Abstract
Acinetobacter baumannii, a notable drug-resistant bacterium, often induces severe infections in healthcare settings, prompting a deeper exploration of treatment alternatives due to escalating carbapenem resistance. This study meticulously examined clinical, microbiological, and molecular aspects related to in-hospital mortality in patients with carbapenem-resistant A. baumannii (CRAB) bloodstream infections (BSIs). From 292 isolates, 153 cases were scrutinized, reidentified through matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF-MS), and evaluated for antimicrobial susceptibility and carbapenemase genes via multiplex polymerase chain reaction (PCR). Utilizing supervised machine learning, the study constructed models to predict 14- and 30-day mortality rates, revealing the Naïve Bayes model’s superior specificity (0.75) and area under the curve (0.822) for 14-day mortality, and the Random Forest model’s impressive recall (0.85) for 30-day mortality. These models delineated eight and nine significant features for 14- and 30-day mortality predictions, respectively, with “septic shock” as a pivotal variable. Additional variables such as neutropenia with neutropenic days prior to sepsis, mechanical ventilator support, chronic kidney disease, and heart failure were also identified as ranking features. However, empirical antibiotic therapy appropriateness and specific microbiological data had minimal predictive efficacy. This research offers foundational data for assessing mortality risks associated with CRAB BSI and underscores the importance of stringent infection control practices in the wake of the scarcity of new effective antibiotics against resistant strains. The advanced models and insights generated in this study serve as significant resources for managing the repercussions of A. baumannii infections, contributing substantially to the clinical understanding and management of such infections in healthcare environments.
Carbapenem-resistant Acinetobacter baumannii (CRAB) bloodstream infections (BSIs) are associated with increased treatment challenges and higher mortality rates.
Patients in intensive care units, especially those on mechanical ventilators, with recent antibiotic use, immunosuppression, or who have undergone invasive procedures, are at heightened risk for CRAB BSI and its consequent mortality.
Prompt use of in vitro effective antibiotics is generally expected to improve survival in BSIs, although the survival benefit of appropriate empirical antibiotic therapy in those with CRAB BSI remains a contentious and debated topic.
The appropriateness of empirical antibiotic therapy and specific microbiological data, beyond carbapenem resistance, do not significantly impact survival outcomes.
Mortality related to CRAB BSI is more influenced by underlying comorbidities and impaired immune responses than by the choice of antibiotics at the time of BSI.
Machine learning-based algorithmic prediction systems demonstrate strong performance in mortality prediction and should be integrated into clinical decision-making processes.
This study underscores the significant mortality risks linked to CRAB BSI, emphasizing that prevention, rather than treatment, is a more prudent strategy. It also demonstrates that machine learning algorithms can effectively predict mortality, providing essential tools to improve clinical decision-making.
Introduction
Acinetobacter baumannii is one of the most important causes of hospital-acquired infections with a high rate of antibiotic resistance in healthcare settings for decades. Despite not being highly virulent, it has become a significant threat, leading to approximately 250,000 deaths in 2019. 1 It is a common cause of bloodstream infections (BSIs) in critically ill patients who are on mechanical ventilators, have a recent history of antibiotic use, immunosuppression, or have undergone invasive procedures, especially in intensive care units (ICUs). 2 Amidst the ongoing severe acute respiratory syndrome-Coronavirus-2 pandemic, which has heightened the susceptibility of critically ill patients, especially those struggling with respiratory distress stemming from viral pneumonia, A. baumannii outbreaks have emerged among COVID-19 patients receiving care in ICUs, hinting at a plausible connection between these phenomena.3–5
Traditionally, carbapenems used to be the preferred choice for treating A. baumannii infections. However, the landscape has shifted, with resistance rates now increasing by 50% in ICUs in the United States. 6 A recent study conducted in Türkiye revealed a staggering 94% resistance among A. baumannii isolates obtained from BSIs, particularly noteworthy due to the alarmingly high rate of colistin resistance. 7 Exploring the mechanisms driving this resistance has proven to be a complex puzzle, which in turn presents significant challenges for physicians.
Carbapenem-resistant A. baumannii (CRAB) stands as a harbinger of treatment setbacks and amplified mortality rates. This grim reality has been validated through a slew of reported incidents, magnifying CRAB-related incidences of treatment futility and escalating mortality rates, a phenomenon supported by a growing body of evidence. 8 Some studies suggest that outcomes are further compromised by inappropriate empirical antibiotic therapy, compounded by underlying medical conditions such as renal failure and neutropenia.9,10 While there is some contention regarding the exact impact of effective empirical antibacterial treatment on 14-day mortality rates, its importance within the context of multivariate analysis remains shrouded in ambiguity.9,11,12 Nevertheless, the practice of initiating early and appropriate empirical antibacterial treatment persists as a cornerstone of medical care.
The aim of this study was to thoroughly analyze the clinical factors intricately tied to in-hospital mortality in patients with CRAB BSI, alongside the microbiological and molecular attributes of corresponding isolates recovered from blood cultures. Employing sophisticated techniques, notably supervised machine learning algorithms, the study constructed complex models for predicting mortality by incorporating a wide array of diverse clinical characteristics. This novel approach aimed to explore the nuanced relationship between clinical factors and mortality in CRAB infections, offering insights into antibacterial effectiveness through advanced data analysis.
Patients and methods
Clinical data collection
This retrospective study aimed to identify the factors influencing the outcomes of CRAB BSI at a university hospital in Ankara, Türkiye. The study involved a review of 292 isolates of BSI stored at −80°C between January 2010 and January 2016, along with the corresponding clinical data of adult patients who had at least one positive blood culture for CRAB and exhibited clinical manifestations and symptoms suggestive of BSI, including fever, leukocytosis, hypotension, and signs of inflammation. The data were retrieved by reviewing electronic medical records (Figure 1). Only the first episode of BSI was considered for patients with recurrent bacteremia. Patients with inadequate or conflicting records, those with polymicrobial BSI, and those infected with carbapenem-susceptible A. baumannii were excluded. Patients who received less than 48 h of treatment, were not assessed by an infection specialist, or were not considered to have healthcare-associated infections were also excluded.
