Abstract
Richard Feynman postulated in 1948 that the path of an electron can be best described by the sum or functional integral of all possible trajectories rather than by the notion of a single, unique trajectory. As a consequence, the position of an electron does not harbor any information about the paths that contributed to this position. This observation constitutes a classical endpoint observation. The endpoint assay is the desired type of experiment for high-throughput screening applications, mainly because of limitations in data acquisition and handling. Quite contrary to electrons, it is possible to extract information about the path of a protein using endpoint assays, and these types of applications are reviewed in this article.
Introduction
O
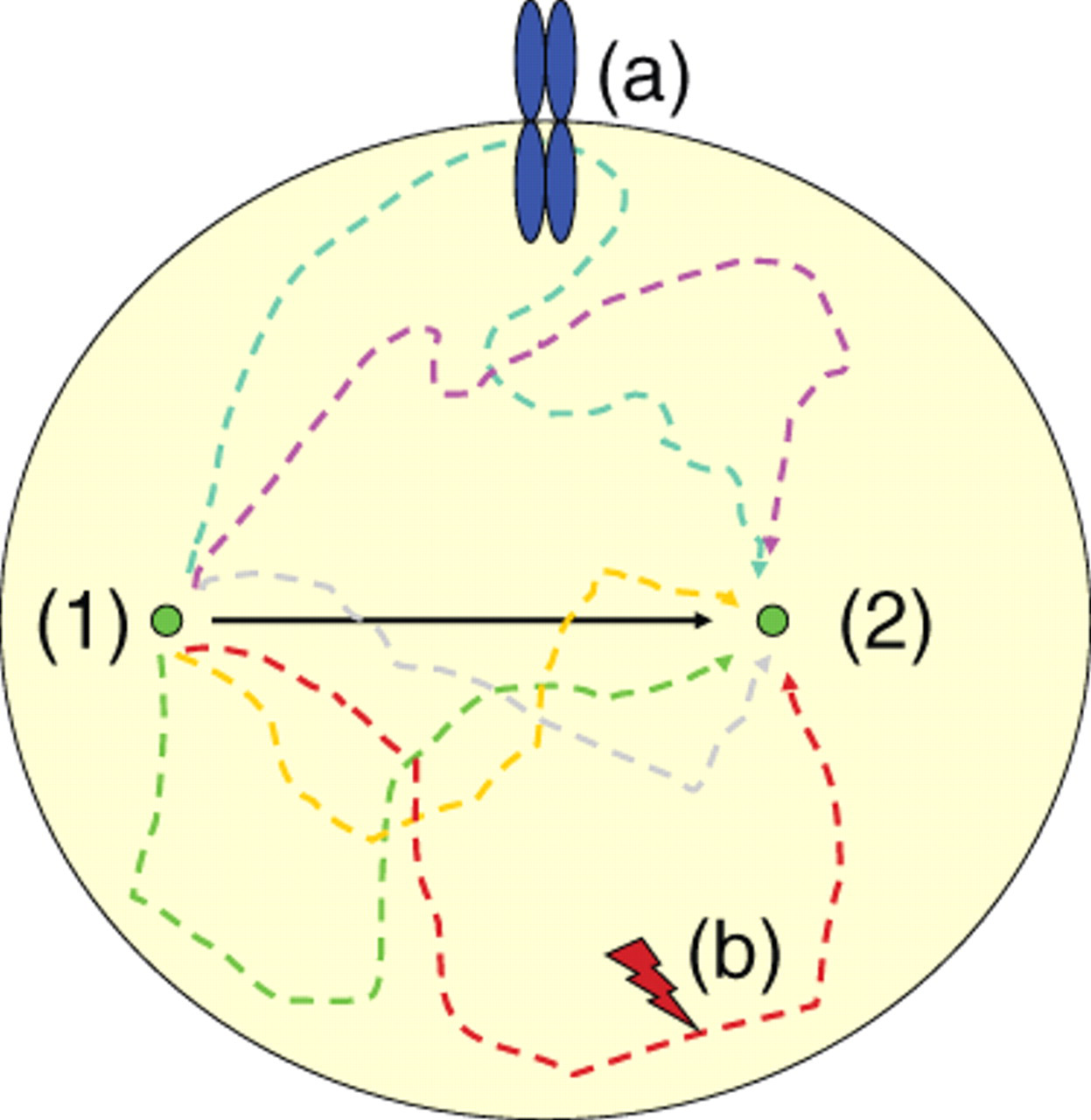
The Feynman trajectories. A fixed-endpoint observation of protein localization (2) does not report the path that the protein has taken from its starting point (1). The state of the protein may have changed by events such as receptor interaction and phosphorylation (a) or protease cleavage (b) that cannot be recorded by mere observation of (2). The identity of a protein at (2) is depending on the path and requires different assay systems to monitor trafficking than imaging.
