Abstract
The mechanical properties of human brain tissue remain far from being fully understood. One aspect that has gained more attention recently is their regional dependency, as the brain’s microstructure varies significantly from one region to another. Understanding the correlation between tissue components and the mechanical behavior is an important step toward better understanding how human brain tissue properties change in space and time and to develop highly spatially resolved constitutive models for large-scale brain simulations. Here, we analyze the correlation between human brain tissue components quantified through enzyme-linked immunosorbent assays (ELISA) and material parameters obtained through an inverse parameter identification scheme based on a hyperelastic Ogden model and multimodal mechanical testing data for eight regions of the brain. We use neural networks as a metamodel to save computational costs. The networks are trained on finite element simulation outputs and are able to replace the simulations in the initial optimization step. We identified strong dependencies between mechanical properties and Iba1 associated with microglia cells, collagen VI, GFAP associated with astrocytes, and collagen IV. These results advance our understanding of microstructure-mechanics relations in human brain tissue and will contribute to the development of highly spatially resolved microstructure-informed constitutive models.
1. Introduction
The brain is one of the most complex organs of the human body. While its biological functions are generally well known, the mechanical behavior is still not fully understood. Computational models that are able to predict strains and stresses occurring in the tissue under mechanical loading are highly relevant for a multitude of applications ranging from surgery planning [1] to the prediction of cortical folding [2–4]. Griffiths et al. [5] showed that the spatial resolution of the material parameters used in mechanical models of the whole brain has a high impact on the results and therefore the accuracy of these models. The human brain shows a highly complex structure [6, 7]. Rather than using constitutive models that rely on constant parameters for different regions, it is therefore desirable to develop models that take into account the local tissue composition. Current constitutive models for human brain tissue are mainly phenomenological [8–14], although there have been more recent approaches that take into account the cell density [15], the concentrations of multiple components [16], or more sophisticated microstructural information like axon directions [17].
In a previous study, we have used enzyme-linked immunosorbent assays (ELISAs) quantifying the concentration of cellular and extracellular tissue components listed in Table 1 [16] as input to train a neural network (more specifically a constitutive artificial neural network, CANN) predicting the viscoelastic response of brain tissue during cyclic compression/tension and compression relaxation. Through a so-called relevance analysis of the neural network, we were able to assess which of the tissue components were most relevant for the predicted viscoelastic response. In another previous study, we have inversely (through finite element simulations) identified hyperelastic parameters [18] that capture the mechanical behavior under three modes, compression/tension and torsional shear, simultaneously. Thus, we are now able to investigate the correlation of the measured tissue components under even more complex loading conditions. In this study, we now combine and further analyze our previous results to investigate direct correlations between individual tissue components and mechanical parameters, namely, shear modulus and nonlinearity, of human brain tissue from different anatomical regions. We identify individual parameter sets using (1) only compression/tension data (comparable to the approach in Linka et al. [16]) and (2) all loading modes, i.e., compression/tension and torsional shear, to assess a possible loading-mode dependency of composition-mechanics correlations.
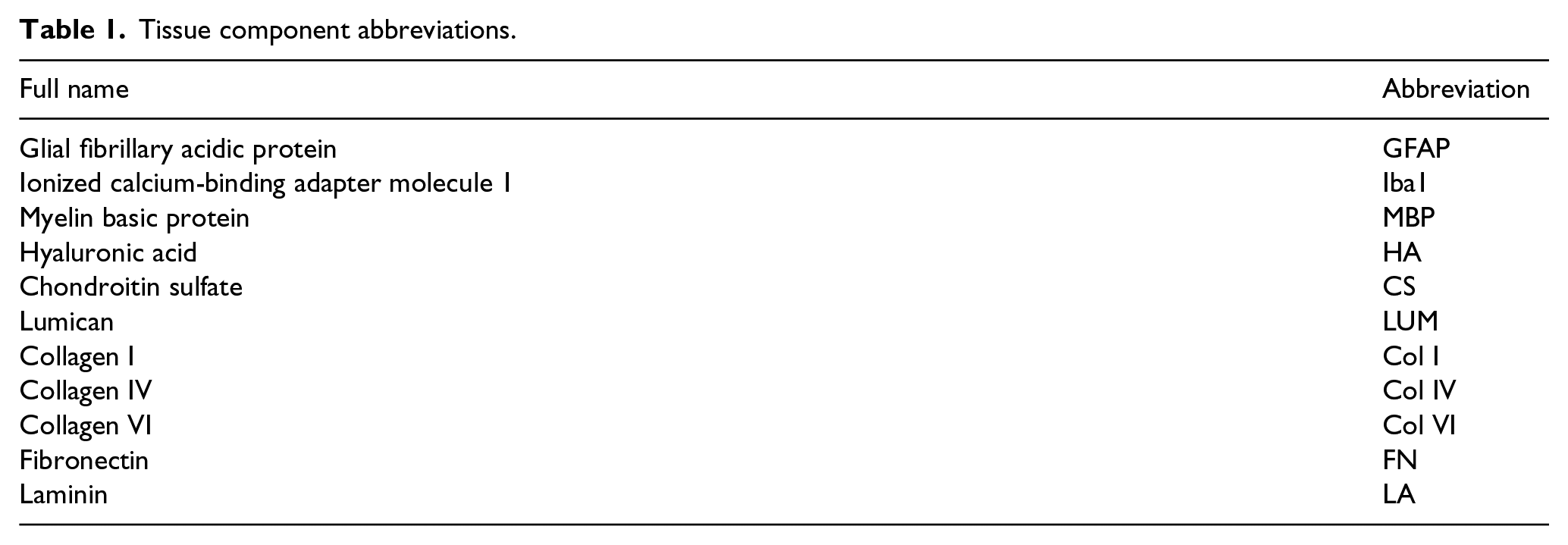
Tissue component abbreviations.

