to tolerate higher concentrations of patulin, via an unknown mechanism, suggests this natural product is not generally cytotoxic.
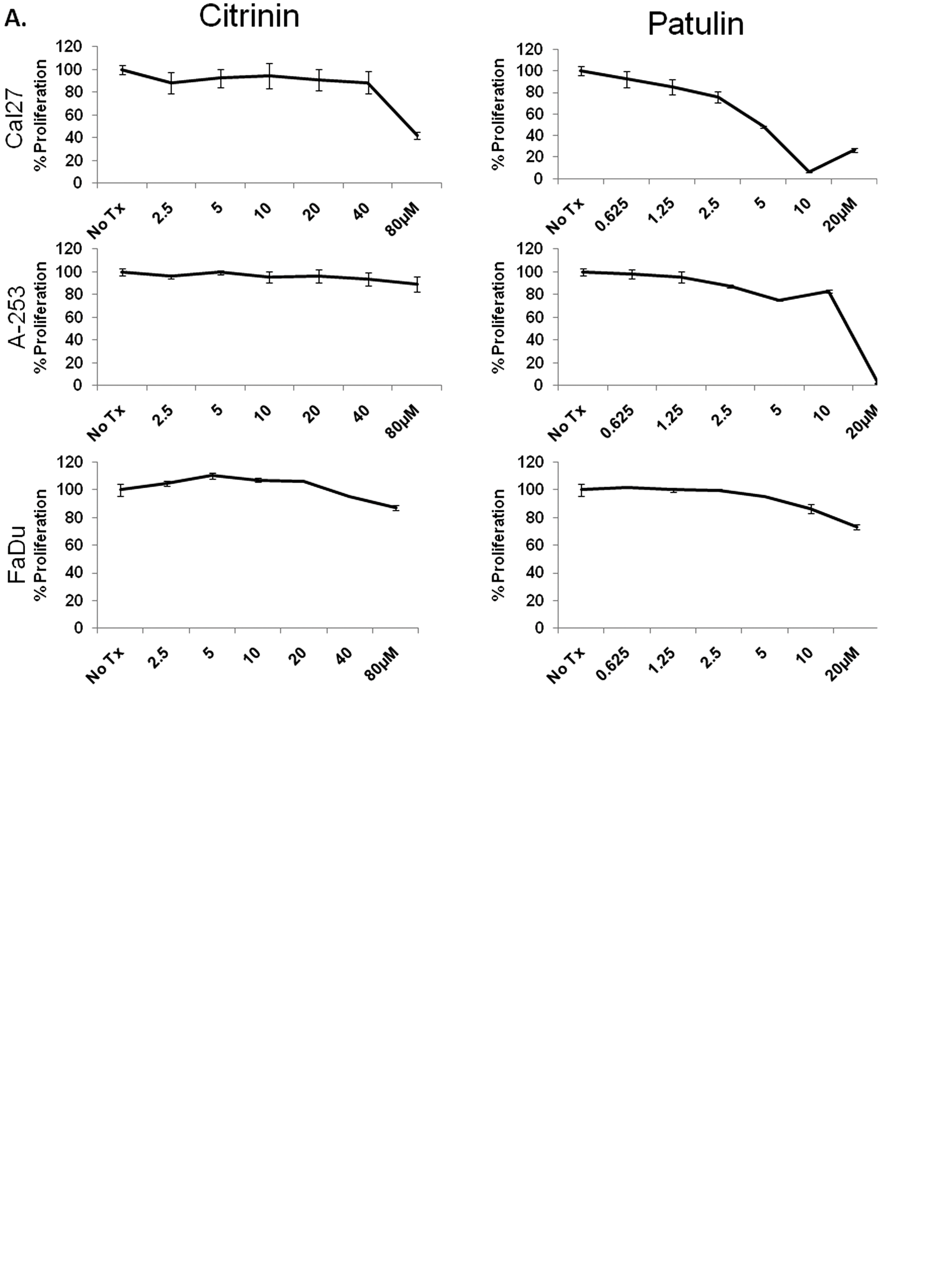
Patulin inhibits proliferation of oral squamous cell carcinoma cells in an oxidative stress–dependent manner. (
Previous studies have revealed that patulin can generate reactive oxygen species (ROS) in Chinese hamster ovary (CHO) cells, peripheral blood lymphocytes, and several human cancer cell lines including HL60 and HEK293.36,37 In light of our previous studies demonstrating that ROS can induce ER stress and the UPR, 2 we sought to determine whether patulin could induce oxidative stress in OSCC. UMSCC1 and UMSCC23 cells were treated with increasing doses of patulin in the presence or absence of the cell-permeable antioxidants tiron or N-acetylcysteine (NAC) for 24 h. Whereas tiron is well regarded as a specific inhibitor of superoxide, NAC is a precursor to glutathione and can increase cellular pools of glutathione and scavenge a variety of ROS, including superoxide. Both cell lines continued to proliferate after 24 h, even at very high concentrations of patulin when cultured with NAC but not tiron ( Fig. 4D and E ). Importantly, supplementing culture medium with essential and or nonessential amino acids could not protect UMSCC1 or UMSCC23 from patulin treatment (data not shown), indicating that the protection provided by NAC was likely the result of its broad free radical scavenging ability and not amino acid repletion.
