Abstract
The authors describe a novel high-throughput screening platform that provides rapid, reliable, quantitative assessment of biofilm formation and removal on engineered surfaces. Unlike traditional biofilm assays based on plate readers, this assay platform is based on high-content screening, which allows for multiplexing to simultaneously quantify the number of bacterial adhesions per unit area and the viability of adhered cells using fluorescent dye combinations. This platform is fully automated and has a throughput of more than 10,000 wells per day. The authors used this platform to examine the influence of different assay buffer systems on bacterial adhesion, viability, and removal on cross-linked polyvinyl alcohol coating films synthesized directly onto the bottoms of 384-well plates. The results indicated that water chemistry, bacteria cell type, and film chemistry combine to govern biofilm formation. In general, both reversible and irreversible bacterial adhesion increased with the extent of cross-linking in coating films, which correlates strongly with coating film cross-linking degree and hydrophobicity, which is closely related. The high-throughput platform offers a powerful tool for rapid evaluation of fouling-resistant coating films in addition to elucidation of fundamental mechanisms governing bacterial adhesion.
Introduction
B
Herein, we focus on the specific problem of biofilm formation on water treatment membranes, which costs the global water industry over $1 billion anually. 10,11 The key step in biofilm formation is the initial attachment of bacteria cells to a substrate surface (aka primary adhesion). 12,13 Primary adhesion is governed by cell-substrate physicochemical interactions and is largely reversible (over short time periods) by simple physical and chemical perturbations (e.g., hydraulic flushing or chemical cleaning). Irreversible adhesion of bacteria cells is initiated by secretion of exopolysaccharide molecules, which anchor and shelter adhered bacteria, alter substrate surface properties, and possibly make biofilm formation more favorable.
Many assay systems have been established to screen for compounds and surfaces of favorable properties (e.g., Alamar blue, 14 adenosine triphosphate [ATP]–driven luminescent assays, 15 and the Calgary biofilm device 16,17 ). However, all of these assays rely on either the metabolically active portion of the biofilm bacteria or the culturable part, 18,19 and therefore conclusions drawn from these assays should be approached with caution. 19 The only assay that currently measures biomass is the crystal violet assay, 20 which is not amenable to automation nor possesses the sensitivity to monitor primary adhesion events.
Here, we introduce an assay system that relies on high-content screening (HCS) to measure initial bacterial adhesion as well as extent of biofilm formation and removal over time. Using dye combinations such as SYTO-9 and propidium iodide, the HCS assay quantifies total adhered cells as well as viability of adhered cells and allows for direct screening of biofilm modulation by surface modification and incorporation of antimicrobial compounds. Finally, this assay is fully automated, making it useful for high-throughput screening (HTS). The platform is demonstrated herein for 2 different bacteria cells in 3 representative water chemistries (seawater, freshwater, and wastewater effluent) and 4 cross-linked polyvinyl alcohol (PVA) film chemistries—giving a total of 24 combinations.
Materials and Methods
Preparation of PVA coating films in 384-well microplates
Mowiol® PVA 6-98 with average molecular weights of 47,000 g/mol and 98.0% to 98.8% hydrolyzed was purchased from Sigma-Aldrich (St. Louis, MO) for the formation of coating films on the bottom in the wells of microplates. Succinic acid (Sigma-Aldrich) was used as the cross-linking agent.
PVA powder was dissolved under stirring in de-ionized water (DI) water at 90°C. Unless otherwise specified, the PVA concentration was 0.10 wt%. Next, PVA solutions were cooled to room temperature and the cross-linking agent was added along with 2 M HCl as catalyst under continuous stirring to produce the PVA casting solution. Cross-linker concentration was selected to produce a theoretical cross-linking degree of 20% unless otherwise specified. The theoretical cross-linking degree was defined by
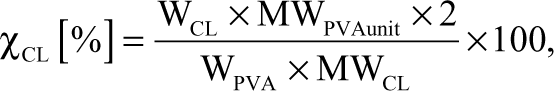
where WCL, WPVA, MWPVAunit, and MWCL represented the weight of the cross-linking agent, the weight of PVA, the molecular weight of 1 PVA unit (—CHOH-CH2—), and the molecular weight of the cross-linking agent, respectively.
Then, 50-µL PVA solutions (see preparation procedure in supplementary material) were dispensed into each well of a black/micro-clear-bottom 384-well plate (cat. 781091; Greiner Bio One, Frickenhausen, Germany) using an automated dispenser (Multidrop 384; Thermo Fisher Scientific, Williston, VT). Then, the aqueous casting solution was allowed to evaporate/air-dry at room temperature overnight (24 h) and followed by 5 curing steps at 100°C of 2 min duration to cross-link the PVA coating films.
