Abstract
The generalizability of artificial intelligence (AI) models is a major issue in the field of AI applications. Therefore, we aimed to overcome the generalizability problem of an AI model developed for a particular center for pneumothorax detection using a small dataset for external validation. Chest radiographs of patients diagnosed with pneumothorax (n = 648) and those without pneumothorax (n = 650) who visited the Ankara University Faculty of Medicine (AUFM; center 1) were obtained. A deep learning-based pneumothorax detection algorithm (PDA-Alpha) was developed using the AUFM dataset. For implementation at the Health Sciences University (HSU; center 2), PDA-Beta was developed through external validation of PDA-Alpha using 50 radiographs with pneumothorax obtained from HSU. Both PDA algorithms were assessed using the HSU test dataset (n = 200) containing 50 pneumothorax and 150 non-pneumothorax radiographs. We compared the results generated by the algorithms with those of physicians to demonstrate the reliability of the results. The areas under the curve for PDA-Alpha and PDA-Beta were 0.993 (95% confidence interval (CI): 0.985–1.000) and 0.986 (95% CI: 0.962–1.000), respectively. Both algorithms successfully detected the presence of pneumothorax on 49/50 radiographs; however, PDA-Alpha had seven false-positive predictions, whereas PDA-Beta had one. The positive predictive value increased from 0.525 to 0.886 after external validation (p = 0.041). The physicians’ sensitivity and specificity for detecting pneumothorax were 0.585 and 0.988, respectively. The performance scores of the algorithms were increased with a small dataset; however, further studies are required to determine the optimal amount of external validation data to fully address the generalizability issue.
Although several artificial intelligence (AI) models have been developed for the detection of pneumothorax using chest radiographs, the clinical use of these models is restricted since their generalizability has not been evaluated, and their performance is not comparable with that of physicians.
This study aimed to overcome the generalizability issue by developing an AI model for one center performing external validation using a small dataset from another center and comparing the results of the models with those of physicians.
The primary model, PDA-Alpha, achieved high sensitivity and specificity for the external test dataset; however, the false positivity rate for PDA-Alpha was high.
The second model, PDA-Beta, developed via external validation of PDA-Alpha, achieved high sensitivity and specificity and an improvement in the false-positive rate.
Since pneumothorax detection algorithms can screen for pneumothorax on all chest radiographs, even patients presenting with chest pain or shortness of breath who are not suspected of having pneumothorax at the first examination can be screened.
Introduction
Pneumothorax is defined as the presence of air in the pleural space between the pleura. Pneumothorax may resolve spontaneously depending on its size, or it may result in severe clinical conditions, such as respiratory failure, hypotension, shock, and respiratory arrest. Early diagnosis and intervention are important to prevent morbidity and mortality. Although thoracic computed tomography (TCT) is the gold standard method for diagnosing pneumothorax, chest radiography is the first choice as it is easily accessible, inexpensive, and has low radiation exposure. However, the diagnostic efficiency of radiographs could be reduced owing to the poor inter-rater agreement for the detection of pneumothorax 1 or insufficient experience in chest radiograph evaluation. 2
Artificial intelligence (AI) can be used to address diagnostic problems. Several AI studies on the detection of pneumothorax using chest radiographs have been reported.3,4 However, the clinical use of these AI models is restricted since the generalizability of these models was not evaluated, and their performance is not comparable with that of physicians. In addition, although radiographs are generally evaluated by clinicians in outpatient settings, these AI studies included radiologists.
The generalizability issue of AI arises from the reduction in the accuracy of AI models due to the use of data from other centers. AI systems using radiographs have generalizability issues owing to the use of different devices, technicians, and imaging protocols. There is no standardized method for overcoming this problem. Developing a unique model for each center or retraining an existing model with large datasets requires time and labor. Moreover, the amount of data required for external validation is unknown.
Therefore, this study attempted to overcome the generalizability issue of an AI model developed for one center through external validation using a small dataset from another center and compared the results of the models with those of physicians from three clinical areas working with pneumothorax (emergency medicine, thoracic surgery, and chest diseases).
Materials and methods
Dataset construction
In this multi-center retrospective study, we propose the use of an AI-based pneumothorax detection algorithm (PDA) to perform accurate detection of pneumothorax using chest radiographs and to address the generalizability issue of AI. Since the generalizability problem is that the performance of an algorithm decreases when it is used outside the center where it was developed, we used datasets from two different centers to develop two different PDAs.
The Ankara University Faculty of Medicine (AUFM) dataset was used to develop the primary model, PDA-Alpha. To evaluate whether the generalizability problem can be addressed using a small dataset from a second center, a secondary model, PDA-Beta, was developed through external validation of PDA-Alpha using the Health Sciences University (HSU) dataset. Both models’ performance scores were assessed using the HSU test dataset. The datasets were created by five chest disease specialists, two chest surgeons, and one radiology physician working at AUFM and HSU. The Ethics Committee of AUFM approved the study protocol (I7-393-20). All data were anonymized.
TCT or pulmonary computed tomography (CT) angiography of patients with pneumothorax with International Classification of Diseases, tenth revision (ICD-10) codes (J93, J95, and S27) were evaluated retrospectively. 5 The evaluation covered the period from January 2012 to December 2019 and was performed using the AUFM hospital information management system (HIMS). Chest radiographs of patients with pneumothorax detected on TCT at admission were obtained in .png format. Patients whose first radiological examination was TCT and those who underwent TCT before obtaining a radiograph were excluded from the study. In addition, the radiographs acquired at admission of patients who were admitted to the AUFM Chest Diseases outpatient clinic due to respiratory symptoms and radiographs of those who did not have pneumothorax on TCT were obtained to create the control group between October and December 2019. The AUFM dataset consisted of 648 pneumothorax and 650 non-pneumothorax chest radiographs.
The HSU validation dataset included only 50 pneumothorax radiographs. As the study aimed to assess the feasibility of addressing the generalizability issue with limited data, cases without pneumothorax were not included in the validation dataset. The HSU test dataset included 50 pneumothorax and 150 non-pneumothorax radiographs. The HSU validation and test datasets were obtained by physicians using pneumothorax ICD-10 codes from the HSU HIMS and covered the period from February to August 2020. For the non-pneumothorax images, patients who had presented at the clinic with complaints of chest pain and/or shortness of breath during the specified time period were retrospectively evaluated. Among these patients, those who were not diagnosed with pneumothorax in their examinations had their initial chest radiographs collected.