Patulin induces apoptosis in OSCC cells
The ability of patulin to inhibit the proliferation of OSCC cells in culture prompted us to examine whether it was cytotoxic. CAL 27 and UMSCC1 (patulin-sensitive) and FaDu (patulin-resistant) cells were treated with 5 and 10 µM patulin, and the activation of the initiator caspases 3 and 7 was examined after 4, 8, and 16 h with a luminescent assay. Consistent with the notion that patulin-mediated reduction in proliferation was due to increased cell death, CAL 27 and UMSCC1, but not FaDu cells, displayed a 4- to 7-fold increase in caspase 3/7 activation after 8 h ( Fig. 5A – C ). To better understand the mechanism of patulin-induced apoptosis, the expression profile of a panel of BH3-only proapoptotic BCL2 family members was examined in UMSCC14A cells. Cultures were treated with 2.5 µM patulin for 6 h. qRT-PCR) analysis of Trizol-harvested mRNAs revealed an upregulation of NOXA, PUMA Bim, Bid, and NBK, which are consistent with apoptotic genes associated with UPR-mediated apoptosis ( Fig. 5D ).
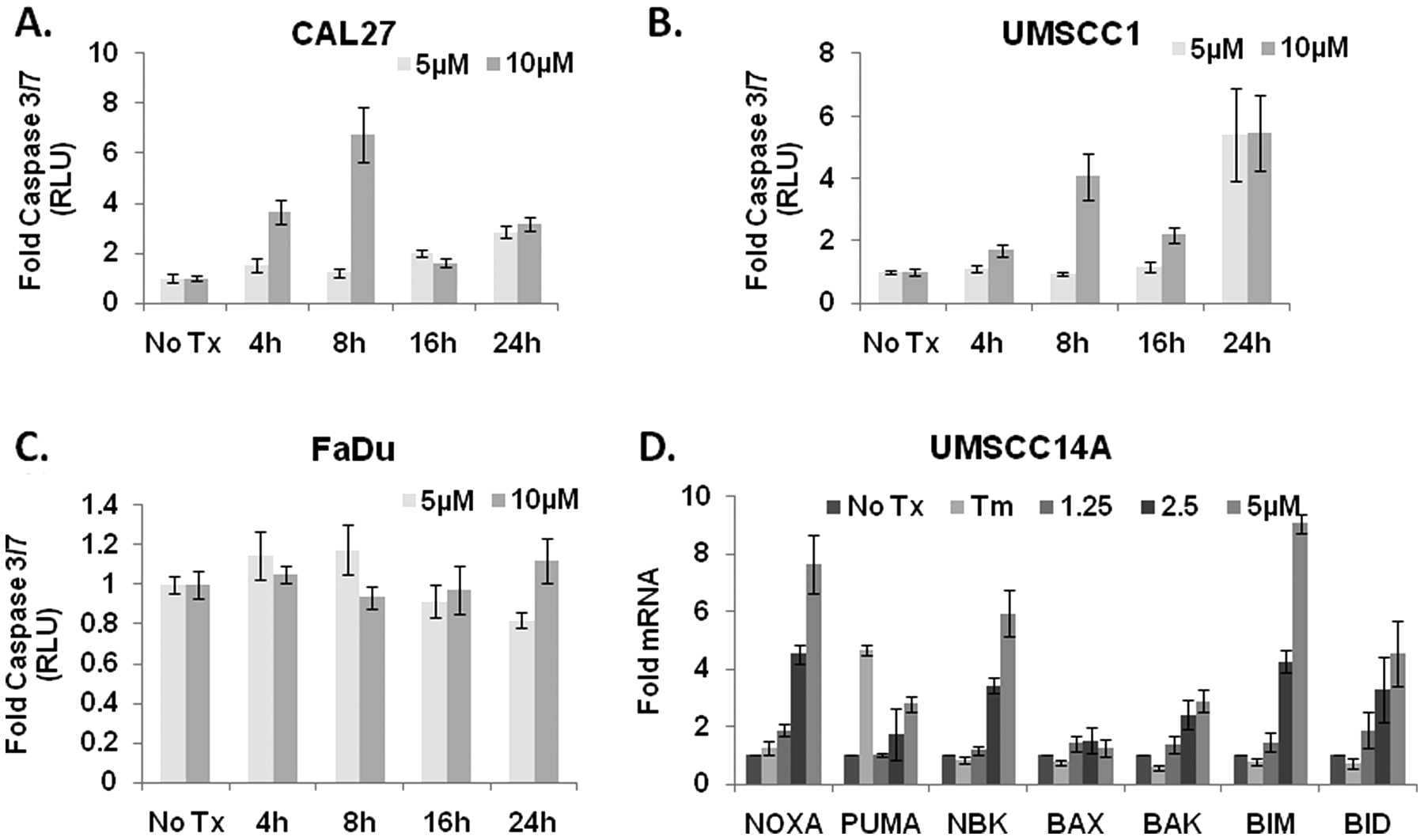
Patulin induces apoptosis in oral squamous cell carcinoma cells. (
Discussion
The recent introduction of affordable, user-friendly screening technologies has facilitated the development of sensitive cell-based and biochemical high-throughput assays within the realm of academia. The application of HTS in an academic setting offers promise for the development of novel therapeutics and encourages the continued investigation of compounds that are not likely to succeed in the clinic as molecular tools to address fundamental biological questions. In the current study, a cell-based HTS strategy was designed to identify natural products that could specifically activate the apoptotic arm of the UPR from prefractionated extracts. Using two cell lines that individually reported on the apoptotic or adaptive stress responses facilitated the rapid identification of bona fide UPR subpathway-specific