Water chemistry
Three classes of aquatic chemistries were explored to establish the role of water quality on bacterial adhesion and removal: freshwater, wastewater effluent, and seawater. Each aquatic chemistry is represented by a complex but organic-free electrolyte. Freshwater was prepared to 1-liter solution by addition of 2.61 mg NaCl, 58.53 mg MgCl2·6H2O, 171.83 mg CaSO4·2H2O, 39.93 mg NaHCO3, 5.35 mg K2SO4, and 17.36 mg Na2SO4·10H2O; wastewater was prepared to 1-liter solution by addition of 73.88 mg NaCl, 88.21 mg NaHCO3, 980.85 mg Na2SO4·10H2O, 30.52 mg KCl, 258.52 mg MaSO4·7H2O, 288.16 mg CaCl2·2H2O, 61.42 mg NH4Cl, and 71.94 mg NaNO3; and seawater was prepared to 1-liter solution by addition of 24.317 g NaCl, 4.4629 g MgCl2·6H2O, 0.1549 g NaHCO3, 0.6794 g KCl, 6.214 g MaSO4·7H2O, 1.3493 g CaCl2·2H2O, 0.0774 g NaBr, 0.023 g H3BO3, 0.0026 g NaF, and 0.0218 g SrCl2·6H2O. The pH value of all water chemistries were adjusted to 7.0. All chemicals (Fisher Scientific, Pittsburgh, PA) were reagent grade or better.
Preparation of bacteria for adhesion
Model microorganisms used in this study included
Plate processing
Of this bacterial suspension in various water chemistries, 50 µL was added to each well of a 384-well plate using a Biomek FX with 96 pipetting manifold (Beckman Coulter, Indianapolis, IN), and the microplates were then incubated for 24 h at room temperature to allow bacterial adhesion and growth of biofilms on the surface of coating films. Before plate processing, dyes (LIVE/DEAD® BacLight™ Bacterial Viability Kits) were dissolved in 30 mL sterile DI water per the manufacturer’s recommendations to a final concentration of 12 µM Syto-9 and 60 µM PI. For the adhesion assay, only SYTO-9 was used, whereas for the parallel assessment of viability assay, SYTO-9 and propidium iodide (PI) were used together. Then, 10 µL of dye solution was added per well and incubated for 20 min at room temperature. The microplates were then imaged and washed 3 times with 100 µL DI water using an ELX405 Microplate Washer (Bio-Tek Instruments, Inc., Winooski, VT) and reimaged. For dispensing a height of 14.6 mm, an x-offset of 0.454 mm, a y-offset of 0.229 mm, and a flow rate of 3 were used, which translates to 273 µL/s. For the aspiration, a final height of 3.81 mm, a delay of 500 ms, an x-offset of 0.686 and a y-offset of 0.229, a manifold speed of 4.7 mm/s, and a vacuum of about 15 in hg were used. The inner diameter was 0.02 in for the dispense manifold and 0.033 in for the aspirate manifold.
Image acquisition and data processing
Microplates were acquired on an ImageXpressmicro (Molecular Devices, Sunnyvale, CA) using green fluorescent protein (GFP) and TRITC zero pixel shift filters (Semrock, Rochester, NY) with image-based focusing. We used a Nikon SFlour 10× lens with a numerical aperture of 0.3 and an exposure of about 150 ms, resulting in 66% saturation of the image on the CoolSNAP HQ camera at 2 × 2 binning to acquire 1 field per well. The resulting 16-bit images (696 × 520 pixel, 1 pixel = 1.29 × 1.29 micron) were processed using MetaXpress 1.74R software (Molecular Devices). Using the multiple-wavelength cell scoring module, this analysis software allowed for counting the total number of adhesions as well as viability: the total number of adhesions in the field of view resulted from the SYTO-9 channel, whereas the number of adhesions containing bacteria with compromised membranes was counted using the PI channel. The following settings were used for both channels in the multiple-wavelength cell scoring analysis module: the approximate minimum width was set to 10 µm, the approximate maximum width was set to 30 µm, and the intensity above background was 1000 gray levels.
Results and Discussion
Screening for biofilm adhesion and removal on PVA-modified surfaces
In this study, we approached biofilm formation from an engineering point of view. Our experiments were designed to evaluate what kind of surface is most resistant to biofilm formation and from which type of surface biofilms are most easily dislodged. As a model surface, we used cross-linked PVA hydrogels, which have been investigated for some time for biofilm resistance in the applications of interest here.
