Abstract
The coronavirus disease 2019 (COVID-19) pandemic, which emerged in late 2019, has caused millions of infections and fatalities globally, disrupting various aspects of human society, including socioeconomic, political, and educational systems. One of the key challenges during the COVID-19 pandemic is accurately predicting the clinical development and outcome of the infected patients. In response, scientists and medical professionals globally have mobilized to develop prognostic strategies such as risk scores, biomarkers, and machine learning models to predict the clinical course and outcomes of COVID-19 patients. In this contribution, we deployed a mathematical approach called matrix factorization feature selection to select the most relevant features from the anonymized laboratory biomarkers and demographic data of COVID-19 patients. Based on these features, developed a model that leverages the deep stacking neural network (DSNN) to aid in clinical care by predicting patients’ mortality risk. To gauge the performance of our suggested model, performed a comparative analysis with principal component analysis plus support vector machine, deep learning, and random forest, achieving outstanding performances. The DSNN model outperformed all the other models in terms of area under the curve (96.0%), F1-score (98.1%), recall (98.5%), accuracy (99.0%), precision (97.7%), specificity (97.0%), and maximum probability of correction decision (93.4%). Our model outperforms the clinical predictive models regarding patient mortality risk and classification in the literature. Therefore, we conclude that our robust model can help healthcare professionals to manage COVID-19 patients more effectively. We expect that early prediction of COVID-19 patients and preventive interventions can reduce the mortality risk of patients.
⇒ Coronavirus disease 2019 (COVID-19) was the cause of severe acute lung damage, with clinical symptoms ranging from asymptomatic to moderate, with cough, fever, and muscular discomfort, to bilateral pneumonia with chronic lung failure and multi-organ dysfunction, which could ultimately lead to death.
⇒ Several machine learning and deep learning algorithms have been developed and tested based on laboratory variables and CT images to predict the death risk of COVID-19 patients.
⇒ There is a need to identify the most important clinical and laboratory features contributing to the risk of death in COVID-19 patients and to develop clinical prediction models that can integrate and analyze these parameters in real time to predict patient outcomes.
⇒ The metric factorization feature selection (MFFS) method was utilized for significant COVID-19 feature selection.
⇒ Exploring the potential of deep stacking neural networks (DSNN) model in predicting the mortality risk of COVID-19 patients.
⇒ Our analysis shows that patients at higher mortality risks had coexisting conditions, including multi-organ dysfunction syndrome (MODS) and factors such as age structure and comorbidities.
⇒ This contribution will significantly support the healthcare system in making the right decision and providing the appropriate facilities for all coronavirus patients; such preventative measures enhance patient survival rates.
⇒ The clinical care application model can be used in hospitals to identify which patients require more attention.
⇒ The clinical predictive model can also be used to predict the mortality risk of patients with other diseases such as heart, kidney, lung, and breast cancer.
Introduction
The novel coronavirus disease 2019 (COVID-19) pneumonia disease was first appeared in Wuhan, China, in late 2019 and later declared a pandemic by the World Health Organization (WHO). 1 It spreads through respiratory droplets in the air breathed when an infected person coughs or sneezes, talks, and breathes. The infectious virus causes massive devastation to public health, social order, educational systems, and economic systems. Consequently, the global infection and mortality rates are around 768 million and 6.9 million, respectively. In this situation, the existing healthcare system was insufficient to cope with such calamitous challenges. Therefore, advanced clinical technologies, such as artificial intelligence (AI) and machine learning (ML), must be industrialized to manage and identify high-risk patients with COVID-19. 2
Moreover, the majority of COVID-19 patients convalesce without intervention, but some necessitate intensive care unit (ICU) admission. The COVID-19 infection is usually elevated in the pediatric and adolescent cohorts, and the geriatric population and the subjects with concurrent pathologies were also susceptible to adverse outcomes.3–5 Currently, safe and effective vaccines are accessible in hospitals, but COVID-19 persists worldwide due to its constant genetic evolution, including the B.1.1.7 and B.1.351 variants, which pose significant challenges for the healthcare system regarding mortality risk. 6 Thus, advanced AI and ML technologies are essential to advance the healthcare system’s scalability, cost-effectiveness, and quality performance. In the meantime, AI and ML can help develop a prognostic model for COVID-19 that can predict the risk of severe outcomes and guide the optimal treatment strategies for patients.
Literature review
Since then, researchers and medical professionals worldwide have made significant efforts to effectively prognosis and minimize the proliferation of COVID-19 through various measures, including social isolation, wearing masks, routine hand washing, self-quarantine, and vaccination. Researchers have recently leveraged AI, ML, and deep learning (DL) algorithms to develop a predictive model for prompt diagnosis and prognosis.7,8
AI approaches have proven effective in predicting disease spread rates, delineating illness duration, determining the peak of infectious diseases, elucidating the course of COVID-19 transmission, and classifying the patients in healthcare systems. Traditional methods involving statistical and classical mathematical algorithms have fallen short in addressing the exigencies of the current pandemic landscape.9–13 Moreover, the ML and DL approaches might be significant for dealing with prognosis and diagnosis. 14 In the literature, the procedures and the treatment of the coronavirus were carried out by Zhang et al. 15 Chen et al. 16 studied the prevention and novel treatment of COVID-19. Holland et al. 17 set up a particular medical kit for frontline workers against the coronavirus virus. Burdick et al. 18 used an extreme gradient boosting (XGBoost) classifier model to predict invasive mechanical ventilation of COVID-19 patients within 24 h of an initial phase, employing data from the first 2 h after admission. A detailed explanation of the potential role of biomarkers in COVID-19 patients can be found in Malik et al. 19 In the study of Ramón et al., 20 five ML models were deployed to predict the mortality rate of COVID-19 patients, including k-nearest neighbors(KNN), Gaussian nave Bayes (GNB), decision tree (DT), and support vector machine (SVM), where XGB achieved better performance in terms of accuracy (92%). Yadaw et al. 21 proposed ML models, including XGBoost, Logistic Regression (LR), SVM, and Random Forest (RF), to predict the death risk of COVID-19 patients. The dataset (n = 3842) was collected between March 9, 2020 and April 9, 2020, from the Mount Sinai Health System in New York City, NY, USA. In terms of performance, XGBoost shows an excellent outcome with an accuracy of 92.0%. In their study, J. L. Guadiana-Alvarez et al. 22 utilized DL and ML algorithms to forecast the fatality risk of COVID-19 patients. The study included a sample size of 1503 patients with 48 features. The performance of the DL model was benchmarked against other models such as RF, XGBoost, LR, artificial neural network (ANN), and SVM. The DL model demonstrated superior performance with a maximum probability of correct decision (93%), accuracy (95%), precision (91%), recall (100%), and F1-score (93%).
Alakus et al. 23 developed retrospective DL models for COVID-19 patients using laboratory findings from patients examined at Hospital Israelite Albert Einstein in Sao Paulo, Brazil, and patients tested for SARS-CoV-2 in early 2020. The initial dataset consisted of 5644 patients with 111 laboratory test results. After filtering, the final dataset included 18 laboratory findings from 600 patients. In the balanced dataset, 520 patients tested negative for PCR test results and 80 tested positive. The predictive model demonstrated a maximum performance of 86.66% accuracy, 91.89% F1-score, 86.75% precision, 99.42% recall, and 62.50% area under the curve (AUC) for identifying COVID-19 patients. Moulaei et al. 24 used ML algorithms to predict the death rate of coronavirus; around 1500 patients’ datasets were collected from Ayatollah Taleghani Hospital, affiliated with Abadan University of Medical Sciences. Moreover, they consider dyspnea, ICU admission, and oxygen treatment the most predictive factors out of 38. Results showed that smoking, alanine aminotransferase, and platelet count were the least reliable predictors of mortality from COVID-19. Based on the experimental results, RF has the highest performance in terms of accuracy (95.03%), sensitivity (90.70%), precision (94.2%), specificity (95.10%), and ROC (99.97%). In addition, there are few studies of specific ML and DL algorithms to predict mortality risk in COVID-19 patients.
Our clinical predictive model is a much more sophisticated approach for minimizing the generalization error rate of one or more generalizers. Deep stacking neural networks (DSNN) work by reducing the biases of the generalizers concerning a given learning set. This minimization proceeds by generalizing in a second space, whose output is the accurate prediction. DSNN architecture is more advanced with classifiers, and it is utilized for different purposes such as preventing network intrusion, intrusion detection wireless, and indicated footballer position.25–28
Since most ML and DL models are handy for a big dataset, these models often struggle with small datasets and generalization well. To overcome these challenges, we propose to use integrated DL with stacking generalization systems that can leverage multiple algorithms. Then this system is employed to enhance the prediction accuracy of COVID-19 patients in hospitals. Our study offers the following contributions:
The clinical predictive DSNN model was implemented to predict mortality risk in COVID-19 patients based on laboratory markers and demographics instead of CT images or radiograph data.
The matrix factorization feature selection (MFFS) mathematical system was deployed for the significant feature selection.
A dataset of COVID-19 patients from Hospital Universitario HM Madrid, Spain, was utilized to validate the novel predictive model. The experimental results showed that DSNN outperformed state-of-the-art algorithms.
The efficacy of the DSNN model was corroborated by two distinct experiments, one with 24 significant features and another with nine paramount COVID-19 features.
The management of COVID-19 patients at advanced levels was ensured by biomarker functionality in patients with systemic manifestations of the disease.
Multiple organ dysfunction syndromes (MODS) and other comorbidities associated with COVID-19 in mortality risk were studied.
