Abstract
The development of cell-based assays for high-throughput screening (HTS) approaches often requires the generation of stable transformant cell lines. However, these cell lines are essentially created by random integration of a gene of interest (GOI) with no control over the level and stability of gene expression. The authors developed a targeted integration system in Chinese hamster ovary (CHO) cells, called the cellular genome positioning system (cGPS), based on the stimulation of homologous gene targeting by meganucleases. Five different GOIs were knocked in at the same locus in cGPS CHO-K1 cells. Further characterization revealed that the cGPS CHO-K1 system is more rapid (2-week protocol), efficient (all selected clones expressed the GOI), reproducible (GOI expression level variation of 12%), and stable over time (no change in GOI expression after 23 weeks of culture) than classical random integration. Moreover, in all cGPS CHO-K1 targeted clones, the recombinant protein was biologically active and its properties similar to the endogenous protein. This fast and robust method opens the door for creating large collections of cell lines of better quality and expressing therapeutically relevant GOIs at physiological levels, thereby enhancing the potential scope of HTS.
Keywords
Introduction
C
Thus, dramatically different strategies have been developed to ensure the fast generation of homogeneous populations of GOI expressing cells. For example, the laser-enabled analysis and processing (LEAP) technology relies on elimination of nonexpressing clones. 7 Another promising approach is targeted integration of the GOI at a predefined chromosomal location. This strategy has the advantage of inserting a single copy of each transgene into the same chromosomal environment. Thus, a very small number of clones need to be characterized, the vast majority of them being identical. Recombinases, such as yeast Flp recombinase that allows the integration of a circular plasmid into a chromosome, can be useful tools when specific sequences are present on both DNA partners. 8 Therefore, targeted integration requires the prior integration of a target site into the chromosome. The recombinase-based Flp-In system commercialized by Life Technologies (Invitrogen, Carlsbad, CA) allows for the integration into a landing pad inserted into the target cell. 9,10 Recently, a dual system based on the PhiC31 and R4 integrases was used to integrate GOIs into pseudo att target sites into human cell lines, CHOS Chinese hamster ovary cells, strain S, and human embryonic stem cells. 11,12
The other option is to target foreign DNA integration via homologous gene targeting (HGT). HGT is usually inefficient in mammalian cell lines, but it is greatly stimulated if a DNA double-strand break (DSB) occurs in the vicinity of the targeted locus. Such DSBs can be delivered using meganucleases, sequence-specific endonucleases that recognize large DNA target sites (>12 bp). 13 These proteins, because of their exquisite specificity, can cleave a unique chromosomal sequence without affecting the global genome integrity. 14 Therefore, meganucleases represent potent stimulators of HGT and have been used to trigger targeted recombination events in a variety of organisms and cell types. 14-18 Here, we describe the cGPS CHO-K1 cell line, a CHO-K1-derived cell line that was engineered to optimize the targeted integration of transgenes.
To validate this system, a reporter gene (LacZ) and 4 relevant pharmacological target proteins (hATX, hMT1, hMT2, and hHCN4) were expressed using targeted and random strategies and characterized. Escherichia coli β-galactosidase, a robust cytosolic protein used in a vast number of expression systems, was used to set up the procedure and compare our results to other published expression systems. The human secreted enzyme, autotaxin (hATX), is a motility factor belonging to the ecto-nucleotide pyrophosphate phosphodiesterase (PDE) family. Recently, a second activity was described for ATX, lyso-phospholipase D (PLD). 19 Autotaxin and lysophosphatidic acid (LPA) have been associated with early cancer progression and cancer cell motility. 20 To understand the catalytic activity of hATX, access to a mammalian source is essential. Several strategies have been developed for ATX expression, including transient transfection of mammalian cells and Escherichia coli production; however, all of these procedures were rather difficult and never resulted in high protein yield. 21-23 The melatonin receptors hMT1 and hMT2 are also major pharmacological targets because they are implicated in circadian rhythm regulation. 24 Finally, hyperpolarization-activated cyclic nucleotide-gated potassium channel 4 (hHCN4) is a human variant of a sarcolemmal ionic channel belonging to the hyperpolarization-activated cyclic nucleotide-gated channel family, and 4 isoforms (HCN1-4) have been identified. 25 These voltage-dependent channels comprising a tetramer of 6-transmembrane domain proteins are activated in the hyperpolarized range of potential and generate an inward current carried by both sodium and potassium HCN channels. They are involved in important electrophysiological processes such as automatism and cell excitability regulation in the heart or in the nervous system. 26 These voltage-dependant channels are considered attractive pharmacological targets for several pathologies 27 and represent good models for the validation of cGPS CHO-K1 cells in patch-clamp automated tests, which require not only faithful integrity of the channels but also a large number of channels in the plasma membrane. 28