22
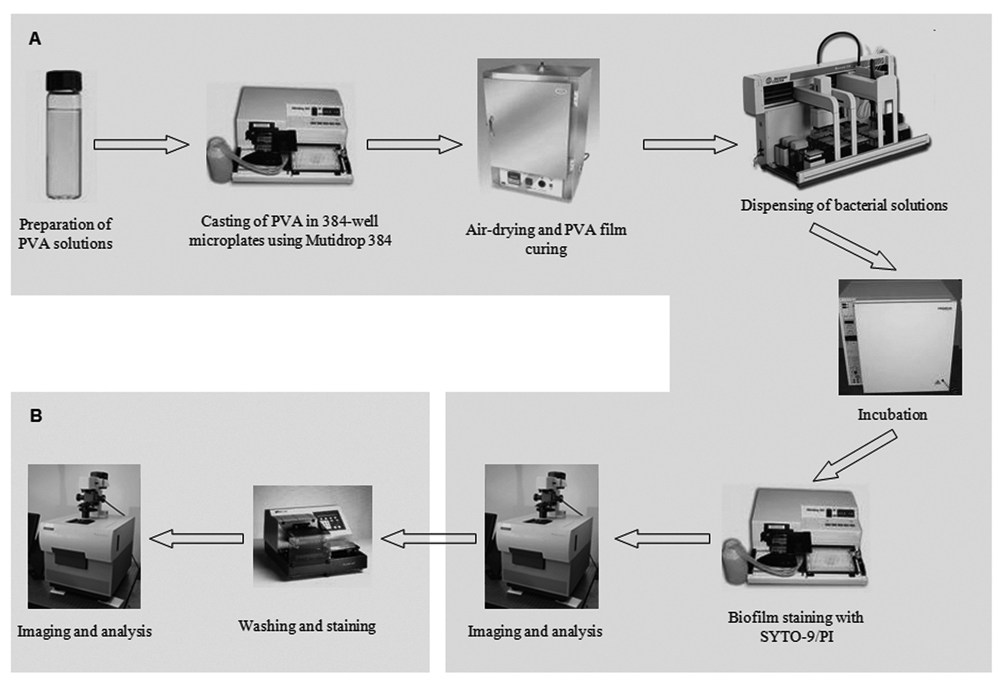
Schematic workflow for the high-throughput synthesis of polyvinyl alcohol (PVA) films and subsequent high-throughput assay for evaluating biofilm-retardant properties of the resulting surfaces. Part
The first step was preparation of the PVA-modified surfaces. Historically, combinatorial synthesis and testing of modified surfaces was a relatively slow process typically relying on 24-well plates or custom-made devices with even lower throughput rates. 23 We have sped up the production of surface-modified plates by using a noncontact dispenser that enabled casting PVA films directly onto the bottoms of 384-well optical microplates. Tens of thousands of wells with varying surface chemistries can be easily assembled in a single day using this configuration. The resulting plates can be stored until needed, adding to the flexibility of this system. The actual assay itself can be divided into 2 steps: (1) a homogeneous biofilm assay using HCS detection of biofilm formation and (2) a nonhomogeneous biofilm assay targeted at biofilm removal.
For initial validation of the assay, we used standard, TC-treated (nonmodified) black 384-well plates with a clear bottom. The assay began with the addition of the bacteria to the 384-well plates using a Biomek FX liquid handler. We used
Our first experiment in
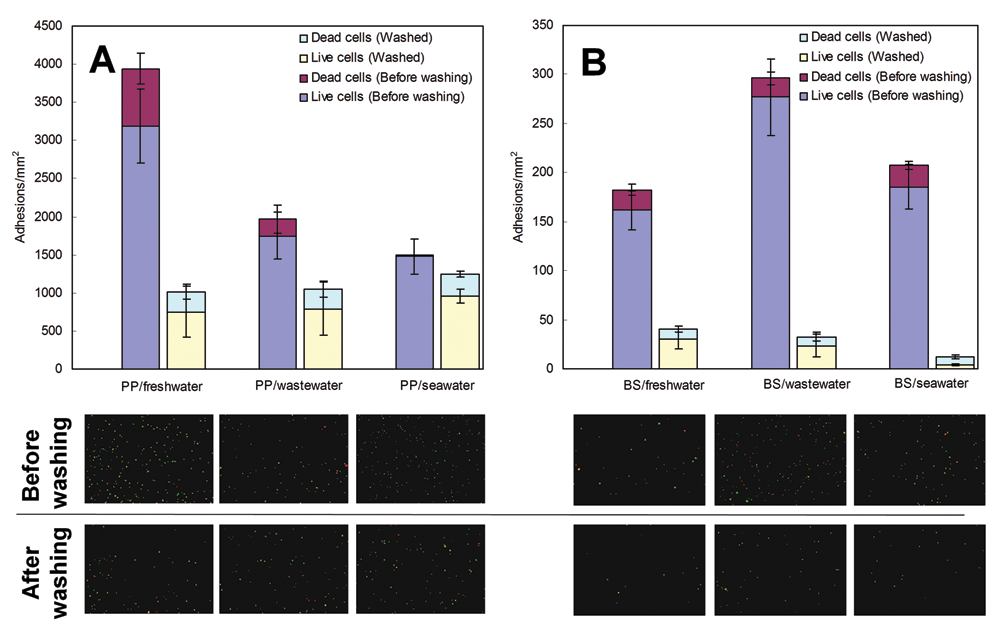
Influence of water chemistry on biofilm formation and removal results: (
The water chemistry has also exerted a significant influence on biofilm formation. Most studies use LB or TSB media, which bear little relevance to any practical conditions in which biofilm-forming bacteria are normally found, thus making translational work much more difficult. We anticipate that in the future, assay buffer systems will be used that resemble more the natural environment.
