Abstract
Background:
Cyclooxygenase 1 (
Results:
Conclusion:
rs2071746 in
Keywords
Introduction
Nowadays, atherothrombotic diseases have become the leading cause of death in elderly Chinese. Aspirin is still the cornerstone of treatment and prevention for these diseases. 1 However, epidemiological surveys 2 have discovered that 0.4% to 70.1% patients are poor response to aspirin application, known as aspirin resistance (AR). Platelet aggregation measured by platelet function tests cannot be effectively inhibited after taking aspirin, known as laboratory AR. Prospective analysis in clinical events has shown that patients with laboratory AR have an increased risk of adverse clinical outcomes. 3 –6 Therefore, investigating the mechanism of laboratory AR may aid in the clinical use of aspirin. The Framingham heart study found that the heritable factors are the key determinants for platelet aggregation. 7 We hypothesized that the genetic susceptibility may be the key to aspirin sensitivity (AS), despite other acquired determinants.
For aspirin-resistance genetic screening, the methods used to describe the phenotype must be reliable. Aspirin is a nonselective and irreversible inhibitor of cyclooxygenase (
Hemoxygenase-1 (HO-1) is another enzyme that has been shown to be involved in the antithrombotic effects of aspirin.
10
Therefore, we proposed that genetic variants of
Methods
Patients
For this study, information and DNA samples were obtained from consecutive patients who presented between April 2008 and June 2010 and took part in physical examination every year in Chinese PLA General hospital, Beijing 309 hospital, and Navy General Hospital in Beijing with coronary artery disease (CAD), stroke, and peripheral arterial occlusive disease (PAOD) who had been receiving regular aspirin therapy (75-160 mg daily) for at least 4 weeks. Inclusion criteria were ≥65 years of age. Exclusion criteria included the use of clopidogrel, ticlopidine, dipyridamole, or other nonsteroidal anti-inflammatory drugs, heparin or low-molecular-weight heparin; acute vascular events; platelet count <150 000 000/L or >450 000 000/L; and hemoglobin <8 g/dL. Any patients that admitted noncompliance or intermittent compliance were excluded from the analyses. The study was approved by the Scientific and Ethics Review Board of the First Geriatric Cardiology Division, Chinese PLA General Hospital (Beijing, People’s Republic of China). All patients provided written informed consent before inclusion in the study.
Following the screening procedures, 431 patients on aspirin therapy were enrolled. By the LTA diagnostic criteria detailed below, there were 36 patients with AR, 164 patients were partial AR or semi-AR, and 231 patients with AS. These 3 groups are comparable.
Blood Sampling
Blood samples were obtained from patients between 7
Light Transmittance Aggregation
Platelet aggregation was assessed in platelet-rich plasma at 37°C by LTA. Samples were centrifuged at 800 rpm for 5 minutes to obtain native platelet-rich plasma. The platelet count was assessed using a standard cell counter. Platelet-poor plasma was obtained by the centrifugation of remaining blood at 4000 rpm at room temperature for 8 minutes. Aggregation was measured with a ChronoLog Aggregometer (Chronolog, Havertown, Pennsylvania). Aggregation was expressed as the maximal percentage change in light transmittance from baseline after the addition of AA (0.5 mmol/L) and ADP (10 μmol/L), with platelet-poor plasma used as the reference. On the basis of this platelet aggregation assay, the definition of AR was the aggregation of ≥70% with 10 μmol/L ADP and of ≥20% with 0.5 mg/mL AA. 9,11 Aspirin sensitivity was indicated by neither of these criteria being met; AR by both criteria being met; only one of the 2 criteria met was deemed partial AR or semi-AR.
Single-Nucleotide Polymorphism Detection
DNA samples were extracted from peripheral blood obtained from all the patients with AR and AS by a standard phenol–chloroform method. The SNP detection was performed by the use of the Sequenom MassARRAY iPLEX platform (Sequenom, California and USA). 12 The assay consists of an initial locus-specific polymerase chain reaction (PCR), followed by single-base extension that uses mass-modified dideoxynucleotide terminators of an oligonucleotide primer. The PCR primers and single-base extension probes were designed by the Assay Design 3.1 software from Sequenom. The genes and their respective haplotypes as well as the primer pairs and probes are listed in Table 1 . The SNPs were genotyped by the Sequenom MassArray system according to the instructions of the manufacturer. The PCR was performed using iPlex chemistry according to the recommendations of the manufacturer as found in the MassArray iPlex standard operating procedure. Data analysis was performed with the MassArray Typer software version 4.0.
