Abstract
In this study, the authors have generated a tapeworm Taenia solium genomic DNA expression library where foreign peptides/proteins were fused to N-termini of M13 cpVIII and expressed at a high copy number on the phage surface, and they showed that this library may be used in bioselection against antipathogen immune sera, allowing the identification of disease-related antigens recognizing antibodies present in clinical samples. They isolated 2 phage clones expressing T. solium–derived antigens specifically reacting with antibodies present in plasma and cerebrospinal fluid samples of neuroimaging-confirmed neurocysticercosis patients. The described antigen discovery strategy may be used for the direct identification of antigens useful for host-pathogen interaction studies as well as for the development of molecular vaccines and diagnostics.
Introduction
N
Phage display technology relies on a physical link between the selected or engineered peptide/protein expressed on the bacteriophage surface (most commonly M13 phage) and the phage DNA encoding them. 12 In this system, foreign molecules are usually expressed as fusions with either M13 minor coat protein cpIII (3-5 copies per phage) or with the major coat protein cpVIII (2700 copies). The generation of large phage libraries expressing random peptides, antibody fragments, genomic-DNA fragments, and cDNAs and their application in biopanning procedure allow rapid identification of phage-displayed ligands and receptors involved in protein-protein, antigen-antibody, DNA-protein, enzyme-substrate, and other biologically important interactions. 13-15 Phage-displayed random peptide as well as genomic DNA expression libraries derived from pathogens are becoming important sources for the identification of genes coding for proteins or their domains involved in pathogenesis and host-pathogen interactions as well as for isolation of antigens with diagnostic or vaccine candidate properties. 16-18
In the present study, we have generated a T. solium genomic DNA expression library where foreign peptides/proteins were fused to N-termini of M13 cpVIII and expressed at a high copy number on the phage surface. To isolate phage clones expressing pathogen-derived antigens with immunodiagnostic and/or immunotherapeutic potential for NCC, the library was used in biopanning and enzyme-linked immunosorbent assay (ELISA)–based screening assays with mouse anti–T. crassiceps hyperimmune sera. Two clones were selected for evaluation of their immunoreactivity with a panel of cerebrospinal fluid (CSF) and plasma samples from patients with confirmed NCC. The described antigen discovery strategy can be used for the direct identification of antigens useful for host-pathogen interaction studies as well as for molecular vaccine and diagnostics development.
Materials and Methods
Construction of a phage display library
All molecular biology procedures were carried out using standard protocols or as recommended by manufacturers. Restriction enzymes, DNA isolation/purification kits, T4 DNA polymerase, T4 DNA ligase, calf intestine alkaline phosphatase (CIAP), and M13KO7 helper phage were obtained from Invitrogen (Carlsbad, CA), Qiagen (Valencia, CA), or GibcoBRL (Rockville, MD). The oligonucleotides were synthesized at GibcoBRL. The T. solium genomic DNA of cysticerci dissected from skeletal muscle of infected pigs was isolated using the QIAamp DNA Kit (Qiagen) and used as described. 19 Briefly, about 10 µg of chromosomal DNA was fragmented randomly by sonication 5 times for 20 s at maximum power with a sonicator Aquasonic 50T (VWR Scientific Products, West Chester, PA). The DNA fragments (100-600 bp) were made blunt ended using T4 DNA polymerase, purified by the Concert Rapid PCR purification System (GibcoBRL), and used in a ligation reaction with 2 µg of SnaBI digested and dephosphorylated pG8SAET vector DNA, using T4 DNA ligase. 20 The ligated DNA was column purified and used to transform Escherichia coli TG1 cells by electroporation using the Gene Pulser II System (Bio-Rad Laboratories, Hercules, CA). Eight electroporations were performed, and the transformed TG1 cells were plated on LB-Amp plates to determine the diversity of the library. The resultant phagemid library was rescued/amplified using M13KO7 helper phage, purified by double PEG/NaCl (20% w/v polyethylene glycol l-8000; 2.5 M NaCl) precipitation and resuspended in Tris-buffered saline (TBS). The phage yield was 1012 colony-forming units (cfu) per milliliter of culture medium.