The data for the remaining biofilm adhesions after washing were on the order of 1000 adhesions per mm2 and did not correlate to the initially adherent amount of
For the purpose of optimizing surfaces, it is important that an assay can pick up subtle changes in biofilm formation—changes that are difficult to measure using a plate reader–based biofilm assay. We reasoned that the hydrophobicity of PVA films should modulate the amount of biofilm growing on it. To put our assay to the final test, we cast PVA films with different cross-linking degrees into the bottom of 384-well plates. The different extent of cross-linking resulted in different hydrophobicities of the resulting films: contact angle measurements revealed that more cross-linked films were more hydrophobic (data not shown).
We used these plates in our assay system, and the results are summarized in
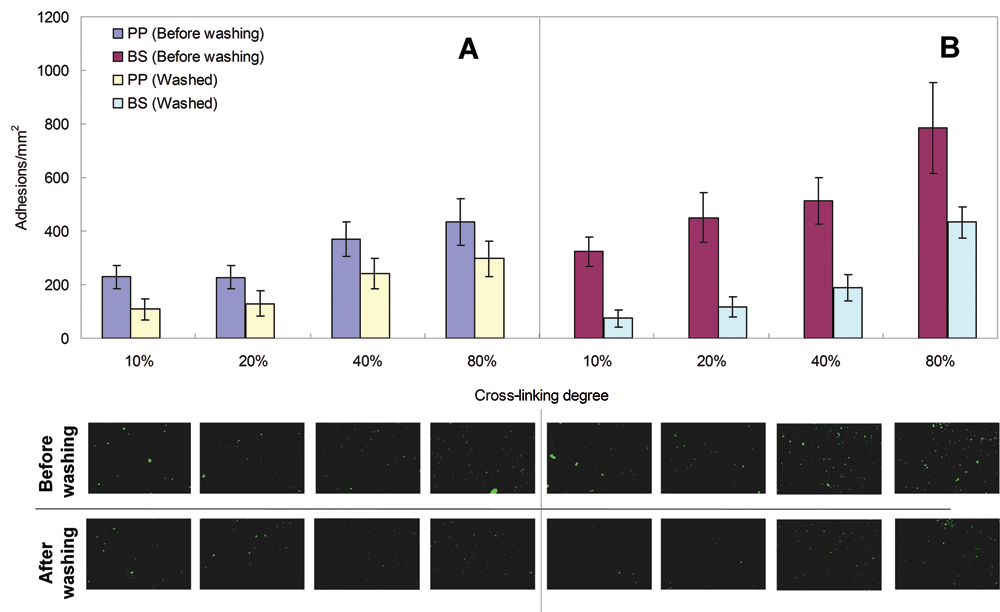
Influence of polyvinyl alcohol (PVA) cross-linking degree on biofilm formation in freshwater media for (
Conclusions
A HCS-based high-throughput assay was developed and applied to assess biofilm growth, viability, and removal from synthesized polymer-coating films. Validation of the assay was carried out on standard 384-well plates using a multiplexed SYTO-9/PI (live/dead) staining with 2 different bacterial species—
Footnotes
Acknowledgements
The authors are grateful for financial support for this research provided by a UC Discovery Grant (#GCP07-10239A) from the UC Industry–University Cooperative Research Program with industrial matching funds provided by NanoH2O, Inc.
EMVH has a financial interest in one of the project co-sponsors, NanoH2O, Inc., through stock ownership and a consulting agreement.
