Abstract
Background:
A highly efficient approach to select microRNA (miRNA) targets is a key to develop a miRNA-based therapeutic approach to cardiac ischemia–reperfusion (I/R). To reverse the change induced by disease, I/R in this case, is the traditional strategy to develop therapeutic drugs. However, examples show that it will not always serve the purpose. In this study, we demonstrate an additional approach of selecting miRNA targets with therapeutic potential following cues from cardioprotection-induced changes rather than by reversing disease-induced changes in cardiac I/R.
Methods:
Isolated perfused rat hearts subjected to I/R were treated with 50 μmol/L sodium hydrosulfide (NaHS) or 10 nmol/L urocortin 2 (UCN2). Cardiac miRNA regulations were determined by miRNA array. Functional screening of selected miRNA mimics, assessed by WST (2-(4-Iodophenyl)-3-(4-nitrophenyl)-5-(2,4-disulfophenyl)-2H-tetrazolium, monosodium salt) activity and lactate dehydrogenase (LDH) release, was performed in H9c2 and neonatal rat ventricular myocytes (NRVMs) with hypoxia/reoxygenation. RNA-induced silencing complex (RISC)-loaded miRNAs caused by mimic transfection were quantified following argonaute-2 immunoprecipitation. Gene regulations of 1 selected miRNA were determined by quantitative polymerase chain reaction and Western blot.
Results:
Treatment with NaHS and UCN2 significantly improved cardiac function and reduced LDH release. The miRNA array indicated a panel of commonly up- and downregulated miRNAs. Among them, 10 upregulated miRNAs with antiapoptotic and antiautophagy potentials were selected for further screening. Mimics of miRNA-221, -150, and -206 were protective in both H9c2 and NRVM. RISC-loaded miRNAs were up by ∼20-fold above. To further prove the feasibility of this approach, miRNA-221 was studied. It reduced I/R-induced caspase 3/7 activity and LC3-II (microtubule-associated protein 1 light chain 3). Measuring genes predicted to regulate apoptosis and autophagy, miRNA-221 mimic decreased Ddit4, TP53inp1, and p27 at both messenger RNA (mRNA) and protein levels, and reduced mRNA of Bak1 and Puma and proteins of Bim and Bmf.
Conclusion:
Mimicking miRNA changes caused by cardioprotective agents, combined with functional screening, enables investigators to efficiently identify novel miRNAs with therapeutic potential in cardiac I/R.
Keywords
Introduction
Since the discovery of the first microRNA (miRNA) 20 years ago, it has been confirmed that more than 1000 miRNAs are ubiquitously expressed across different species and different cell types. 1,2 The seed region of miRNA nucleotides binds to complementary messenger RNA (mRNA) sequences in the 3′-untranslated region assembling into a complex known as miRNA RNA-induced silencing complex (RISC), through which mRNAs are negatively regulated by translational repression or target mRNA degradation. 3,4 As up to 90% of mammalian genes are regulated by miRNAs, this nexus is a “hot spot” for studies of gene regulation. 5 The following features of miRNA suggest they are promising therapeutic targets in acute cardiac disease including ischemia–reperfusion (I/R). (1) Expression of miRNA is rapidly dysregulated in cardiovascular diseases including myocardial infarction (MI). 6 (2) Each miRNA regulates multiple mRNAs posttranscriptionally providing the opportunity to influence the expression of entire networks of genes with potentially powerful integrated overall effects upon the responses to injury including inflammation, fibrosis, and apoptosis. 7 (3) MicroRNAs are small and their sequences are well conserved between animal species and humans. Therefore, the outcomes yielded from animal studies may well be translatable into clinical applications. Already plentiful existing data demonstrate the therapeutic potential of miRNA-based interventions. 8,9
MicroRNAs are differentially regulated in different disease states. To reverse the change induced by disease is a traditional strategy to develop therapeutic drugs. However, it will not always serve the purpose. For example, during cardiac hypertrophy and heart failure, while both miRNA-1 and miRNA-133 are downregulated, overexpression of miRNA-1 prevents hypertrophy but overexpression of miRNA-133 does not. 10,11 MicroRNA dysregulation following I/R injury is a time-dependent process. In response to ischemia or I/R, cardiac cells including cardiomyocytes, cardiac fibroblasts, and endothelial cells undergo variable miRNA dysregulation. These changes can be apparent as early as 15 minutes after coronary artery ligation and may last for hours, days, or months. 12 This dysregulation may comprise a mix of protective and deleterious effects. Following very brief ischemia or reperfusion, such changes in miRNA expression might be relevant to early cell survival. A prolonged ischemia and/or reperfusion will cause permanent injury to the heart, and changes in miRNA expression may be pertinent to apoptotic status and other chronic processes including cardiac fibrosis and remodeling. Dysregulation of any given miRNA may vary in direction and main effect through the course of the natural history of a disease. For example, in different studies, miRNA-1 has been reported to be up- and downregulated following ischemia. 12,13 There are contradictory reports for miRNA-21 as well. 14,15 Therefore, to complement the traditional strategy of seeking miRNA therapeutic targets by reversing the dysregulated miRNA in disease, it is necessary to develop supplementary selection strategies that better pinpoint the timing and direction of beneficial change in expression of individual miRNAs. The miRNA response to cardioprotective agents may assist in identification of potential therapeutic miRNA targets.
