Abstract
Shiga toxin–producing Escherichia coli (STEC) has emerged as important enteric foodborne zoonotic pathogens of considerable public health significance worldwide. The aim of this study was to determine the prevalence of the top seven STEC serotypes and to identify these serotypes in samples of animal source foods and vegetables. A total of 294 samples including 84 meat samples, 135 milk and dairy product samples and 75 green leafy vegetable samples were tested. The samples were harvested in mTSB-broth pre-enriched with novobiocin and then were tested by amplifying 16S shiga toxin (stx1/2), and eae genes using multiplex polymerase chain reaction (m-PCR) assay. A total of 260 (88.4%) samples were positive for E. coli and 29 (11.1%) of them were positive for shiga toxin, and eae genes. The positive samples were cultivated on CHROMAgar STEC and the colonies were evaluated for top seven STEC by m-PCR. The top seven STEC serotypes were detected in 27 (93.1%) of the samples: the STEC O111 serotype in 11 (40.7%) beef samples, STEC O45 in 3 (11.1%) chicken, STEC O145 in 6 (22.2%) parsley, 3 (11.1%) lettuce, 1 (3.7%) spinach, and 1 (3.7%) cheese, and STEC O103 in 2 (7.4%) lettuce samples. None of the samples was found positive for STEC O26, O121, and O157 serotypes. This study highlights the fact that the top seven STEC group poses a great risk in terms of food safety and public health in both animal source foods and vegetables.
INTRODUCTION
Although most Escherichia coli strains which exist in the normal intestinal flora of human and warm-blooded animals are harmless, some strains are pathogenic (CDC, 2016). The majority of these pathogenic types are contained in the Enterohemorrhagic E. coli (EHEC) family, and are also called shiga toxin-producing E. coli (STEC) or verotoxin-producing E. coli (VTEC) (Barlow and Meller 2010; CDC 2016). According to recent classifications, E. coli O157 and non-O157 (O26, O45, O103, O111, O121, and O145) serotypes are co-evaluated and expressed as the top seven STEC (USDA, 2021).
Top seven STEC serotypes can cause hemorrhagic colitis (HC) (Lee et al., 2009), hemolytic uremic syndrome (HUS) (Brusa et al., 2021; Madic et al., 2011), thrombotic thrombocytopenic purpura (TTP) (Dambrosio et al., 2007), and permanent disability in people (O’Hanlon et al., 2005; Ojo et al., 2010). In the United States, serogroups within the top seven STEC have been reported to cause serious infections in people (CDC, 2017). In fact, 71% of STEC related infections are composed of non-157 STEC serotypes included in the top seven STEC group (Wang et al., 2013). One of the most common causes of acute renal failure in children in Europe is HUS caused by STEC, and deaths occur in 10% of cases (Padhye and Doyle, 1992). In the USA, it was reported that 5.441 cases of STEC infections, including 2.323 O157 (0.72 cases per 100 000 population) and 3.104 non-O157 cases (0.97 cases per 100 000 population) CDC, 2016). Similarly, in the European Union (EU) it was reported that 6.378 cases (1.82 cases per 100 000 population) of STEC infection in 2016 (EFSA, 2017).
Top seven STEC serotypes are important foodborne pathogens of zoonotic character. Direct contact with infected people who have consumed contaminated foods and water, with animals, or with the environment are important routes of transmission of these species (CDC 2016). Ruminants, especially cattle, are important reservoirs for the spread of STEC strains to the environment (Gyles, 2007). It has been reported that outbreaks in humans caused by STEC usually occur due to the consumption of water or food contaminated with cattle feces (Erickson and Doyle 2007, Persad and LeJeune, 2014). Fresh leafy vegetables (lettuce, parsley, spinach, etc.) are also among foods with a high risk of contamination with STEC (Hou et al., 2013; Verhaegen et al., 2016).