Materials and Methods
cGPS CHO-K1 system
In the cGPS CHO-K1 system (Cellectis bioresearch, Romainville, France), the hygromycin resistance gene was under the transcriptional control of the human EF1α promoter. Roughly 1 kb of additional EF1α sequence, corresponding to the first 2 untranslated exons and first intron, was present between the promoter and hygromycin resistance gene. A cleavage site for the natural endonuclease I-CreI was inserted 16 bp upstream of the ATG in the hygromycin resistance gene. A promoterless neomycin resistance gene was positioned 94 bp downstream of the SV40 polyadenylation sequence, terminating the hygromycin resistance gene. The single copy of the construct was stably integrated into the CHO-K1 cell genome. The phenotype of the cGPS CHO-K1 cell line is hygromycin resistant, G418 sensitive. The integration matrix contained 2 DNA fragments homologous to the landing pad described above (0.8 kb upstream from the I-CreI cleavage site and 0.53 kb downstream). The downstream homologous sequence represents the first 530 bp of the neo gene coding sequence. Both homology arms are separated by (1) the puromycin resistance gene, which lacks a promoter in the plasmid; (2) a multiple cloning site, allowing the cloning of a full expression cassette; and (3) the pSV40 promoter. This plasmid cannot induce a puromycin and neomycin resistance phenotype on its own; the puroR gene is promoterless, and the neoR gene lacks 243 base pairs at its 3′ end. The I-CreI coding sequence was cloned into pcDNA6.2/V5-DEST (Invitrogen) using Gateway® technology according to the manufacturer’s recommendations to create the meganuclease expression vector.
GOI cloning
The human melatonin receptors (hMT1 and hMT2), hATX variant 2 (hENPP2), and hHCN4 coding sequences were isolated by reverse transcriptase polymerase chain reaction (RT-PCR) of human brain or heart mRNA (Clontech, Palo Alto, CA) and subcloned into the NheI and NotI sites of the integration matrix (Cellectis bioresearch) and pcDNA3.1(+) vector (Invitrogen). Detail of constructions is available upon request.
Establishment of stably transfected cell lines
CHO-K1 cells (CCL61, American Type Culture Collection, Manassas, VA) were maintained in Ham-F12 medium supplemented with 10% (v/v) fetal calf serum, 2 mM L-glutamine, 500 U/mL penicillin, and 500 µg/mL streptomycin. The cGPS CHO-K1 cells were maintained in Ham-F12K medium (Kaighn’s modification) supplemented with 10% (v/v) fetal calf serum, 2 mM L-glutamine, 500 U/mL penicillin, 500 µg/mL streptomycin, and 600 µg/mL hygromycin. Nucleofection of cGPS CHO-K1 and CHO-K1 cells was performed according to the manufacturer’s instructions using the Nucleofector machine (Amaxa, Cologne, Germany). Adherent CHO cells were harvested at day 3 of culture, washed twice in cold phosphate-buffered saline (PBS), trypsinized, and gently resuspended in T nucleofection solution to a final concentration of 2 × 106 cells/100 µL. Then, 1 µg of integration matrix/GOI vector and 5 µg of meganuclease expression vector were mixed with 0.1 mL of the cGPS CHO-K1 cell suspension, transferred to a 2.0-mm electroporation cuvette, and nucleofected using program U-023 and an Amaxa Nucleofector apparatus. Similarly, CHO-K1 cells were nucleofected with 5 µg of plasmid pcDNA3.1/GOI. Immediately after nucleofection, 0.5 mL of prewarmed CHO-K1 or cGPS CHO-K1 medium was added to the cuvette. Cells were transferred to a Petri dish containing 5 mL CHO medium using a plastic pipette provided in the Amaxa kit and cultured in a humidified 37°C, 5% CO2 incubator. After 24 h, CHO-K1 cells were cultured in medium containing 800 µg/mL G418 (Invitrogen) for 12 days. After G418 selection, single colony clones were picked and seeded in 96-well plates containing complete medium supplemented with 0.4 mg/mL G418. After 24 h of transfection, cGPS CHO-K1 cells were incubated at 37°C under 5% CO2 in Ham-F12K medium supplemented with 10% (v/v) fetal calf serum, 2 mM L-glutamine, 500 U/mL penicillin, 500 µg/mL streptomycin, and 0.6 mg/mL G418 (Invitrogen). Ten days after G418 selection, single colony clones were picked and seeded in 96-well plates in complete medium supplemented with G418 at 0.6 mg/mL and at 10 µg/mL puromycin (Sigma-Aldrich, St. Louis, MO). Six to 7 days later, double-resistant clones were amplified in complete medium supplemented with the 2 selection agents. Ten antibiotic-resistant cell clones were isolated and analyzed for each stably transfected cell line.