The goal of this report is to demonstrate the feasibility of selecting miRNA targets with therapeutic potential guided by cues from cardioprotection-induced changes rather than purely by reversing disease-induced changes in cardiac I/R. This study covers (1) screening using miRNA array technology comparing miRNA profiles in an I/R model with or without cardioprotection; (2) miRNA functional screening using cardiac H9c2 cells and primary neonatal rat ventricular myocytes (NRVMs) to select miRNAs that protect against I/R injury in vitro. In this study, 10 miRNAs were selected from 20 upregulated miRNAs, which should be enough to support our hypothesis; (3) characterization of miRNA transfection induced regulation of RISC-loaded miRNA; and (4) elucidation of underlying protective mechanisms in the regulation of apoptosis and autophagy by detecting changes in predicted targets at mRNA and protein levels by quantitative polymerase chain reaction (qPCR) and Western blot, respectively. As a proof-of-principle example, the underlying mechanisms of miRNA-221’s effects were further studied. Together, this work demonstrates a helpful supplementary strategy for the selection of miRNA targets by mimicking the changes induced by cardioprotection rather than confining attention to reversing changes caused by cardiac ischemic injury. This augmented approach to target selection facilitates discovery of potential therapeutic miRNA targets for in vivo studies to assess miRNA-based intervention in cardiac ischemic injury.
Materials and Methods
This study was approved by the Institutional Animal Care and Use Committees of the National University of Singapore and complied with the Guide for the Care and Use of Laboratory Animals published by the National Institutes of Health (NIH Publication No. 85-23, Revised 1996). All rats were housed in a temperature-controlled room (21°C ± 2°C) with 12 hours light and dark cycle. Water and diet were available ad libitum. Forty-five male Sprague-Dawley rats (280 ± 30 g) were used in this study for miRNA array in an isolated heart perfusion model. The NRVMs were isolated from 1- to 3-day-old neonatal rats. The H9c2 cells (a rat cardiac myoblast cell line) purchased from American Type Culture Collection (Manassas, Virginia) were used in this study at 25th to 30th passage. MicroRNA mimics, inhibitors, their controls, transfection reagents, and miRNA stem-loop qPCR primers and reagents were purchased from Life Technologies (Life Technologies, Thermo Fisher Scientific Inc, Singapore). Chemicals and reagents were purchased from Sigma-Aldrich Co, LLC, Singapore if not otherwise indicated.