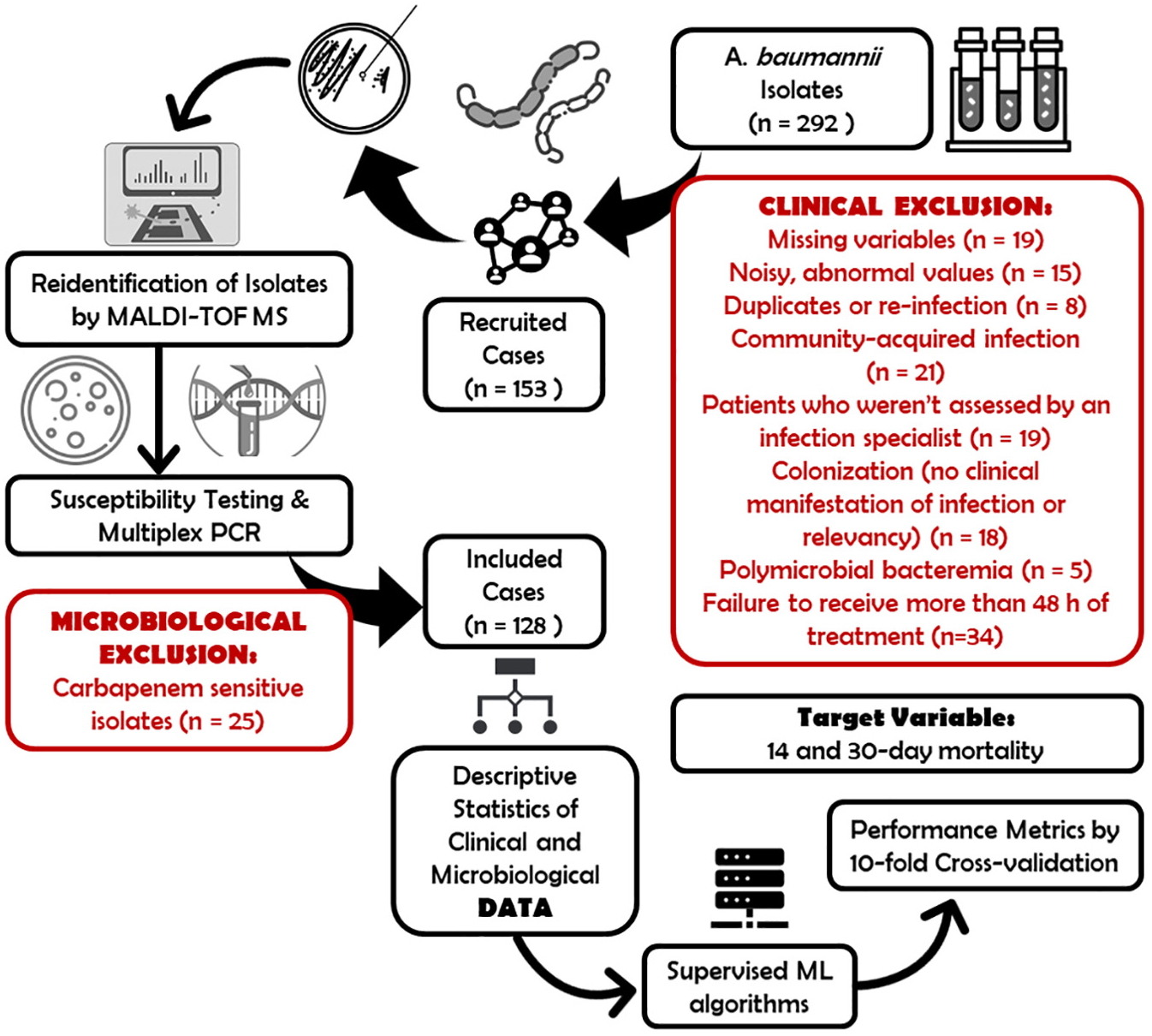
Illustration of the patient recruitment and methodology.
The collected patient data included age, sex, comorbidities (such as arterial hypertension, diabetes mellitus, coronary artery disease, heart failure, cirrhosis, chronic kidney disease, pulmonary diseases, malignancy, and rheumatological disease), dates of first admission and ICU admission, blood culture details, information on empiric and definitive antibiotic initiation, discharge and death dates, recent surgeries, invasive and noninvasive procedures (e.g., intubation, central vein, and urinary catheterization), and complete blood count and basic laboratory data at the time of admission. Diabetes mellitus was determined based on known diagnoses or the use of antidiabetic medications. Coronary artery disease and heart failure were diagnosed based on objective evidence by a cardiologist. The estimated glomerular filtration rate was calculated using the Chronic Kidney Disease Epidemiology Collaboration (CKD-epi) formula, and the results were interpreted by the Kidney Disease | Improving Global Outcome (KDIGO) 2012 clinical guideline. 13 Active cancer was defined as a diagnosis of cancer (excluding non-metastatic non-melanomatous skin cancer and non-metastatic papillary thyroid cancer) or any cancer treatment within the previous 6 months, including evidence of recurrences or metastases. Charlson-comorbidity scores were used to assess comorbidities.
Detailed information on antibiotic doses and timing was recorded. “Appropriate antimicrobial use” referred to treatment with at least one antimicrobial agent that displayed in vitro activity against CRAB for at least 48 h. The study considered “prior antibiotic” as the administration of any systemic antibacterial therapy for at least 48 h, from 4 weeks before the culture date to 1 day before the culture date. “Recent history of corticosteroid use” was defined as the use of systemic corticosteroids (with a potency equivalent to at least prednisolone 0.5 mg/kg per day) for at least 5 days within 3 months. “Recent history of cancer chemotherapy” referred to treatment with conventional chemotherapeutic agents within 3 months, excluding targeted, hormone-targeted therapies, tyrosine kinase inhibitors, and monoclonal antibodies.
All positive blood cultures had to match their clinical significance, and decisions regarding significance were made by intensive care physicians, internists, or consultant infectious disease specialists based on Systemic Inflammatory Response Syndrome (SIRS) criteria. However, for this study, sepsis was re-classified according to the definitions recommended by The Third International Consensus Definitions for Sepsis and Septic Shock. 14 Organ dysfunction was represented by an increase in the Sequential Organ Failure Assessment (SOFA) score by 2 points or more. Cases where organ dysfunction could not be represented in this way were not considered sepsis and were excluded. The quick SOFA (qSOFA) score was not addressed in this study as it was considered a useful tool for identification and suspicion, but not a definitive diagnosis. Septic shock was defined in cases where vasopressor therapy was required to maintain a mean arterial pressure of 65 mmHg and/or serum lactic acid levels were higher than 2 mmol/L despite initial aggressive fluid therapy.
Bacteria identification
Bacterial identification as A. baumannii complex was performed using the BD Phoenix™ (Becton and Dickinson Diagnostic Systems, Sparks, MD, USA) and automated VITEK®2 Compact (bioMérieux, Marcy-l’Etoile, France). The isolates were stored at −80°C and then reinoculated onto blood agar and the identification of the isolates was confirmed by using MALDI-TOF-MS (VITEK® MS RUO and the SARAMIS™ software; bioMérieux) as A. baumannii.15,16
Determination of antibiotic resistance genes
DNA isolation
A single colony of the isolate was inoculated into 2 mL of brain-heart infusion broth for 18 h at 37°C. After harvesting by centrifugation at 12,000g for 5 min, the bacterial pellet was resuspended in Tris-EDTA (TE) solution (10 mM Tris-HCl, pH 8, 1 mM EDTA). The cells were washed and centrifuged three times with 750 µL TE. The bacterial pellet in 500 µL TE was boiled for 20 min and then centrifuged. The supernatant containing the DNA was collected and stored at −20°C until use.