In the Argentine Food Codex (Brusa et al. 2017), analysis is requested in minced meat, ready-to-eat foods, sausages, and vegetables for the top seven E. coli O157, O26, O103, O111, O145, and O121 serotypes. In the EU, it has been stated that ‘25 grams of sprouts should not contain top seven STEC (O157, O26, O103, O104, O111 and O145)’. In this regard, the EU declared that it is necessary to report STEC isolates of animal, food, feed, and human origin (Commission Regulation (EU) No 209/2013). According to the Turkish Food Codex (2011), only E. coli O157, which is in the STEC serogroup, should not be present in 25 g/ml of some animal foods and packaged raw vegetables. Although there are some experimental studies (Bozatli et al. 2021; Celik et al. 2021; Dikici et al. 2015) on this topic in Turkey, no study has been found on the prevalence of the top seven STEC serotypes. In this study, it was aim to determine the prevalence of the top seven STEC serotypes in samples of animal source foods and vegetables.
MATERIALS AND METHODS
Sample collection
From June 2018 to August 2019, a total of 294 samples including 84 meat samples (53 beef, 31 chicken), 135 milk and dairy product samples (100 raw milk, 35 white cheese) and 75 green leafy vegetable samples (22 lettuce, 34 parsley, 19 spinach) were collected from several butchers, markets, charcuteries, and local markets from the Balıkesir Province, Turkey. The samples were transported under refrigerated conditions (4–8 °C) to the Food Hygiene and Technology Laboratory, Faculty of Veterinary Medicine, Balıkesir University and analyzed on the same day.
Preparation of pre-enrichment for m-PCR analysis
Twenty-five g/ml of each sample were cultivated in 225 ml of mTSB-broth pre-enriched with novobiocin (mTSB broth, 1.09205, Merck, Darmstadt, Germany) and incubated at 37 °C for 18–24 h (ISO/TS 13136, 2012). Aliquots of 1.5 ml from the enriched broth were mixed with glycerol (20%) and stored at − 80°C until used.
DNA extraction
Bacterial DNA was extracted from the mTSB broth using a commercially available DNA extraction kit (PrepSEQ™ Rapid Spin Sample Preparation Kits, Thermo Fisher, Waltham, MA USA) following the manufacturer's recommendations. Briefly, 750 μL of the mTSB broth was transferred to the spin columns in the kit and centrifuged at 15.000 × g for 3 min. The supernatant was removed and 50 µL of lysis buffer (5 µL of proteinase K in meat samples) was added. The tubes were incubated for 12 min at 97°C and cooled to room temperature. They were then centrifuged at 15.000 × g for 1 min and 250 μL of nuclease-free water was added. DNA extraction was completed by centrifugation at 15.000 × g for 2 min and stored at −80 °C until using.
The samples were initially tested by amplifying 16S, stx1/2, and eae genes for identification of E. coli and shiga toxin genes by m-PCR. The specific primers for E. coli (16S-VMP5-6) and shiga toxin (stx1/2) and eae genes were used (Monday et al. 2007). The m-PCR was carried out at a final mixture volume of 25 µL. The mixture consisted of 1 µl of template DNA (100 ng), 12.5 µl of master mix (Thermo Scientific, K0171), and 0.25 μM of each primer. m-PCR was performed using the following protocol: after an initial denaturation at 95 °C for 15 min, the next 35 cycles were carried out at 95 °C for 30 s (denaturation), 59 °C for 20 s (annealing) and 72 °C for 52 s (extension) in a thermal cycler (TCA0096, Thermo Scientific, Finland). The final extension was at 72°C for 1 min. (Monday et. al., 2007). To determine the reliability of the results and detect any external contamination, all m-PCR samples were processed in duplicate. As the positive control, DNA isolated from E. coli O157:H7 (ATCC: 35150) was supplied by the Department of Food and Hygiene, Faculty of Veterinary Medicine, Balıkesir University. The amplification products were subjected to electrophoresis in 1.5% agarose gels at 100 V for 1 h and visualized under UV light.