Affinity selection of phage clones expressing T. solium–derived antigens by biopanning and ELISA screening
Six-week-old female BALB/cAnN mice, bred in our animal facilities, were immunized with T. crassiceps soluble antigens (TcAg) as described previously. 21 Hyperimmune sera from 3 mice were pooled, diluted 1:200 in phosphate-buffered saline (PBS) containing 0.1% bovine serum albumin (BSA), and used in a biopanning assay. Phage biopanning was carried out essentially as described in our previous studies. 21 Briefly, the wells of Nunc Maxisorp microplates were coated with goat antimouse IgG (0.5 µg/well; Zymed, South San Francisco, CA), washed with PBS/Tween 0.05%, and blocked with PBS/2% BSA. Then, mice sera described above were added to 4 wells, and after washing with PBS-Tween, 1010 phage particles per well from the genomic DNA phage library were added. After incubation for 4 h at 4°C, the plate was washed with cold PBS-Tween, and bound phage particles were eluted using glycine-HCl (0.2 M, pH 2.2) and neutralized by adding Tris-HCl (1 M, pH 9.1). The eluted phages were plated on LB-Amp plates, and individual colonies were rescued/amplified using M13KO7 helper phage and used in ELISA screening. Briefly, the bacterial colonies were used to inoculate 96-well cell culture plates (Costar, Corning, NY) containing 150 µL of 2-YT-Amp. Plates were incubated for 4 h at 37°C, and helper phage was added to each well. After 30 min of incubation without shaking and 30 min of incubation with shaking, plates were centrifuged and supernatants were removed. Then, 150 µL of 2-YT-Amp-Kan was added to the cells, and the plates were incubated overnight at 37°C. The supernatants from plates containing the phage were used in ELISA screening with mice anti–T. crassiceps sera as described below. Also, DNA was purified from these clones, and the DNA inserts were amplified by PCR using pG8SAET vector-based primers and analyzed on agarose gel.
Cerebrospinal fluid and plasma samples
Seventeen CSF and 22 plasma samples from patients with NCC confirmed by CT and MRI were obtained at the Instituto Nacional de Neurologia y Neurocirugia, Mexico City. All samples were positive in ELISA with T. solium cyst fluid antigens (TsAg). CSF samples from 18 patients with encephalitis and plasma from 7 healthy individuals were used as negative controls.
Enzyme-linked immunosorbent assay
ELISAs were carried out as previously described. 5,6,19 Briefly, 1010 cfu of amplified phage was added to microplates (Nunc), incubated for 1 h at 37°C, and washed with PBS and PBS-Tween 0.1%. After blocking with PBS–2% BSA, CSF and plasma samples diluted at 1:40 and 1:100 in PBS–1% BSA, respectively, were added to the wells and incubated overnight at 4°C. Horseradish peroxidase (HRP)–conjugated antihuman IgG diluted in PBS–1% BSA (1:5000) was added to the wells and incubated 1 h at 37°C. The plates were washed, and ABTS single solution (Zymed) was added. The OD readings at 405 nm were registered using the Opsys MR Microplate Reader (DYNEX Technologies, Inc., Chantilly, VA). The Ig subtypes present in plasma samples from NCC patients were determined using recombinant phage clones and HRP-mouse antihuman IgG1, IgG2a, IgG3, IgG4, and IgE Abs (Zymed). All samples were tested in duplicate, and the reported data are representative of 3 independent experiments. Similarly, the plates were coated with TcAg (1 µg/well), and ELISA was performed using samples from NCC patients as described above. For ELISA screening, the plates were coated with phage particles and incubated with mice hyperimmune sera, and HRP/mouse anti-IgG conjugate was added to the wells.
DNA sequencing
The DNA sequences of the inserts of phage clones were determined using automated the ABI Prism 310 Genetic Analyzer (Applied Biosystems, Foster City, CA), miniprep-purified double-stranded DNA, and a pair of pG8SAET vector-based primers 5′(ggaaaaacatttattcaattcgtaaactag) and 3′(ggtttatcagcttgctttcgaggtgaatttc). The DNA and deduced amino acid sequences were analyzed with the ExPASy Molecular Biology server and BLAST database. The DNA and amino acid sequences of isolated clones were submitted to the GenBank database.
Statistical analysis
The correlation between variables was verified by using the chi-square test for contingency tables.
Results
Generation of T. solium genomic DNA phage display library and biopanning with hyperimmune mouse sera
To generate the genomic DNA phage library, T. solium genomic DNA was randomly fragmented by sonication (100- to 600-bp DNA fragments) and cloned in pG8SAET phagemid vector as 5′ gene fusions with M13 gVIII. A genomic expression library with the complexity of about 1.25 × 107 independent clones was generated, and the rescue/amplification of the library resulted in a phage library of 1012 clones.