Polymerase chain reaction
All isolates were screened for OXA-like genes (blaOXA-23, blaOXA-24, blaOXA-58, and blaOXA-51) and for other carbapenem resistance genes (blaOXA-48, blaKPC, blaNDM-1, blaVIM, blaIMP) with multiplex PCR.17–19 The primers were given in Supplemental File 1. Amplified products were visualized on 2% agarose gel electrophoresis. Escherichia coli VIM-2, Klebsiella pneumoniae IMP-13, K. pneumoniae KPC-2, E. coli NDM-1, and K. pneumoniae NCTC 13442 for OXA-48 were used as positive control isolates for PCR.
Antimicrobial susceptibility testing
Antimicrobial susceptibility testing of A. baumannii isolates was performed against meropenem, tigecycline, and colistin by Etest® (bioMérieux) and against amikacin (30 μg), ampicillin-sulbactam (10/10 μg), piperacillin-tazobactam (30/6 μg), cefepime (30 μg), ceftazidime (10 μg), trimethoprim-sulfamethoxazole (1.25/23.75 μg), ciprofloxacin (5 μg), and tobramycin (10 μg) by disk diffusion method. All results were interpreted according to The European Committee on Antimicrobial Susceptibility Testing (EUCAST) breakpoint and Clinical and Laboratory Standards Institute interpretive criteria for disk diffusion method.20,21Pseudomonas aeruginosa ATCC 27853, E. coli ATCC 25922, and E. coli NCTC 13846 were used as quality control strains. Minimal inhibitor concentrations (MICs) for tigecycline were interpreted based on the United States Food and Drug Administration breakpoint criteria for tigecycline when testing Enterobacterales (susceptibility at ≤2 mg/L, intermediate at 4 mg/L, and resistance at ≥8 mg/L). 22 CRAB isolates from patients that were included in the clinical outcome analysis were further confirmed for colistin susceptibility by broth microdilution (BMD) according to the EUCAST recommendations. 23 The isolates that were non-susceptible to ≥1 agent in ≥3 antimicrobial categories were defined as multi-drug-resistant (MDR) and the isolates that were non-susceptible to ≥1 agent in all but ≤2 antimicrobial categories are defined as extensively drug-resistant (XDR). 24
Statistical analysis
Continuous variables were represented using median values and interquartile range while categorical variables were presented as percentages. To analyze the data, the chi-squared (χ2) test or Fisher’s exact test was applied to categorical variables, and the Mann-Whitney U test was used for continuous variables. Two-sided significance testing was employed, and p-values less than 0.05 were considered statistically significant. Any missing variables within an acceptable rate (<20%) were imputed using random forest (RF) imputation. Descriptive statistics and data imputation were performed on the raw dataset using various packages (psych, randomForest) in the R 4.2.2 programming language.
Machine learning algorithms
Prior to the development of classifier models for predicting outcomes of death within 14 and 30 days, feature selection was undertaken to eliminate unnecessary, redundant, or non-informative features, utilizing well-established methods like information gain (IG) and gain-ratio (GR) attribute evaluation methods, resulting in the selection significant features. Subsequently, five machine learning algorithms: RF, Naïve Bayes (NB), support vector machine (SVM), neural network (NN), and logistic regression (LR), were leveraged to construct the adverse event prediction models, each fine-tuned with algorithm-specific settings. For instance, a polynomial model was adapted for SVM, RF utilized 200 decision trees, and the NN model was configured with 250 neurons in its hidden layers, employing a rectified linear unit activation function and the Adam optimization algorithm.
The developed models were analyzed using Orange 3.34.0 Software (the University of Ljubljana, Ljubljana, Slovenia) to generate confusion matrices, compute algorithm performance, and render plots. The “Test and Score” module enabled the identification of suitable validation techniques, with stratified tenfold cross-validation being the method of choice. This entailed dividing the dataset into 10 segments, training models on nine, and testing on one, cycling through all segments as test sets. The predictive efficacies were assessed via performance testing, with the average of the 10 test outcomes considered as the conclusive result for each algorithm. Performance metrics derived from the confusion matrix, including sensitivity (recall), specificity, accuracy, receiver operating characteristics, and F1 statistics, were utilized for the evaluation of all developed models.
For depicting survival rates, highlighting significant feature variances, and evaluating the influence of empirically appropriate antibiotherapy, Kaplan-Meier plots were illustrated, and log-rank tests were executed.
Results
Patient characteristics
A total of 292 isolates with corresponding patients were identified during our study period (Figure 1). Among them, 34 patients had insufficient or incomplete medical records, and eight cases were adjusted for reinfection or data duplication. By further excluding 21 cases where blood culture positivity occurred within 48 h of hospital admission and lacked prior hospitalizations, five cases of poli-bacteremic BSI, 18 cases lacking significant signs of septicemia or acute SOFA score changes, 19 individuals who were not consulted by the infection department before death or discharge, and 34 patients who succumbed prior to receiving empiric antibiotic treatment for 48 h, a total of 153 patients were recruited. Their corresponding isolates were subsequently reinoculated and reidentified. Among these, 25 isolates were meropenem susceptible as determined by the gradient test. Since meropenem-susceptible cases were not pertinent to our study’s focus, the final cohort comprised 128 patients and their initial episodes of CRAB BSI.
Demographics and clinical data
The study cohort was predominantly male (n = 78, 60.9%), with a median age of 64 years (±23) years. Two most prevalent comorbidities were active cancer and arterial hypertension. A majority of patients had been admitted to the ICU before the onset of bacteremia, and the median time between admission and bacteremia occurrence stood at 17 days (±27.75). As detailed in Table 1, the comparison studies revealed that except for active cancer, chronic kidney disease, and chronic heart failure; certain comorbidities did not exhibit statistically significant differences with either endpoint. Hematologic malignancies were more common in patients with 14- and 30-day mortality, whereas having a solid tumor did not show any significant links to mortality. Notably, the rate of severe neutropenia, potentially tied to hematologic malignancies or recent chemotherapy exposure, displayed a marked increase in patients with 14- and 30-day mortality. For a comprehensive overview, detailed percentages and numerical counts for each comorbidity, alongside corresponding p-values, can be found in Table 1.
Demographic and clinical characteristics of patients stratified by 14- and 30-day mortality.