Protein Translocation in Cellular Signaling
Protein translocation is one of the most fundamental processes in cellular physiology. Most proteins undergo at least one cycle of translocation—for instance, cotranslational translocation across the endoplasmic reticulum (ER) membrane. Other proteins remain compartmentalized but are trafficked within these compartments such as in the case of vesicular trafficking along microtubule tracks. Translocation does not necessarily require membrane compartments. For instance, cytoplasmic proteins such as signaling molecules are frequently recruited by other localized protein complexes. One example involves early activation of signal transduction pathways by ligand-induced phosphorylation of membrane-resident tyrosine kinases. Phosphorylation of the receptor results in recruitment of cytoplasmic adapter molecules such as Grb2 or Shc to the membrane, leading to activation of a signaling cascade that can be visualized by fluorescence microscopy ( Fig. 2 ). 3,4 Translocation as a consequence of activated signaling is quite common and includes among others recruitment to the membrane, nuclear shuttling (e.g., transcription factors), or secretion (e.g., activation of prehormones such as IL1b by caspase-1-mediated proteolytic cleavage).
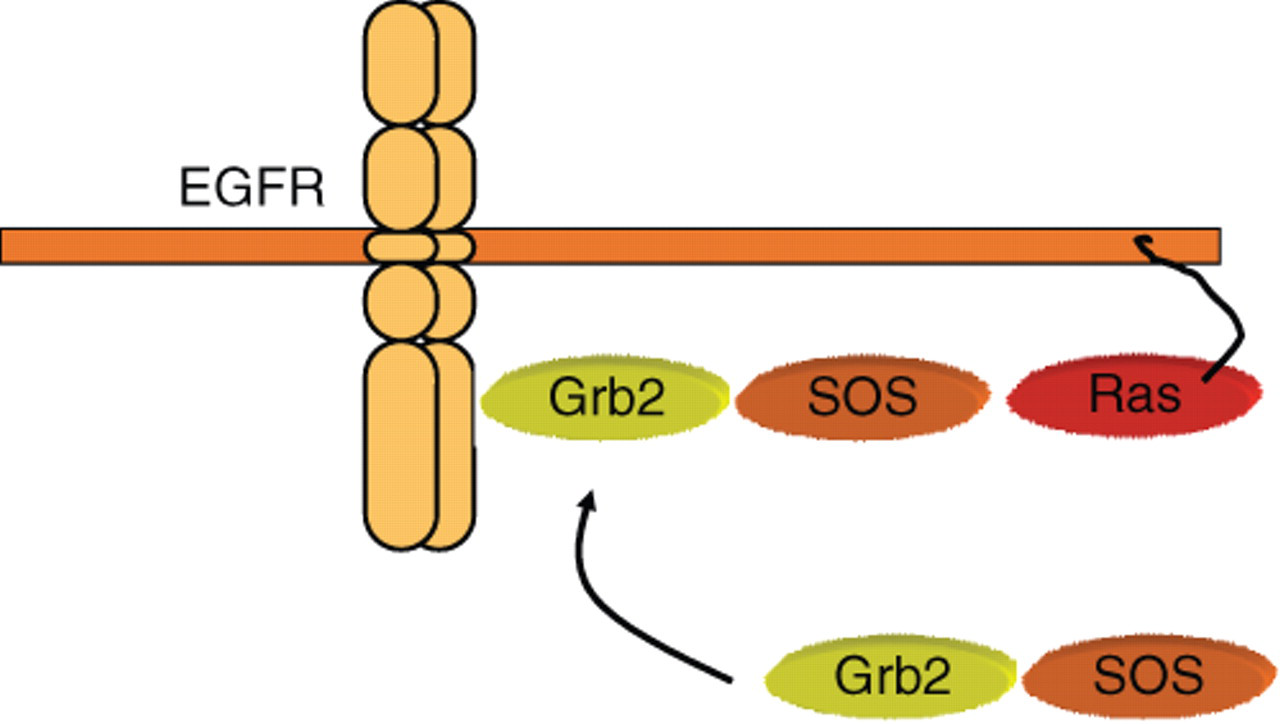
Examples of cellular signaling events that undergo translocation. Grb2 is a cytoplasmic adapter molecule constitutively associated with the Ras guanosine triphosphate (GTP) exchange factor SOS. Upon activation of a receptor tyrosine kinase such as the EGFR, Grb2 is recruited to phosphotyrosine residues at the cytoplasmic tail of the receptor, resulting in membrane localization of SOS, leading to activation of membrane-proximal RAS and subsequent signal transduction along the mitogen-activated protein (MAP) kinase cascade.