The clinical predictive model’s performance was evaluated with accuracy, recall, F1-score, precision, specificity, AUC, and maximum probability of correction decision (MPCD).
To our knowledge, no study has yet utilized DSNN algorithms to predict the fatality risk of COVID-19 patients based on laboratory tests and demographic results. This study may motivate other researchers to test the model in the laboratory using different biomarkers, which also can be used for other diseases.
The rest of the study is structured as follows. After reviewing the related works, the methodology for early database characterization of COVID-19 patients was discussed in the section “Materials and methods.” We then present the ML and DL application models and performance metrics in the section “Results.” Section “Discussion” details the experimental setup, results, and discussion. Finally, we conclude the study with our findings in the section “Conclusions and future study.”
Materials and methods
The proposed methodology is illustrated in Figure 1 with a schematic diagram. The approach includes several steps: (i) describing and cleaning the dataset, (ii) using the MFFS-based approach for selecting significant features, (iii) applying data splitting techniques, and (iv) developing, training, and testing the proposed model. The expounded details are elucidated in the following subsections.
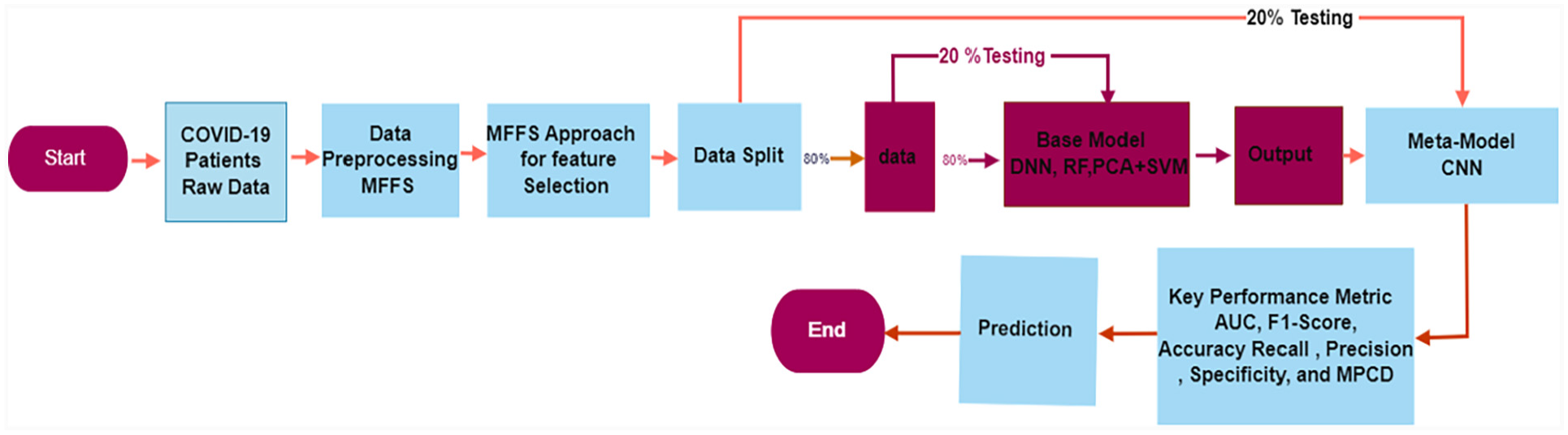
Schematic exemplification of the clinical care model.
Clinical dataset description
This study leveraged big data analytics techniques to analyze a large-scale clinical dataset of 2307 PCR-confirmed COVID-19 patients from Hospital Universitario HM Madrid, Spain. The raw dataset comprised 92 features and included a varied patient population of 1377 males and 930 females. The clinical data have shown that the majority 71% of patients had favorable outcomes and were discharged from the hospital. However, a significant proportion of 16% of patients had fatal outcomes and succumbed to the disease. The remaining 13% of patients were still receiving treatment in the hospital at the time of data collection. Patient identity (ID) was maintained using a unique identifier to ensure data privacy and security.
Data cleaning and feature selection technique
In the data cleaning phase of our study, we faced several technical challenges with our raw dataset, such as class imbalance, inconsistent units of measurement, redundant information, a mix of numerical and categorical data types, and missing values. To fix all these problems in the dataset, we applied various data preprocessing techniques such as resampling to balance the classes, feature selection to remove redundant information, unit conversion to standardize the units of measurement, encoding to convert categorical data into the numerical format, and the KNN imputation scheme was deployed to predict the missing values based on the similarity or distance between the existing records. We applied specific filtering criteria to our raw dataset to ensure quality and relevance. We only included records of patients who had a PCR test positive and whose outcomes were known (either discharged or deceased). In addition only included the successive aged ≥
However, some of these features still did not meet our inclusion criteria. Therefore, applied the MFFS approach to refine the dataset features further and identify the most significant ones for predicting the fatality risk of COVID-19 patients in advance. Wang et al.
29
proposed the MFFS scheme for important attribute selection. This method assumes all features exist on a linear manifold within the Euclidean space
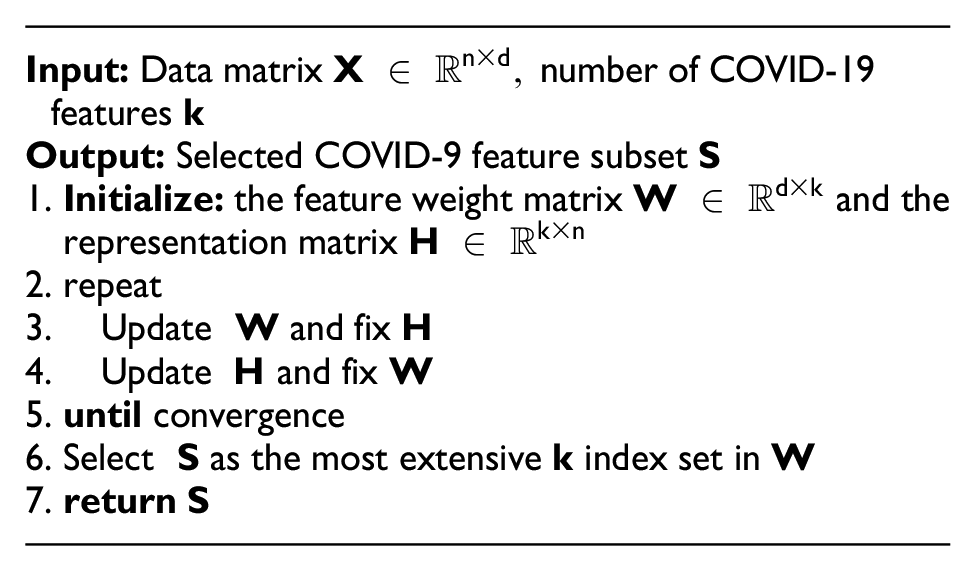

where
After implementing the MFFS technique, there was a significant reduction possible in the number of COVID-19 features. Out of the 40 features, only 24 were retained based on high rank. Of these 24 features, 17 belong to laboratory markers, 6 comorbidities features, and the age of patients. Features 1–18 were considered numerical data
MFFS approach deployed to select significant features in COVID-19 patients: |S|= feature important score;
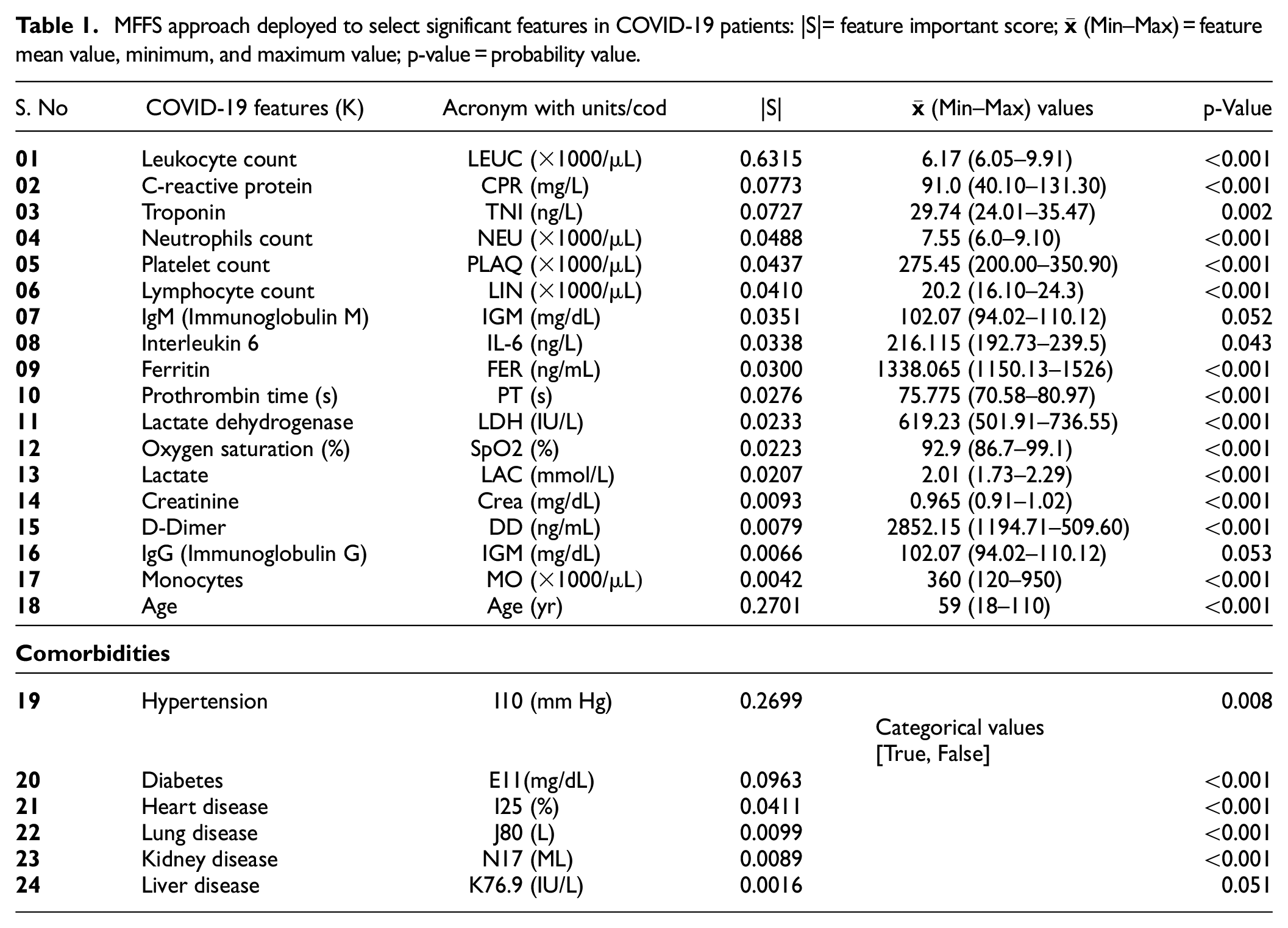
In addition, the statistical approach t-test and p-value have been utilized further to verify the significance of the selected 24 features, and a t-test was deployed to compare the means of two sets of features and determine whether their differences were statistically significant. The p-value represents the probability that the difference between the two sets of features is due to chance. If the value of
Moreover, to optimize the DSNN model for minimal features and maximal performance, we conducted another experiment with only nine key features. These features were chosen based on their statistical significance and clinical relevance: leukocyte, age, hypertension, C-reactive protein, troponin, neutrophils, interleukin, platelet count, and lymphocyte. These features reflect the immunopathological response and the comorbidities of organ dysfunction that lead to COVID-19. Among all the delineated features in Table 1, only leukocytes, age, and hypertension have the highest score
Experimental setup
The entire experimental process is illustrated in Figure 1 and consists of three main steps: (1) data processing and cleaning, (2) development of the proposed model, and (3) model evaluation and prediction.