These three datasets included a total of 1548 chest radiographs according to the inclusion and exclusion criteria (Figure 1). All pneumothorax cases in the datasets were confirmed by TCT. Non-pneumothorax cases included radiographs showing emphysema, atelectasis, bronchiectasis, pleural effusion, reticulation, consolidation, nodule/mass, pneumonia, and pulmonary edema, in addition to normal radiographs. Both posteroanterior and anteroposterior radiographs were included.
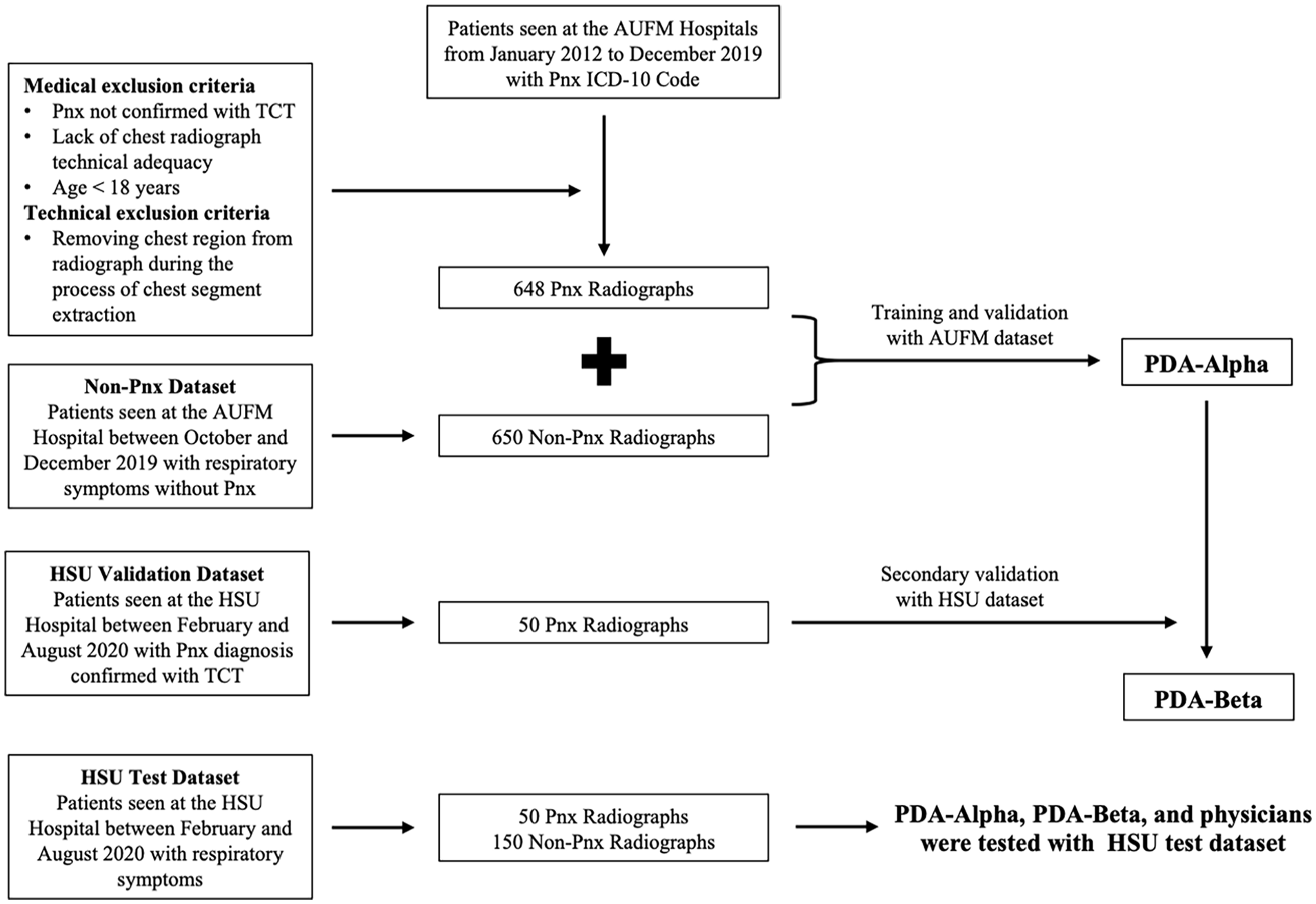
Study flow diagram.
Technical characteristics of the datasets: In AUFM clinical practice, chest radiographs are taken in the posteroanterior view with the patient standing and in full inspiration. The exposure factors used for this view are around 100–110 kV and 4–8 mAs. For the anteroposterior view, the patient is positioned in a supine position, and the exposure factors are also around 100–110 kV and 4–8 mAs. The source-to-image distance is set to 180 cm for all posteroanterior radiographs and to 100 cm for the supine anteroposterior view. At HSU, the patient position and source-to-image distance are the same as at AUFM. However, the exposure factors are slightly different, around 100–110 kV and 2–4 mAs at HSU. AUFM uses the Dynamic X-Ray Dynalift 55S, while HSU employs the Siemens Multix Pro for radiographic imaging.
AI technique
The model development process was divided into two phases. In the first phase, the chest segment was extracted from the radiographs using image processing. During the second phase, a deep neural network was trained to detect pneumothorax in the extracted chest segment.
Chest segmentation with image processing
In addition to the chest, chest radiographs also included the neck, shoulder, abdomen, and patient-related information. Image processing was used to remove these regions from the original radiographic image for noise reduction in the deep neural network. Thus, a deep neural network trained using the chest region only was developed. The chest-segment extraction process consisted of four steps. Representative radiographic images and the corresponding outputs of each step are shown in Figure 2. The steps were as follows:
The input radiographic image (Figure 2(a)) intensity was enhanced via gamma correction with a gamma value of 2 (Figure 2(b)). 6
The presence of light pixels surrounded by darker pixels in the original radiographic image resulted in the formation of gaps in the enhanced image. The gaps were filled using the connect-component method (Figure 2(c)). 7
After filling the gaps, the regions surrounding the chest were filled with white color using a modified version of the flood fill algorithm. 8 The algorithm was applied to the left-, top-, and right-edge pixels. The ordinary flood fill algorithm connects and colors pixels with similar brightness. When an ordinary flood fill algorithm is applied to radiographic images, it generally does not color the entire region outside the body. Therefore, after applying the ordinary flood fill algorithm, the colored regions were enhanced by postprocessing. During postprocessing, each colored pixel was replaced with a colored square of size (k/10) × (k/10), where k is the minimum height and width of the input radiographic image (Figure 2(d)).