Many previous works use closed-form analytical solutions for constitutive models [8, 19], including our previous study [16]. However, the necessary assumption of homogeneous deformation does not hold when specimens are glued for tensile tests due to the thereby-introduced non-slipping boundary conditions and introduces an error [12, 20–22]. Finite element simulations, on the contrary, are able to resolve the inhomogeneous deformation state and therefore achieve a higher accuracy compared to homogeneous analytical solutions. Nevertheless, the use of finite element simulations comes with significantly higher computational costs, which is especially a problem in parameter identification tasks, where the simulation model is called multiple times. To address this issue, [23] explored different machine learning approaches for accelerating soft material parameter identification. They found that replacing the finite element simulations within the iterative optimization framework with neural network metamodels achieved excellent results. Therefore, we present an improved parameter identification scheme, where we initially replace the computationally expensive finite element model of the experiments with a surrogate model in terms of a neural network. The presented approach has potential applications beyond the task presented here, as it can be easily adapted to models implementing other constitutive models. This becomes especially interesting for more complex models capturing the time-dependent and/or biphasic nature of tissue, for example, poro-viscoelastic models.
2. Methods
2.1. Human brain samples
We obtained seven whole human brains including the cerebrum, cerebellum, and brainstem from three female and four male body donors who had given their written consent to donate their body to research. Details about the body donors and transport as well as storage of the brains are given in Hinrichsen et al. [18]. We received the brains between 9 and 46 h post mortem and directly cut them into 1-cm-thick coronal slices that we kept refrigerated at
2.2. Human brain tissue composition
The ELISAs to quantify the concentration of different cellular and extracellular components in human brain tissue were only performed for brains 1-5. For brains 3-5, we extracted samples directly after cutting the brains into slices to minimize the post mortem degradation of proteins before the samples were frozen and stored at
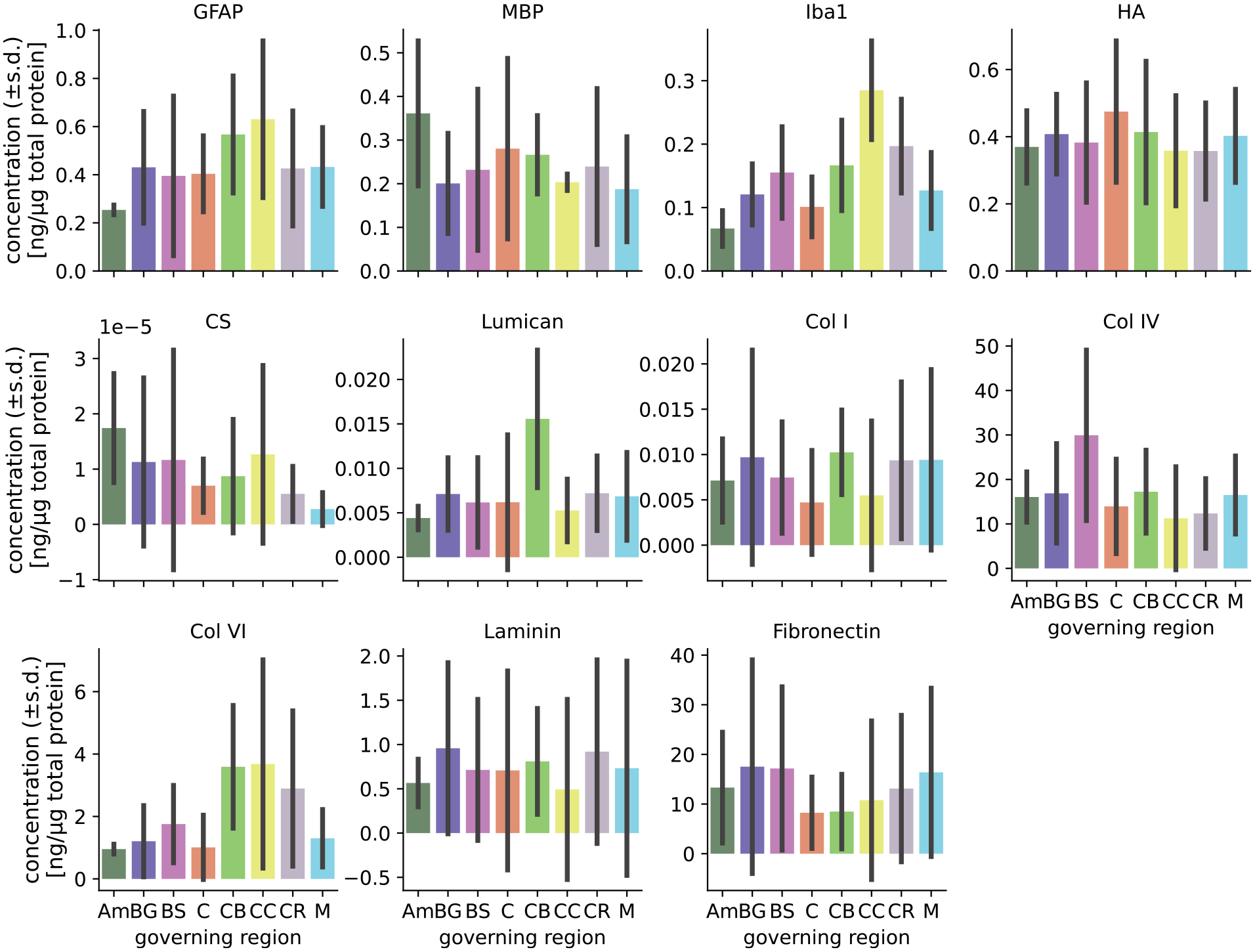
Tissue composition (quantified in Linka et al. [16]) in different brain regions.