To isolate phage clones specifically recognizing Abs in the hyperimmune sera, a 2-step selection strategy consisting of biopanning and ELISA screening was applied. Then, 4 × 1010 phage particles from the library were biopanned against the hyperimmune sera from mice immunized with TcAg. Forty individual clones were randomly picked from the library after the first round of selection and used in an ELISA screening with the mouse sera. This resulted in isolation of 11 positive clones. PCR and NcoI/BamHI enzyme digestion analysis showed that all clones were carrying 100- to 400-bp DNA inserts (data not shown). Three of 11 clones were selected based on their highest reactivity in ELISA with 5 CSF samples from NCC patients and further characterized. The DNA and the deduced amino acid (aa) sequence analysis of these clones revealed that 2 clones expressing a 54 aa peptide (clone EK26, GenBank accession number AY699707) were identical, and the clone designated EK5 (GenBank accession number AY699706) carried a 63 aa peptide. Interestingly, for the correct expression of identified open reading frames, a –1 shift at 3′ of the DNA insert in EK5 and a +1 shift at both ends of the insert in EK26 are required. This is not unusual since the same phenomenon was regularly observed, possibly as a result of ribosomal slippage, in studies where the expression system and vector used in our study were analyzed. 19,22 Sequence similarity search in the public sequence databases revealed a homology (36% of identical and 68% of positive aa in a 41 aa long stretch) between EK5 and a hypothetical protein of intestinal parasite Giardia lamblia (G. lamblia ATCC 50803; hypothetical protein GL50803_112258, GenBank), whereas EK26 did not show any significant homology to reported sequences.
Immunoreactivity of phage clones
For the evaluation of the antigenic properties of isolated phage clones, their immunoreactivities were tested in an ELISA using 17 CSF and 22 plasma samples from patients with NCC confirmed by CT and MRI. As shown in
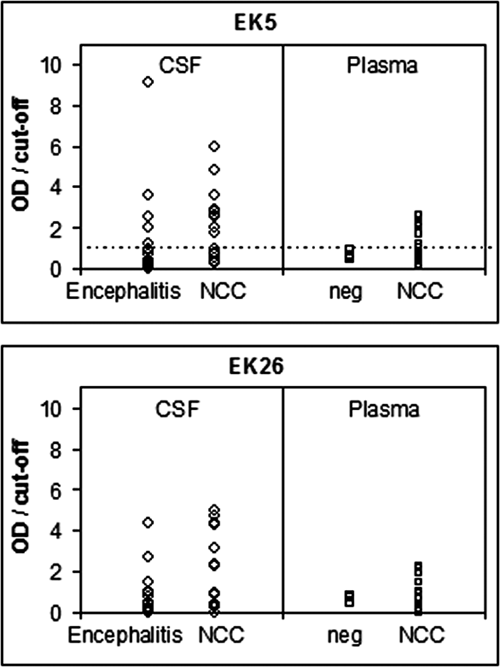
Enzyme-linked immunosorbent assay binding of phage clones EK5 and EK26 to antibodies present in cerebrospinal fluid (CSF) samples from 17 patients with confirmed neurocysticercosis and 18 patients with encephalitis and plasma samples from 22 patients with neurocysticercosis and 7 control healthy individuals. Each sample was also tested against M13 control phage, and a final OD value is a difference between phage-bearing peptide and control M13. Data are expressed as OD/cutoff values, so values above 1 (dashed line) are positive cases and values below 1 negative cases. The cutoff values were determined as the mean + 2 SD of negative controls. NCC, neurocysticercosis.
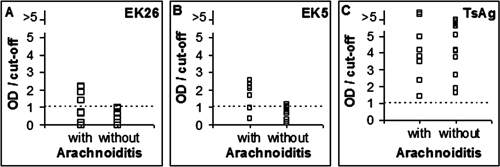

Enzyme-linked immunosorbent assay binding of phage clones EK26 and EK5 and TcAg to antibodies present in plasma samples from patients with and without arachnoiditis (
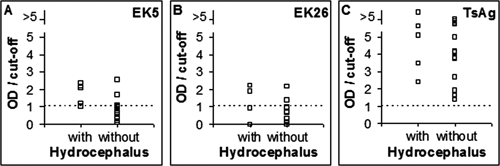
Enzyme-linked immunosorbent assay binding of phage clones EK5 and EK26 and TcAg to antibodies in plasma samples from patients with and without hydrocephalus (
The IgG subtyping showed that the predominant subtype present in plasma samples from NCC patients was IgG1 (data not shown).