Details of the Primers and Probes for the Candidate Genes.
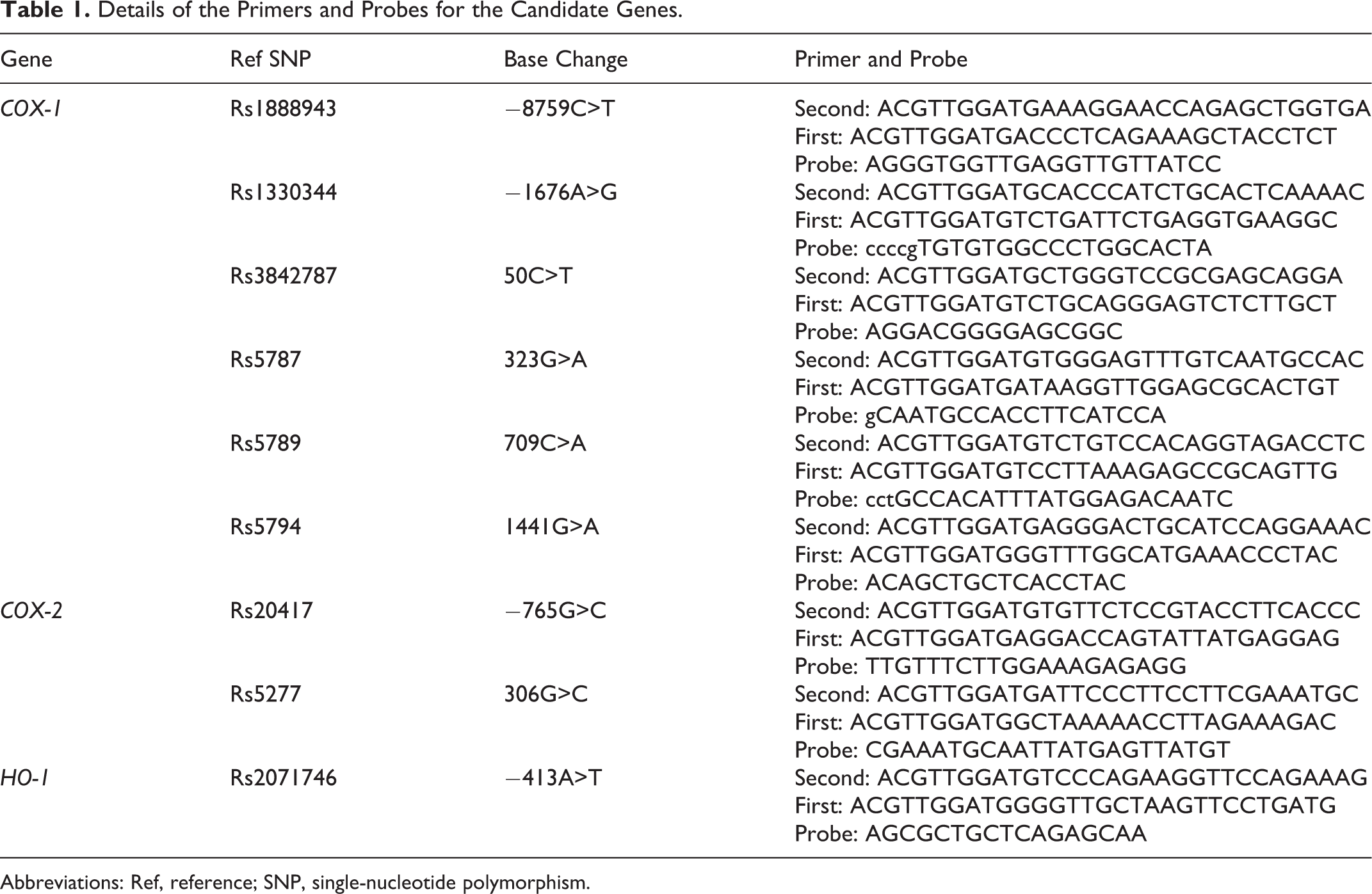
Abbreviations: Ref, reference; SNP, single-nucleotide polymorphism.