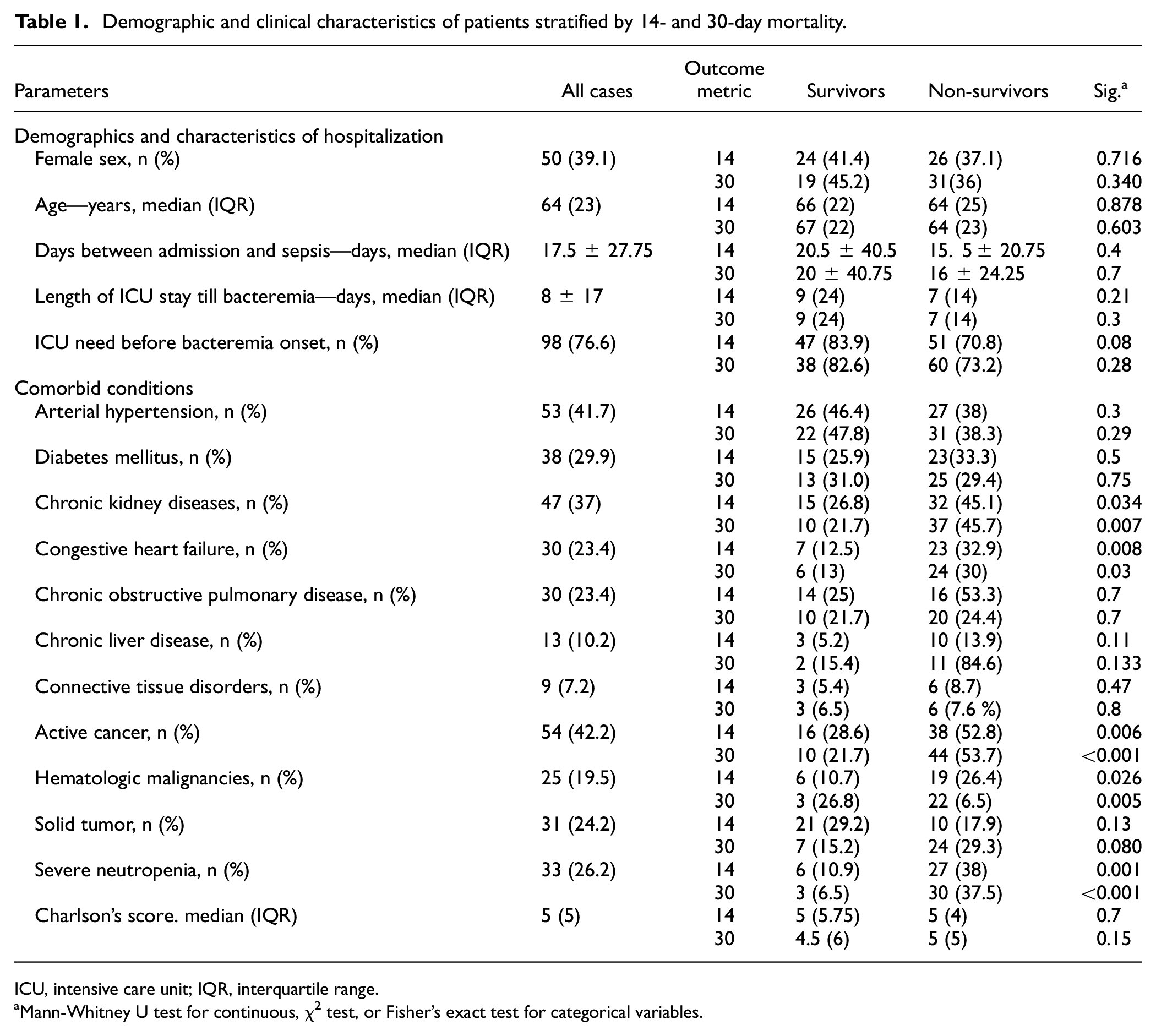
ICU, intensive care unit; IQR, interquartile range.
Mann-Whitney U test for continuous, χ2 test, or Fisher’s exact test for categorical variables.
Regarding specific treatments and interventions, the need for mechanical ventilation at the time of bacteremia and central venous catheterization was higher in the non-survivors group compared to survivors (Table 2). Similarly, recent antibiotic treatments, present in 94 patients (75.8%), were more common in the non-survivors group. No significant differences were noted in the use of recent antifungal therapies and gastric acid suppressors between survivors and non-survivors; however, higher rates of recent chemotherapy or corticosteroid use were observed in non-survivors.
Features of invasive interventions, specific medication background, and appropriateness of empiric antibiotherapy categorized by 14- and 30-day mortality.
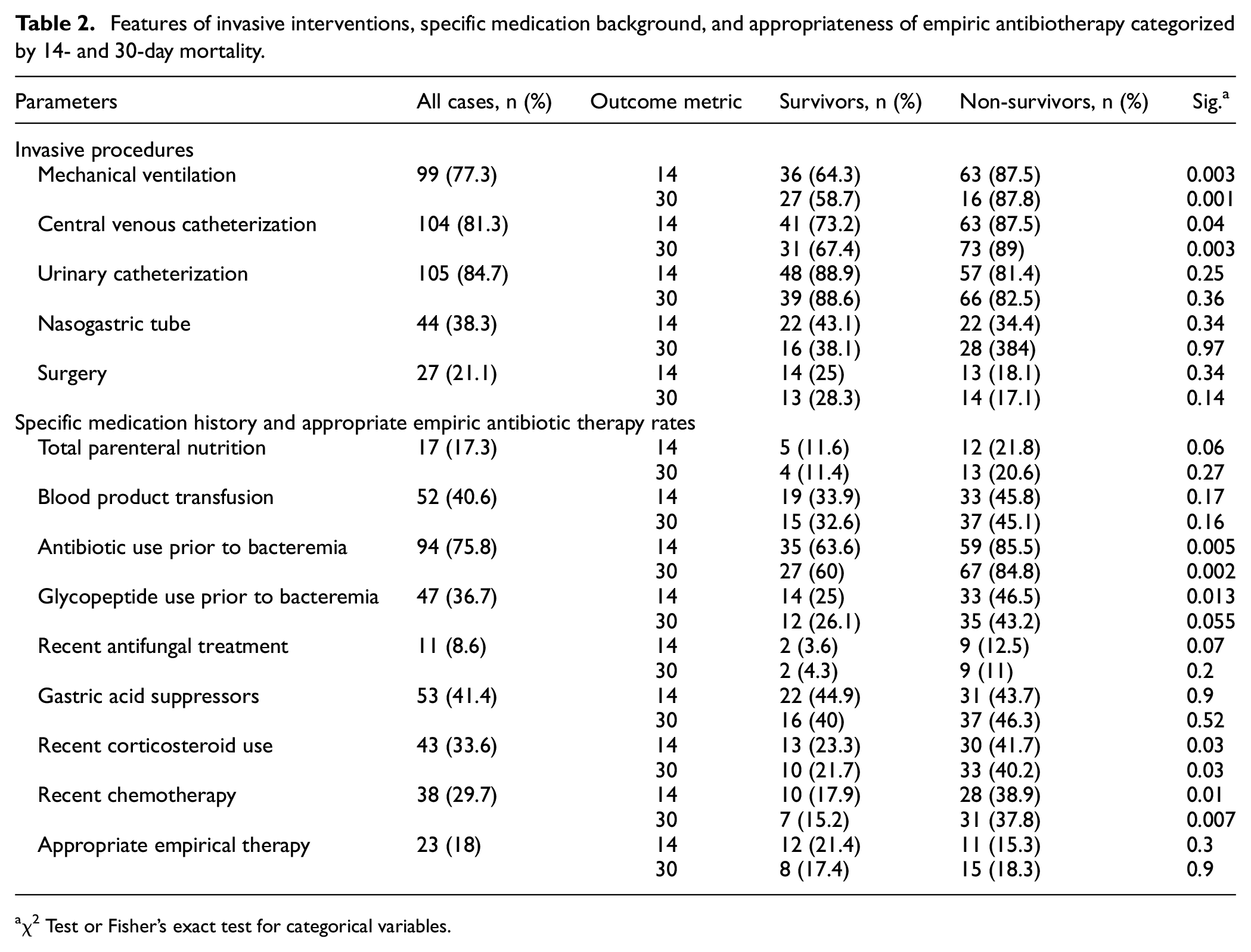
χ2 Test or Fisher’s exact test for categorical variables.