β-galactosidase enzymatic activity
The cGPS CHO-K1/lacZ clones expressing β-galactosidase protein were detached from the Petri dish by 0.05% trypsin/EDTA and resuspended in F12-K medium supplemented with 10% fetal bovine serum. Cells (200,000) from each clone were centrifuged at 1200 rpm for 5 min. The supernatant was aspirated and 100 µL PBS added to each pellet. The cells were incubated for exactly 10 min at 37°C in an atmosphere of 95% air/5% CO2. Next, the cells were loaded with 100 µL of 2 mM Fluorescein Di β-galactopyranoside (FDG; Sigma-Aldrich) for exactly 1 min at 37°C in an atmosphere of 95% air/5% CO2. After loading, 200 µL PBS supplemented with 2% fetal bovine serum was added and the vials left on ice until cytometric evaluation. Cells were analyzed on an EPICS XL/MCL flow cytometer (Beckman Coulter, Roissy, France) using a 488-nm ion-argon laser. The emitted fluorescence (emission wavelength 520 nm) was collected on the fluorescence 2 channel. Basal fluorescence was measured in 10,000 cGPS CHO-K1 cells per assay. Alternatively, trypsinized cells were resuspended in complete F-12K medium at a density of 50,000 cells/mL. Five thousand cells were aliquoted in triplicate in a white 96-well plate (PerkinElmer, Waltham, MA). β-Glo reagent (100 µL; Promega, Madison, WI) was added to each well. After a 30-min incubation, the plate was read on a luminometer (Viktor, PerkinElmer). Results are expressed as the ratio of signal to background (parental cGPS CHO-K1 cells).
Autotaxin enzymatic assay
CHO-KI/hATX or cGPS CHO-K1/hATX clone cells were washed twice with PBS and then 3 times with serum-free F12-K medium supplemented with 1% glutamine to remove the serum (2 mL per well and per wash for a 6-well plate). The cells were then incubated in serum-free F12-K medium (1 mL/well) for 6 h at 37°C in a humidified atmosphere containing 7% CO2. After incubation, the conditioned medium (CM) was separated from the cells, centrifuged to eliminate cell debris, and dialyzed overnight against 10 L of 20 mM HEPES (pH 7.4), 6 mM D(+)-glucose, 1 mM CaCl2, and 1.2 mM MgSO4 using Spectra-Por 1.7-mL/cm tubing (Pierce Chemicals, Interchim, Montluçon, France). After dialysis, the CM was concentrated roughly 15-fold using an Amicon Ultra 10,000 (Millipore, Billerica, MA). Concentrated conditioned media (CCM) were aliquoted and stored at −20°C until use.
The phosphodiesterase activity of recombinant ATX was measured with pNppp substrate using a modification of Razzell and Khorana’s method. 29 The recombinant ATX proteins were used as an enzyme source. Samples (10 µL) were incubated in 96-well plates at a final volume of 100 µL containing 5 mM p-nitrophenyl phenylphosphonate and 50 mM Tris-HCl (pH 9.0) buffer. After 30 min at 37°C, the reaction was stopped by adding 100 µL of 0.1 M NaOH. The production of p-nitrophenol was measured by following the absorbance at 410 nm using a Pherastar plate reader (BMG, Offenburg, Germany) and subtracting the blank. The initial rates of pNP formation, which were measured after 30 min of incubation using light absorbance at 410 nm, were plotted against increasing substrate concentrations and fit according to the Michaelis-Menten equation. Nonlinear regression analyses of the kinetic data with calculation of the Vmax and Km were performed using the statistical software package Prism 4.0 (GraphPad Software, La Jolla, CA).
Radioligand saturation with intact cells
CHO-K1 or cGPS CHO-K1 cells expressing hMT1 or hMT2 receptor were resuspended in 50 mM Tris/HCl (pH 7.4), 1 mM EDTA, and 5 mM MgCl2 and aliquoted into 96-well polypropylene plates (13,000 cells/well). [125I]-2-Iodomelatonin (5 pM to 1.5 nM) was added to determine the total binding, and control wells contained an additional 1 µM melatonin to determine nonspecific binding. Incubation was performed at 37°C for 2 h in a total volume of 250 µL. Cells were then transferred to unifilter GF/B plates (PerkinElmer) with a FilterMate cell harvester (PerkinElmer) and washed 3 times with 1 mL ice-cold Tris (50 mM). Microscint 20 (40 µL/well; PerkinElmer) was added before sealing the plates. The radioligand associated with the filter plates was evaluated by scintillation counting using TopCount (PerkinElmer). Experiments were conducted in triplicate, and data were expressed as femtomole of radioligand-specific binding sites (total minus nonspecific) per milligram of total protein. Graphic representations and data analyses were generated using PRISM 4.03 (GraphPad Software).