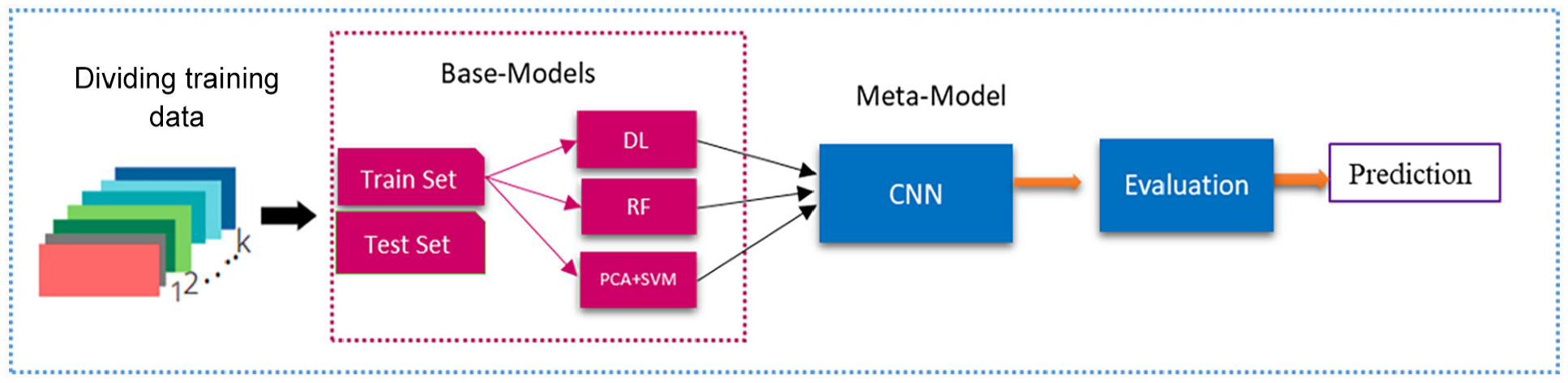
The deep stacking neural network architecture.
Machine learning and DL application models
DL is a subfield of machine learning, which itself is a branch of AI. DL models use multi-layered neural networks to learn and make predictions or decisions without being explicitly programmed to perform these tasks. 34 Voluminous research in healthcare has demonstrated the varied applications of AI in healthcare-related sustainable development goals. The implementation of AI has commenced for a wide variety of health problems, including tuberculosis and malaria, the diagnosis and prognosis of patients afflicted with COVID-19, diagnosing cancer by mammography, identification of brain tumors via magnetic resonance imaging, and predicting Alzheimer’s disease.35–38 The advancement of AI transformation may guide medical professionals to provide more effective care and treatment closer to the patients. AI never controls patients’ illnesses and gives them superhuman disease management skills. 39 To select the most robust model for predicting the inflammatory and immune responses of COVID-19 patients, we applied and compared four different models: PCA with SVM, RF, DL, and DSNN. These models were trained and tested using laboratory test results and demographic data of COVID-19 patients.
PCA with SVM
The PCA is a well-established statistical tool deployed to examine the correlations between items and variables to determine principal components with a significant correlation. Deploying the few fundamental principal components changed a high-dimensionality dataset into a low-dimensional dataset, lowering the dataset’s dimensionality. As a result, the number of principal components equals the number of eigenvalues of the correlation matrix, with a value always greater than 1. SVM is an ML algorithm for classification and regression analysis. When SVM combines with PCA, it can improve the model’s performance by reducing the features’ dimensionality and removing noise from the data. 40 In this contribution, PCA with SVM was deployed as a base model to classify COVID-19 patients and the fatality risk prediction strategy for minimizing preventable deaths from the public health perspective.
Random forest
It is an ensemble approach that joins multiple decision trees. Because of randomness, it offers much greater performance than a single decision tree and can reduce model variance. The RF models easily handle all kinds of regression and classification issues because it has less sensitive and requires fewer parameters to configure.
41
It performs better when the number of variables is significantly greater than the number of observations. Let us consider the input variable
Phase 1: Supposed that a bootstrap sample
Phase 2: Develop a random forest tree
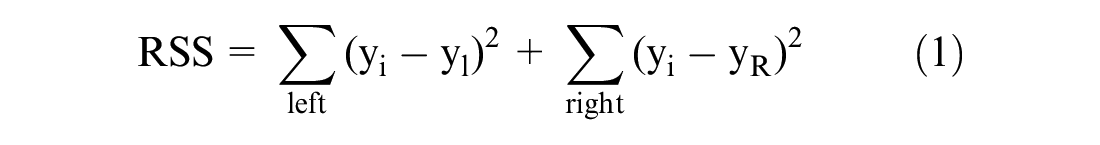
where
Deep learning
Recently, DL architectures have garnered great interest across myriad disciplines due to their unrivaled representational capabilities, particularly in computer vision, but they are also expanding into other fields, including medical diagnosis, prognosis, and predictions. DL uses a hierarchy of several layers to understand different representation levels. 43 DL is a neural network with three primary layers: input, hidden, and output. Mathematically presented as in equation (2).
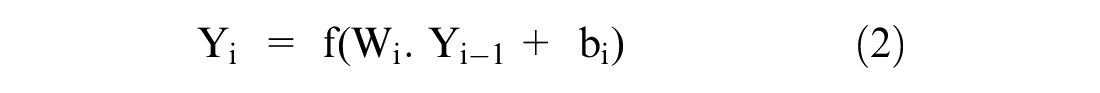
The above equation shows the output of the layer in the network, where f, Wi, b, and Yi–1 are, respectively, defined as the activation function, weight, bias, and output of the previous layer. In addition, the network has multiple hidden layers, each subject to the output of the last layer; this ultimately generates the output variable Yi as COVID-19 patients’ mortality risk.
Deep stacking neural network
A DSNN is a type of neural network architecture. It is inspired by DL and the idea of stacked generalization. It combines several different models (or heterogeneous ensembles) to create a more robust and accurate model.
28
The complete architecture of the DSNN model is seen in Figure 2. Also, in the main unit of DSNN, the hidden layer and output layer were sigmoidal nonlinear and linear, respectively. The linear property ensures greater competence and optimization for the output network’s weight. The DSNN architecture includes the fundamental structure of single and multi-layered units; the functional neural network unit consists of two training sets of weights and one hidden layer. The lowest unit’s topology consists of the linear input and hidden output layers with linear and nonlinear groups. It is significant to observe that the fine-tuning model of DSNN weight is implemented in each unit rather than generally through all layers. Total training samples in activity matrix H are known in the hidden layer. Hereafter, the upper weight matrix U can be efficiently learned. Suppose that the training vector X = [x1, …, xi,…, xN] in which each vector xi is represented as xi = [x1, …, xji, …,

where
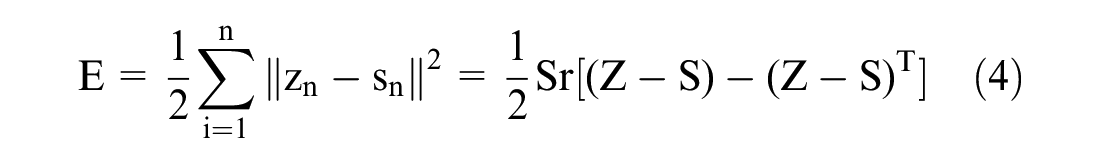
for output,
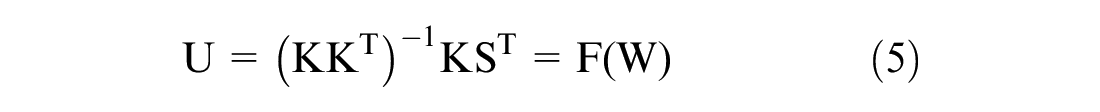
Deploying Lagrangian multiplier44,48 to resolve the optimizing problem for
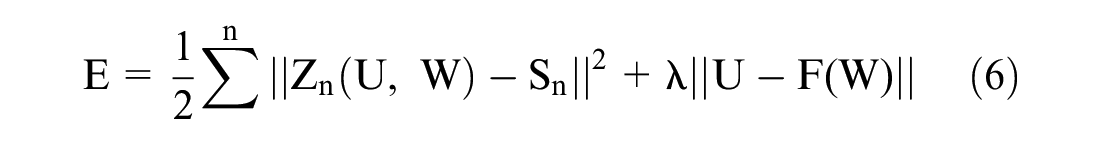
To fine-tune the W output weight matrix, Batch-mode gradient descent45,49 was used; it can be calculated as in equation (7).