Region abbreviations.

2.3. Mechanical testing
The mechanical experiments were completed within 96 hours post mortem. We used a biopsy punch to extract cylindrical samples of
2.4. Mechanical modeling
We approximate the mechanical behavior of the tested human brain tissue with a finite element (FE) model implementing a compressible Ogden-type hyperelastic constitutive model. The strain energy density is split into an isochoric part

The formulation for the isochoric part is a reformulated version of the Ogden model [24] in terms of the classical shear modulus

with the isochoric principal stretches

proposed by Ogden [24], where

taken from the linear regime, although we would like to note that this definition of
2.5. Inverse mechanical parameter identification
To quantify the mechanical behavior of the tested specimens, we identify the Ogden material parameters by minimizing the error between the simulated values

which is adapted from Gavrus et al. [25]. Since the output of the model depends on the shear modulus
To speed up the costly inverse parameter identification procedure, we use a neural-network-based metamodel. This approach has been used successfully to identify material parameters for blood clot and ventricular myocardium [23]. We train separate models for each of the three loading modes: compression/tension, shear up to 15% strain, and shear up to 30% strain. All of the networks consist of an input layer, an output layer, and hidden layers. We decided to set the number of hidden layers to two in order to strike a balance between model complexity and training efficiency [27].
The input layer is made up of three neurons, corresponding to the three input values: the two material parameters and the stretch or shear values. The output layer is structured with just one neuron that generates a continuous variable representing the stress.
We apply Bayesian optimization with a Gaussian processor to the remaining hyperparameters to explore the best neural network architecture.
In the hidden layers, we examine two hyperparameters: the number of neurons and the activation function. We set the minimal number of neurons for both hidden layers to 64 and the maximal number to 512. The search space is then explored using a step size of 32. The ReLU function is commonly used as activation function. In our search for the optimal activation function, we also consider ELU and tanh alongside ReLU [28]. The same activation function is used for the two hidden layers as well as the input layer. For the output layer, we use a linear activation
We chose adaptive learning rate optimization using Adam as our optimizer. The tuner selects the most suitable learning rate from three available options: 0.0001, 0.001, and 0.01.
Each model undergoes a separate tuning process. We implement hyperparameter tuning using the
We let the tuner run for 10 trials with two executions per trial and choose the validation loss or mean absolute error of the validation data as objective function. The function
We split the data into three different subsets using the function
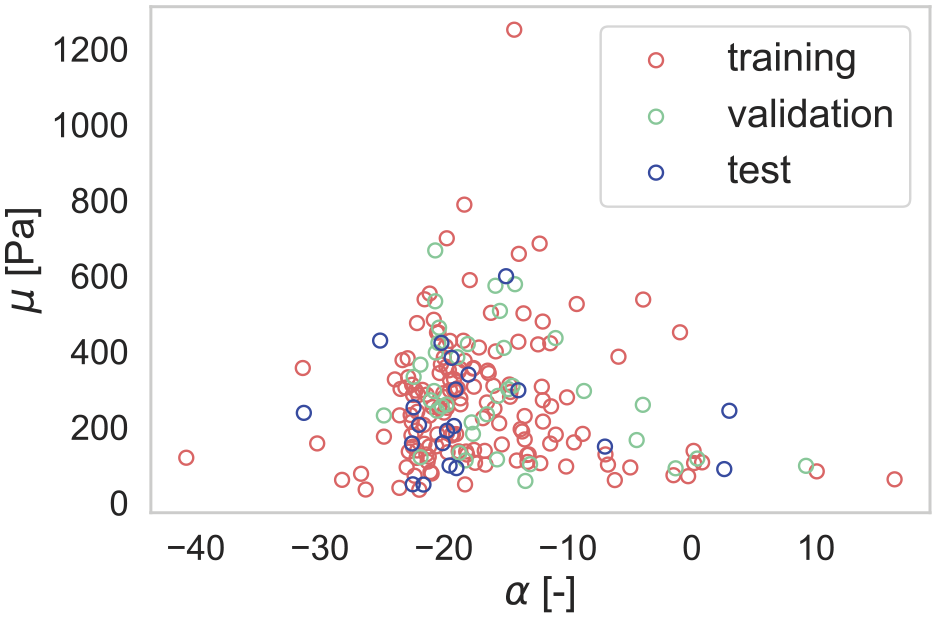
Distribution of material parameters, divided into training, validation, and test data.
After selecting the optimal hyperparameters, the model is trained again using these hyperparameters to produce a final model that is capable of making accurate predictions on new, unseen data. The training and validation data are equal to the one utilized for hyperparameter tuning. The models for compression/tension were trained for 6000 epochs and for the shear models, we conducted training for 1000 epochs.
Once the training is complete, the model is evaluated on the purely unseen data in the test set.
The inverse parameter identification initially uses the metamodel (the trained neural networks). Once an optimum is found, indicated by the fulfillment of one of the convergence criteria, the optimization is started again using the FE model. This provides a good compromise between computational cost and accuracy.