Cellular dielectric spectroscopy
CHO-K1 or cGPS CHO-K1 cells expressing hMT1 or hMT2 receptor were plated at a density of 80,000 cells/well on MDS Analytical (Concord, Canada) 96-well assay plates with embedded electrodes and incubated at 37°C in 6% CO2 for 24 h, in the presence of 100 ng/mL pertussis toxin when specified. Prior to the measurements, the cells were washed 3 times with Hank’s balanced salt solution, 0.1% bovine serum albumin (BSA), and 20 mM HEPES (pH 7.4) and left to equilibrate at 28°C for 30 min. The impedance measurement was performed on an MDS Analytical CellKey system with a baseline recorded for 5 min before the online addition of melatonin and further kinetic measurements for 15 min. The resulting data were expressed as the maximal signal corrected for the baseline and represented as a percentage. Experiments were performed in duplicate.
Patch-clamp recording of hHCN4 current
CHO-K1 or cGPS CHO-K1 cells expressing the hHCN4 ion channel were seeded in 35-mm Petri dishes in culture medium with 5% fetal calf serum and 50 ng/mL nocodazole to stop cell division. The cells were used within 1 to 3 days. Patch-clamp experiments in the whole-cell configuration were carried out at the controlled temperature of 35 ± 1°C (Cell MicroControls). The extracellular bathing and recording solution was a modified Tyrode’s solution containing (in mM) the following: NaCl 110, KCl 30, MgCl2 0.5, CaCl2 1.8, HEPES 10, and glucose 20, adjusted to pH 7.4 with NaOH. The borosilicate pipettes were filled with an intracellular solution containing (in mM) the following: KCl 130, NaCl 10, MgCl2 0.5, EGTA 1, and HEPES 5, adjusted to pH 7.2 with KOH. Membrane capacitance was canceled and access resistance was compensated to approximately 70%. The currents, recorded by a RK-400 (Bio-Logic, Claix, France) or Axopatch 200-B (Axon Instruments, Sunnyvale, CA) amplifier, were filtered at 1 KHz, digitized at 5 KHz, and analyzed with pClamp 9.0 software (Axon Instruments). The hHCN4 current was elicited every 6 s by a 5-s hyperpolarizing pulse to −110 mV from a holding potential of −30 mV and deactivated by a depolarizing pulse at +20 mV for 600 ms. The current was measured as the difference between the instantaneous current at the beginning of the hyperpolarizing pulse and the value of the inward current at the end of the pulse. The membrane capacitance value was measured as the value corrected on the amplifier. Values were expressed as mean ± SEM. From 10 selected clones, 10 cGPS CHO-K1 exhibited an immunofluorescence signal, whereas only 2 of the 10 CHO-K1 were positive.
SDS-PAGE separation and Western Blotting
Sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) 4% to 12% was performed according to Laemmli 30 followed by Sypro Ruby staining and Western blotting detection using chicken anti-autotaxin antibody 22 followed by a horseradish peroxidase (HRP)–conjugated anti-chicken antibody (Sigma-Aldrich) before chemiluminescence detection of the immunocomplexes.
Results
Stable cell line generation
We developed the cGPS system that allows for the targeted integration of a GOI in a predefined locus in CHO-KI cells. A landing pad containing a hygromycin resistance gene under the control of an EF1α promoter-intron fusion, a recognition site for the I-CreI meganuclease, and a promoterless neomycin resistance gene was integrated into the genome of CHO-KI cells (
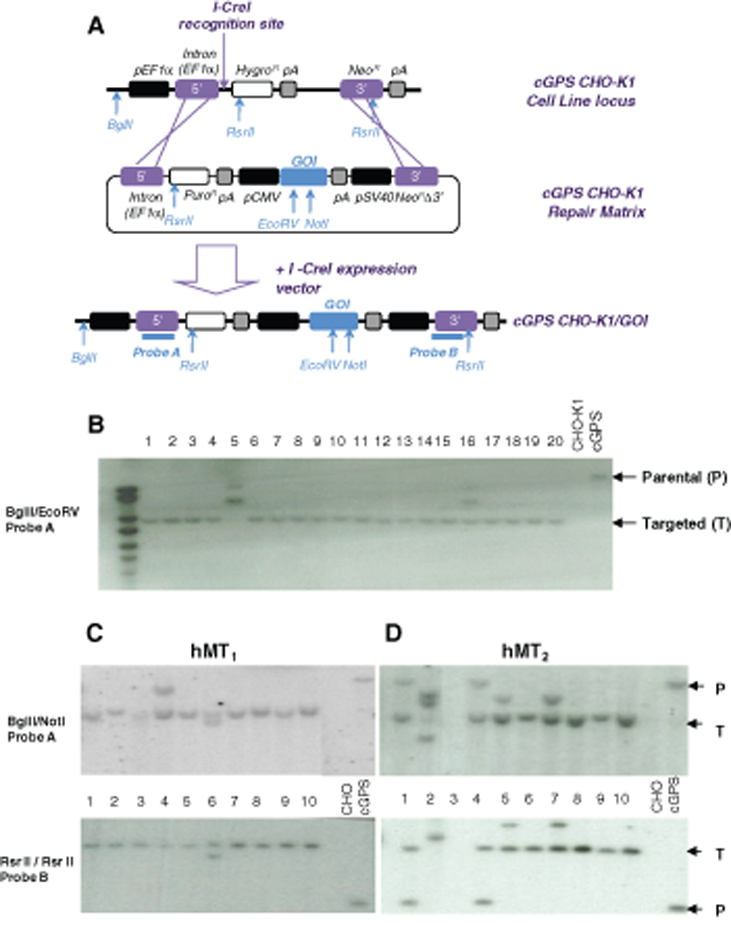
cGPS principle overview and molecular characterization of clones. (
We also used a classical random integration method: GOIs were cloned into pCDNA3.1(+). The resulting plasmids were transfected into CHO-KI cells, and stable transformants were selected on G418. Clones were obtained in about 44 days. With the cGPS strategy, HGT of the GOI was induced with the I-CreI meganuclease, and clones were selected for G418 and puromycin resistance with a short 28-day protocol. Isolated clones were then analyzed at the genomic level, for protein expression and/or activity and for expression stability.