In many cases, the subcellular localization of a signaling molecule determines the activity of the protein. This has been demonstrated by tagging proteins with specific localization signals to render the corresponding signaling cascade constitutively active. One example is the targeting of PI3K to the plasma membrane by a CAAX box, leading to constitutive activation of AKT1 signaling. 5,6 The most common localized signaling events that lead to translocation are phosphorylation of a localized target (such as tyrosine kinase receptors), production of membrane-resident phosphatidylinositols (resulting in recruitment of pleckstrin homology [PH] domain containing proteins), exposure of nuclear localization or export signals (leading to nuclear import or export), protein cleavage (such as cleavage of Notch, leading to nuclear translocation of the Notch intracellular domain), or attachment of a sorting motif. I shall consider these as functional translocations (i.e., changes in localization that occur as a consequence of a specific cellular activity). Therefore, if localization were a consequence of activity, would it be possible to deduce the activity of a protein by observation of its localization? If so, the observation of a fixed-endpoint localization of a protein can infer prior activation of the underlying specific cellular activity.
One example is receptor-mediated recruitment of green fluorescent protein (GFP)–tagged Grb2 to the plasma membrane, which requires previous activation of the receptor tyrosine kinase. The presence of Grb2 at the membrane thus indicates that activation of a localized tyrosine kinase has taken place. In terms of a cost-effective endpoint assay, GFP-Grb2 recruitment to the membrane is much easier to record than complex kinase assays or in-cell immunoblotting and can be easily accomplished on an imaging platform. This type of approach to study cellular signaling on an image-based fixed-endpoint platform has become increasingly popular. For instance, as a means to de-orphanize G-protein-coupled receptor (GPCR) translocation of GFP-tagged beta-arrestin has been used. 7-12 Similar assays have been described to study PI3K activation using the GFP-tagged PH domain of AKT1. 13,14 A number of high-throughput screens have been performed based on functional translocation of signaling proteins, including GLUT4, 15 beta-arrestin, 16 GFP-Foxo, 17 Arabidopsis PIN1, 18 GFP-p53, 19 and GFP-LC3. 20 In all of these cases, the cellular activity can be deduced from localization of a reporter gene. Translocation assays are powerful techniques to generate qualitative information but have limited potential for quantitative analysis. Improvements in imaging software such as Cell Profiler or CellClassifier 21-23 enable to some degree the quantitative analysis of fluorescence intensities or localization (e.g., the number, size, and shape of spots) and may help overcome this problem in the future.
Inducible Reporter Activation—Reporter Switches
To monitor stimulation-induced activation of a reporter gene with endpoint assays, enzymatic switches can be used. Ideally, the inducible activation is an irreversible process because frequent switching between on- and off-states complicates interpretation of these data. In principle, an irreversible modification to the reporter can be achieved by 1 of 3 different modifications: (1) a simple on/off switch that is active only in response to stimulation, (2) reconstitution of the full enzyme by bimolecular fragment complementation (split proteins), or (3) stimulation-induced rapid degradation.
Attempts to specifically activate a reporter by a localized event include calcium sensors, 24 an adenosine triphosphate (ATP) sensor, 25 a pH sensor, 26,27 and an actin strain sensor. 28 All of these examples use a fluorescent protein variant to monitor changes in intensity or emission of fluorescence, mainly because changes in the spectral characteristics are easier to interpret than variations in enzymatic activities. One advantage of this type of approach is the ease of recording. In addition, fluorescence-based switches offer the possibility to perform time-resolved imaging, thus offering the opportunity to gather a lot more information about subcellular compartmentalization and trafficking. Drawbacks of these reporters are the relatively low dynamic range of fluorescent protein intensities, spectral overlap, and background emission. Enzymatic reporter switches such as a luciferase-based sodium channel sensor 29 offer advantages in terms of quantitation but have limited potential to deduce previous changes in localized protein activities.