2.6. Statistical methods
We use the Kendall rank correlation coefficient
Component concentrations were obtained for specimens from five human brains and averaged over the eight regions listed in Table 2, while the mechanical parameters were calibrated based on region-wise averaged experimental data from a total of seven brains (including the five brains). Generally, each ELISA specimen was extracted adjacent to a corresponding mechanical specimen. Still, there is also a small number of ELISA specimens for which no mechanical data were available due to problems during testing or the subsequent data processing. The mechanical parameter data set contains more specimens than the ELISA tissue composition data. Figure 3 visualizes the distribution of specimens over the considered brain regions and the overlap between the two data sets. The individual distributions of specimens from the two data sets (tissue composition and mechanics) over the tested brains and regions are shown in Figures 9 and 10. We also calculated correlation coefficients for data not averaged over regions, where only the specimens denoted with ’both’ in Figure 3 were used–comparable to the study in Linka et al. [16]. The correlation coefficients as well as the corresponding
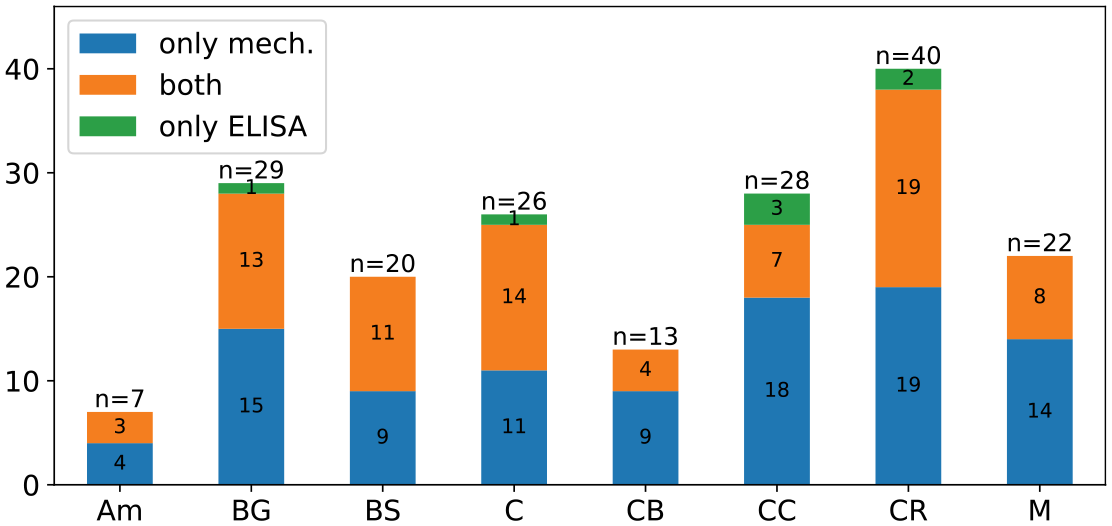
Distribution of ELISA and mechanical samples over the eight regions. Both denotes mechanical samples for which an adjacent ELISA sample was taken.
3. Results
3.1. Performance of surrogate model