Statistical Analysis
Comparisons of the continuous variables were made using the analysis of variance (ANOVA) test or Wilcoxon rank sum test, dependent upon the distribution of the data. Normal data are presented as mean ± standard deviation, data of skewness distribution are presented as median (
Results
Characteristics of the Patients
The study analysis included 431 patients on aspirin therapy. Baseline clinical characteristics are shown in Table 2
. Based on the criteria of the AA-induced aggregation and ADP-induced aggregation, 231 (53.60%) patients were shown to be sensitive to aspirin, while 164 (38.05%) patients were classified as semi-AR and 36 (8.35%) patients were classified as AR. These 3 groups were comparable. No significant differences were found among the 3 groups with respect to age, sex, body mass index (BMI), smoking habits, alcohol, hypertension, CAD, cerebrovascular disease, PAOD, use of calcium-channel blockers, platelet count, Blood urea nitrogen (BUN), creatinine, glucose, cholesterol, high density lipoprotein (HDL), low-density lipoprotein (LDL), HCY, hs-CRP, ATIII activity, PAC-1, and CD62P. In contrast, a significant difference was observed in those patients who take angiotensin-converting enzyme inhibitors/angiotensin receptor blockers (
Demographic Details for the Patients.
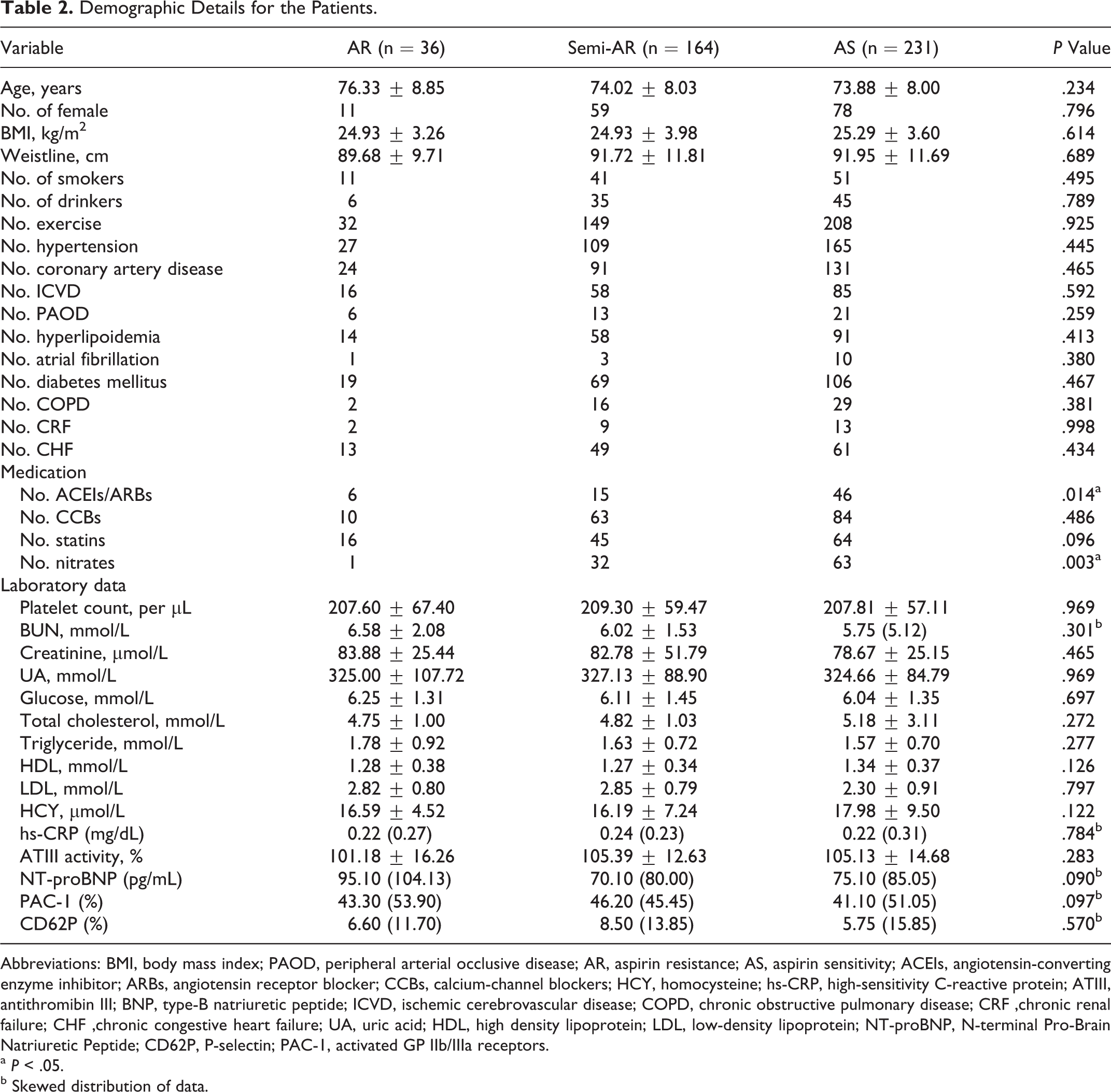
Abbreviations: BMI, body mass index; PAOD, peripheral arterial occlusive disease; AR, aspirin resistance; AS, aspirin sensitivity; ACEIs, angiotensin-converting enzyme inhibitor; ARBs, angiotensin receptor blocker; CCBs, calcium-channel blockers; HCY, homocysteine; hs-CRP, high-sensitivity C-reactive protein; ATIII, antithromibin III; BNP, type-B natriuretic peptide; ICVD, ischemic cerebrovascular disease; COPD, chronic obstructive pulmonary disease; CRF ,chronic renal failure; CHF ,chronic congestive heart failure; UA, uric acid; HDL, high density lipoprotein; LDL, low-density lipoprotein; NT-proBNP, N-terminal Pro-Brain Natriuretic Peptide; CD62P, P-selectin; PAC-1, activated GP IIb/IIIa receptors.
a
b Skewed distribution of data.
Allelic and Genotype Frequencies
The allelic and genotypic frequencies of COX-1_rs1888943, COX-1_rs3842787, COX-1_rs5787, COX-1_rs5789, COX-1_rs5794, COX-2_rs5277, and COX-2_rs20417 variants did not differ significantly among all the patients with