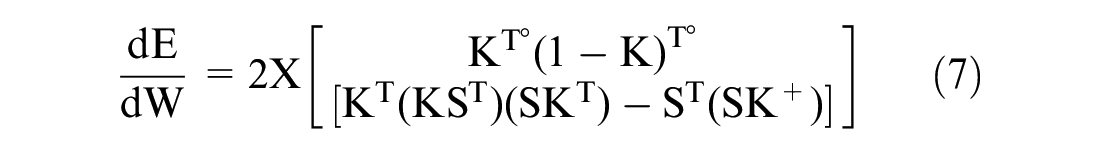
where
Key performance measures
The proposed models were evaluated using performance metrics. COVID-19 patients who were discharged or survived were classified as class 1, while those who deceased were classified as class 0. The performance metric helps us understand whether the model can discriminate between the two classes. It provides metrics such as true positive (TP), false positive (FP), true negative (TN), and false negative (FN). The KPM used were recall, precision, specificity, the AUC, F1-score, accuracy, and MPCD.22,46
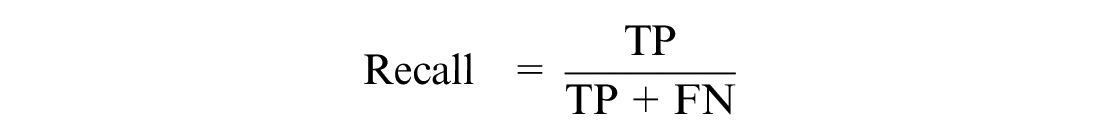
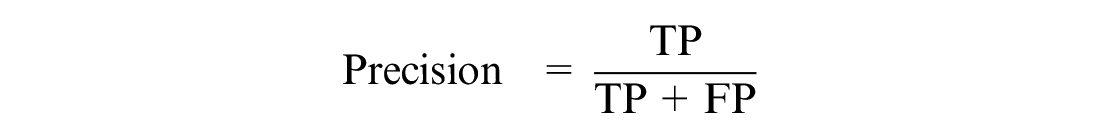
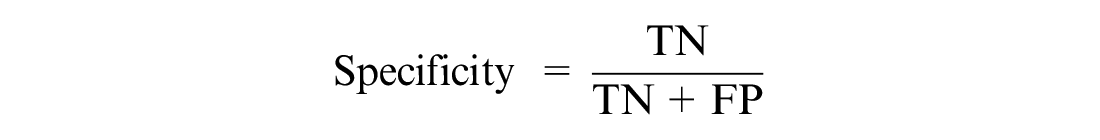
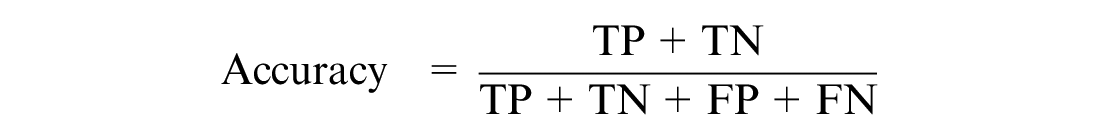
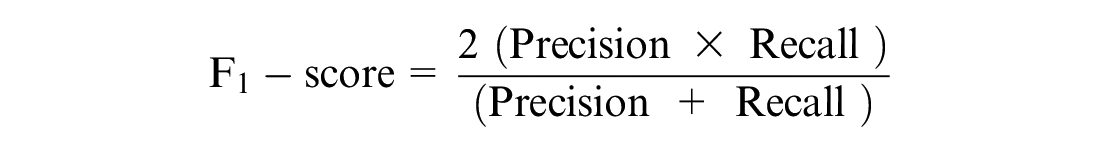
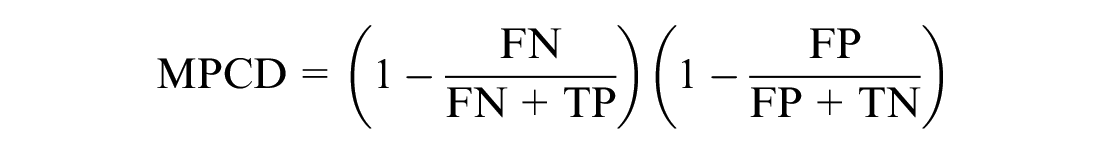
where
Mean Absolute Error (MAE) is the average of the absolute difference between the predicted and actual values. The formula for MES is:
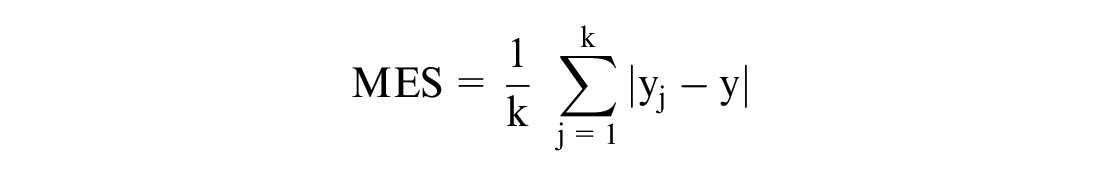
where
Results
This section commences by elucidating the descriptive statistical analyses of the important factors that exacerbate the mortality risk of COVID-19 patients; it also presents the main results of the proposed model.
Symptom-based characterization of COVID-19 patients
There is considerable interest in employing symptoms to prioritize COVID-19 diagnostics and establish symptom-based management. However, this approach may be unsuccessful in identifying patients who require testing for COVID-19 infection. The criteria to determine who should undergo RT-PCR testing vary from region to region and ultimately depend on hospital resources and public health goals. The classification of patients based on COVID-19 early symptoms can be divided into three categories. (i) General symptoms: One epidemiological patient history, two clinical symptoms, one etiological indication of COVID-19 infection, and patients with pneumonia indications on CT scans or PCR are required. (ii) Mild symptoms: Fever, cough, fatigue, body aches, headaches, sore throat, and chest distress, required isolation. (iii) Severe symptoms: At rest, the
Biomarkers-based classification criteria for COVID-19 patients
A characteristic that can be scientifically assessed as an indicator of normal biological and pathological processes is a biomarker. Biomarkers have become an important diagnostic method in managing COVID-19 patients, the etiological organism for COVID-19. 19 In hospital practice, biomarkers ensure that resources are utilized as effectively as possible to improve clinical treatment and avoid serious complications. 48 COVID-19 is a multisystem disease caused by a diffuse systematic process with a complicated combination of immunological inflammatory and coagulated cascades. The genetics and acquired variations in the host immune system further affect the host with wide heterogeneity in the clinical picture, course, and outcomes. 19 The COVID-19 biomarkers offer valuable information such as detection and classification to set prognosis criteria, timely diagnosis, manage high-risk cohorts, formulate hospital admission and discharge criteria, rationalize therapies, and forecast mortality risk.