Images with several white regions were obtained as a result of the earlier steps. These regions correspond to the area outside the chest. The chest region was considered to be a rectangular area in the middle of the radiographic image. Therefore, the right and left sides of the white area on the top of the image could not include the chest. Similarly, the regions above and below the white area on the sides of the image could not include the chest. In this step, the pixels corresponding to the chest area were extracted by eliminating these areas. The pixels between the bottom-most pixel of the top white regions, the right-most pixel of the left white regions, and the left-most pixel of the right white regions were chosen as the chest segment boundary. The original radiographic image inside the boundary was then extracted and resized to 600 × 600 pixels to obtain the end results of image processing (Figure 2(e)).
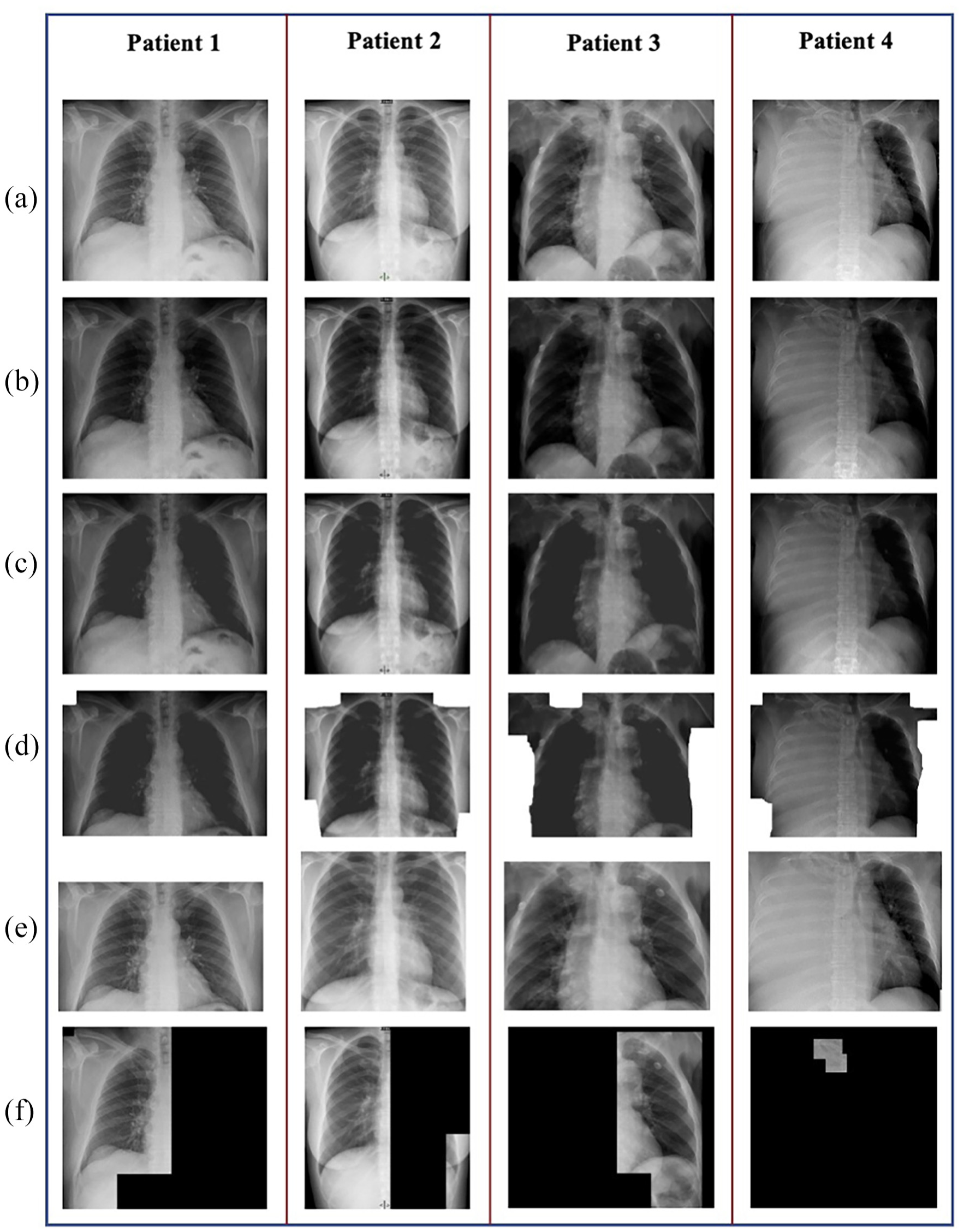
Sample images from the chest segmentation process steps. (a) Input radiographic image. (b) Image enhanced with gamma correction. (c) Image after connect-component method. (d) The image is colored with our modified version of the flood fill algorithm. (e) Extracted chest image. (f) Lung image extracted from part c using the Otsu algorithm and 8-connected algorithm.
The chest segment extraction process was applied to images from the AUFM and HSU validation datasets to remove the regions beyond the chest. However, it also removed the chest regions in two images, which had to be excluded. The remaining images were used to train the deep neural network. Hence, PDA-Alpha was trained with 1298 images, and PDA-Beta was trained with 1348 images.
Image processing was only used to detect the chest region, and the original lung image with bones was fed to the deep neural network. However, the literature review revealed that some studies removed the bones from the chest using image processing and used the remaining lung image without bones to detect pneumothorax. 7 This was accomplished by applying the Otsu algorithm and an eight-connected algorithm to the image obtained at the end of the second step (Figure 2(c)) of our chest extraction process. 9 This method was also attempted in the present study. However, steps 3 and 4 of the chest segment extraction process were included in the final study as the abovementioned process also removed lung regions at times. Such images were excluded from the analysis, resulting in datasets with fewer data points. Some of the images resulting from the aforementioned process are shown in Figure 2(f).
Training the deep neural network to detect pneumothorax
In the second step of the PDA, the deep neural networks were trained using images extracted via image processing. The problem was considered a binary classification problem with classes “not PNX” and “PNX.” Two types of artificial neural networks were trained: transformer and convolutional neural network (CNN). In recent years, transformers have been widely used in computer vision tasks. In 2020, Dosovitskiy et al. proposed a novel architecture called a vision transformer that was trained on a large image dataset and achieved competitive scores comparable with those of CNN-based models. 10 Vision transformers and CNNs pre-trained on ImageNet 11 were trained. CNN models were preferred as they outperformed vision transformers.