Ischemia–Reperfusion Protocol in Isolated Heart Preparations
Hearts were perfused according to a standardized protocol as described previously. 16 In brief, rats were anesthetized with 50 mg/kg sodium pentobarbital intraperitoneally, hearts were rapidly harvested, and then perfused on a modified Langendorff apparatus (PL3508B22-V Langendorff System, ADInstruments, Advance Tech Pte Ltd, Singapore) with modified Krebs-Henseleit buffer at 37.5°C and gassed continuously with a mixture of 95% O2 and 5% CO2 to maintain pH at 7.4 ± 0.1 (SG98 ELK SevenGo Duo RDO pH/RDO meter, Metter-Toledo Pte Ltd, Singapore). Hearts were perfused at 75 mm Hg constant pressure while beating spontaneously. Cardiac function was monitored throughout perfusion. After equilibration, hearts were subjected to 20 minutes of global zero flow ischemia and 60 minutes of reperfusion and received (1) no treatment (control), (2) 50 μmol/L sodium hydrosulfide (NaHS), or (3) 10 nmol/L urocortin 2 (UCN2). Drugs were given 15 and 30 minutes before ischemia, respectively. Coronary effluent was collected at the early and end of reperfusion. Hearts were freeze-clamped at the end of perfusion to extract RNA for miRNA array analysis. Hearts undergoing normoxic protocols or exposed to 20 minutes ischemia only, with or without treatments, were also collected (n = 5 each group).
MicroRNA Array Assessment
RNA from −80°C stored cardiac tissue powder was extracted with TRIzol Reagent (Invitrogen, Life Technologies, Singapore) followed by DNase I treatment (Ambion, Life Technologies, Singapore). MicroRNA profiling was performed with LNA™-modified oligonucleotide (Exiqon, Denmark) probes (Sanger miRBase ver 12.0) according to the manufacturer’s instructions. In brief, a total of 1 μg RNA per sample was processed for 3′-end-labeling with Cy3 fluorescent dye (miRCURY LNA miRNA Hi-Power labeling kit, Exiqon, Denmark), hybridization with miRNA probes on the slide (miRCURY LNA microRNA array kit, 7th generation-human, mouse & rat, Exiqon, Denmark), and washing. The microarray chips were scanned using InnoScan700 (Innopsys, Carbonne, France). MicroRNA expression is positively correlated with fluorescent intensity. Fluorescent intensity with a raw reading less than 300 was considered as noise and filtered, and 75th percentile global normalization was used to normalize data (GeneSpring GX, Agilent Technologies, Singapore). MicroRNAs with an absolute fold change ≥2.0 (I/R-NaHS and I/R-UCN vs I/R-control) were selected.
Hypoxia/Reperfusion in H9c2 and NRVM
The H9c2 cells were cultured according to a standard protocol. Ischemia–reperfusion was induced by 15-hour hypoxia (<0.2% O2) with modified Dulbecco modified Eagle medium (DMEM): serum free and reduced glucose of 1 g/L (vs 4.5 g/L in normal DMEM), followed by 2 hours normoxic culture (“reperfusion”). The NRVMs were isolated from 1- to 3-day old rats as described previously. 17 Briefly, ventricles were minced and digested in 0.3 to 0.5 mg/mL collagenase type II (Worthington Biochemical Corp, i-DNA Biotechnology Pte Ltd, Singapore) for 3 to 4 times. Cell suspensions were collected and combined for Percoll (GE Healthcare, SciMed [Asia] Pte Ltd, Singapore) density gradient centrifugation to separate cardiomyocytes from cardiac fibroblasts. Purified cardiomyocytes were seeded in a 1:4 mixture of M199 and DMEM containing 15% fetal bovine serum. After overnight attachment, fresh medium was replaced with 50 µmol/L cytosine arabinoside-C; 100 units/mL penicillin, 100 µg/mL streptomycin, and 250 ng/mL amphotericin B were presented during the entire culture process. I/R was induced by 6-hour hypoxia (<0.2% O2) in modified Esumi buffer 18 : 137 mmol/L NaCl, 12 mmol/L KCl, 0.49 mmol/L MgCl2, 0.9 mmol/L CaCl2•2H2O, 4 mmol/L 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid, 20 mmol/L sodium lactate, pH 6.2 followed by 2-hour “reperfusion” with normoxic culture.