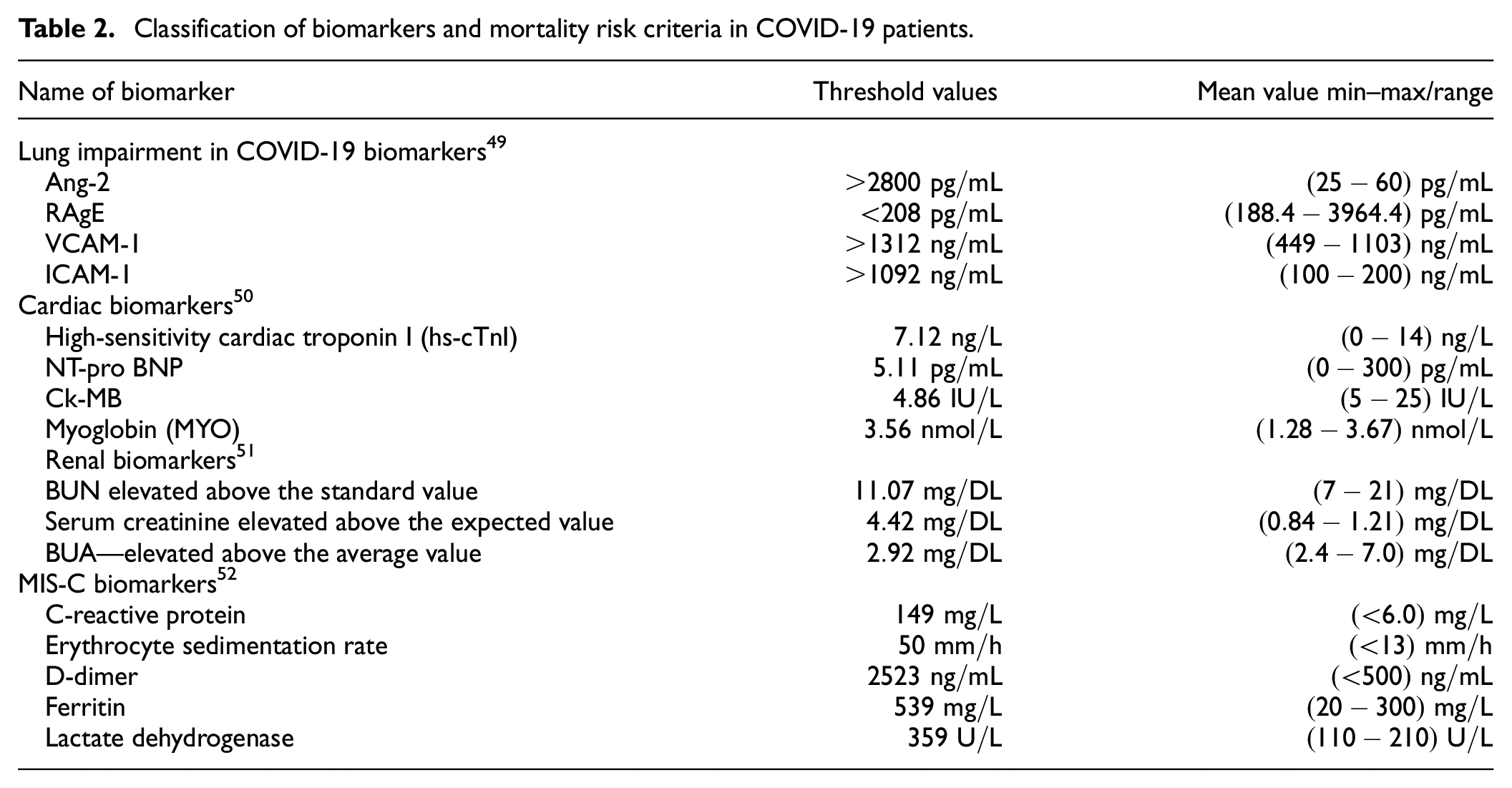
Also, the use of COVID-19 biomarkers in children is highlighted, particularly to forecast multisystem inflammatory syndrome in a child (MIS-C). Eventually, biomarkers can be employed as predictive tools for doctors to identify and effectively manage patients more likely to suffer adverse outcomes from COVID-19. We aim to use biomarkers results and predict the death risk of COVID-19 patients and classification criteria using the DSNN model. The predictive criteria for biomarkers derived from various literature sources are shown in Table 2.
Classification of biomarkers and mortality risk criteria in COVID-19 patients.
COVID-19 association with multi-organ dysfunction syndrome
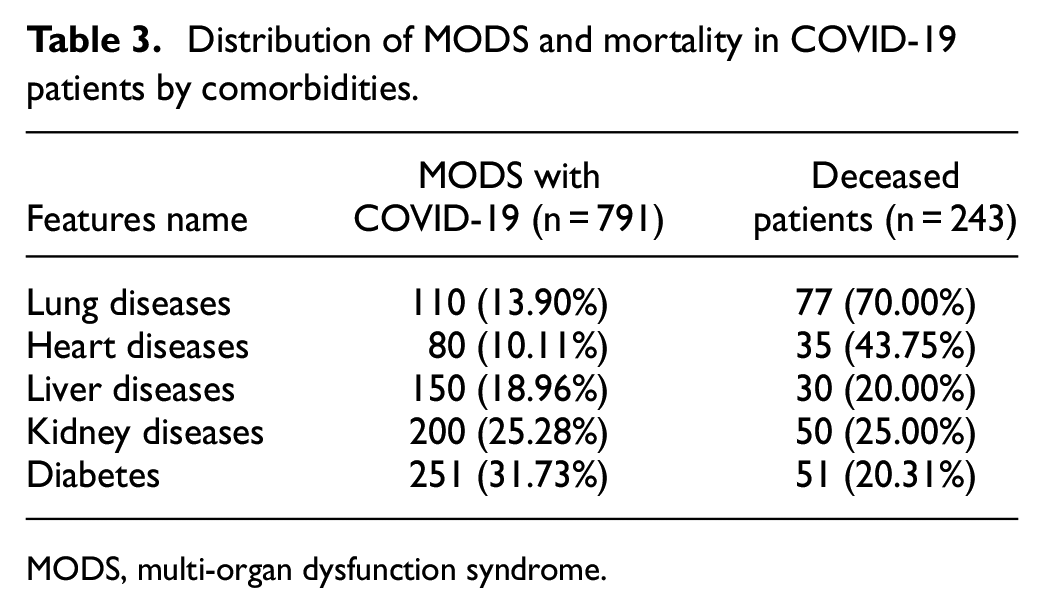
Although there have been a lot of basic and clinical studies on MODS, much work remains to be done to keep patients safe. 53 Dataset information reveals that MODS is one of the critical factors in increasing the risk of death among COVID-19 patients. Table 3 shows the distribution of MODS and mortalities in COVID-19 patients; of the 1463 COVID-19 patients in the dataset, 791 were diagnosed with MODS. In addition, 110 of these 791 patients had lung disease, and 77 died. Heart disease was another risk factor for mortality; 35 of 80 patients with this disease had a fatal outcome. The liver disease affected 150 patients, of whom 120 were successfully treated and discharged from the hospital. Kidney disease can lead to dangerous complications; only 150 of 200 patients with this condition survived. When coronavirus infects diabetic patients, the outcome can be worse; according to data analysis, 51 of 251 patients died. According to the data analysis, lung infection contributed most to the mortality rate of COVID-19 patients. This study examined the impact of MODS on the fatality risk of COVID-19 patients. The results showed that COVID-19 patients with MODS had a higher mortality risk than those without MODS. Especially, lung diseases showed a higher mortality rate among COVID-19 patients, and these findings suggest that the fatality of COVID-19 patients depends on various factors, and MODS is one of them.
Distribution of MODS and mortality in COVID-19 patients by comorbidities.
MODS, multi-organ dysfunction syndrome.
COVID-19 with other comorbidities
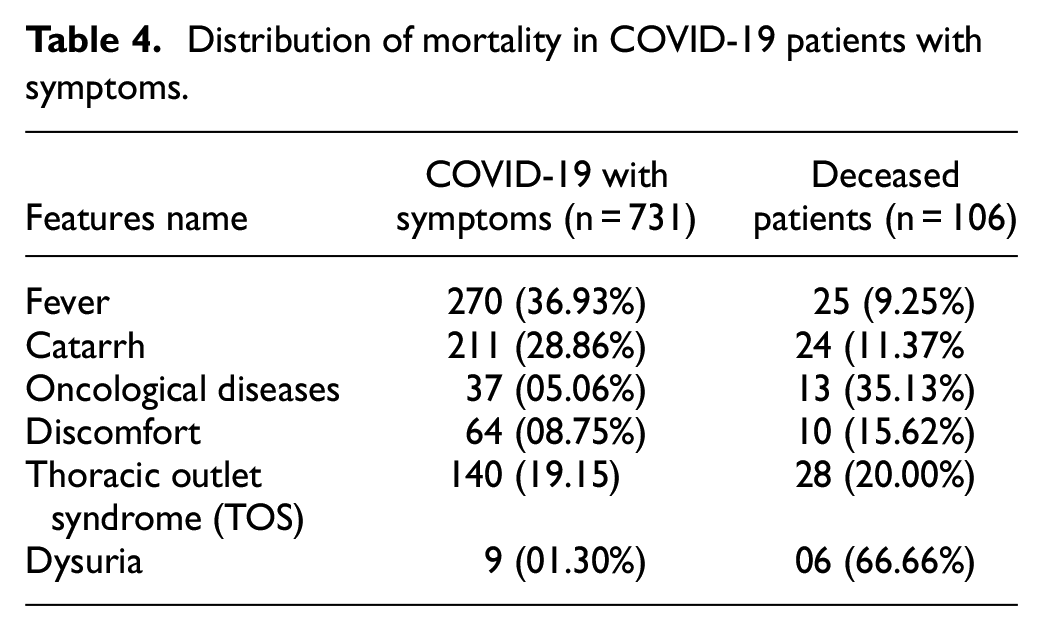
The mortality risk of COVID-19 patients was also influenced by other comorbidities such as fever, catarrhal table, oncological diseases, discomfort, thoracic outlet syndrome (TOS), and dysuria. Table 4 presents the statistical facts about the mortality risk of COVID-19 patients along with comorbidities or symptoms: according to data analysis, 9.25% of 270 patients with fever died. Catarrh is a condition that involves phlegm and mucus in the throat and nose. Influenza, colds, and allergies can cause it. COVID-19 mortality and catarrh are unclear. Catarrh by itself is usually not a serious condition. However, it can be a sign of a COVID-19 infection, which can lead to complications and increase the chance of death. 54 According to the data records, 11.37% of the 211 patients who had catarrh with COVID-19 died. Oncological disease, commonly referred to as cancer, presents a multifaceted association with COVID-19. In general, patients diagnosed with oncological conditions may exhibit increased vulnerability to contracting COVID-19 and may experience greater severity and fatality rates after infection, 55 35.13% of 37 patients deceased with the oncological disease, 15.62% of 64 patients passed away with general distress. TOS compresses nerves, veins, and arteries between the base of the neck and upper chest. It affects shoulder, arm, and hand pain, numbness, and weakness, 56 20% of 140 patients died with TOS, and 66.66% of nine patients with dysuria died as reported in the dataset. Hence, it is apparent that the co-occurrence of preexisting medical conditions with COVID-19 significantly influences the mortality rate of COVID-19.