VGG19, InceptionV3, and ResNet50 were used as the CNN architecture backbones.12–14 ResNet50 was selected as the final backbone as it outperformed the other models. The designed CNN is shown in detail in Figure 3. ResNet50 uses RGB (red, green, and blue) images as input. However, since radiographic images are grayscale, dimensional inconsistencies occurred between the radiographic images and ResNet50. This problem was addressed by converting each radiographic image to a new RGB image by setting all channels equal to the grayscale value. In other words, we repeated each radiographic image three times and created an appropriate input for ResNet50. The final model was trained for 40 epochs with stochastic gradient descent and 0.9 momentum. Cosine-annealing warm restarts were used as the learning rate scheduler. During training, each input was provided to the model after data augmentation, which consisted of the following operations:
Adding random white and black curves of varying lengths and degrees
Adding salt and pepper noise to the image
Rotating the image between −20° and 20°
Zooming the image between 0.9 and 1.1
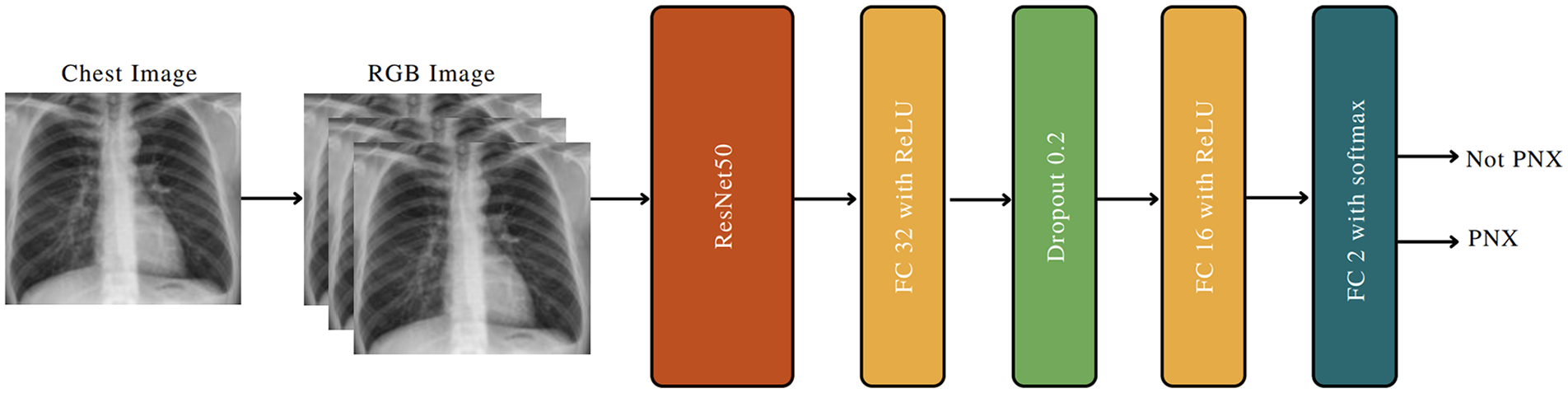
CNN architecture designed to detect pneumothorax from chest images.
The PDA training process consisted of image processing, data augmentation, and backpropagation. The CNN model was trained on the AUFM dataset and the combined dataset (AUFM dataset and HSU validation dataset) to obtain PDA-Alpha and PDA-Beta, respectively. To measure the PDA performance, each dataset of the HSU test dataset was passed through image processing, and the CNN prediction was obtained. Finally, the predictions were evaluated using true labels. These processes are shown in detail in Figure 4. The same image processing method and CNN were used in both the training and test phases. The hyperparameters of the training process and the hardware components used within the study are provided in Supplemental Material Table T1 and T2, respectively.
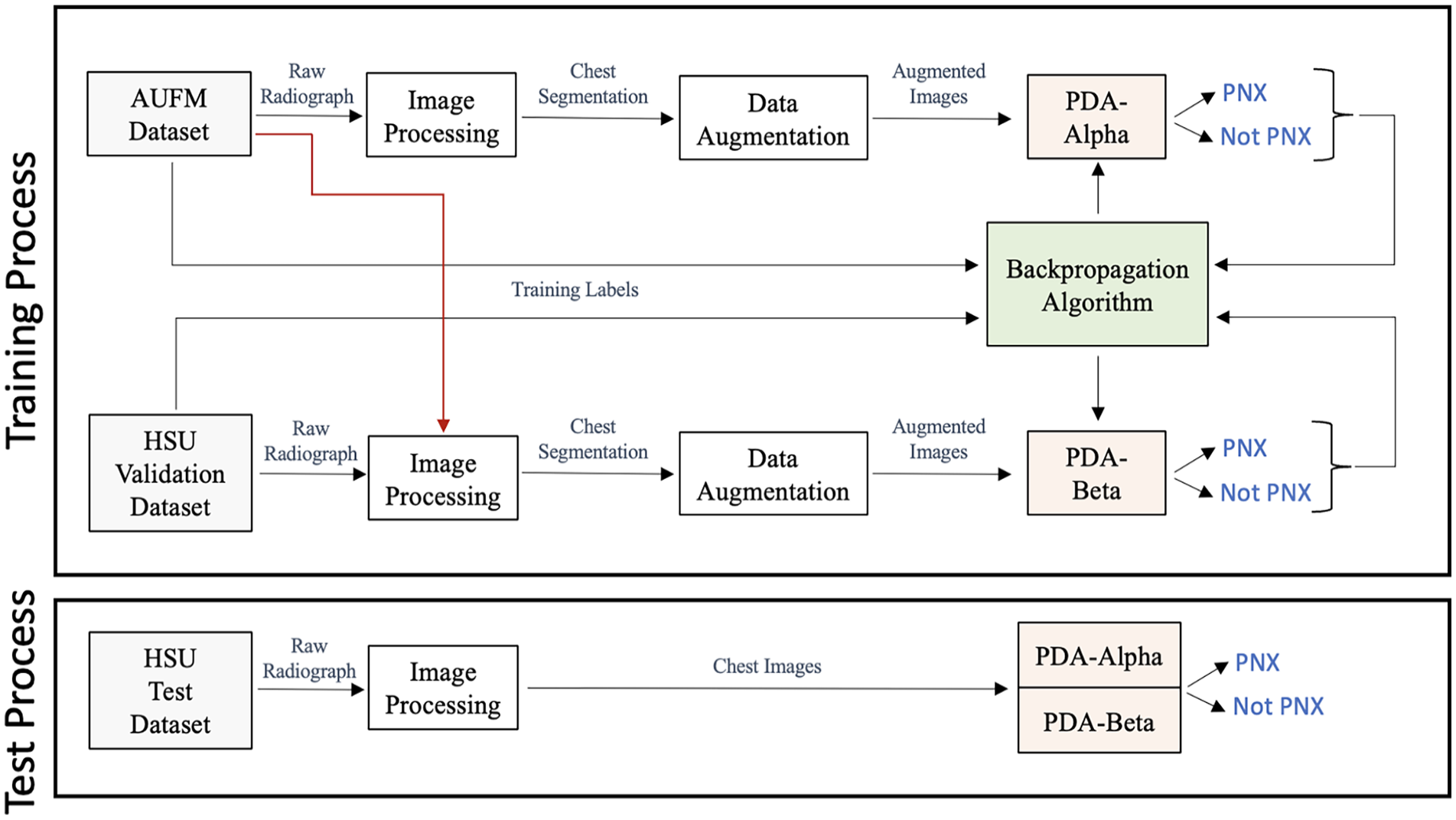
Training and test stages of PDA-Alpha and PDA-Beta.