To evaluate the neural-network-based metamodels’ generalization capability, we initially set aside 10% of the data that were not utilized during hyperparameter tuning and training. With this new, unseen data we tested our networks and assessed the results using the coefficient of determination. Table 3 presents the mean values for the coefficient of determination, along with the corresponding standard deviations, for the test set. The compression/tension model attained a mean coefficient of 0.9234 with a standard deviation of 0.1230. The torsional shear models both achieved excellent results with a mean coefficient of determination greater than 0.98 and a standard deviation below 0.05. Figure 4 illustrates the best achieved fits during testing for each loading mode, measured by the coefficient of determination.
Mean

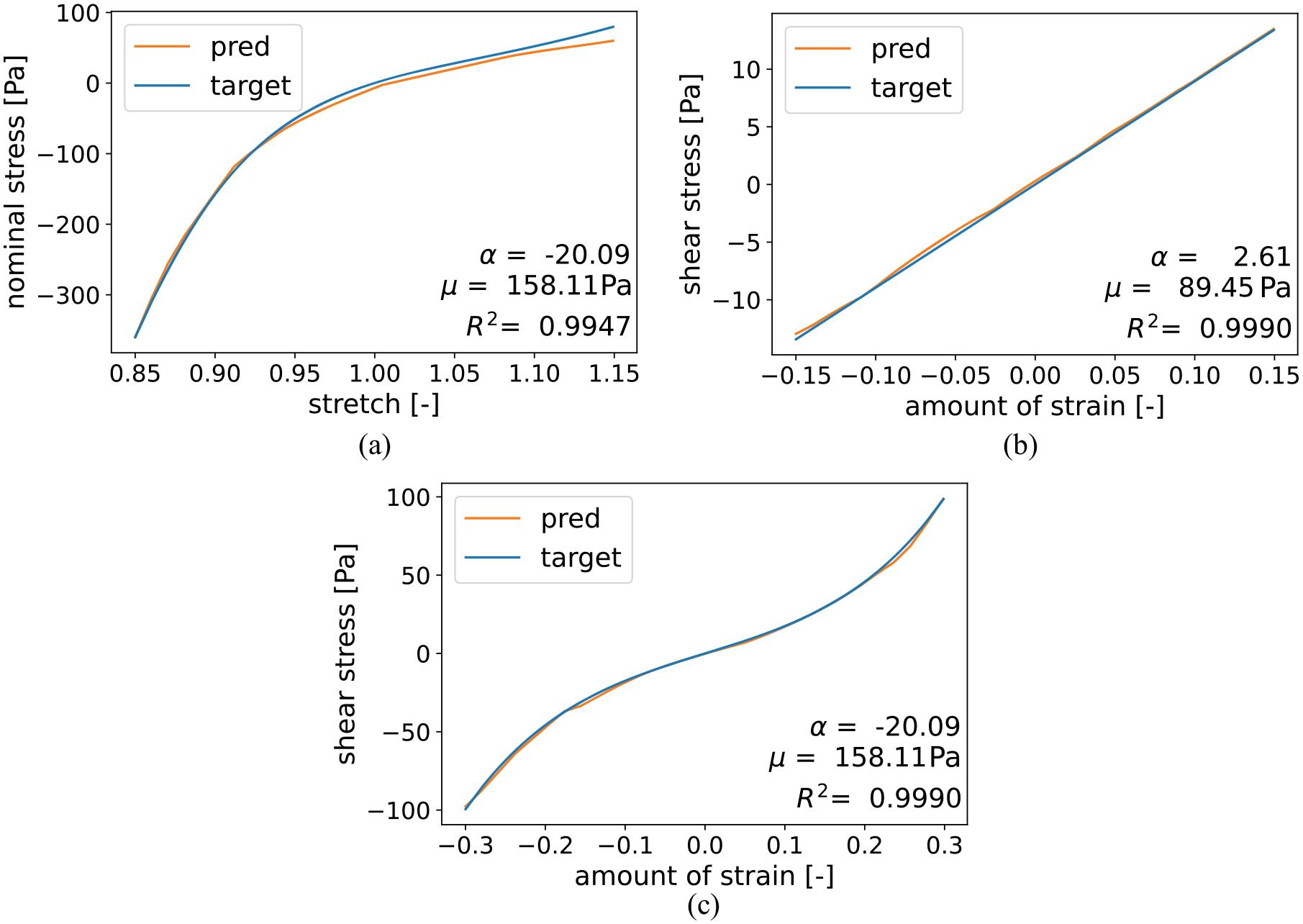
The best achieved fits during testing for each loading mode with corresponding coefficient of determination. The shown modes are compression/tension up to 15% strain (a) as well as torsional shear up to an amount of shear of 0.15 (b) and 0.3 (c).
Figure 5 shows an exemplary history of an optimization in terms of the parameters
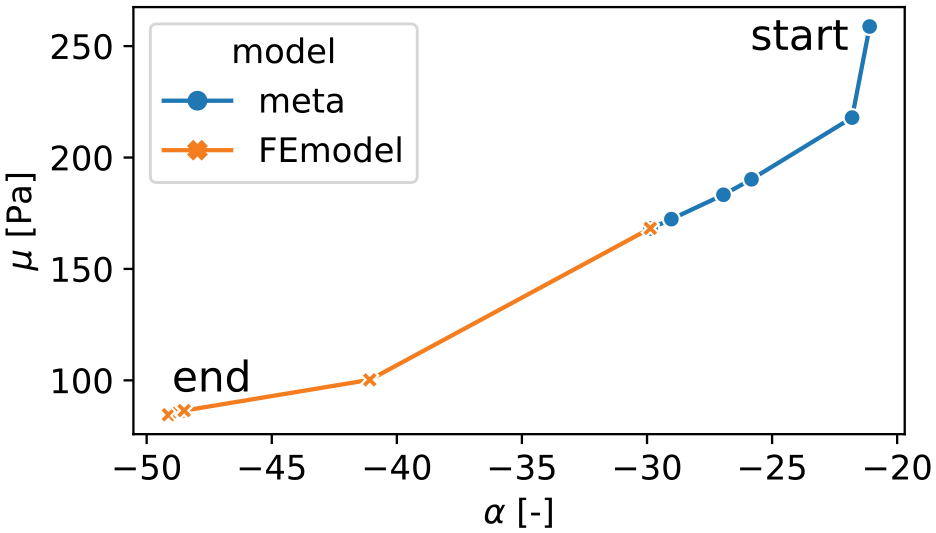
Exemplary history of the parameters
3.2. Mechanical parameters