Phenotypic and genotypic analysis of top seven STEC serotypes
The samples that were positive with stx1/2 and eae genes by m-PCR were cultured on CHROMAgar STEC medium (ST162 (B), Paris, France) at 37°C for 18–24 h. Three-five mauve colonies on the CHROMAgar STEC medium were suspected of being STEC and were analyzed for the top seven STEC by m-PCR assay. For this purpose, wzx O26, wzx O45, wzx O103, wzx O111, wzx O121, wzx O145 and wzx O157 the gene-specific primers were used (DebRoy et al., 2005; Monday et al., 2007). m-PCR outcomes that had a molecular size of 528 bp were accepted to be positive for E. coli O157:H7, those with 320 bp were E. coli O103, those with 255 bp were E. coli O45, those with 418 bp were E. coli O145, and those with 829 bp were E. coli O111. A 100-bp DNA ladder (NL1403, GeneRuler ThermoFisher, USA) was utilized as a size marker.
RESULTS
In this study, E. coli was detected in 260 (88.4%) of 294 samples by 16S (VMP5-6) amplification. In addition, 29 (11.1%) samples of the 260 E. coli positive samples were determined as STEC through amplification of the stx1/2 and eae genes. A total of 29 samples that were positive for stx1-2 or eae genes according to the m-PCR result made from the enrichment broth were inoculated into CHROMAgar STEC. At least 3-5 colonies grown on CHROMAgar STEC were selected and serotyping was performed for the top seven STEC. Of the 29 samples with stx1/2 and eae genes, seven STECs were detected in 27 samples. It was determined that 12 (22.6%) of 53 beef samples, 3 (9.67%) of 31 raw chicken samples, 1 (2.85%) of 35 white cheese samples and 13 (17.3%) of 75 vegetable samples carried at least one of the stx1/2 and eae genes. Of these, the top seven STEC were detected in 27 (93.1%) samples: the STEC O111 serotype in 11 (40.7%) beef samples, STEC O45 in 3 (11.1%) chicken samples, STEC O145 in 6 (22.2%) parsley, in 3 (11.1%) lettuce, in 1 (3.7%) spinach, and 1 (3.7%) in cheese sample, and the STEC O103 serotype in 2 (7.4%) lettuce samples (Fig 1). None of the samples was found positive for STEC O26, O121, and O157 serotypes (Table 1). Two samples (1 beef, 1 parsley) were shiga- toxin gene positive, but they were not identified as top seven STEC serogroups by m-PCR based on the ISO/TS 13136:2012.
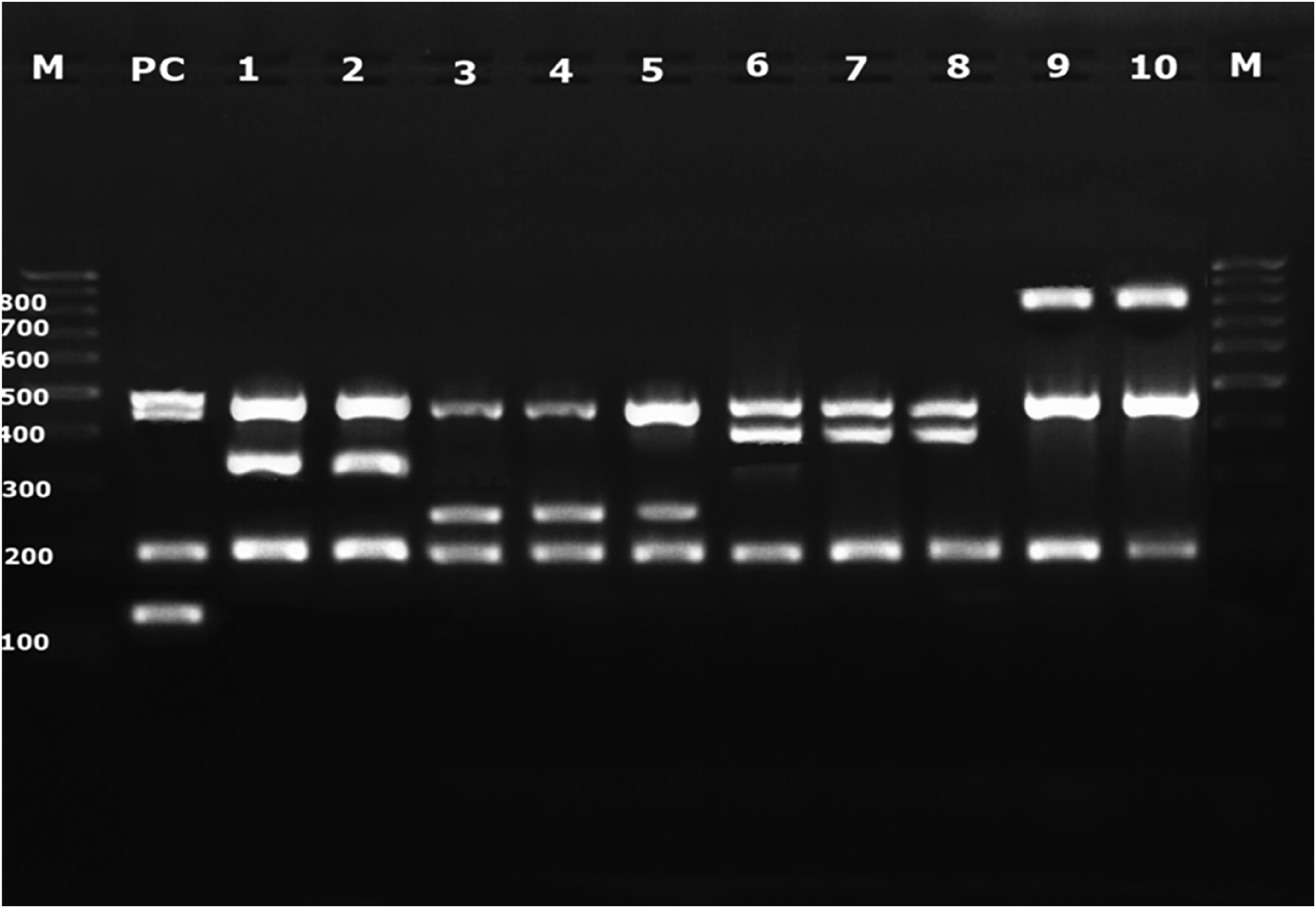
Multiplex polymerase chain reaction assay for detecting of top seven STEC targeting the wzx gene. M: DNA Ladder; PC: Positive control (E. coli 0157:H7, ATCC :35150); 1, 2: E. coli O103; 3, 4, 5: E. coli O45; 6, 7, 8: E. coli O145; 9, 10: E. coli O111.
Identification of top seven STEC serogroups in samples of animal source foods and vegetables.
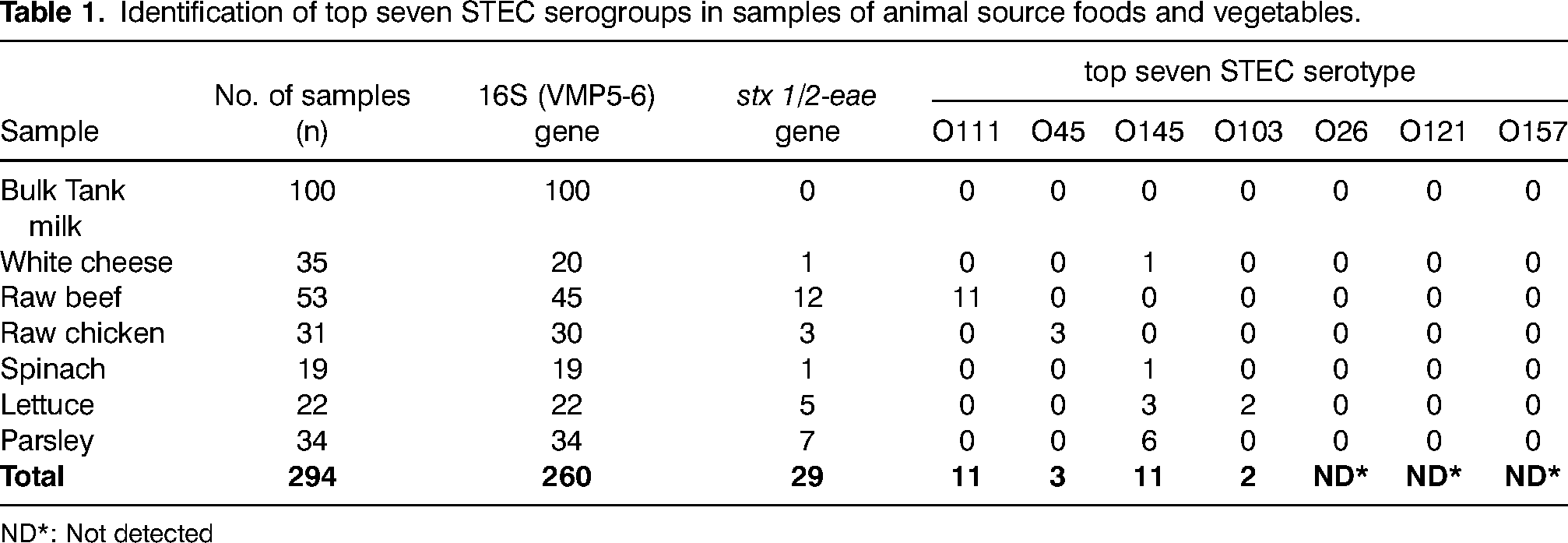

ND*: Not detected
DISCUSSION
STEC can be transmitted to humans through the consumption of contaminated foods such as raw or undercooked beef, raw milk, and water contaminated with faeces, fruits and vegetables. Since even low levels of STEC in foods can cause serious foodborne infections, detection methods are needed that are sensitive enough to identify a few STEC cells in foods (Verhaegen et al., 2016). Therefore, enrichment cultures are needed. By increasing the concentration of target bacteria, enrichment cultures both increase the sensitivity of the method and ensure the viability of the bacterial cells with positive results (Amagliani et al., 2018). Secondly, ISO/TS 13136 (2012) is a method developed specifically for the detection of the top seven STEC strains by focusing on the stx1 and stx2 virulence