True-positive and false-positive radiographs were demonstrated for both models using heatmaps (Supplemental Material F1). It was observed that the more intense red color was concentrated in the areas where there was a decrease in lung markings outside the visceral pleural line.
Test procedure
The HSU external test set was provided to 20 voluntarily participating physicians to compare and interpret the results of the developed models. These physicians were selected from the emergency (n = 5), thoracic surgery (n = 5), and chest disease (n = 10) departments, who, unlike radiologists, were responsible for detecting pneumothorax and implementing the necessary intervention. Only radiographs without any clinical information were presented to the participants. To simulate a real-life setting, the participants were not informed that it was a pneumothorax study, and they were instructed to detect pathologies on the chest radiographs. They could mark more than one pathology on the prepared form and were given 15 days to complete the task. The results were subsequently anonymized and statistically evaluated.
Statistical analysis
Receiver operating characteristic (ROC) curves were used to describe the diagnostic performance of the models. An area of 0.50 implied that the variable added no information. The areas under the ROC curves (AUCs) and 95% confidence intervals (CIs) for all variables were calculated using the method described by Hanley and McNeil. 15 The Cohen’s kappa coefficient and its 95% CI were used to evaluate the agreement between methods and observers. In cases where multiple raters were involved, the generalized kappa statistic was preferred. The sensitivity, specificity, positive and negative predicted values, accuracy, and Wilson CIs were calculated for the diagnostic performance of the models. The comparison of proportions between the two groups was evaluated using the z-test. The prevalence of pneumothorax was accepted as 5% in patients presenting with chest pain or shortness of breath. 16 The optimal thresholds for the prediction of pneumothorax by PDA-Alpha and PDA-Beta were determined using the Youden Index. MedCalc version 18.11 and R version 4.2.1 were used for statistical analysis. Statistical significance was set at p < 0.05.
Results
PDA-Alpha was initially subjected to internal testing with 15% randomly obtained from the AUFM dataset. The test dataset consisted of 77 pneumothorax and 117 non-pneumothorax radiographs. A ROC curve was generated (Supplemental Material F2). The AUC for PDA-Alpha was 0.997 (95% CI: 0.959–0.999). Supplemental Material T3 shows the confusion matrix of the model. The sensitivity and specificity were 0.987 (95% CI: 0.930–0.998) and 0.974 (95% CI: 0.927–0.991), respectively (Supplemental Material T4).
The HSU test dataset, which included 200 radiographs (50 pneumothorax and 150 non-pneumothorax), was used to evaluate the generalizability and the performance of PDA-Alpha, PDA-Beta, and physicians.
PDA-Alpha vs PDA-Beta
This testing phase was conducted to assess the generalizability of PDA-Alpha with data from another center and determine the effect of a small intervention on the performance of PDA-Beta.
PDA-Alpha and PDA-Beta classified radiographs into two categories: pneumothorax and not pneumothorax. A ROC curve was generated (Figure 5), and the thresholds for PDA-Alpha and PDA-Beta to predict pneumothorax were 0.336 and 0.981, respectively. The AUCs for PDA-Alpha and PDA-Beta were 0.993 (95% CI: 0.985–1.000) and 0.986 (95% CI: 0.962–1.000), respectively. Notably, the difference between these two AUCs was not statistically significant (p = 0.619). As illustrated in the confusion matrices, although PDA-Alpha successfully detected 49/50 pneumothoraxes, there were seven false-positive predictions. By contrast, PDA-Beta successfully predicted 49/50 pneumothoraxes with only one false-positive prediction (Table 1). In the performance metrics, the positive predictive value increased from 0.525 to 0.886 after external validation (p = 0.041) (Table 2). Cohen’s kappa values for PDA-Alpha and PDA-Beta were 0.897 (95% CI: 0.828–0.967) and 0.973 (95% CI: 0.937–1.000), respectively, indicating almost perfect agreement for both.
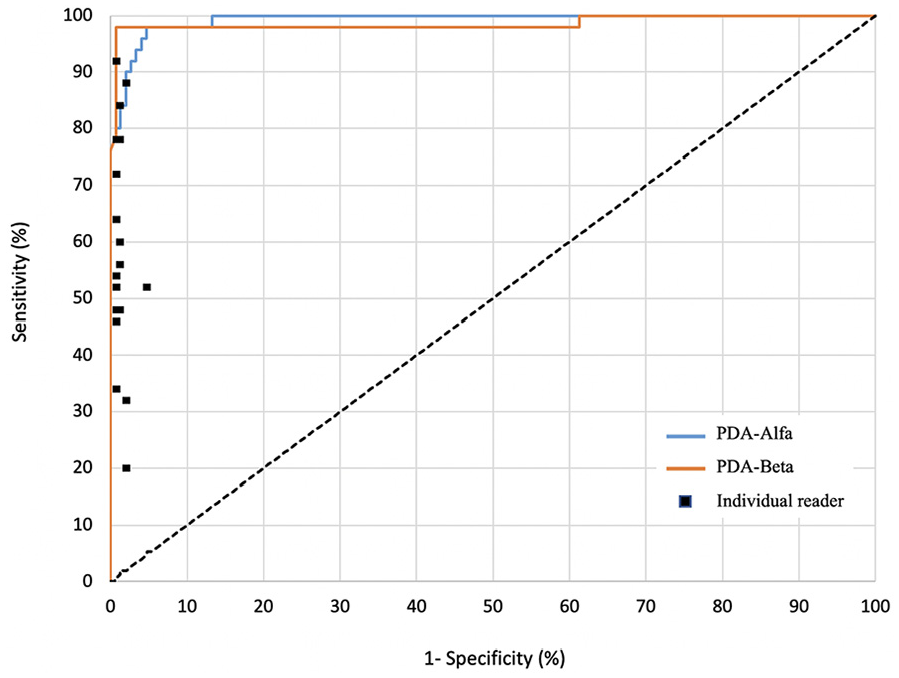
Receiver operating characteristic curves for PDA-alpha and PDA-beta compared with physicians for pneumothorax detection.
The confusion matrices of PDA-Alpha and PDA-Beta.