We fit the FE model with the hyperelastic one-term Ogden model to experimental data from seven human brains published in Hinrichsen et al. [18], distinguishing between the eight regions defined in Table 2. By fitting first only the compression/tension data and, second, all modes including the two torsional shear modes (up to an amount of shear of 0.15 and 0.3) simultaneously, we obtain two different parameter sets. This allows us to later compare whether the different tissue components contribute only to the tension/compression behavior or to the combined mechanical response in all loading modes including shear. An example result for the simultaneous fit of all three loading modes is shown in Figure 6. The fit of the compression/tension data in Figure 6(a) shows the ability of the model to capture the pronounced compression/tension asymmetry that is characteristic for brain tissue. Meanwhile, there is a larger deviation in shear up to 0.15 in Figure 6(b), while the shear mode up to a shear of 0.3 (Figure 6(c)) shows a much better result. The resulting values for the shear modulus
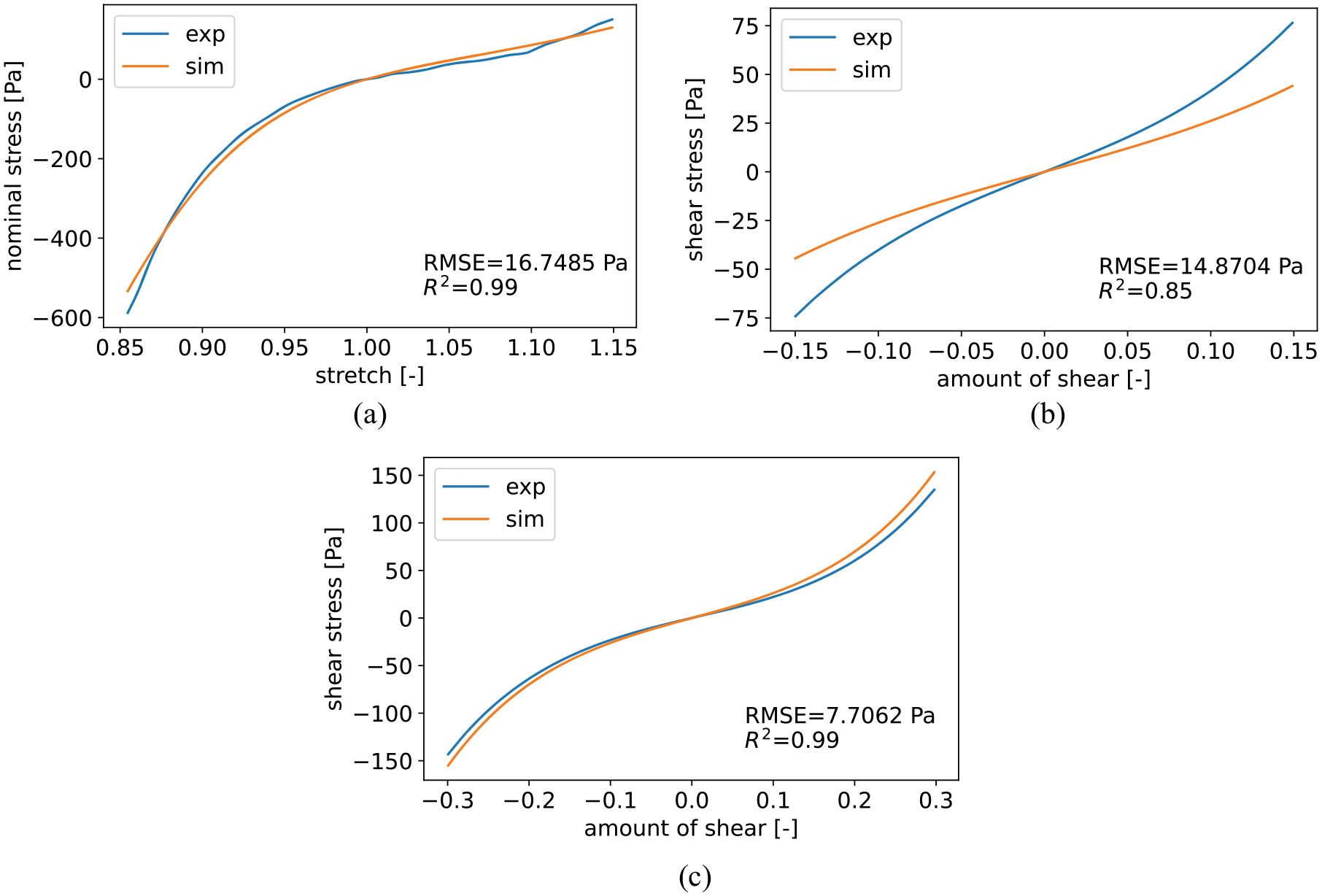
Exemplary fit for the Amygdala region (Am) with material parameters
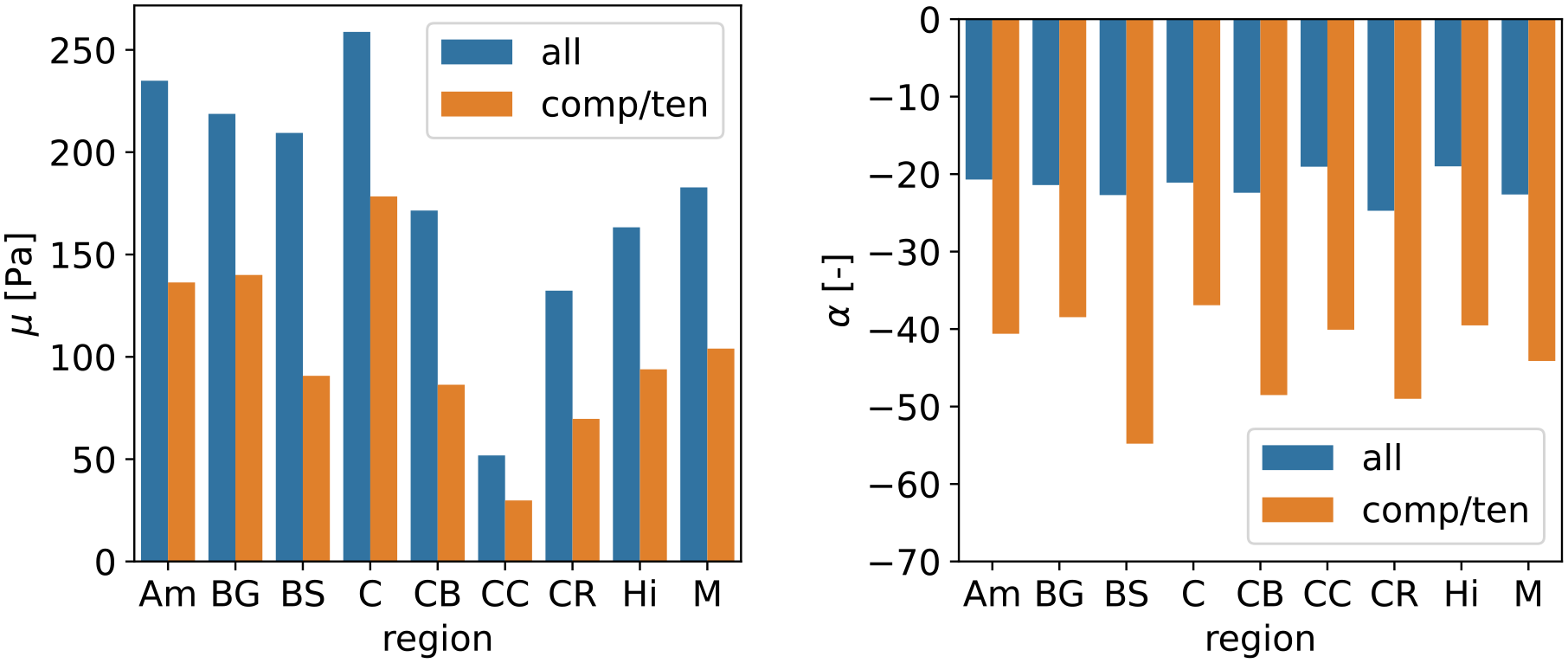
Mechanical parameters
3.3. Correlating mechanics with tissue composition
The simultaneous availability of parameters characterizing the mechanical behavior of human brain tissue in different regions as well as quantitative information on tissue components allows us to investigate their relationship. The linear regression plots for the shear modulus
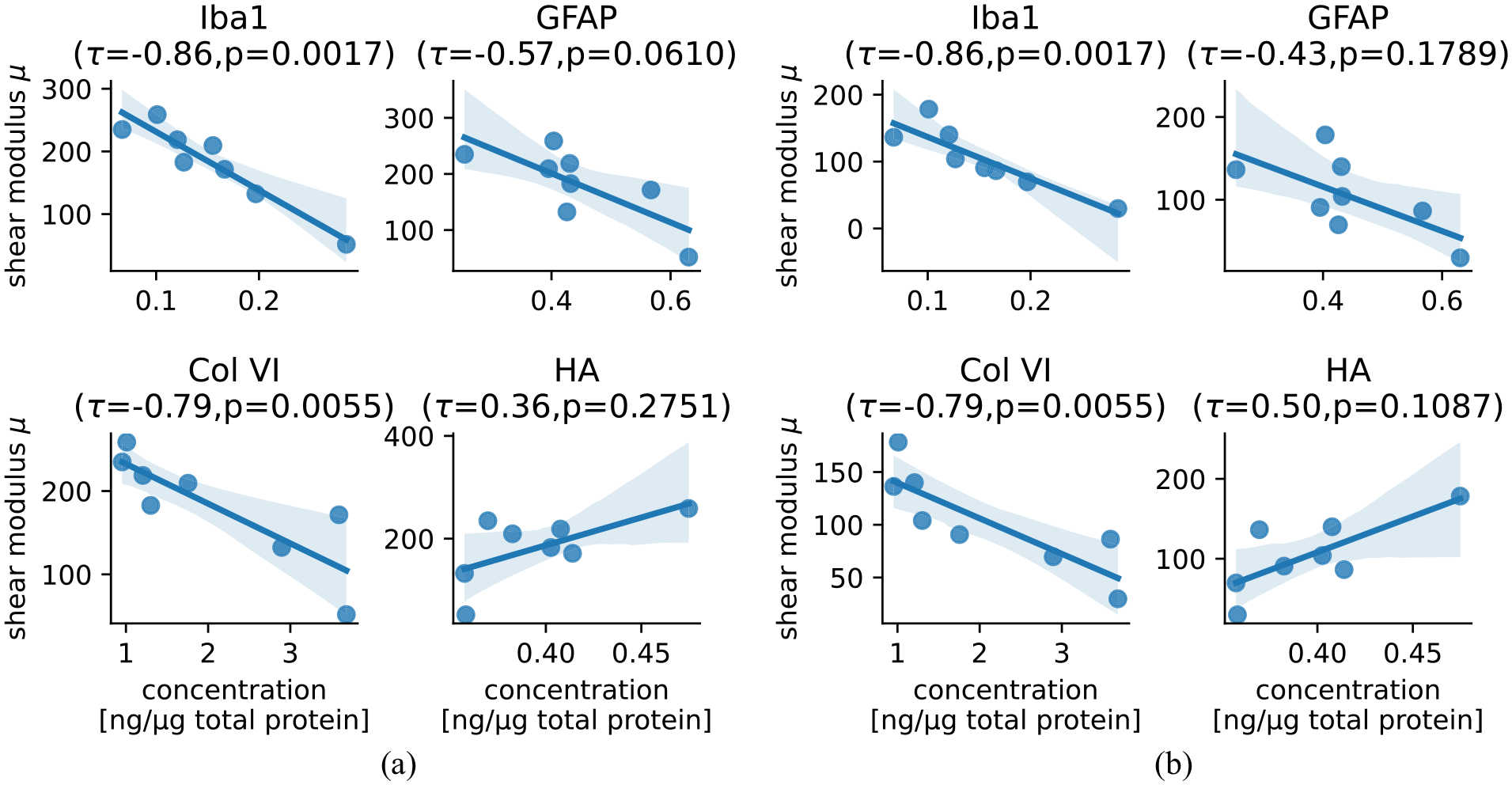
Correlation of relative component concentrations for Iba1, GFAP, Collagen VI, and hyaluronic acid (HA) with the shear modulus
4. Discussion
In this study, we aimed to identify microstructural components of human brain tissue that influence the macroscopic mechanical behavior of the tissue. This was achieved by correlating mechanical parameters for the one-term hyperelastic Ogden model, the shear modulus
4.1. Performance of surrogate model
To mitigate the high computational cost of the finite element simulation used in our inverse parameter identification scheme, we have used neural networks as surrogate models. The models were trained on the output of the finite element simulation and achieved satisfactory results, as demonstrated in Figure 4. We have proposed a procedure that starts the parameter identification with the surrogate model until the optimization converges and then switches back to the original finite element model to achieve even higher accuracy. The example history in Figure 5 shows that the surrogate model leads to an optimum that is closer to the final optimum than the initial parameters. Thus, our study shows that the surrogate model is an easy-to-use tool to reduce the computational time of an inverse parameter identification. Similar results were obtained for the identification of mechanical parameters for blood clots and right ventricular myocardium [23] or aortic wall [31]. However, it should be noted that the neural networks require a sufficient amount of training data, which highly depends on the individual model to be replaced.