As a result, 19 patients received empirical colistin, among whom four were found to be infected with colistin-resistant strains. Additionally, three patients who did not receive empirical colistin were treated empirically with antibiotics proven effective in vitro—two with amikacin and one with trimethoprim-sulfamethoxazole. Fifty-seven patients had died by the time culture results became available or definitive therapy commenced, with five of them having received empirical colistin coverage. Irrespective of colistin use, 12 patients were found to have received appropriate empirical antibiotics, and 36 were started on inappropriate definitive antibiotics. To sum up and refocus on the main theme, Supplemental File 2, illustrates the appropriate antibiotics utilization rates along with their corresponding timestamps on the timeline.
Table 3 summarizes the mortality rate differences regarding to accepted origin of bacteremia. These rates are fairly comparable, however, the occurrence of septic shock is significantly higher in non-survivors for both 14- and 30-day mortality outcomes.
Origins of bacteremia, occurrence of septic shock, and susceptibility to colistin, categorized by 14- and 30-day mortality.
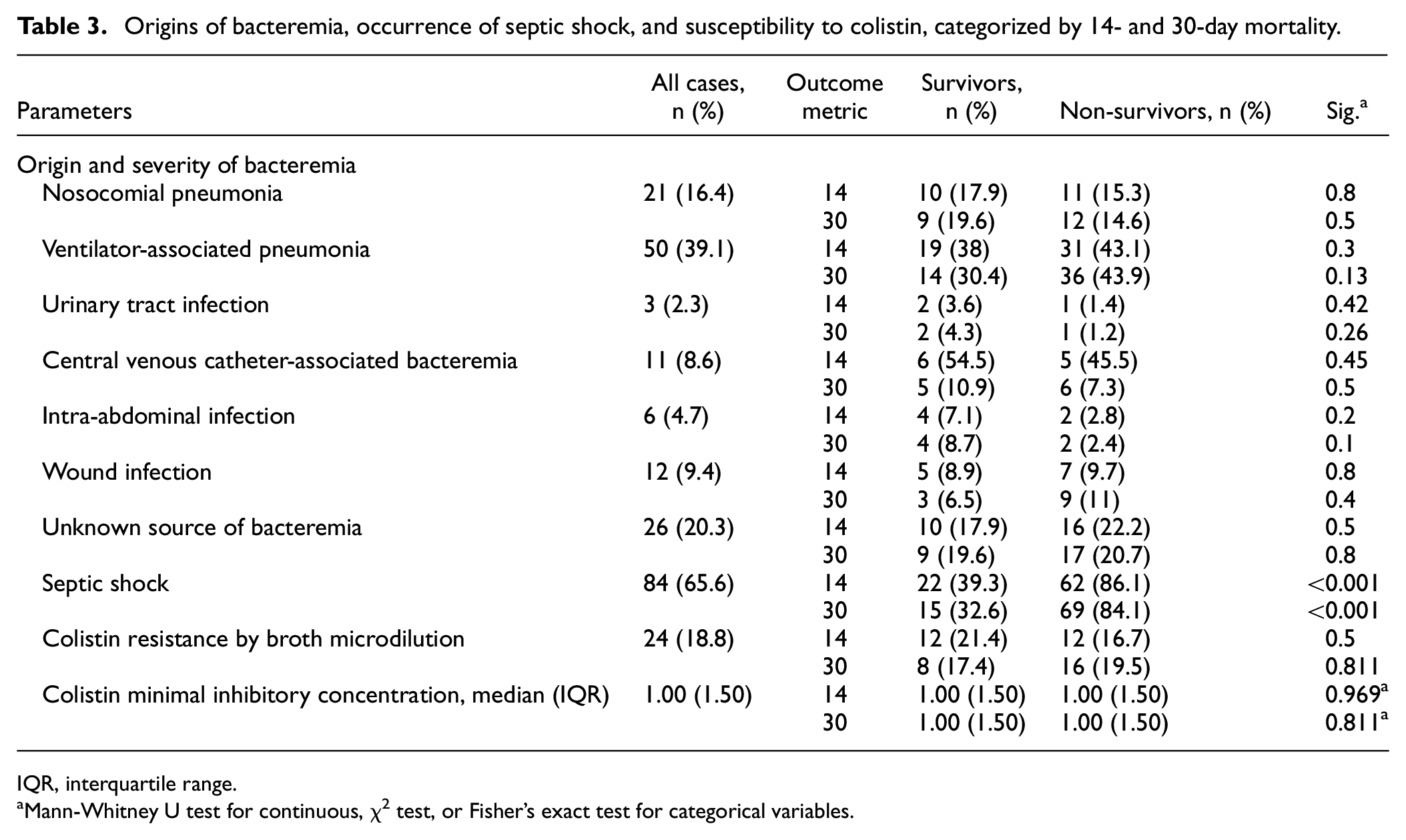
IQR, interquartile range.
Mann-Whitney U test for continuous, χ2 test, or Fisher’s exact test for categorical variables.
Bacterial identification, antimicrobial susceptibility, and molecular features
The identity of 153 isolates was confirmed as A. baumannii by using MALDI-TOF-MS (Vitek MS RUO and the SARAMIS software; bioMérieux) and by blaOXA-51 positivity. As previously stated, 25 instances were excluded since the isolates were found susceptible to meropenem. The antibiotic resistance rates were 64.8% (83/128) for amikacin and tobramycin, 91.4% (117/128) for trimethoprim-sulfamethoxazole, 98.4% (126/128) for ampicillin-sulbactam and piperacillin-tazobactam, 96.9% (124/128) for ceftazidime, 99.2% (127/128) for ciprofloxacin and 100% for cefepime by disk diffusion method. All isolates were resistant to meropenem and tigecycline. Isolates were further tested by BMD against colistin. Twenty-four (18.8%) out of 128 CRAB strains were resistant to colistin (MIC50/MIC90 = 1/24 mg/L, MIC range = 0.125–256 mg/L). Only one isolate showed resistance to colistin, identified by a MIC value of 4 mg/L via gradient testing, while, through BMD testing, a MIC of 2 mg/L indicated susceptibility in this strain. No statistically significant difference was observed between survivors and non-survivors in terms of colistin resistance rate and the median MIC of colistin determined by the BMD test (Table 3).