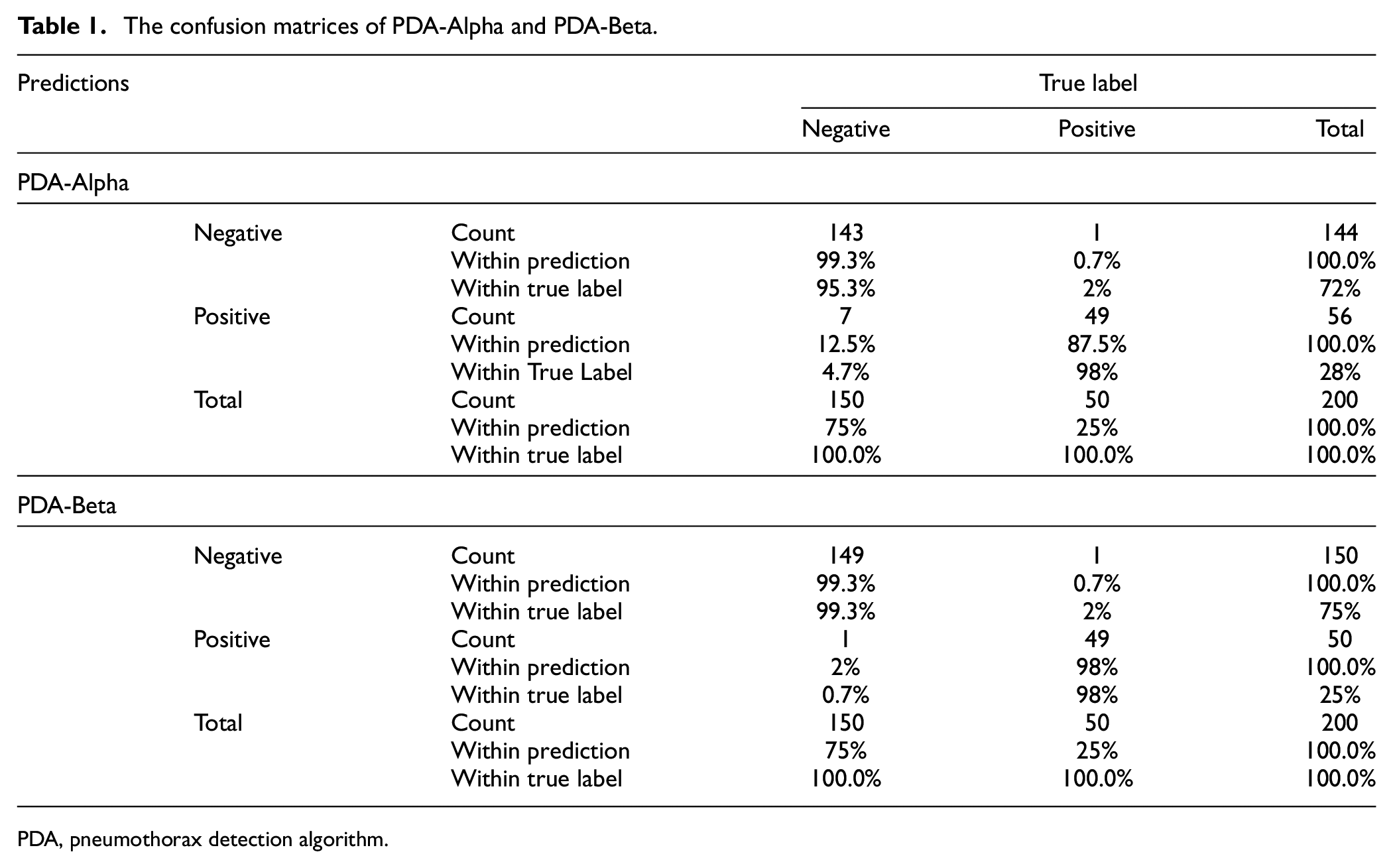
PDA, pneumothorax detection algorithm.
The performance metrics of PDA-Alpha and PDA-Beta.
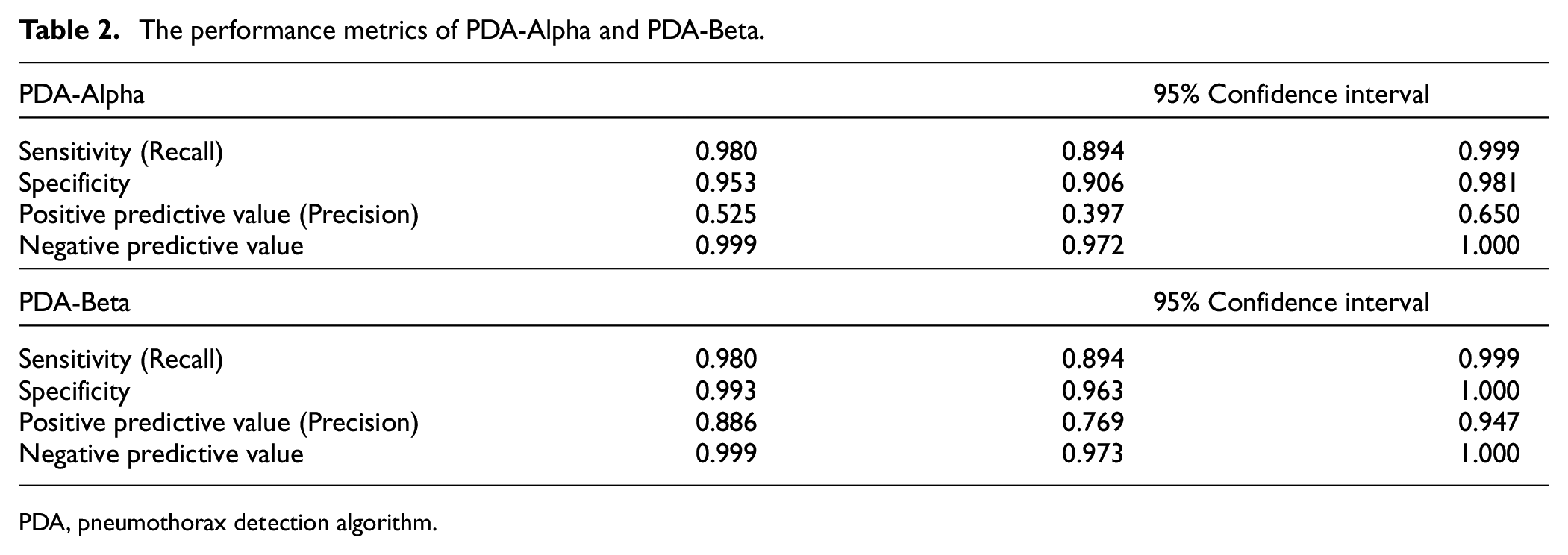
PDA, pneumothorax detection algorithm.