hits. An additional advantage provided by the use of such an assay (cell-based) is the mitigation of challenges associated with hit compound permeability and ATP-dependent efflux once it passes the membrane. We identified methoxycitrinin and five additional congeners, including one not previously described; however, when purified, very high concentrations were required to induce the UPR, and they were not appreciably toxic.
In an attempt to find more potent mycotoxins, we obtained the structurally related natural products ochratoxin A and patulin to determine their corresponding biological activity profile in this assay system. Patulin treatment increased the expression of CHOP, GADD34, ATF3, ATF4, and BiP and led to XBP1 splicing in a dose-dependent fashion, indicating that it could induce the UPR in OSCC cells. Low-dose patulin-induced stress was robust enough in five of the six OSCC cell lines examined to overwhelm the adaptive capacity of the UPR and induce apoptosis. Patulin was also observed to increase transcripts for CHOP and induce significant cell death in normal human epidermal keratinocytes after 24 h at a rate that was similar to malignant cells. Although these experiments indicated that patulin did not exhibit selectivity for malignant cells, the fact that FaDu and UMSCC23 could tolerate higher doses indicates that patulin is not generally cytotoxic. The mechanism by which these two cell lines were resistant to patulin treatment is not known. It is possible that a variety of mutational events in genetically unstable cancer cells could alter the ability of CHOP or ATF4 protein to be expressed following stress or that lower than average basal levels of stress or even aneuploidy contributed to the resistance. In addition, it is plausible that slower doubling times and reduced protein synthesis rates in culture could limit their dependence on the UPR for survival and narrow the therapeutic window of any apoptosis-inducing compounds.