Multiplex PCR detected blaOXA-23 in all isolates. Three bacterial strains tested positive for the presence of the blaVIM, and an additional three strains were found to be positive for blaIMP. None of the isolates exhibited the presence of blaNDM, blaKPC, blaOXA-24, blaOXA-58, and blaOXA-48 genes. Following the analysis of antibiotic susceptibility information, it was determined that out of the CRAB isolates, 120 (93.8%) were classified as XDR, with the remaining 8 isolates (6.3 %) showing MDR phenotype.
Machine learning models for 14- and 30-day mortality
A total of eight ranking features recruited from IG and gain-ratio selection methods were selected for the 14-day mortality prediction models, and nine for the 30-day mortality models (Figure 2). The importance of features varied depending on the ranking method used (as described in the Patients and methods section) and the most crucial feature regardless of the endpoint and ranking method was the “septic shock” variable. Using the IG algorithm, severe neutropenia and the length of stay were found to be the next two most important features for 14-day mortality, while neutropenic day counts with mechanical ventilator support need were the next two important features for 30-day mortality. Neutropenia and neutropenic day counts were also among the highest information-giving features after the septic shock feature for 14- and 30-day mortality prediction models using the GR method. The presence of appropriate empirical antibiotic therapy and microbiological data, including antibiotic susceptibility profiles, colistin MIC values, and the existence of carbapenemase genes were not encountered as mortality predictors.
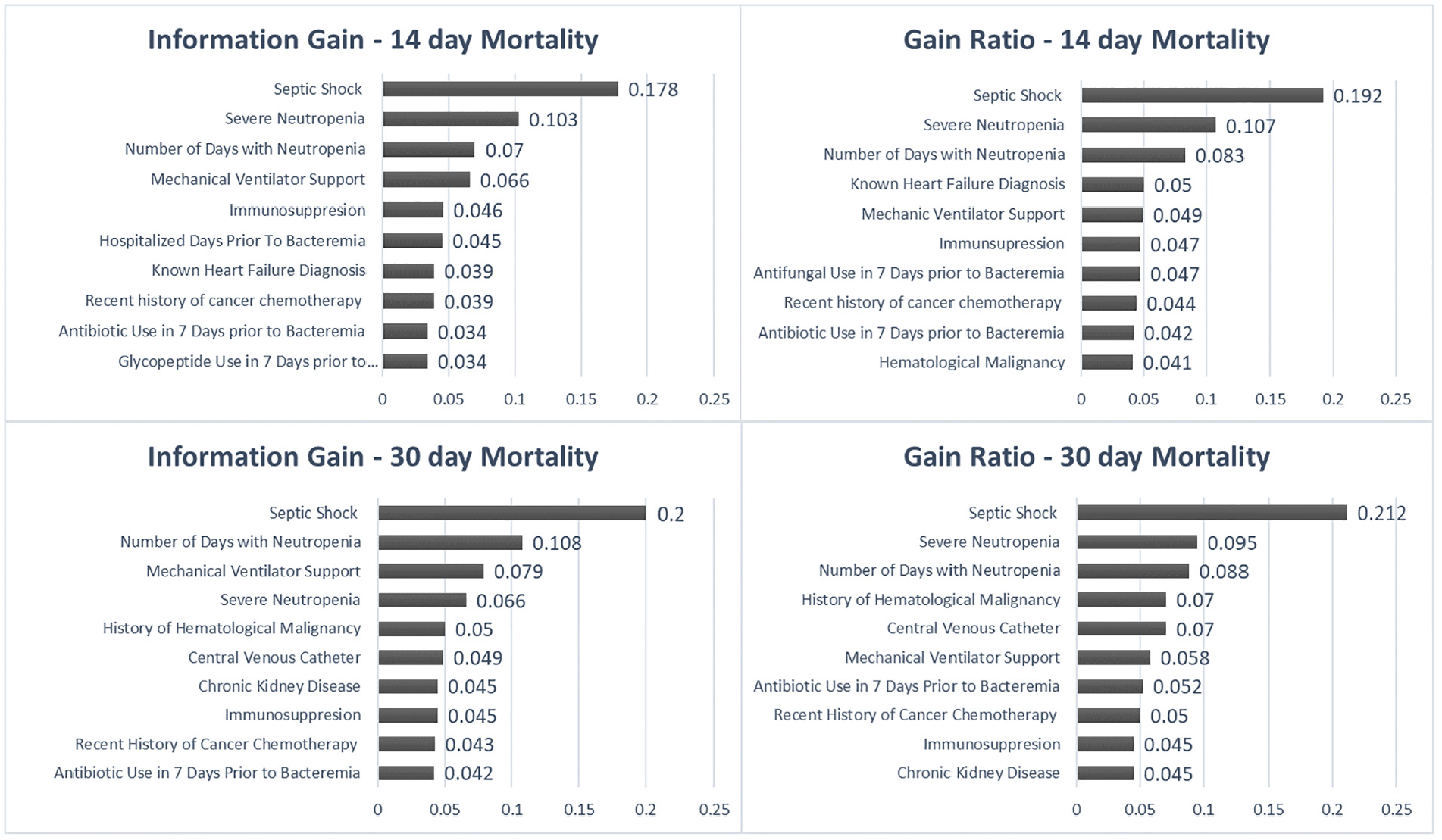
Ten most important features predicting outcome in the processed dataset.
Five machine learning algorithms were used to predict both 14- and 30-day mortality rates (Figure 3). For 14-day mortality prediction, the NB model had the highest area under the curve (AUC; 0.822) and specificity values (0.75). The RF classifier model had the highest recall (sensitivity) value (0.85). NN, LR, and SVM models showed modest performance metrics. For 30-day mortality prediction, the RF model had the highest recall, F1 statistics, and AUC values (0.82, 0.83, and 0.854, respectively), while the LR model had the highest specificity (0.815). The NB, SVM, and NN models had similar AUC, accuracy, recall, F1 statistics, and specificity values. The predictive performance of the 14- and 30-day ML models is further illustrated in the Area Under Receiver Operating Characteristics (AUROC) graphs as shown in Figure 4.