Discussion
Neurocysticercosis is the most frequent parasitic disease of the human nervous system. 1,3,5 The diagnosis of neurocysticercosis is based on CT and MRI, which are expensive and often not accessible in endemic areas. 4 Also, there is no available vaccine for the prevention of disease. Thus, there is a clear need for the search for T. solium–related recombinant antigens to create an antigen cocktail offering a highly sensitive and specific immunodiagnostic assay for NCC as well as effective vaccine for the prevention of the disease.
In our study, a T. solium genomic DNA phage library was created in the phagemid vector in such a manner that the recombinant peptides/proteins are expressed as N-terminal fusions with the pVIII capsid protein in M13 bacteriophage. We used the M13 cpVIII-based instead of the cpIII- based phagemid system because it is much more efficient for the selection of ligand-binding phages, as has been shown by isolating Staphylococcus epidermidis cell receptor genes. 23
Despite the existence of previous reports describing the feasibility of using genomic DNA phage display libraries to isolate ligand-binding proteins or their domains, there are few reports describing the identification of antigens from phage-displayed genomic DNA libraries using pathogen- specific sera or monoclonal antibodies. 18,24,25 Thus, pathogen-related antigens have been isolated using mono- or polyclonal antibodies generated against Mycoplasma hyopneumoniae, 16 Mycoplasma mycoids, 17 and H5N1 avian influenza virus. 26 Here, we show that a phage display T. solium genomic DNA library can be used in bioselection against antipathogen immune sera, allowing the identification of antigens recognizing antibodies present in clinical samples. We isolated 2 phage clones expressing T. solium–derived antigens specifically reacting with antibodies present in CSF and plasma samples of neuroimaging-confirmed NCC patients. To improve the accuracy of diagnostic phage-based ELISA, it may be necessary to create, as mentioned above, an antigen cocktail containing several phage clones. Alternatively, due to the presence of anti-M13 antibodies in human sera, 27 the generation of antigens identified in this study as phage-free peptides/proteins probably will result in more sensitive and accurate ELISA. 28
Interestingly, the observation that phage EK26 may preferentially recognize antibodies present in plasma from NCC patients with arachnoiditis and that phage EK5 may preferentially recognize Abs present in plasma from NCC patients with hydrocephalus indicates that identified molecules may serve as potential molecular markers for these clinical manifestations, which are the two most severe forms of NCC and usually require therapy with a combination of corticosteroids, antiparasitic drugs, and/or surgical procedures.
Although we assume that immunodiagnostic tests cannot replace neuroimaging for the clinical management of NCC, 4 they may provide the only tool for diagnosis of infection in endemic areas. The genomic phage library screening is particularly suitable for identification of antigens from pathogens with complex life cycles and limited availability of certain parasite developmental stages, as is the case with Toxoplasma gondii 21 and T. solium. In a recent study, we have isolated NCC-specific T. crassiceps phage clones from a T. crassiceps phage- displayed cDNA expression library after bioselection against hyperimmune mouse sera. 29 One of the isolated clones expressed N-terminal 16 aa peptide of thioredoxin from which 13 aa were identical to the N-terminal region of thioredoxin from Echinococcus granulosus. This selection technique may find application as an alternative genome-wide proteomic approach for the elucidation of protein function since the rapid accumulation of DNA sequence data from genomic projects has led to an increasing demand for powerful tools for functional analysis. 15 Interestingly, the sequence of EK5 showed a significant homology with a protein from G. lamblia, the most frequently isolated intestinal parasite worldwide. The lack of homology between sequences of the clones isolated in this study and reported T. solium and T. crassiceps sequences was not unexpected since there is scarce information about sequences of these parasites yet.
It is important also to emphasize that filamentous phage have clear advantages over conventional recombinant antigens or purified antigenic fractions used in immunodiagnostic or vaccines tools since their large-scale production is simple and highly cost-effective, permitting rapid purification of homogeneous and stable antigen preparations.
Finally, we have reported for the first time the feasibility of using a genomic phage display library to isolate pathogen-derived antigens with potential diagnostic value through bioselection with immune sera. The described antigen discovery strategy can be used for the direct identification of antigens useful for host-pathogen interaction studies as well as for molecular vaccine and diagnostics development.
Footnotes
Acknowledgements
The authors thank M. E. Munguia and G. Acero for technical assistance.