Molecular evidence for targeted integration in cGPS CHO-K1 clones
To monitor targeted integration events in the cGPS system, 20 cell clones obtained with the lacZ construct and 9 or 10 clones obtained with the other cassettes were analyzed by Southern blot. Targeted integration was analyzed by 1 probe monitoring the upstream regions of the cGPS landing pad and a second probe downstream of the construct for hMT1 and hMT2. The parental cGPS locus and the GOI (LacZ, hMT1, or hMT2) targeted cGPS loci, with relevant restriction sites and probe locations, are shown in
Summary of Integration Analysis
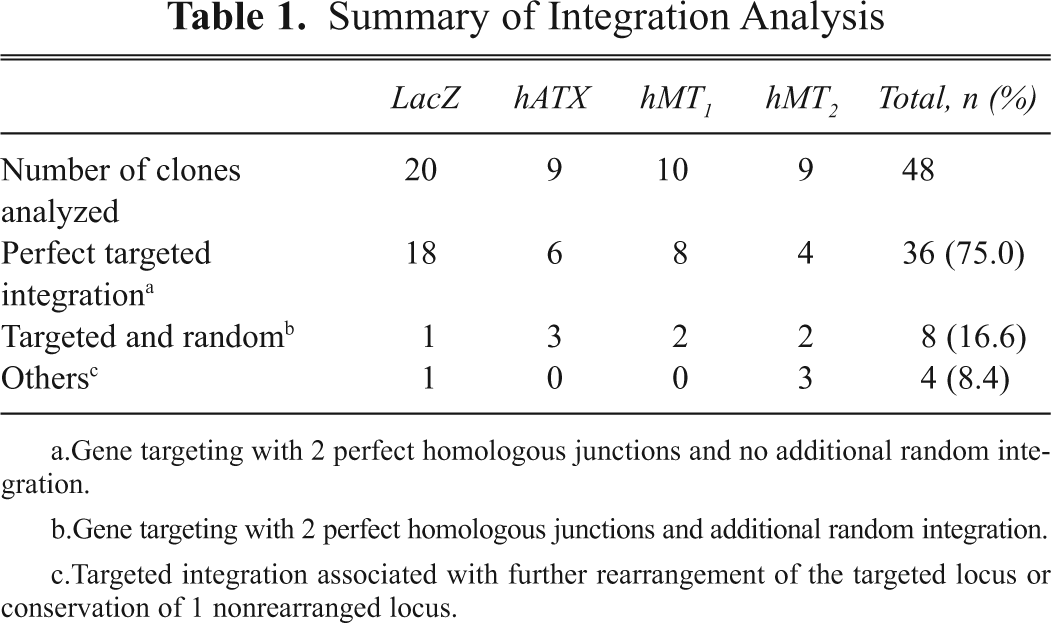
Gene targeting with 2 perfect homologous junctions and no additional random integration.
Gene targeting with 2 perfect homologous junctions and additional random integration.
Targeted integration associated with further rearrangement of the targeted locus or conservation of 1 nonrearranged locus.