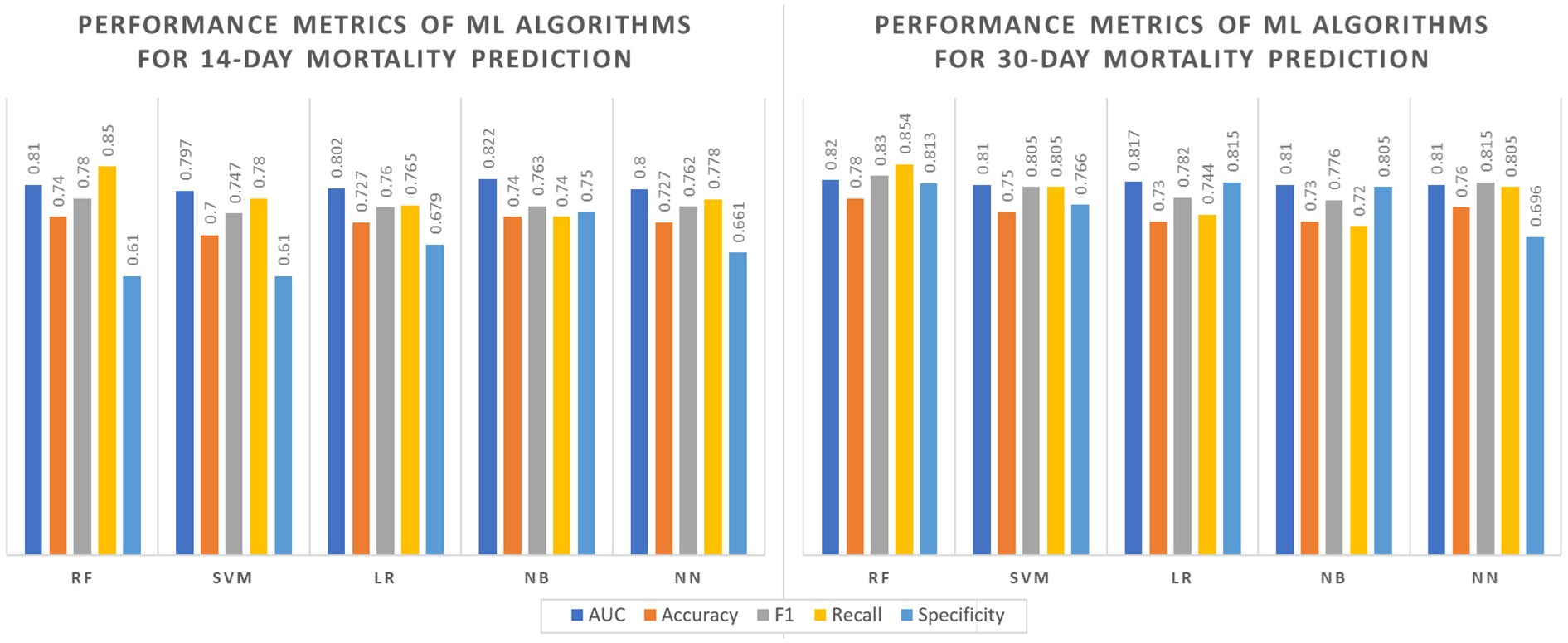
Comparisons of machine learning algorithms’ performance metrics for target variable prediction.
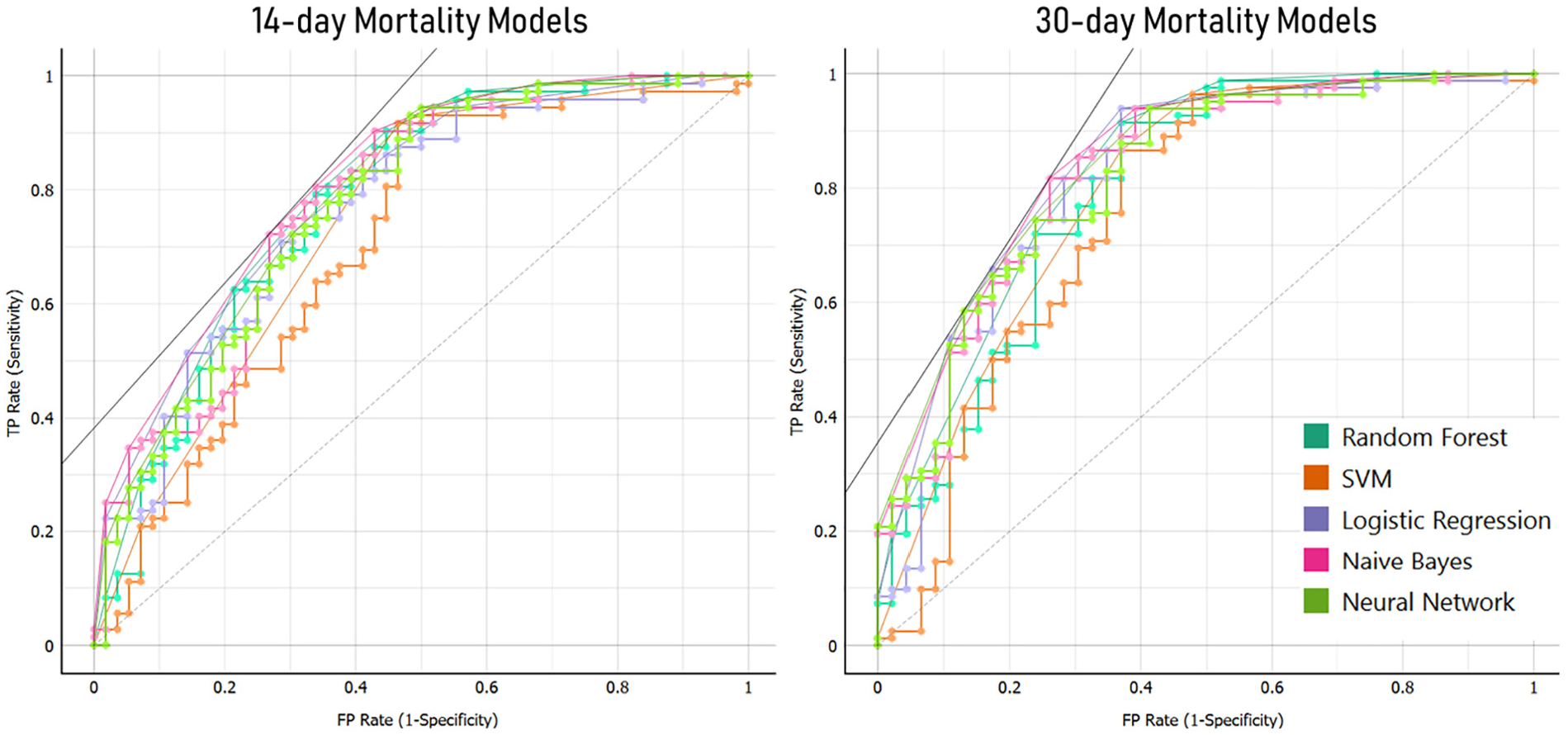
AUC plots of ROC for machine learning models predicting 14- and 30-day mortality.
Kaplan-Meier graphics were used to visualize the effect of the highest-ranking categorical variables, namely septic shock, severe neutropenia, and one medium-level ranking feature, hematologic malignancy (Supplemental File 3). The impact of appropriate empirical antibiotic therapy was also illustrated by Kaplan-Meier, showing no significant effect on endpoint prediction.
Discussion
The current investigation revealed that the presence of shock, severe neutropenia, age, number of days hospitalized before bacteremia, and active cancer condition held notable prognostic significance for both 14- and 30-day mortality. The predictive value of malignancy in mortality may vary depending on malignancy subtypes (solid vs hematological), recent chemotherapy history, and the depth of neutropenia. Machine learning models and log-rank analyses in this study identified neutropenia and hematological malignancy as significant influencers for both 14- and 30-day mortality rates. Unlike clinical attributes, the results did not indicate any connection between microbiological data and outcomes. This discrepancy might stem from an uneven distribution of data distribution concerning antibiotic susceptibilities or the considerable uniformity of OXA-like genes among the included isolates.