genes and the eae adhesion factor genes. However, the limitation of this rapid screening method remains limited to a small number of pathogenic STEC serogroups.In this study, the prevalence of top seven STEC species was investigated in a total of 294 samples, including 219 animal source foods (raw milk, cheese, beef, and chicken) and 75 green leafy vegetable samples (spinach, lettuce, and parsley). O45, O103, O111, and O145 serotypes of the top seven STEC were detected from different food samples; but O157, O26 and O121 serotypes could not be detected in any of the samples (Table 1). It was determined that 12 (22.6%) of 53 beef samples included at least one of the stx1/2 or eae genes, with 11 (91.6%) of them were STEC O111. Momtaz et al. (2013) analyzed 340 beef samples and found 67 (19.7%) STEC serotypes (21 samples were O157, 11 were O26, 3 were O103, 7 were O111, 8 were O145, 3 were O45). In addition, 53 (79.1%) of the 67 STEC positive samples were determined to be top seven STEC. In another study, a total of 18 (35.2%) E. coli strains isolated from 51 ground beef samples were determined as top seven STEC serogroups (11 samples were O121, 6 were O45, and 1 was O145) (Magwedere et al., 2013). Bai et al. (2015) stated that 19 (9.9%) of 191 beef samples contained STEC, but none of them were in the top seven STEC serogroups. There are differences between the findings of this present study and those of other studies (Bai et al., 2015; Magwedere et al., 2013; Momtaz et al., 2013). The reason for this difference can be explained by the absence of a standard method for the detection of non-O157 STEC serotypes in foods, especially meat and meat products, with genetic and biochemical diversity within themselves (Bai et al., 2015).