Distribution of mortality in COVID-19 patients with symptoms.
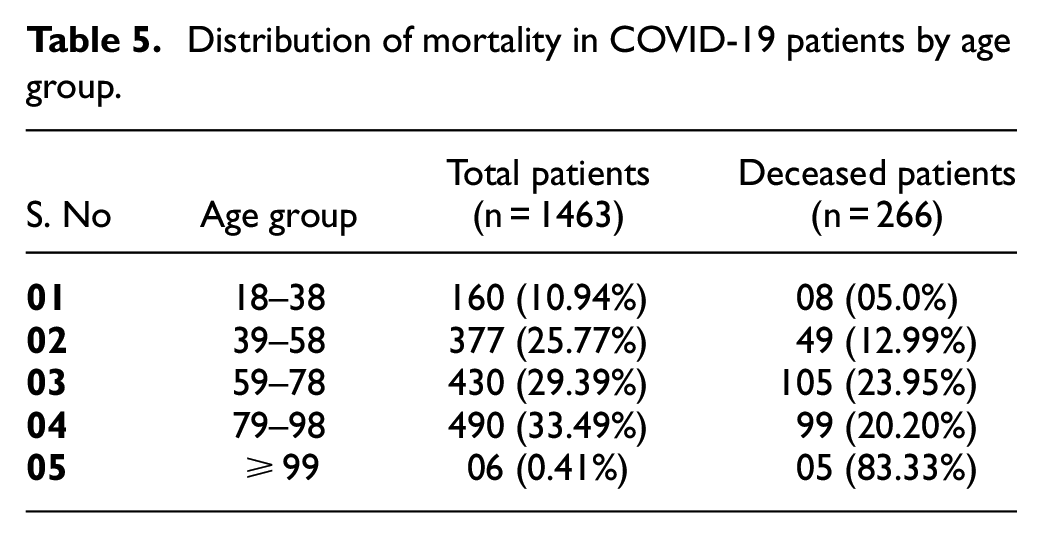
Notably, the study included 1463 patients who were infected with COVID-19. Among them, 266 succumbed to the disease, from 18 to 110 years old. Another cause of death was classified into MODS and other comorbidities. Some patients had both MODS and comorbidities, while others had only one of them. Among the 266 fatalities, 243 were related to MODS, either by itself or in conjunction with other comorbidities, and 106 were attributed to other comorbidities, either alone or in conjunction with MODS. Therefore, the total number of deaths was equal to the combination of MODS-related and comorbidity-related deaths without counting the patients who had both conditions twice. In other words, the union of MODS and other comorbidities deaths equal to the total death representing in Table 5.
Distribution of mortality in COVID-19 patients by age group.
COVID-19 with age structure
The study by Rymkiewicz 57 reported that the increased mortality rate of COVID-19 in geriatric populations is attributed to immunosenescence. The descriptive statistical analysis in Table 5 indicates that COVID-19 distresses all age groups, but not equally. Geriatric populations have a higher mortality rate compared to working folks. The infection (morbidity) rate is typically higher in younger individuals, while mortality is higher in the geriatric population.
Key results from the proposed model
This subsection delineates the experimental outcomes and furnishes a discourse on the findings, which entailed devising and implementing clinical prediction classifiers, including PCA + SVM, RF, DL, and DSNN. In this contribution, two experiments were executed; both utilized identical models and conformed to the same performance standards. However, the number of features deviated between the two experiments. So, in the first experiment, 24 features were utilized for all proposed classification algorithms to generate mortality predictions and assess the performance of the clinical prognostic models, as the outcomes illustrated in Table 6. Figure 3 compares the performance of the proposed clinical predictive models, where the DSNN model outperforms other models in all seven metrics, as evidenced by the higher bar height. This study applied PC + SVM to prognoses COVID-19 patients, and the performance profile of this classifier can be visualized in Figure 3, including F1-score (0.972), AUC (0.914), recall (0.969), accuracy (0.972), precision (0.975), specificity (0.950), and MPCD (0.899). The RF classifier demonstrated strong performance across all seven indicators, as proved by the AUC (0.869), F1-score (0.953), recall (0.950), accuracy (0.975), precision (0.957), specificity (0.907), and the MPCD (0.887). The DL algorithm was implemented using and obtained excellent results across multiple metrics, as indicated by the AUC (0.941), F1-score (0.966), recall (0.960), specificity (0.965), accuracy (0.971), precision (0.973), and MPCD (0.920). The DSNN model outperformed all other application models, achieving an AUC (0.960), F1-score (0.981), recall (0.985), accuracy (0.990), precision (0.977), specificity (0.970), and MPCD (0.934). In addition to that, the clinical care model exhibited a high-performance indicator in terms of accuracy achieved on both the training and test sets, which attained 0.990 and 0.985, respectively, and also visualized model predictive accuracy and loss function in Figure 4, the training and testing accuracy and loss function curves at some stage closely aligned, implying that our model looks stable and generalizing well to new data.
Summary of the outcomes of clinical predictive models with 24 attributes.
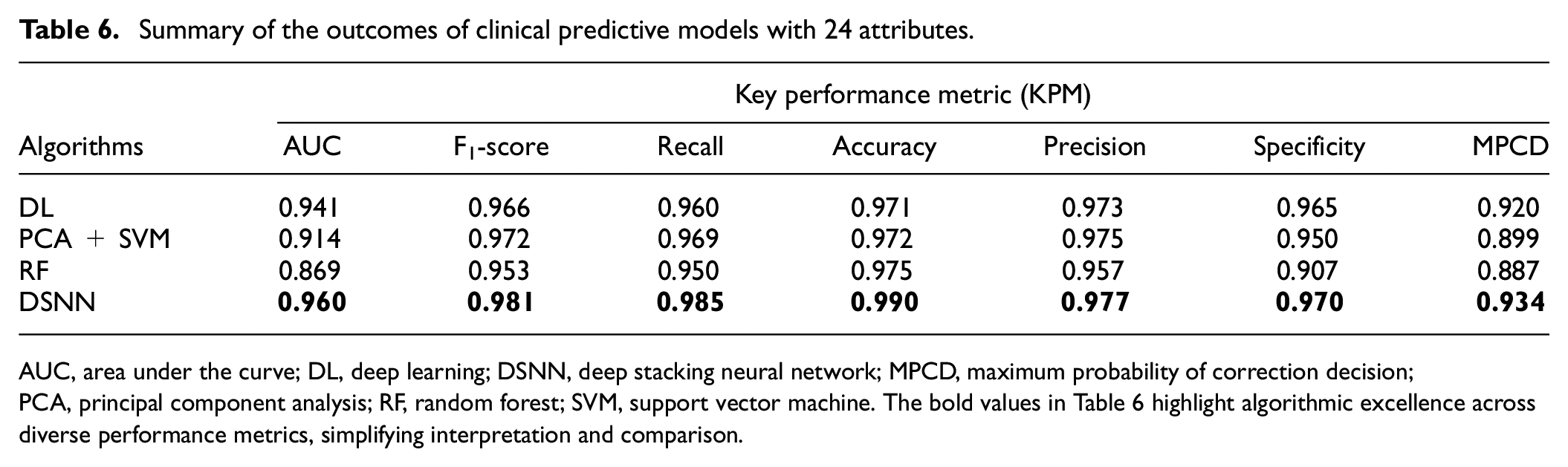
AUC, area under the curve; DL, deep learning; DSNN, deep stacking neural network; MPCD, maximum probability of correction decision; PCA, principal component analysis; RF, random forest; SVM, support vector machine. The bold values in Table 6 highlight algorithmic excellence across diverse performance metrics, simplifying interpretation and comparison.