Transfection of miRNA Mimics and Inhibitors into H9c2 and NRVM
Transfections were undertaken according to the manufacturer’s instructions. In brief, the transfection complex was formed by incubating each individual miRNA mimic or inhibitor or its control with lipofectamine at a ratio of 20 pmol:1 μL in antibiotic-free DMEM. Cells were incubated with the transfection complex for 3 hours, then topped up with normal growth medium and incubated for a further 21 hours. Transfection was terminated by medium change; 25 nmol/L doses of mimics or inhibitors were transfected 24 hours before hypoxia. Additionally, miRNA fold changes correlated with different concentrations of 0.2, 1, 5, and 25 nmol/L were tested. Cy3-labeled miRNA mimic control was used to assess miRNA transfection efficiency.
MicroRNA Functional Screening
Ten miRNAs commonly upregulated by both NaHS and UCN2 with antiapoptotic and antiautophagy potentials were screened for therapeutic potential in hypoxia/reperfusion. H9c2 and NRVM were seeded in 96-well plates at 60% and 90% confluence, respectively. Following transfection, cells were subjected to I/R, then lysed for WST-1 (Abcam, Singlab Technologies Pte Ltd, Singapore) assessment. The WST-1 measures cellular mitochondrial dehydrogenase activity as an indicator of the number of metabolically active cells. 19 Hypoxia culture medium was collected for measurement of lactate dehydrogenase (LDH; Sigma-Aldrich Co. LLC, Singapore) release, an indicator of cell death. 20
Argonaute-2 Immunoprecipitation
Argonaute 2 (Ago2), the miRNA-specific binding protein in the RISC, was pulled down by Anti-ago2 antibody (monoclonal 11A9, Sigma-Aldrich Co. LLC., Singapore). 21 In brief, H9c2 transfected with negative mimic control and specific miRNA mimics were lysed and preincubated with 2.5 µg Ago2 antibody, or rat immunoglobulin G (IgG) as nonspecific control overnight, and then further incubated with 20 µL Dynabeads (Life Technologies, Thermo Fisher Scientific Inc, Singapore) for 3 hours. Beads were washed 5 times with buffer: 300 mmol/L NaCl, 50 mmol/L Tris-HCl, 1 mmol/L MgCl2, and 0.1% (v/v) NP-40, at pH 7.4. Beads binding Ago-2 were washed out with protein lysis buffer or TRIzol for protein or RNA extraction, respectively. The input (total cell lysate), unbound, and specific bound proteins and RNA were collected for Western blot and qPCR analysis.
Caspase 3/7 Analysis
Activities of caspase 3 and -7, indicators of apoptosis, were measured using luminescent assay (The Caspase-Glo 3/7 Assay, Promega, Singapore). Following 15 hours of hypoxia and 2 hours of reoxygenation, cell lysis were incubated with reagents following instruction and Luminometer readings were taken (EnSpire 2300 Multilabel Reader, PerkinElmer, Singapore). Data were presented as relative to I/R control.
MicroRNA Stem-Loop qPCR and mRNA Real-Time qPCR
RNA from H9c2 and NRVM was extracted with TRIzol reagent (Invitrogen, Life Technologies, Thermo Fisher Scientific Inc, Singapore) followed by DNase I treatment. MicroRNA expression was determined by Stem-loop qPCR (TaqMan MicroRNA Assay, Life Technologies, Singapore; CFX96 Real-Time PCR Detection System, Bio-Rad Laboratories Pte Ltd, Singapore). Data were normalized to U6B as the reference gene and Normoxia control using ΔΔCT method or normalized to U6B only using ΔCT method. The mRNA expression was determined by universal reverse transcription and real-time PCR (iScript Reverse Transcription Supermix for RT-qPCR, iTaq Universal SYBR Green Supermix, Bio-Rad Laboratories Pte Ltd, Singapore) and normalized by TATA-box binding protein (Tbp) as a reference gene. Both miRNA and mRNA expressions were analyzed using the ΔΔCT method. The forward/reverse mRNA primer sequences are summarized in Table 1.
The Sequences of mRNA Primers Used in this Study.

Abbreviations: R, reverse primer sequence; F, forward primer sequence.