Gene targeting with 2 perfectly homologous junctions was observed in 19 of 20 clones with LacZ (
Analysis of the expression of GOIs’
To monitor protein activity levels with hMT1, hMT2, hATX, and hHCN4, 10 cGPS CHO-K1 clones and 10 CHO-K1 clones were randomly chosen and analyzed for the biological activity of the expressed recombinant proteins (
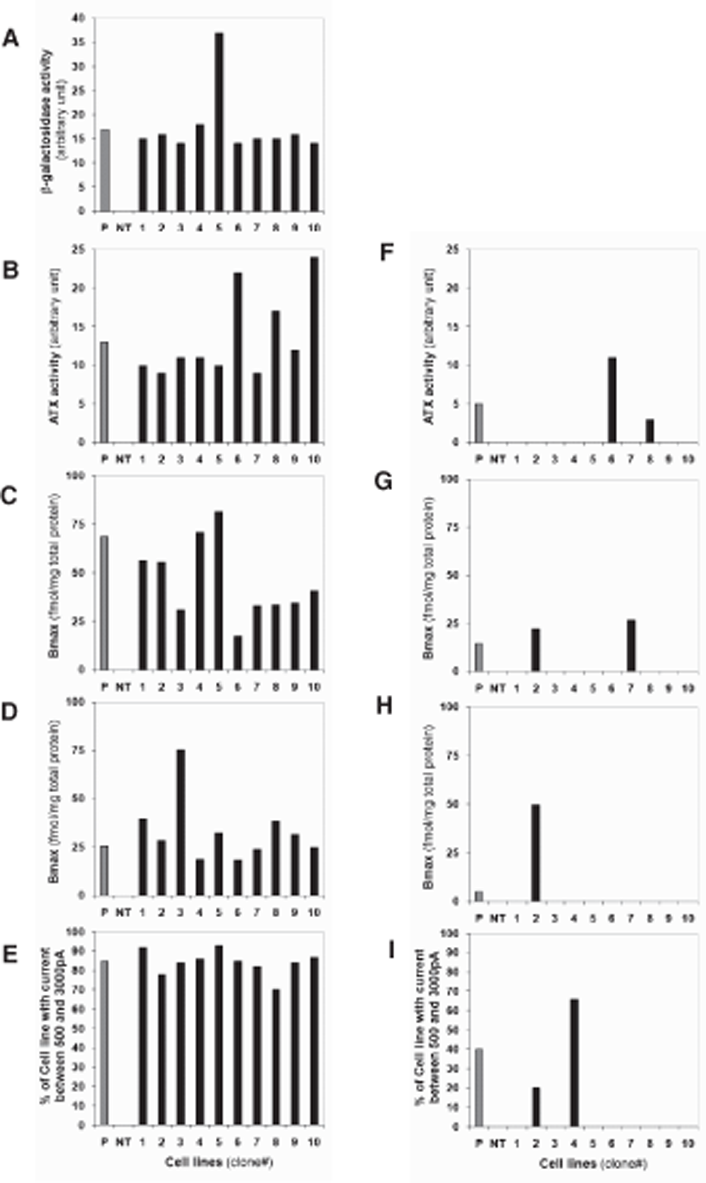
Recombinant protein expression in targeted (cGPS CHO-K1) and random transfection of CHO-K1. β-Galactosidase activity was measured with Fluorescein Di-β-galactopyranoside (emission wavelength 520 nm) using flow cytometry and 10 transfected cGPS CHO-K1 cells (
Functional characteristics of recombinant hATX
The analysis of recombinant hATX by SDS-PAGE and Western blot revealed only 1 protein (110 kDa), which corresponded to the expected protein size of endogenous hATX (
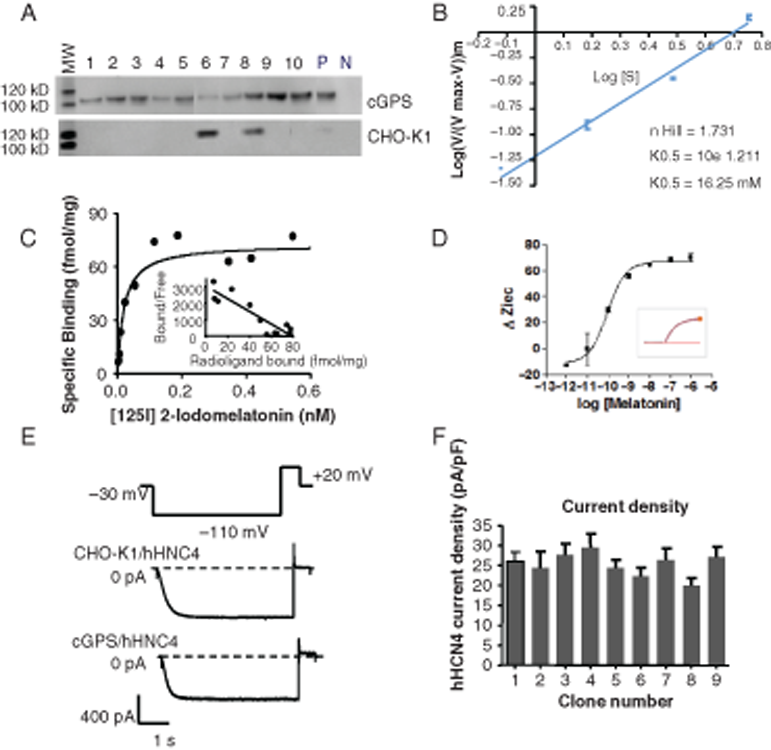
Functional characteristics of recombinant proteins expressed by the cGPS CHO-K1 system. (
Saturation assays and functional characteristics of recombinant melatonin receptors expressed in cGPS CHO-K1 targeted cells
From cGPS CHO-K1/hMT1 and cGPS CHO-K1/hMT2 cells, an analysis of the saturation data using 1-site and 2-site binding hyperbola fits (F-test, GraphPad Prism) revealed the presence of a single high affinity binding site for 2-[125I]-melatonin (
The functional activity of recombinant melatonin receptors hMT1 and hMT2 was studied using cellular dielectric spectroscopy. This technology is based on the change in intercellular impedance, mainly the resistance to an external electric current, of a monolayer of cells.