4.2. Loading mode dependency of material parameters
In addition to our previously reported data set of hyperelastic parameters for the one-term Ogden model calibrated to compression/tension and torsional shear data simultaneously [18], we have now calibrated parameters using compression/tension data only, similar to the original paper presenting the here-used ELISA data [16]. The direct comparison in Figure 7 shows that only compression/tension data leads to a model with higher strain stiffening behavior, characterized by lower shear moduli
4.3. Correlation between tissue components and mechanical parameters
We have used statistical analyses to identify correlations between the identified region specific hyperelastic material parameters characterizing the mechanical behavior and concentrations of specific tissue components. Component concentrations were obtained for five human brains and averaged over regions in Table 2 while the mechanical parameters were calibrated on region wise averaged experimental data from the same brains as well as two additional ones. For the 11 components studied in Table 1, we calculated the Kendall rank correlation coefficient
The ELISA results were taken from Linka et al. [16], where a CANN was trained on these data together with data from compression/tension experiments to predict the viscoelastic response of human brain tissue. The results presented here now complement that study as we (1) obtain parameters from a finite element simulation as opposed to an analytical solution assuming homogeneous deformation as this is not fully valid for the experiments at hand, (2) use a hyperelastic one-term Ogden model, which we calibrate using the quai-static response under compression/tension and torsional shear simultaneously in addition to compression/tension data only, and (3) evaluate direct correlations between mechanical parameters and individual tissue component concentrations. In Linka et al. [16], a so-called relevance propagation through the CANN was used to quantify the influence of the different ELISA components on the model output. Strikingly, the CANN model had showed that fibronectin (FN) had the highest relevance with a value about three times higher compared to the next highest relevance of Iba1. In our analyses, we find the highest correlation with a clearly visible trend for Iba1 (Figure 8) and only low
Differences between the two approaches could be explained by the fact that the CANN model uses a Prony series with a hyperelastic part characterizing the instantaneous, quasi-elastic response, while our parameters characterize the quasi-static response, as they were calibrated using the averaged loading and unloading at a mean loading rate of
Taken together, our analyses presented in this study indicate direct correlations between the Iba1 concentration representing the amount of microglia in the tissue, the GFAP concentration representing the amount of astrocytes, the concentration of Col VI, and the concentration of Col IV with the tissue stiffness. These observations are consistent with other studies. Iba1 is specifically expressed in microglia/macrophages, which have been shown to migrate to areas of higher stiffness [33]. This may indicate that microglia do not actively contribute to tissue stiffness, but rather populate stiffer areas through a mechanosensing mechanism. This hypothesis is consistent with the observation of [34] that cell bodies do not appear to deform under macroscopic tissue loading and thus do not contribute to the mechanical response. Macrophages are activated after central nervous system (CNS) injury [35, 36] and have been shown to exert traction forces in the repair of vascular rupture [37]. The extracellular matrix protein Col VI provides a support structure for cells [38, 39], is highly expressed in a variety of cancers associated with brain tumors [40], and has been linked to neurodegeneration [41]. Col IV is also an extracellular matrix protein that is abundant in CNS scars [42, 43] and has shown increased values after stab injuries of rat brain tissue together with reduced stiffness [44], which is in good agreement with our findings. GFAP is the major intermediate filament of astrocytes [45], which are highly relevant for the interconnectivity of neurons and the vasculature. Astrocytes showed to be sensitive to the mechanical properties of their environment [46, 47]. HA is a component of the extracellular matrix and is of great structural importance [48] . It is also used to tune the stiffness of hydrogels [49] and plays a role in age-related neurodegeneration [50].
5. Conclusion
By combining information on tissue composition and hyperelastic material parameters, we have identified candidates for tissue components that correlate with the macroscopic mechanical behavior of human brain tissue. The strongest influence was found for Iba1 expressed in microglia/macrophages, followed by collagen VI, GFAP expressed in astrocytes, and collagen IV. To quantify the mechanical properties of human brain tissue, we have identified material parameters for a hyperelastic one-term Ogden model through a neural-network-enhanced inverse parameter identification scheme and show that the correlations found hold for parameters calibrated based on compression/tension and torsional shear data as well as compression/tension data alone. This finding is highly relevant for the design of future experimental testing setups. The identified candidate components are particularly interesting for use in future microstructure-informed constitutive models, which have many potential applications ranging from surgical planning [1] and brain injury modeling [51] to models predicting tumor growth [52] or cortical malformations [4]. The reported results contribute to the path toward highly spatially resolved and patient-specific models for human brain tissue. However, further multidisciplinary research efforts are needed to identify the mechanisms active at different time and length scales to accurately model the macroscopic mechanical behavior of human brain tissue over its entire life cycle.
Footnotes
Appendix 1
Acknowledgements
We cordially thank Jasmin Würges for performing the ELISAs and Lisa Stache for providing the human brains. Furthermore, the authors thank those who donated their bodies to science so that anatomical and biomechanical research could be performed. Results from such research can potentially improve patient care and increase mankind’s overall knowledge. Therefore, these donors and their families deserve our highest gratitude.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The support from the German Research Foundation (Deutsche Forschungsgemeinschaft, DFG) through the grant BU 3728/1-1 to S.B. as well as through project number 460333672 CRC1540 Exploring Brain Mechanics (subprojects A01 and A02) is gratefully acknowledged.
Availability of data and materials
The ELISA data have been published previously in Linka et al. and the mechanical data in Hinrichsen et al. The code generated and analyzed during this study is available from the corresponding author on reasonable request.