An alternative tool is the use of split reporter genes 30 : in this case, a signal is generated by interaction of 2 partners each coupled to a split half of the reporter enzyme. One part of this pair may be compartmentalized, thus monitoring productive interactions at a specific subcellular site. One part can be tagged with a localization signal, whereas the other may report functional translocation. Sequence-based localization signals have been identified that target proteins to subcellular compartments, including the nucleus (NLS, NES), the nuclear envelope (CAAX + NLS 31 ), the ER (KDEL, signal peptide [SP]), the Golgi apparatus (SP), mitochondria (SP), and the plasma membrane (SP, TM, GPI, CAAX box). 32-35
Split reporter has been described for GFP variants 36-39 and a variety of enzymes, including Gaussia luciferase, 40,41 Renilla luciferase, 42,43 Photinus luciferase, 44,45 Cre recombinase, 46 TEV protease, 47 ubiquitin, 48 beta-lactamase, 49-51 beta-galactosidase, 52 dihydrofolate reductase, 53 and tRNA synthetase. 54 The most popular of these are fluorescent proteins and luciferases because of the simplicity in their use, their robustness and high sensitivity, and their potential for quantitation. Identification of permissive split sites requires experimental optimization. 30,55 The 2 parts should not have high affinity for each other, which would lead to high background levels of activity. Split reporter genes have been successfully used in HTS applications—for example, to study the release of apoptotic factors from mitochondria using split Renilla luciferase. 56
Compartmentalized Reporter Switches
Another way to report stimulus-induced enzymatic activity can be achieved by removing the observed product from the equilibrium ( Fig. 3 ). In this case, the enzyme may be removed from the site of interaction, for instance, by secretion. The enzymatic activity in a separate compartment, such as the extracellular space, or confined in a subcellular compartment, can be recorded in an endpoint assay and will monitor previous activation of the construct.
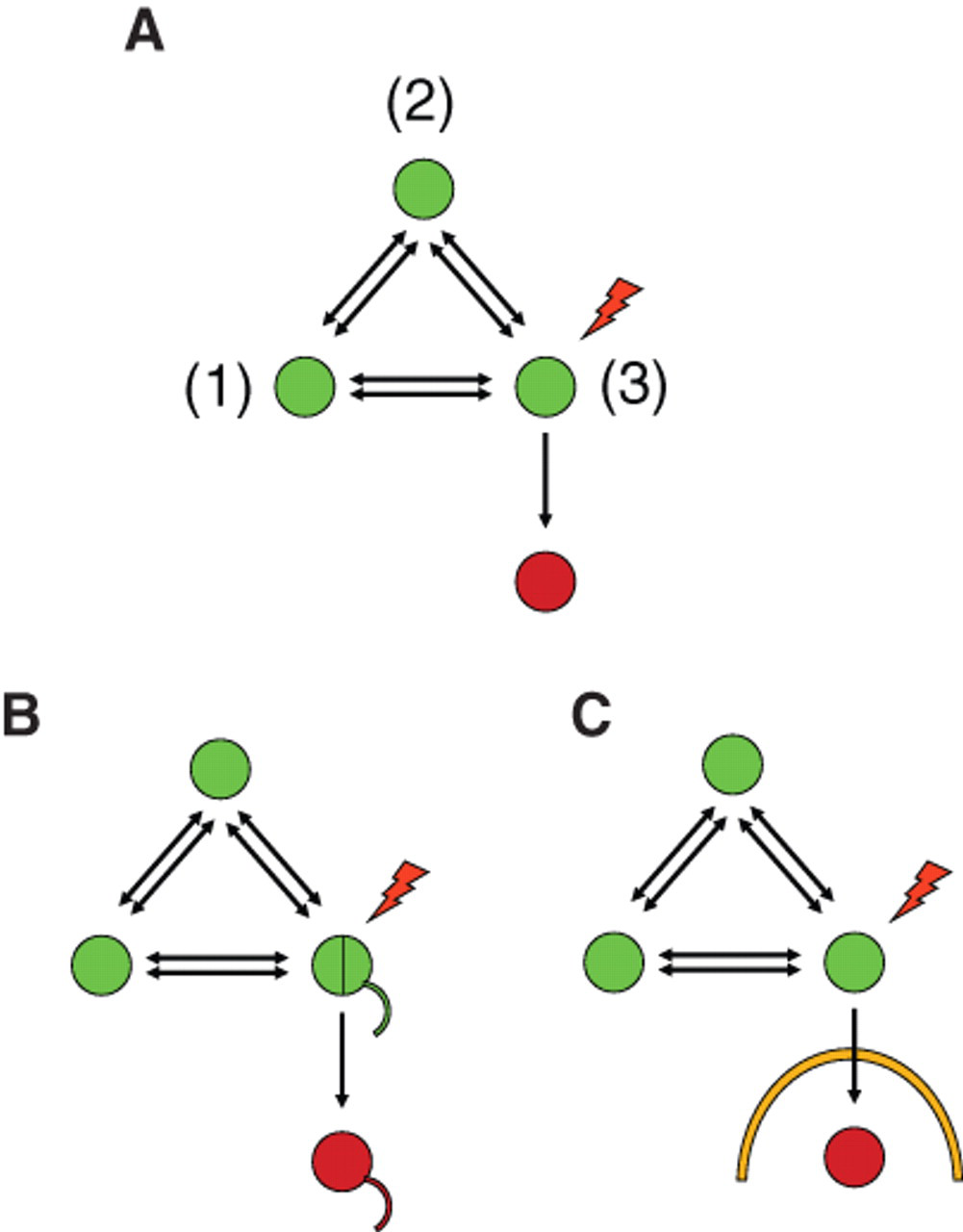
Irreversible compartmentalized reporter gene activities. (
Examples include the use of secreted alkaline phosphatase to monitor Golgi protease activity 57 and amyloid precursor protein (APP) shedding. 58 Secretion of a reporter protein is very attractive because the end product can be noninvasively harvested from cells. This allows multiplexed analysis in combination with other cellular reporters and, if desired, time-kinetic measurements. Another secreted enzyme reporter is Gaussia luciferase (GLUC). GLUC has been used to monitor apoptosis, 59 autophagy, 59,60 ER stress, 61 the strength of signal peptides, 62 growth factor uptake, 63 sodium channel activity, 29 and virus tracking 65 and to report the activation of transcription factors. 66 GLUC has become the primary choice for in vivo imaging based on successful applications to monitor tumor growth. 64,67,68 Recently, improvements to the enzyme kinetics have also enhanced its usefulness for HTS applications. 69