32,33
Upon stimulation of the receptor with 100 nM melatonin, the signaling cascade eventually leads to minute changes in cell adherence and/or cell shape, resulting in a change in the intercellular impedance. Interestingly, the kinetics and values of the impedance signal are dependent on the G-protein subtype involved in the cascade (see Verdonk et al.
32
for examples and theory). We obtained typical Gi-coupling kinetics upon melatonin stimulation of cGPS CHO-K1/hMT1 cells, corroborating that the recombinant hMT1 receptor is Gi-coupled (
Functional characteristics of the recombinant hHCN4 channel expressed in cGPS CHO-K1 targeted cells
Immunostaining of fixed and permeabilized cGPS CHO-K1/hHCN4 cells with fluorescent anti-hHCN4 antibody revealed major expression of the recombinant hHCN4 channel at the plasma membrane (data not shown), whereas no fluorescent signal was detected in cGPS CHO-K1 cells transfected with the vector alone.
However, the patch-clamp technique represents the gold-standard method for recording ionic channel activity and testing drug efficacy on them. In the whole-cell configuration, it allows the recording of transmembrane ionic current generated by all the channels embedded in the sarcolemmal membrane of a single cell. 34 Thus, the size of the signal is directly proportional to the number of channels expressed by the cell. One of the main difficulties encountered with this time-consuming technique is to obtain a maximal number of cells expressing a satisfactory amount of functional channels. This problem is further enhanced with the emerging use of automated-based patch-clamp devices for higher throughput data generation. 28 An ideal cell line should therefore provide a sufficient level of channel expression with the smallest cell-to-cell variation, ensuring a good signal-to-noise ratio of the recordings, a good reproducibility, and a high success rate of the experiments.
hHCN4 expressed in cGPS CHO-K1 and CHO-K1 cells led to the generation of comparable hyperpolarization-activated inward currents with biophysical properties similar to those described in the literature (
Current Measurement in Clones Having Targeted or Nontargeted Integration of hHCN4
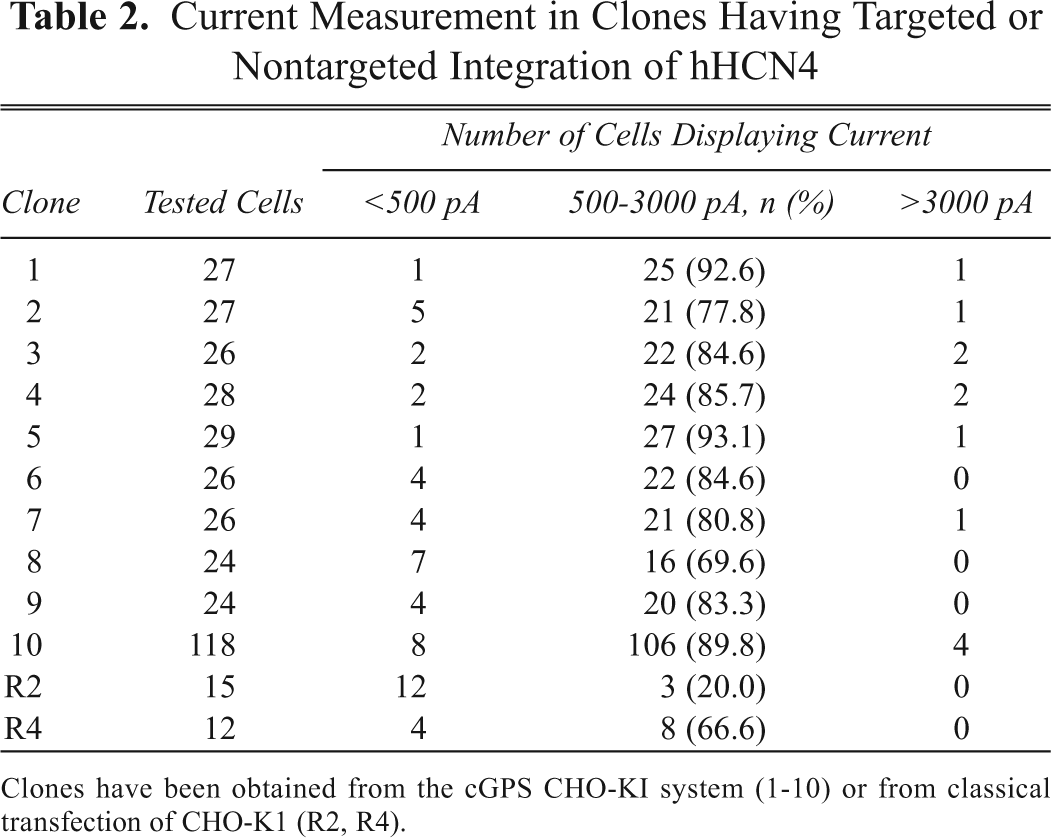
Clones have been obtained from the cGPS CHO-KI system (1-10) or from classical transfection of CHO-K1 (R2, R4).