Physicians vs AI
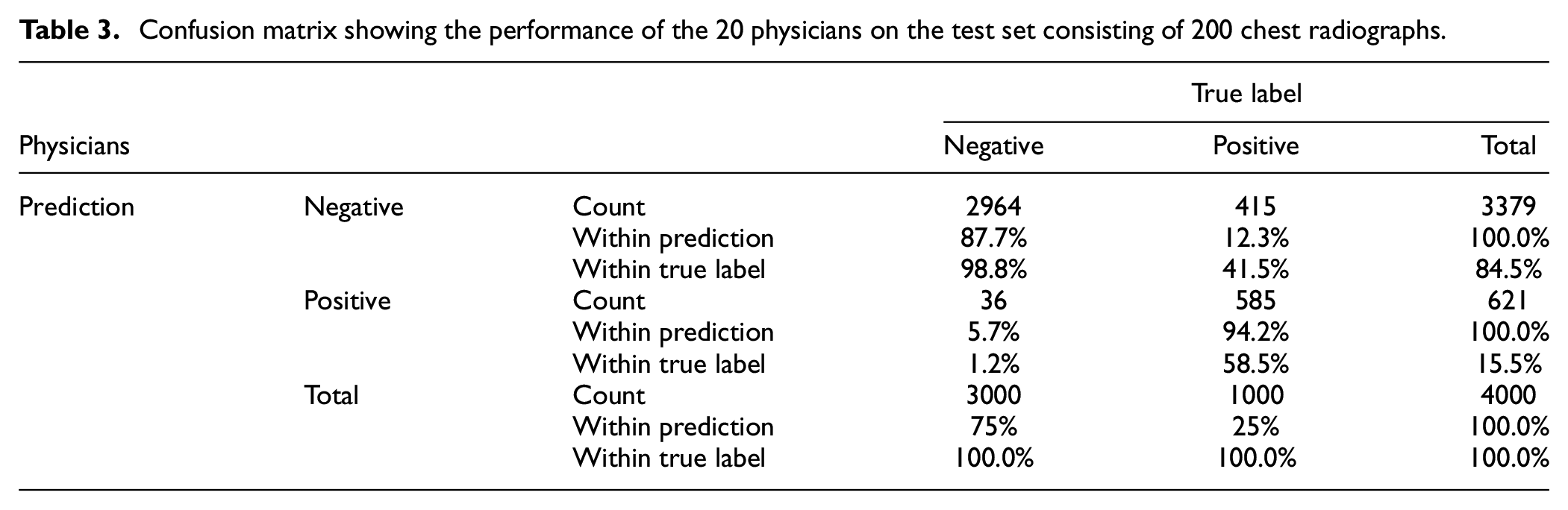
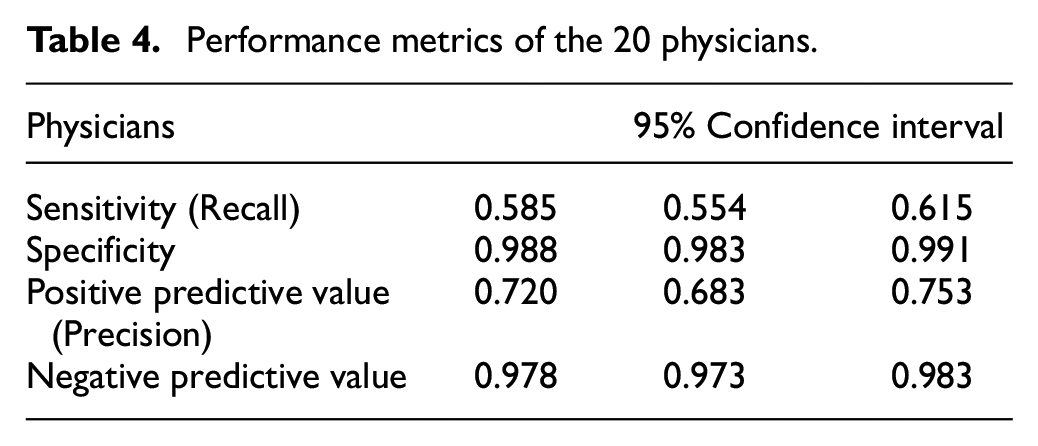
The HSU test dataset was applied to the 20 physicians as well as the AI algorithm. Confusion matrices, showing the individual performance of each physician on the test dataset consisting of 200 radiographs, were separately created. Afterward, the confusion matrices belonging to the 20 physicians were combined into a single matrix (Table 3). The performance metrics for the detection of pneumothorax on radiographs are shown in Table 4. The generalized kappa coefficient was 0.621 (95% CI: 0.611–0.631) for the inter-observer agreement of the 20 readers.
Confusion matrix showing the performance of the 20 physicians on the test set consisting of 200 chest radiographs.
Performance metrics of the 20 physicians.
Discussion
The primary model (PDA-Alpha) achieved high sensitivity and specificity for the external test dataset (0.980 and 0.953, respectively). However, the high false-positivity rate for PDA-Alpha was considered a generalizability problem. Therefore, external validation was performed using a small dataset, and a second model (PDA-Beta) was developed. High sensitivity and specificity were also obtained with the second model (0.980 and 0.993, respectively); in addition, the false-positive rate decreased significantly. Physicians evaluated the same dataset to clarify the results obtained from the models with sensitivity and specificity of 0.585 and 0.988, respectively, for detecting pneumothorax. This is the first study to overcome the generalizability issue of AI through validation with a small dataset obtained from a second center.
Considering the pneumothorax detection performance of the physicians, pneumothorax can be detected with high specificity by physicians; however, pneumothorax may be missed in patients with non-specific symptoms, such as chest pain or shortness of breath, because of the low sensitivity of the physicians. This can easily be addressed by AI. A PDA can evaluate every new chest radiograph added to the HIMS for the presence of pneumothorax. Thus, even patients who are not suspected of having pneumothorax at the first examination can be identified. When this potentially life-threatening pathology is detected in patients with respiratory emergencies, the physician requesting the test will be alerted. As chest radiographs are uploaded to the system within a few minutes and the model can assess them almost instantaneously, there will be no time wasted, enabling prompt progression to further investigations and treatment stages. Since it is impossible to perform TCT in all patients presenting with chest pain or shortness of breath, the significance of AI in the detection of respiratory emergencies using radiographs is clear.