AR, semi-AR, and AS. However, between the patients with AR and AS or with semi-AR, the frequencies of the COX-1_rs1330344 variant alleles, HO-1_rs2071746 variant alleles, and HO-1_rs2071746 variant genotypes differed significantly (Table 3 ).
Distribution of SNPs in HO-1, COX-1 and COX-2 Genes Among the 3 Groups.
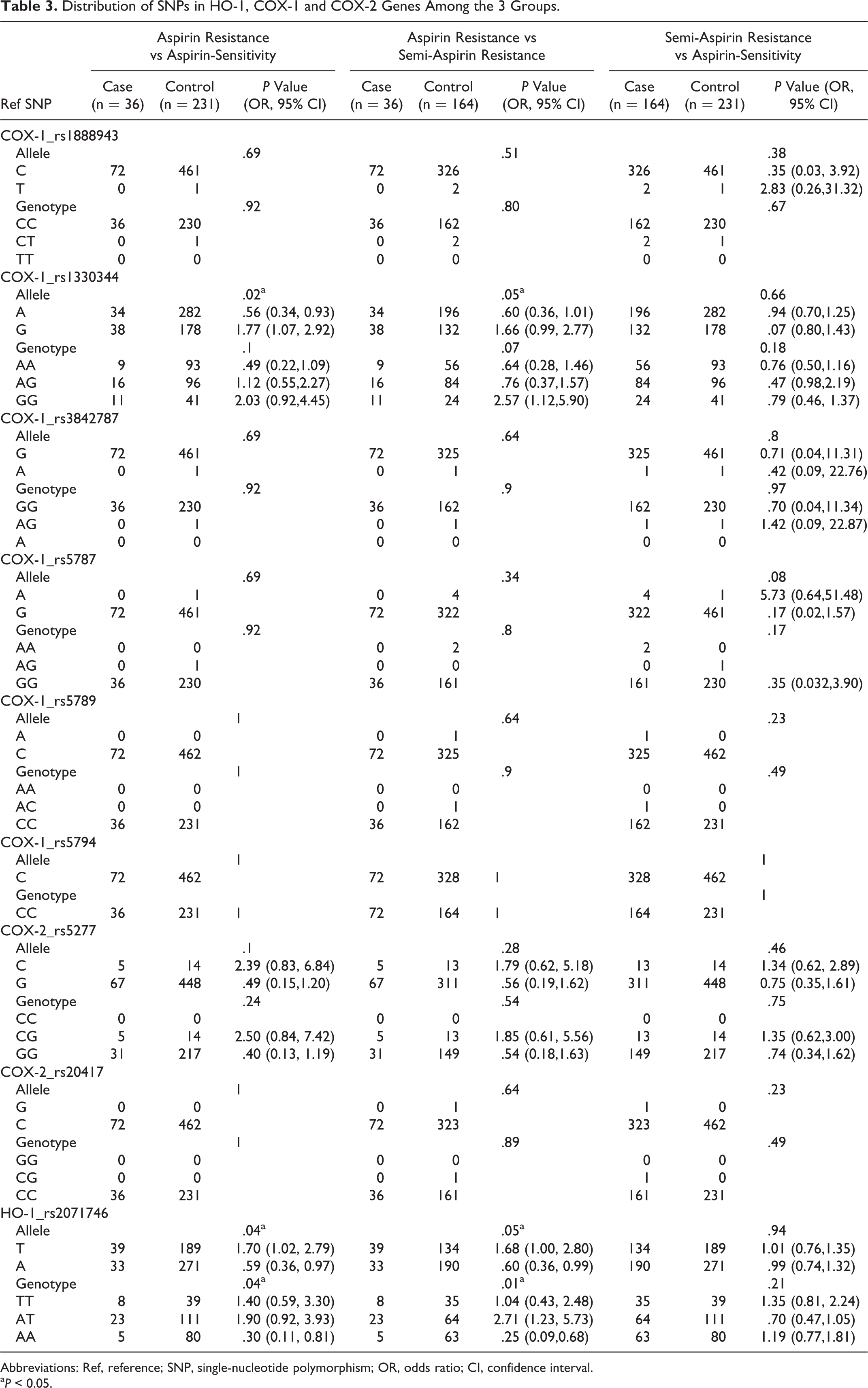
Abbreviations: Ref, reference; SNP, single-nucleotide polymorphism; OR, odds ratio; CI, confidence interval.
a
HO-1 rs2071746 (−413A>T) Associated With AR
The T allele of HO-1 in rs2071746 (−413A>T) was associated with AR, when compared with not only AS patients but also with semi-AR patients (
Genotype of rs2071746 (A>T) Responses to Aspirin Resistance.
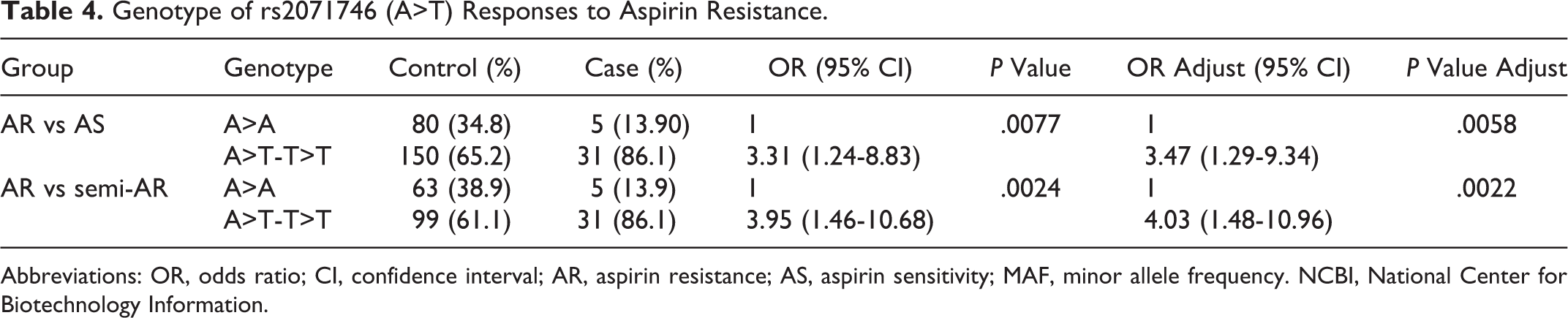
Abbreviations: OR, odds ratio; CI, confidence interval; AR, aspirin resistance; AS, aspirin sensitivity; MAF, minor allele frequency. NCBI, National Center for Biotechnology Information.
COX-1 rs1330344 (−1676A>G) Associated With AR
The G allele of
COX-1 Haplotype Associated With AR
Six SNPs in COX-1 were selected. Both rs 1888943 and rs1330344 are in the promoter region of
Information of Selected SNPs in COX-1 Gene.