Western Blot
Standard procedures were employed for Western blots. In brief, protein concentrations from H9c2 cells lysis were measured (Bio-Rad Laboratories Pte Ltd, Singapore) and normalized with 6-fold loading buffer and dH2O. Samples of 25 μg protein were loaded for electrophoresis through 5% to 14% sodium dodecyl sulfate–polyacrylamide gel electrophoresis gels. Following transfer, polyvinylidene difluoride membrane was incubated with antibodies separately, Bim, Bak1, Puma, Bmf, Ddit4, Tp53INP-1, and p27 (Cell Signaling Technology, Research Biolabs PTE Ltd, Singapore), LC-3, and β-actin (Santa Cruz, Cell Science Pte Ltd, Singapore). β-Actin was used as loading controls to normalize loading.
Statistics
Data were compared for differences by 1-way analysis of variance followed by Bonferroni post hoc analysis or the unpaired 2-tailed t test (Graph Pad Prism, San Diego, California), as appropriate. All values are presented as mean ± standard error of the mean. A P value of less than .05 was considered statistically significant. MicroRNA array data were analyzed with GeneSpring GX (Agilent Technologies). The distributions of miRNA profiling and fold changes are shown by Principal Component Analysis plot (PCA) and Heat map, respectively.
Results
H2S and UCN2-Induced Cardioprotection Against I/R Injury Similarly Regulate Cardiac miRNA
In as little as 20 minutes of ischemia and ischemia followed by reperfusion, multiple miRNAs were dysregulated, with most downregulated (Supplemental Figure 1). We hypothesized that either or both H2S and UCN2 would regulate miRNA in a cardioprotective pattern. Cardiac function and LDH release from the isolated hearts are presented in Figure 1. There were no functional differences between groups at baseline (Figure 1a). However, pretreatment with NaHS or UCN2 significantly improved functional recovery at the end of reperfusion compared to nontreated controls (Figure 1B). Heart rates were not different between 3 groups, for example, 293 ± 12, 290 ± 21, 276 ± 13 bpm, respectively. The LDH release was reduced at the early and late reperfusion. Despite different underlying protective mechanisms, 22,23 the miRNA profiles induced by NaHS and UCN2 were very similar (Figure 2). The PCA plots clearly showed that NaHS and UCN2 treatment shifted overall miRNA profiles in normoxia, ischemia, and reperfusion (Figure 2A). During I/R, NaHS and UCN2 induced very similar changes in miRNA regulation (Figure 2B), with ∼20 miRNAs up- and 20 downregulated (≥ 2.0-fold or ≤ −2.0-fold vs I/R control, Figure 2C).