To collect information about localized events other than signal peptide–mediated secretion, the reporter enzyme has to be present at the desired site in the first instance. There is little information about the subcellular localization of luciferases. Renilla luciferase and Photinus luciferase are believed to be cytoplasmic enzymes, but this awaits clear demonstration. Secreted luciferases such as those from Gaussia princeps or Metridia longo are sorted through the secretory pathway by classical signal peptides. However, in the absence of a signal peptide, GLUC is exported by unconventional secretion from cells. 59 The mechanism of this type of secretion is poorly understood, but it is sensitive to treatment with Brefeldin A. One hypothesis is that dNGLUC translocates from the cytoplasm to the secretory pathway, but this requires further proof. The use of certain anchor sequences such as β-actin enables complete retention of dNGLUC inside cells. Cytoplasmic proteases such as caspases and members of the HtrA family induce the release of dNGLUC from β-actin when specific protease cleavage sites are inserted. 59 dNGLUC leaves the cell by unconventional secretion, and thus it is removed from the site of activity. A subcellular event (such as proteolysis) can be monitored by an enzymatic activity in a separate compartment (such as the extracellular space). 59 This system has been used to monitor apoptosis 59 and autophagy 60 and can be applied to a variety of cytoplasmic proteases. The endpoint (activity present in SN) provides information about a previous enzymatic event (protease cleavage in cytoplasm). This is a simple quantitative assay that is very attractive in terms of high throughput, cost-effectiveness, and robustness. A similar assay to monitor intracellular protease activity has been designed based on a tripartite reporter of a single-chain antibody fused to a Golgi retention signal via a caspase target site linker. 70 Cleavage of the protease site by caspases results in transport of the single-chain antibody to the cell surface, which can then be stained with specific antibodies.
Summary and Conclusions
A lot of information about protein trafficking has been collected over the past decades. Unlike electrons, it is now possible to visualize the path of a protein within cells. Using compartmentalized reporter gene activities, the trafficking path and also cellular events can be observed using fixed-endpoint assays that are not recordable with conventional imaging techniques. As such, there will be an increase in this type of assay in the near future that helps us understand protein homeostasis at a molecular level on a genomic scale and facilitate HTS projects.
Footnotes
Acknowledgements
I thank Brian Seed for insights and helpful discussions and Buzz Baum for critically reading the manuscript.
RK was supported by the MRC.