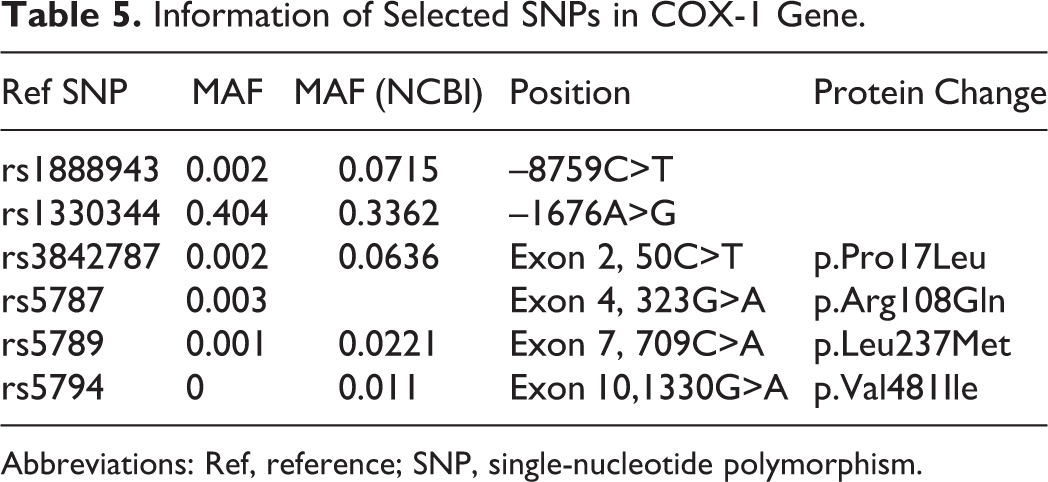
Abbreviations: Ref, reference; SNP, single-nucleotide polymorphism.
COX-1 Haplotype Responses to Aspirin Resistance.
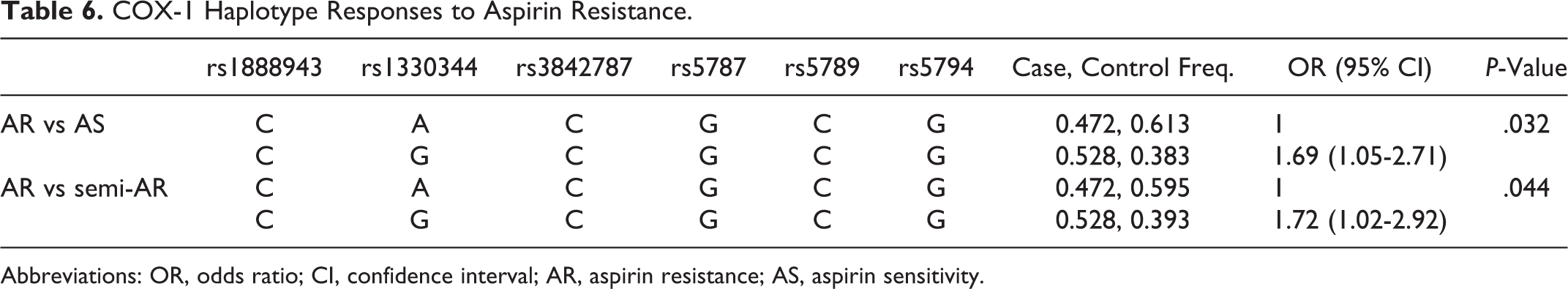
Abbreviations: OR, odds ratio; CI, confidence interval; AR, aspirin resistance; AS, aspirin sensitivity.
COX-2 Genetic Association With AR
The alleles and genotypes of rs5277 and rs20417 showed no differences among the 3 groups (Table 3). The minor allele frequency of rs20417 is also rare in the Chinese Han population.
Discussion
There are plenty of reports about genetic susceptibility to AR. Genetic variants of uridine 5'-diphosphate-glucuronosyltransferase (
Platelet aggregation at the sites of atherosclerotic plaque plays a role in the thrombosis.