Previous studies have reported varying mortality rates for CRAB BSI, ranging from 22% to 62%.9,25–28 However, attributing mortality solely to CRAB BSI is challenging due to the presence of severe comorbid conditions, especially in critical care settings. Many factors have been shown to be associated with mortality in CRAB BSI cases, including underlying disease severity, comorbidity status, mechanical ventilation, age, appropriateness of empirical and definitive treatment, shock presence, immunosuppression, Acute Physiology and Chronic Health Evaluation II (APACHE II) score, Pittsburgh bacteremia score, and prior antibiotic history.25,29–33 Nevertheless, certain aspects of factors influencing CRAB BSI-related mortality remain unclear and subject to ongoing debate.
Interestingly, the prerequisite for ICU admission prior to BSI appears not to correlate with resultant mortality, in our study. Similarly, the relationship between the adequacy of empirical antibiotic treatment and the occurrence of 14- and 30-day mortality events remains unproven. Machine learning algorithms, with their ability to analyze vast amounts of data and discern patterns, further echo this observation. Drawing from this, one might postulate that mortality rates associated with CRAB BSI may be less amenable to change. The lethality of this infection is particularly pronounced in patients with pre-existing conditions such as hematological malignancies, acute neutropenia, chronic renal and cardiac failure, and a compromised immune system due to medication or illness, which is consistent with findings reported in the literature.34–36 In such cases, the factors that determine mortality seem unalterable by the time diagnosis occurs, painting CRAB BSI as a potentially preventable yet challenging-to-treat complication prevalent in patients with severe underlying diseases. This brings us to a pivotal discourse in the field: the role of appropriate empirical therapy in influencing mortality rates. Parallel to this is the debate over the most accurate time interval to assess mortality rates and identify significant predictors. Current literature shows a propensity toward 14- and 30-day mortality rates. However, the latter garners both acclaim and critique, primarily because numerous intervening variables might taint a month-long observation. The inconsistency in findings is also evident as certain studies uphold the assertion that inappropriate empirical therapy stands as an autonomous risk factor for CRAB BSI-related mortality.10,31,32,37,38 while others find no such correlation.11,39 Surprisingly, there is one cohort study that reported appropriate therapy being associated with worse outcomes in bacteremia, possibly due to a selection bias, as severely ill patients or those with high suspicion of XDR bacteremia were more likely to receive empiric colistin therapy, while their milder counterparts were less likely to receive it. 40
Furthering this discourse, it is crucial to assess not only the adequacy but also the selection of empirical treatment regimens. According to the Infectious Diseases Society of America guidelines, high-dose ampicillin-sulbactam is recommended when combined with another agent, even in non-susceptible cases, while colistin monotherapy is specifically discouraged. 41 In our study, a small subset of patients received colistin monotherapy, which does not align with these guidelines. Additionally, a minority were treated with a combination of colistin and sulbactam; however, the sample size was too limited to determine the negative effects of monotherapy conclusively or to elucidate the benefits of this combination. The emergence of novel agents, such as sulbactam-durlobactam, offers potential clinical advantages that may redefine treatment standards for CRAB. 42 However, our study did not reflect these newer approaches, as it involved an older cohort. Nonetheless, these developments emphasize the importance of choosing an appropriate regimen to manage such infections, pointing to a need for updated empirical treatment strategies that reflect microbial susceptibility and patient-specific factors.
The main limitation of this study was its retrospective nature, leading to unavoidable selection bias. Researchers heavily relied on medical records, which, despite meticulous recording, can still introduce confounding factors, such as variations in standardized sepsis treatment, compliance with septic shock protocols, or infection control practices during catheterization. Additionally, changes in recommended antimicrobial dosages and practices over time could also introduce bias. Critical scores like APACHE II and Pittsburgh bacteremia score were not available for all patients, further limiting the analysis. Moreover, only 18 out 128 patients received appropriate empirical antibacterial treatment which makes it significantly difficult to assess the effect of appropriate antibiotic treatment on mortality. Despite its inherent limitations, the study’s robustness emanates from its substantial sample size and the innovative integration of machine learning algorithms for a stratified approach to predicting mortality. This strategy allowed the delineation of factors influencing mortality across various strata. To the best of our knowledge, this marks the second study to date to incorporate machine learning algorithms into prognosticating patient outcomes. The first study, which employed machine learning-based models to predict fulminant sepsis in patients with A. baumannii, encompassed a greater range of numerical variables, including scores and laboratory values, in their algorithms compared to our study. 43 Notably, their models ranked appropriate empirical therapy as the least significant feature, and frequencies exhibited no statistical disparity between survivors and non-survivors, aligning with our findings.
This study also provides preliminary data for rational and useful risk stratification strategies for mortality and emphasizes the importance of infection control practices as a more effective strategy in the absence of promising novel antimicrobials against resistant strains.
Supplemental Material
sj-docx-1-imj-10.1177_10815589241258964 – Supplemental material for Predictive modeling of mortality in carbapenem-resistant Acinetobacter baumannii bloodstream infections using machine learning
Supplemental material, sj-docx-1-imj-10.1177_10815589241258964 for Predictive modeling of mortality in carbapenem-resistant Acinetobacter baumannii bloodstream infections using machine learning by Murat Özdede, Pınar Zarakolu, Gökhan Metan, Özgen Köseoğlu Eser, Cemile Selimova, Canan Kızılkaya, Ferhan Elmalı and Murat Akova in Journal of Investigative Medicine
Footnotes
Author contributions
Pınar Zarakolu, Gökhan Metan, Murat Akova, and Murat Özdede conceptualized and designed the study. Pınar Zarakolu and Özgen Köseoğlu Eser performed susceptibility testing and multiplex PCR. Canan Kızılkaya applied MALDI-TOF MS to strains. Murat Özdede, Cemile Selimova, and Gökhan Metan collected clinical data. Murat Özdede and Ferhan Elmalı were responsible for data curation, and statistical analysis. Murat Özdede utilized feature selection processes and machine learning algorithms. The initial draft was composed by Murat Özdede. All coauthors provided feedback on earlier drafts. The final manuscript was read and approved by all authors.
Declaration of conflicting interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: Gökhan Metan received honoraria for speaking at symposia and lectures organized by Gilead Merck, Sharp, and Dohme (MSD), and Pfizer, as well as a consultation fee from the United Nations Turkey Office. He has also received travel grants from MSD, Pfizer, and Gilead to participate in conferences. Other authors declare that they have no competing interests.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Microbiological analysis including identification, susceptibility testing, and PCR were funded by Institutional Scientific Research Project (Number: 014 D12 101 002-817) and stationary expenditure were provided by research team.
Ethics approval
This study was conducted in accordance with international and national regulations, aligning with the Declaration of Helsinki, the Human Tissue Act 2004, and the Turkish Data Protection Law. Ethical approval was granted by the university’s non-interventional clinical research ethics committee (Reference: GO 17/835, dated: October 24, 2017).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