The majority of HUS cases in the United States are caused by STEC O111. This serotype has been cited as the cause of 3 (42.8%) of the 7 reported non-O157 STEC outbreaks (Brooks et al., 2005). Nobili et al. (2017) reported that although the prevalence of STEC is low, their presence in raw beef and ready-to-eat meat products poses a considerable public health risk. In this research, the STEC O111 serotype was detected in a total of 11 (40.7%) beef samples. The high rate of O111 is considered important for public health in Balıkesir Province. Balıkesir, known as ‘the city that feeds Turkey’, is a critical source for meat and meat products throughout the entire country of Turkey (Bal Onur and Aksoy Biber, 2017).
Although, E. coli was found in all of the bulk tank milk samples (100%) in this study, none of the isolates had the STEC virulent gene. Albonico et al. (2017) found the STEC O45 serotype in only 2 (2.9%) of 67 tank milk samples, as well as O157 and O145 in 1 (1.5%) sample. Another study found the stx gene in 32 (12.5%) of 255 bulk tank milk samples with 4 (12.5%) of these isolates were E. coli O157 and 6 (18.8%) were E. coli O26 serotypes (Trevisani et al., 2014). Automatic milking systems are widely used in dairy cattle farms in Balıkesir and its region, which may explain any STEC serotypes detected in the present study. In addition, the detection of STEC O145 in only 1 (2.8%) cheese sample (Table 1) may be explained as contamination during or after production.
E. coli was isolated from all of the 75 (100%) vegetable samples in this study (Table 1) with 13 (17.3%) of them carrying at least one of the virulent genes. A total of 9 (69.2%) of 13 isolates were determined to be top seven STEC serogroups (1 spinach sample was O145, 2 lettuce were O103, and 6 parsley were O145 serotype). Li et al. (2016) detected top seven STEC in 62 (44.6%) of 139 raw vegetable samples (55 samples were O157, 5 were O45, 1 was O145, and 1 was O111) indicating that the results of the present study seem to be compatible with their results. For the serotypes in this group, water, sediment, and soil are main secondary reservoirs. These species can survive for days or even months in agricultural lands (Jones et al., 1999). Irrigation waters, rains, organic fertilizers, and the feces of wild animals are shown as STEC contamination sources of agricultural lands (Cooley et al., 2007). This situation increases the possibility of contamination of foods (spinach, lettuce, and parsley, etc.) that make contact with the soil or grow close to the soil. In addition, it has been reported that STEC contaminations due to the use of fertilizers or contaminated water-based fertilizers occur in vegetables such as sprouts and lettuce (Verhaegen et al., 2016).
The health of people who consume foods contaminated with STEC is at risk. It has been reported that the top seven STEC serotypes pose increasingly significant threats to public health worldwide (Castro et al. 2017). Adams and Moss (2008) indicated that even a low number of STEC serotypes, between 10 and 100 cells, would be sufficient to establish an infection. Khalil et al. (2015) isolated E. coli O157 in 28 (5.7%) of 486 vegetables samples (378 unpackaged vegetables, 108 packaged organic green leafy vegetables) and 14 (50%) of them had at least one of the virulence genes. In this research, a total of 75 vegetable samples were analyzed and none of them was O157 serotype. The results of the present study differ from the results of Khalil et al. (2015). This difference may be due to the type, number, and the growing conditions of the samples as well as the analysis methods. Transmission of STEC strains to humans has been reported to occur through direct contact with ruminants and the consumption of food (meat, milk and dairy products, or vegetables) or water contaminated with ruminant feces (Ferens and Hovde 2011; Tseng et al. 2015).
CONCLUSION
Although there are studies on the presence of E. coli O157 in food and feces samples in different animals in Turkey (Aslan et al., 2017; Şenses-Ergül et al., 2015; Öksüztepe et al., 2010), no study has been carried out on the prevalence and molecular characterization of top seven STEC serotypes in food samples. In conclusion, this study highlights the fact that top seven STEC serotypes in animal source food and vegetables pose a great risk in terms of food safety and public health. Additionally, consumers should be made aware of this issue and hygiene rules should be strictly followed during the slaughtering, milking, and production stages.
Footnotes
DECLARATION OF CONFLICTING INTERESTS
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
FUNDING
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Scientific Research Project Committee of Balıkesir University, Turkey, (grant number 2018/170).