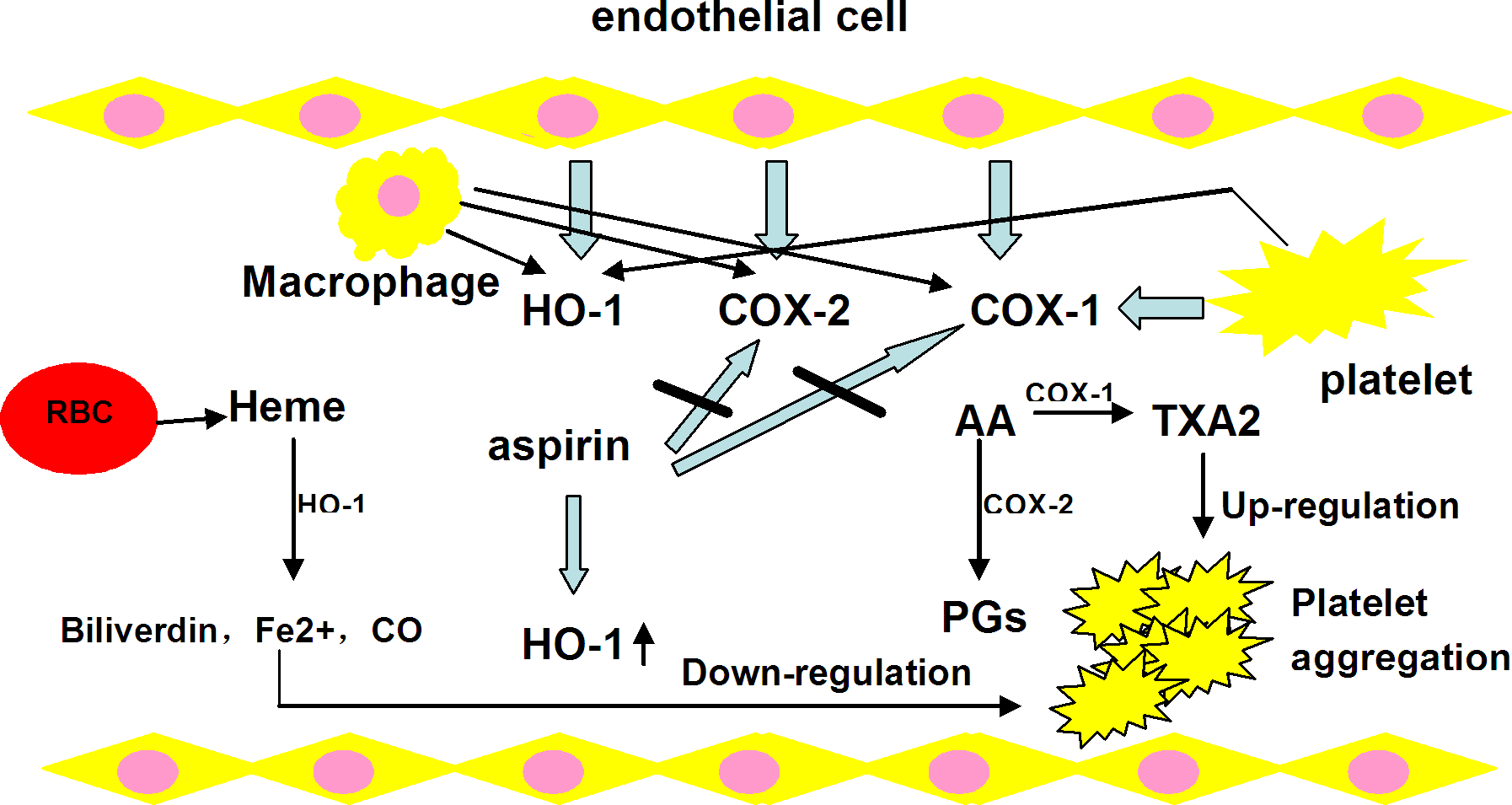
Effects of aspirin via HO-1, COX-1, and COX-2 pathways on platelet aggregation. HO-1 indicates hemoxygenase-1; COX-1/2: cyclooxygenase-1/2.
The highly conserved
Unlike
Similar to
Studies have shown that the major mammalian stress-responsive transcription factors, heat-shock factor, nuclear factor-erythroid 2, AP-1, and the nuclear factor-κB family members, all regulate the
Conclusion
We examined the association of genetic polymorphisms with laboratory aspirin resistance determined by AA-induced LTA combined with ADP-induced LTA. Our study demonstrated for the first time that rs2071746 (−413A>T) gene polymorphism in
Footnotes
Xiao-li Li and Jian Cao equally contributed for this work.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship and/or publication of this article: supported by the Healthcare Fund (07BJZ01) and the National Science and Technology Support Project (2009BAI86B04).