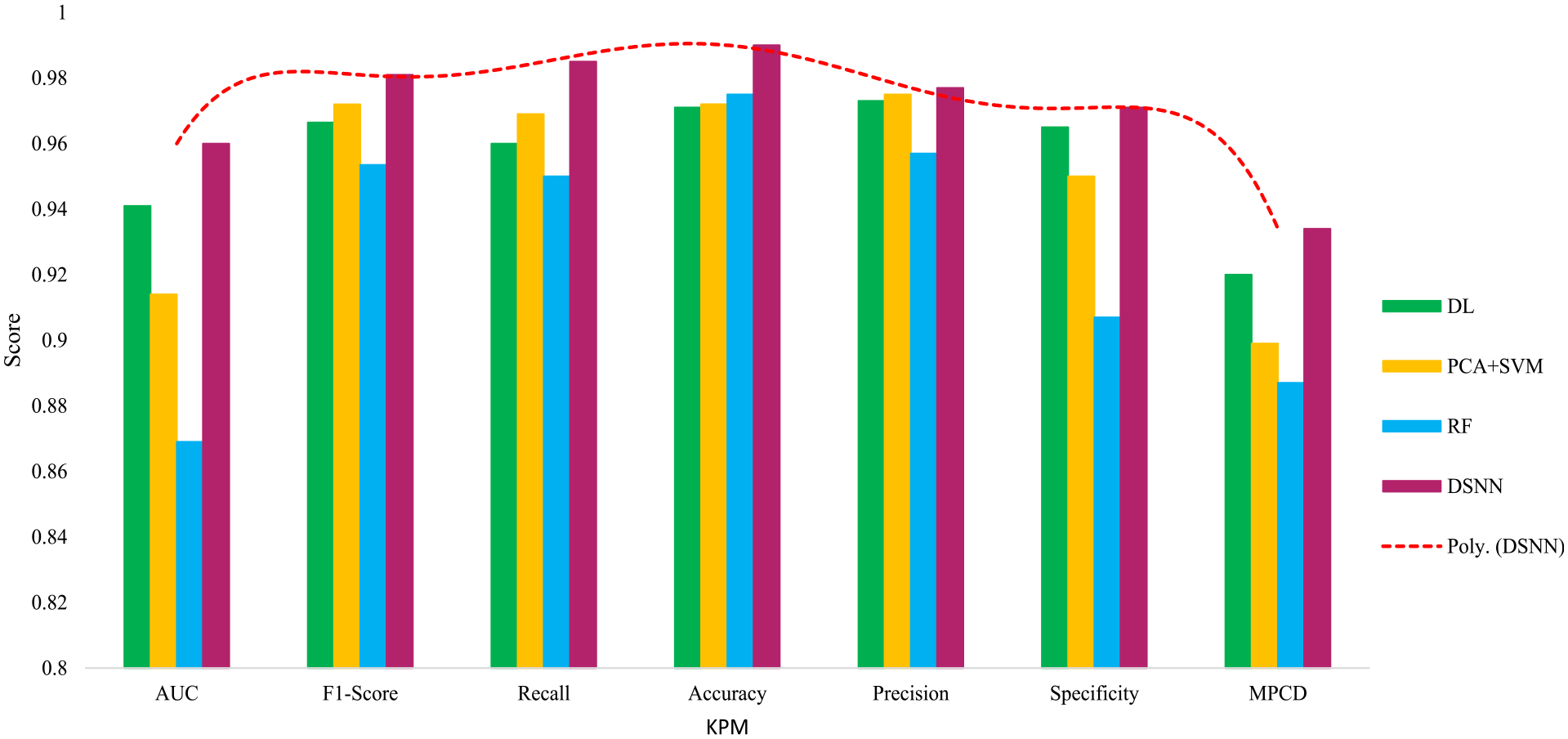
Performance comparison among clinical predictive application models.
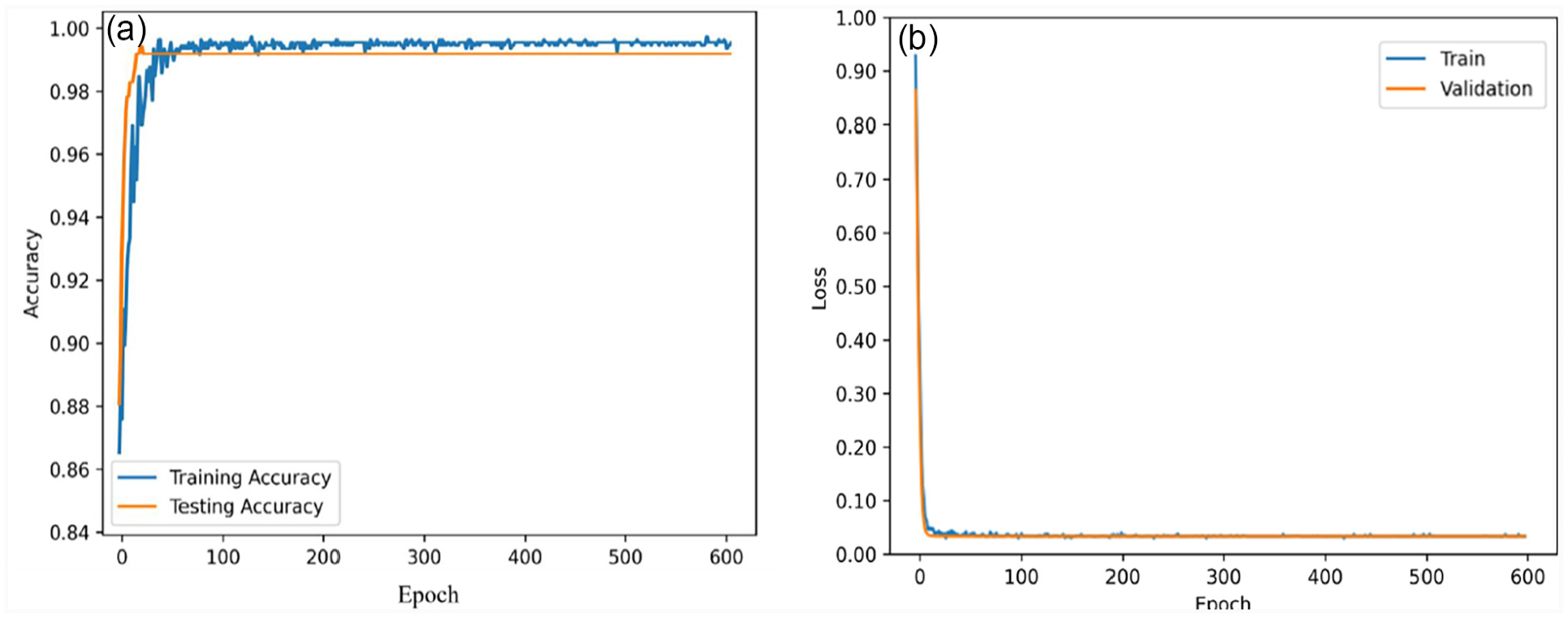
The performance of the DSNN model with 24 features is shown by its (a) accuracy and (b) loss.
In the second experiment, we used a parsimonious set of 09 features for forecasting COVID-19 patients’ mortality risk. This experiment trained and validated the DSNN model to ensure that the proposed model could attain high performance and efficiency with minimum features. The model achieved an AUC (0.971), F1-score (0.934), recall (0.927), training accuracy (0.932), test accuracy (0.925), precision (0.941), specificity (0.966), and MPCD (0.901), also shown in Table 7. Hence, Figure 5 describes the accuracy and loss function of the proposed model with nine features, indicating its capability to forecast the mortality risk of COVID-19 patients accurately.
Summarizes the comparison of our study with existing studies for mortality prediction in COVID-19 patients.
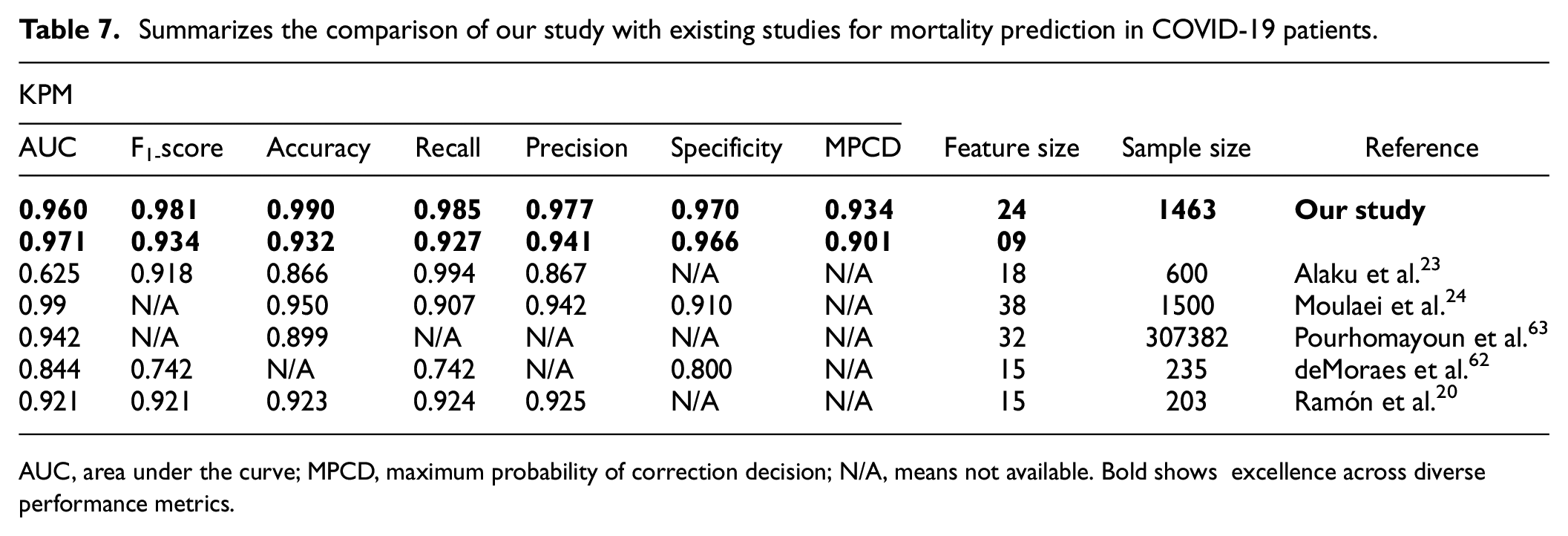
AUC, area under the curve; MPCD, maximum probability of correction decision; N/A, means not available. Bold shows excellence across diverse performance metrics.