Expression stability in cGPS CHO-K1 targeted clones
The β-galactosidase activity in cGPS CHO-K1/lacZ cells was measured 2 ways. First, cGPS CHO-K1/LacZ clones were analyzed by fluorescence-activated cell sorting (FACS) using a fluorescent substrate for β-galactosidase over 20 weeks. This method allows appreciation of the homogeneity of expression in cells belonging to the same clonal population. A symmetric and narrow fluorescent signal was observed (
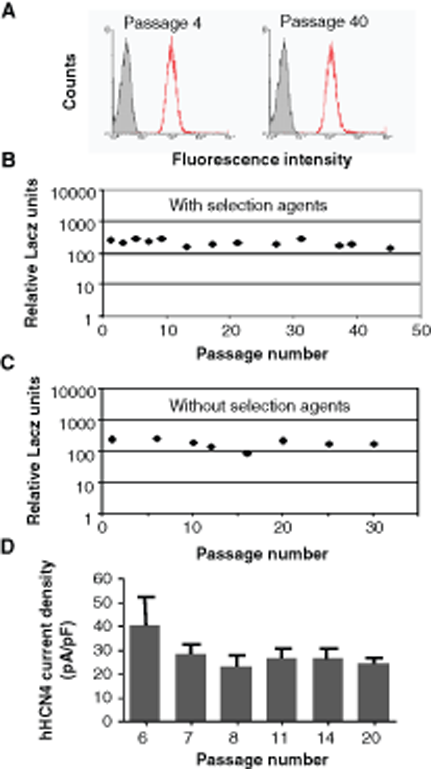
Homogeneity and stability of β-galactosidase and HCN4 expression in cGPS CHO-K1 clones. (
Discussion
Homologous gene targeting is a powerful method for integrating different constructs at the same locus displaying the required properties for transgene expression, but the efficacy of the process is usually low. The use of meganucleases, which strongly stimulate the process, results in the frequencies of targeted events.
14
Furthermore, the double selection process we devised appeared to efficiently identify such targeted events; among 48 clones characterized at the genomic level, all appeared to be a targeted event in Southern blots, as shown by the loss of the parental pattern and/or gain of a specific band corresponding to targeting the landing pad locus. Moreover, 75% were “pure” targeting events with no additional transgene integration and no further rearrangement of the targeted locus, as described previously.
16,31
Compared the classical cell transformation that we conducted in parallel, this process was not only more precise but also faster; the targeted clones were obtained in 28 days instead of 44. Thus, there was no trade-off between the precision of the approach and the time to implement it but rather a double advantage for the cGPS process. This speed is comparable with what has been described with other targeted methods based on recombinases or integrases or with the LEAP approach (see above). However, in contrast with other systems, the cGPS system allows for the specific integration of an expression cassette without the plasmid backbone (
Functional characterization of targeted clones also provided convincing arguments for the potential of cGPS technology. All cGPS targeted clones expressed a functional protein, compared only 17.5% of randomly inserted clones. Furthermore, the expression/activity is generally homogeneous from clone to clone and close to the level of expression in a polyclonal double-resistant population. Given the high percentage of perfect targeting events among clones and the relatively homogeneous expression of the GOI, working with a polyclonal population resulting from double selection with G418 and puromycin seems to be sufficient (
Finally, one of the most important drawbacks of randomly inserted transgenes is the instability of expression after a long period of culture. This aspect is important because a period of cell amplification is necessary before performing the HTS, and cells should be able to sustain a constant level of expression. In most cases of random insertion, the GOI is silenced because of multiple copy insertion, adding another level of complexity. Moreover, having a stable cell sustaining GOI expression without a selection agent during the amplification step is an invaluable advantage before running the cell-based assay. Here, we demonstrate that targeting genes in the cGPS CHO-K1 locus allow for long and sustainable expression, even in the absence of selection drugs.
Altogether, these data demonstrate that the cGPS system provides a fast, convenient, and efficient method of deriving stable clones with genes of therapeutic interest for HTS studies, as well as other research use.