The constant pressure on cancer cells to meet enhanced oxygen and nutrient demands likely depends on enhanced basal activation of the UPR for vascularization and proliferation. OSCCs were selected for the present work because many highly secretory OSCC cell lines and fresh surgical biopsies have been shown to express increased levels of the eukaryotic translation factors eIF4E and eIf2α. This important finding and our own work demonstrating the ability of the proteasome inhibitor bortezomib to induce UPR-dependent cell death in a panel of OSCC cells has driven the hypothesis that malignant cell populations might be preferentially sensitive to UPR-inducing compounds. In addition, epithelial cell-derived tumors may be more susceptible to UPR-inducing compounds because of increased rates of protein secretion. Therapies that target oncogene-driven growth signals are often rendered ineffective because of redundant molecular networks for these intermediates. The continued identification and investigation of agents that interfere with protein folding homeostasis will shed light on whether they may be leveraged against the reliance of cancer cells (vs. nonmalignant adjacent cells) on UPR and escape the need for neoplastic cell targeting. In the present work, we have shown that following patulin treatment, the UPR is activated in numerous OSCC cell lines.
Mycotoxins are metabolites or toxins secreted from filamentous fungi such as Penicillium, Aspergillus, Fusarium, and associated genera. 38 Patulin is a common mycotoxin, contaminating apples and other commercial fruit crops, that has been studied for more than 7 decades, including early work that examined patulin as a potential cancer therapy and a large multicenter trial using it as a nasal rinse to fight the common cold. 39 Patulin has been shown to interfere with glutathiones and induce oxidative damage, resulting in the formation of DNA adducts and strand breaks, and it is toxic to cancer cells.40,41 Consistent with this mechanism, we found that NAC, a precursor to glutathione, protected OSCC cells from patulin, whereas tiron, a superoxide-specific scavenger, was not protective, suggesting a specific pattern of oxidative stress is required for patulin-induced toxicity. We have previously shown that tiron could protect OSCCs from bortezomib, 2 suggesting varied oxidative stress can induce CHOP-dependent cell death and additionally that CHOP is also involved in the production of oxidative stress, suggesting a complex interplay between ER and oxidative stress. 42 It is likely that fungi developed the ability to secrete generally toxic oxidative stress–inducing agents in response to environmental stress as a survival mechanism. 43 Patulin has also been shown to interfere with uptake of amino acids and protein production. 26 Taken together, our results indicate that patulin likely induces an oxidative stress that culminates in the activation of ER stress, the UPR, and eventually cell death.
Despite its interesting effects on the UPR, we do not believe that patulin is a promising lead for anticancer drug development in an unmodified form. Previous work on the mechanism of action of patulin has indicated that it is an efficient and nonspecific trap for cellular thiols, including glutathione.40,41 Close structural analogs of patulin, lacking its inherent electrophilicity, are much less active as cytotoxins, 44 suggesting that the mechanism of action is largely, if not entirely, dependent on reaction with cellular thiols. In general, thiol reactivity is unlikely to be a mechanism of action that will be highly selective for cancer cells. In fact, patulin has been reported to be a general genotoxin and carcinogen, as well as having negative effects on multiple organs and inducing DNA damage and apoptosis upon contact with skin. 45
The results reported here provide an example of the power of HTS to identify natural products that target specific cellular pathways involved in UPR signaling and malignancy. Additional efforts to identify molecular probes and potential new therapeutic agents will undoubtedly be uncovered by further efforts to harness the broad chemical diversity from unique sources of microbial-derived secondary metabolites.
Footnotes
Acknowledgements
The authors wish to thank Luis Guillermo Acosta and Frank Gonzalez (Bioprospecting Unit, INBio). The marine sediment sample was obtained under the permit R-CM-INBio-03-2006-OT.
Portions of this work were supported by National Institutes of Health (NIH) grants DK042394, HL052173, and HL057346, as well as MH084182 and MH089782 (R.J.K.) and DE019678 (A.M.F.). P.G.C. gratefully acknowledges the Spanish Foundation of Science and Technology (FECYT) for a postdoctoral fellowship. Additional aspects of this work were supported by NIH grant U01 TW007404 as part of the International Cooperative Biodiversity Group initiative at the Fogarty International Center (D.H.S. and G.T.-C.) and the H. W. Vahlteich Professorship (D.H.S.).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