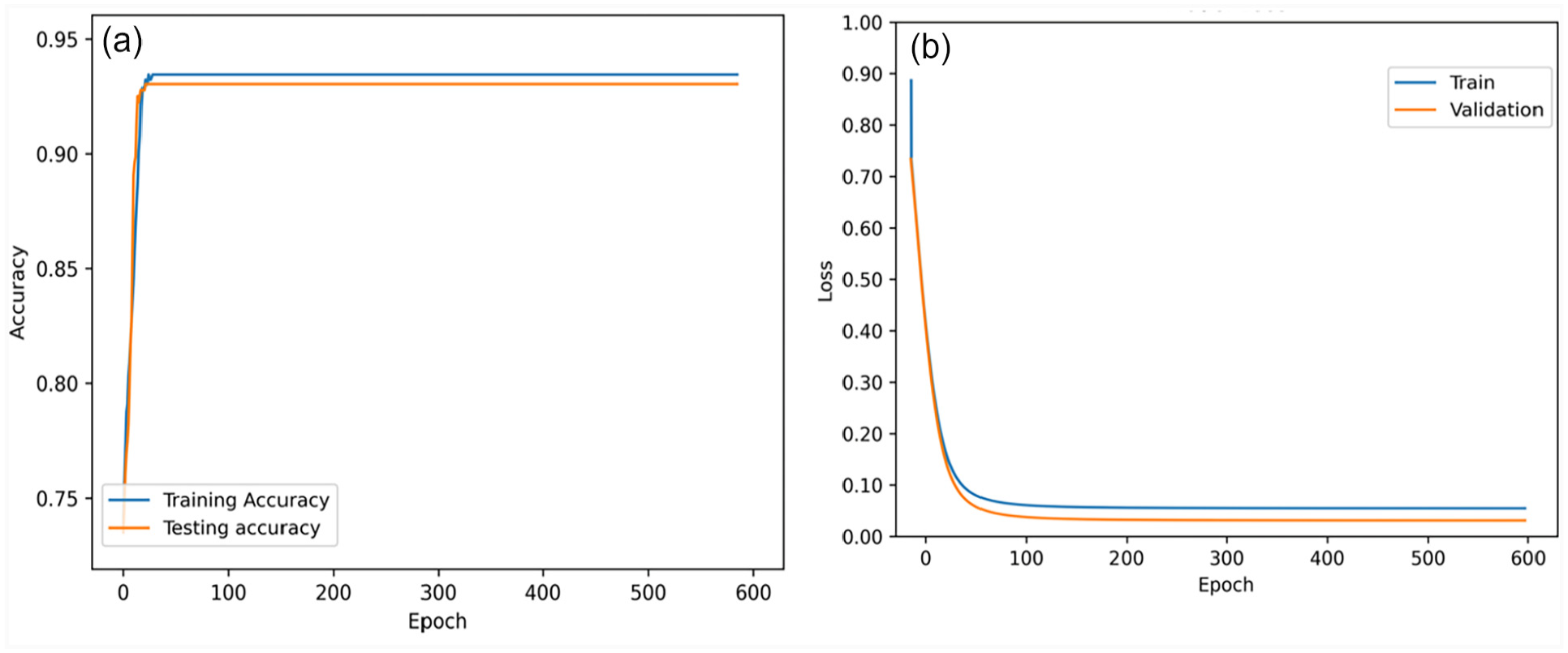
The performance of the DSNN model with 09 features as shown by its (a) accuracy and (b) loss curves.
Discussion
This section discusses the results of the proposed models, their implications, and limitations, and compares them with the existing works on ML and DL models for COVID-19 prognostication. Furthermore, we analyze the performance of each KPM for all proposed models: PCA + SVM, RF, DL, and DSNN, by conducting an experimental study using two distinct feature sets. First, we discuss the outcomes of using the clinical predictive model on a dataset with 24 features, and second, we explore the effects of reducing the feature set to 9 features, as follows.
Accuracy is a study feature that indicates how well experimental parameters correspond to population parameters and ensures the validity and reliability of the results. 58 Our all-predictive models achieved an accuracy greater than 0.970, demonstrating their robustness and performance in predicting fatal risk among COVID-19 patients. In particular, our DSNN model achieved test and train accuracies of 0.985 and 0.990, respectively. Clearly, accuracy serves as a rudimentary metric to evaluate the performance of the model. Therefore, to make a robust prediction model, one more advanced performance indicator has been measured, named precision and recall. Precision refers to the proportion of correct detections reported by a model. In simple words, the ratio between the true positive observation and the total predicted positive observation. The clinical model’s perfect precision would have been 1. 59 Our clinical predictive models obtained outstanding precision scores exceeding 0.950, particularly DSNN being the best with a score of 0.977, it nearly optimal precision. Recall efficiently predicts the true positives of each available category. In the significant classification process, the recall value for the best model is always measured as 1. 59 All four models obtained the best recall value greater than 0.950, while the DSNN model gained 0.985. Instead of using a curve to evaluate the classifier’s performance, we want to use a single scalar metric, or F1-score, and compute the F1-score as the harmonic mean of precision and recall 59 ; and suggested models obtained F1-score greater than 0.950, especially DSNN achieved 0.981. The AUC was applied in the classification task to ascertain the classes that yielded the most accurate predictions. The evaluation criteria for the AUC are delineated in Mandrekar. 60 When the AUC score is 0.50, it means no discrimination ability, while scores ranging from 0.60 to 0.80 are deemed reasonable, scores between 0.80 and 0.90 are considered excellent, and values exceeding 0.90 are exceptional. The RF’s AUC score can be regarded as good, as the range of results fell between 0.8 and 0.90. The AUC values of the other three models were exceptional, and scores exceeding 0.90 are superior. Based on the AUC score performance, all clinical predictive models can be deployed to predict the mortality rate of COVID-19 patients. Finally, we calculate the model loss function, which measures how well the model fits the data and how robust it is to over-fitting. 61 Figure 4(b) shows that the training and validation loss rapidly converge to a minimum within the first 90 epochs, after which they remain stable and closed for several epochs. It implies that our clinical care model is highly generalizable on unseen data.
Moreover, we experimented with only nine features, the clinical predictive model attained high efficacy and performance in prognosticating COVID-19 patients’ mortality risk. It achieved plausible performance with minimal features, exceeding 0.90 for all the KPM, showing that our model is robust for unseen data. All new experimented outcomes are seen in Table 7. The model training and test accuracy, minimal error rates, and loss values also provided evidence of its stability and robustness, as shown in Figure 5. These outcomes designate the prognostic algorithms’ capacity to precisely depict the nonlinear and complex relationships among the parameters and the outcome and its ability to generalize well to unseen data. The DSNN methodology is a distinctive and flexible technique that can effectively handle diverse sparsity arrangements while upholding consistent accuracy. This model exhibits advantages over other extant algorithms regarding accuracy, efficiency, and flexibility. Hence, this research work entirely focused on these five key indicators for model evaluation. The remaining indicator was computed for use in making comparisons. 62
Evaluation of results against existing studies
In this subsection, Table 7 compares our study with existing scholarly work on ML and DL frameworks for the prediction of COVID-19 patients using biomarkers and demographic features. Furthermore, researchers utilize various attributes in their model for training and evaluation. However, since all the authors were considered laboratory markers and demographic features, all the authors adopted different approaches for significant feature selection. Table 7 shows that our DSNN algorithm with 24 features outperforms all the other methods in terms of most KPM. The second experiment with nine features also achieves commendable performance and efficiency in predicting COVID-19 mortality risk. Thus, the framework yields satisfactory outcomes even when utilizing a limited number of features.
The comparative results also demonstrate that our contribution has a larger sample size than most of the other studies, except for Pourhomayoun et al., 63 which has a much larger sample size but lower performance, and it shows that our DSNN framework can handle large and complex datasets better than existing ones. The outcomes also accentuate the gaps in some of the previous research papers, such as missing scores for some KPM or low scores for some KPM. Thus, our study achieved higher performance measures than previous studies with fewer features, indicating its superiority and efficiency in predicting the risk of death in COVID-19 patients based on biomarkers and demographics.
Study limitations
We acknowledge the limitations of our studies. First, our data were acquired from Hospital Universitario HM Madrid, Spain; clinical data do not reflect global COVID-19 patients, which may confine the applicability of our findings to other populations and settings. Second, our studied data were limited to 24 attributes, which may not cover all the relevant factors influencing the mortality risk of COVID-19 patients. Third, our data were subjected to some missing values and outliers, which may impair the quality and validity of our results. Fourth, our framework was trained and tested on a relatively small sample size, which may diminish its statistical significance and accuracy.
Conclusions and future study
COVID-19 was the black swan of the 20th century, resulting in a disastrous global health crisis that endangered human civilization and well-being. This prognostic modeling study has identified the most effective feature selection technique for forecasting the outcome of COVID-19 patients. This paper presents a novel predictive model known as DSNN that can accurately predict the mortality risk of COVID-19 patients. Our experimental results reveal that the higher mortality risks also had MODS and other coexisting medical conditions such as age structure and existing diseases. Compared the DSNN model with the other proposed models, it was proved that the stacking ensemble with the DL model has effectively enhanced prediction accuracy. The expectation is that this clinical care model will become notably beneficial for the healthcare system in making sensible decisions and providing ample facilities for COVID-19 patients; such preventive measures maximize patient survival rates.
In the future, the targeted treatments will be designed; meanwhile, sufficient actual data are expected to be accessible for algorithm validation and a plan to deploy the quantum neural network model for efficient and accurate performance.
Footnotes
Authors’ note
Xu Songhua is also affiliated to Institute of Medical Artificial Intelligence the Second Affiliated Hospital XJTU, Shaanxi, China.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethics approval
The anonymized data were obtained via special request to HM Hospitals (Spain) under their COVID DATA SAVE LIVES initiative, which was approved by their Ethics Committee. Our analysis adhered to HM Hospitals’ data use terms and protection standards.
Data availability statement
The dataset and related outcomes of this research work will be accessible from the corresponding author on a realistic request.