Although PDA-Alpha had high-performance scores, we developed a second model to observe the changes after the model was validated using data from 50 patients from another center. Subsequently, a significant improvement was observed in the false-positive rate, which was the weakest point of the model. In all, 50 patients were used for external validation as it is an easily accessible number, and validation with a higher number of patients might not be practical. However, the effects of our method will have to be re-evaluated for models that include more than two (binary) classifications.
Fallahpoor et al. stated that creating a new dataset to overcome the generalizability problem is non-feasible and time-consuming, and they evaluated the effects of image preprocessing, different CNN models, and different combinations of datasets. 17 They assessed generalizability using various combinations of two large COVID-19 CT image datasets with deep learning, showing that a combination of datasets can improve generalizability. Zech et al. pooled data from three different centers for their pneumonia-detection algorithms and obtained superior results compared with those of the separate datasets. However, despite using pooled data, the external results were still inadequate. 18
The popularity of data-centered studies over model-centered studies is predicted to increase. 19 Algorithms developed with a small amount of high-quality data are expected to replace those developed with open-source big data. Our study substantiates this concept as it was designed with a small amount of high-quality data and achieved good results. At the beginning of the study, an open-source dataset containing over 10,000 pneumothorax radiographs was evaluated by physicians for data quality, and it was found that only 20%–30% of the radiographs were of acceptable quality. 20 Thus, the question arises: are only the performance scores of AI important, or is the development process also important? Developing highly successful models from a dataset consisting of radiographs on which the lungs are not clearly visualized will likely elicit a negative response from physicians.
To enable these algorithms to be shared and adapted with other healthcare centers, it is necessary to demonstrate their reliability within clinical practice. Real-world data should be obtained using a period of 3–6 months in the centers where the models are developed. For this purpose, a new study should be designed to show that, compared to the period when AI was not used, the diagnosis success of AI-enhanced physicians increased, the time it took for the clinician to reach the diagnosis, and therefore the time between the patient’s admission to the hospital and the initiation of treatment, reduced. If the data obtained from this study substantiate the clinical benefits of the model, then transportability would be considered. Beyond technical aspects such as the development of a user interface and deployment protocols, the initial step involves creating a test dataset specific to the relevant healthcare center that intends to employ this model, and the model should be tested. In cases where the results are not deemed acceptable, external validation is conducted in accordance with the methodology highlighted in our study, using a limited number of pneumothorax-containing radiographs from that center. Hence, by ensuring that the model’s performance reaches an adequate level, we advance to the deployment phase. The relative simplicity of radiograph data compared to MRI and CT scans, along with the similarity in imaging techniques and acquired images despite variations in device manufacturers, enhances the adaptability of such AI models, making their integration into diverse clinical settings more feasible.
Strengths
The prejudice of the end user, physician experience, and unfamiliarity with the subject during the deployment phase of AI models are significant barriers. In addition to creating the study design, the construction of the dataset and the testing phase were performed by physicians who believed that improving the diagnosis of pneumothorax was necessary. Compared with studies conducted by engineers or radiologists, the present study is one of very few clinical studies. The importance of this contribution will be better understood since AI models are used more intensively in daily practice.
Limitations
The most important limitation of the present study was that the high success of PDA-Alpha limited our ability to strictly evaluate the effects of external validation when compared with PDA-Beta. Therefore, only the false-positive rate was evaluated. Second, the clinical and technical significance of the two-class model is limited because patients presenting with chest pain or shortness of breath may have one of several pathologies. The physicians were not informed that the present study was a pneumothorax study, and they were instructed to choose among 11 classes during the test phase. It is necessary to create a dataset specific to each class to develop a model for all possible pathologies.
However, it is crucial to underline that the AI model in our study was specifically designed not for the purpose of differential diagnosis among various radiological pathologies but rather to detect pneumothorax, a respiratory emergency, in real time when a chest radiograph is taken. Two important medical contributions of the model are: to prevent chest radiographs containing pneumothorax from being missed and to alert the relevant physician within seconds; to avoid loss of time in the clinician’s workload even though it is easy to detect. Therefore, classification was done between two groups defined as pneumothorax and others. In addition, AI models for differential diagnosis of pathologies on chest radiographs have been previously developed in several studies on open-source large datasets. 20 Developing a similar model would not provide novelty to our study.
Despite the growing focus on the generalizability issue in recent years, it remains one of the most significant challenges in the clinical integration of AI models. Are we able to immediately use an AI model developed anywhere in the world in our own hospital? Or would we be able to use it only after validating it with our own data? How much data are needed? There are no definitive answers to these questions yet.
In conclusion, we created an AI model for pneumothorax detection, assessed its performance using data from another center, and proposed a novel approach to solve the generalizability issue. Furthermore, we emphasized the importance of comprehensive AI research that is focused on model development and approaches to address the challenges in clinical usage.
Supplemental Material
sj-docx-1-imj-10.1177_10815589231208479 – Supplemental material for Can the generalizability issue of artificial intelligence be overcome? Pneumothorax detection algorithm
Supplemental material, sj-docx-1-imj-10.1177_10815589231208479 for Can the generalizability issue of artificial intelligence be overcome? Pneumothorax detection algorithm by Elvan Burak Verdi, Muhammed Yılmaz, Deniz Doğan Mülazimoğlu, Abdussamet Türker, Aslıhan Gürün Kaya, Özlem Işık, Aslı Bostanoğlu Karaçin, Övgü Velioğlu Yakut, Bülent Mustafa Yenigün, Çağlar Uzun, Atilla Halil Elhan, Özlem Özdemir Kumbasar, Akın Kaya, Ayten Kayı Cangır, Cantürk Taşçı, Ahmet Murat Özbayoğlu and Serhat Erol in Journal of Investigative Medicine
Footnotes
Acknowledgements
We would like to express our sincere thanks to the 20 physicians who voluntarily participated in the test, which included 200 chest radiographs, for their support.
Author contributions
Study design: EBV, MY, DDM, AT, AGK, AMÖ, and SE
Literature search: EBV, MY, DDM, AT, and SE
Data collection: EBV, ÖI, ABK, ÖVY, BMY, AKC, CT, and SE
Analysis of data: EBV, MY, DDM, AT, AGK, ÇU, AHE, AK, ÖÖK, AMÖ, and SE
Manuscript preparation: EBV, MY, DDM, AT, AGK, AMÖ, and SE
Review of manuscript: EBV, MY, DDM, AT, AGK, ÖI, ABK, ÖVY, BMY, ÇU, AHE, ÖÖK, AK, AKC, CT, AMÖ, and SE
All authors reviewed and approved the final version of the manuscript before submission. All authors have submitted the ICMJE Form for the Disclosure of Potential Conflicts of Interest.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Data availability statement
The datasets are not publicly available due to privacy and ethical restrictions but are available from corresponding authors upon reasonable request.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