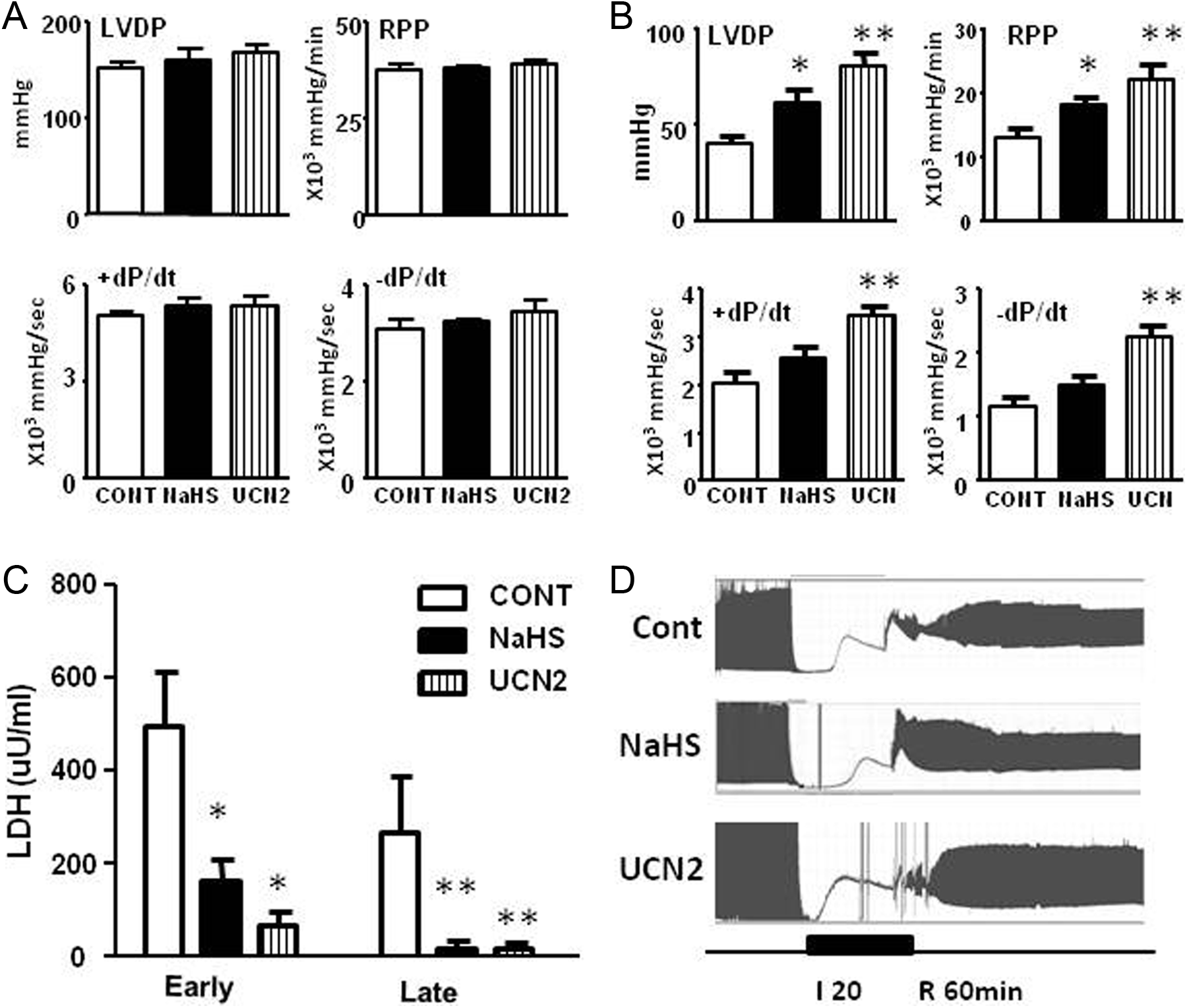
Pretreatment with sodium hydrosulfide (NaHS) and urocortin 2 (UCN2) protects hearts against ischemia/reperfusion injury. Cardiac function is indicated by left ventricular developed pressure (LVDP); RPP (rate pressure product) = LVDP × heart rate; +dP/dt (the rate of increase in pressure over time); −dP/dt (the rate of decrease of pressure over time). A, Baseline cardiac function of nontreated control, NaHS, and UCN2. B, Cardiac functional recovery at the end of reperfusion. C, Lactate dehydrogenase (LDH) release at the early and late of reperfusion. D, Time course of LVDP during perfusions. *P < .05, **P < .01 vs nontreated control (control) by 2-tail unpaired t test. N = 5 each group.

MicroRNA (miRNA) profiling and fold change caused by sodium hydrosulfide (NaHS) and urocortin 2 (UCN2) treatment. A, Principal component analysis (PCA) showing the overall miRNA regulations induced by NaHS and UCN2 treatment under normoxia, ischemia, and reperfusion conditions. Ischemia/reperfusion (I/R) groups are circled. B, Heat-map and (C) miRNAs list showing all up- and downregulated miRNAs for 2-fold or above during reperfusion (vs I/R control).
MicroRNA Overexpression Protective of H9c2 in I/R
Preliminary work indicated that hypoxia for 15 hours followed by 2-hour reperfusion in H9c2 induced significant cell injury as determined by cell morphology, cell viability, apoptosis assessment, WST, and LDH measurements. The transfection efficiency was about 97% (Supplemental Figure 2). Ten miRNAs upregulated by both cardioprotective treatments with potential antiapoptotic and antiautophagy properties were selected for functional screening tests. Among 10 miRNA mimic transfections, 5 (miRNA-221, -150, -206, -145, and -29b) were protective as indicated by increased WST activity and reciprocally decreased LDH release (P < .05 vs I/R-mimic control, Figure 3A and C). To double check the effects of these miRNA interventions, downregulation of the same miRNAs by miRNA inhibitor transfection was performed as well (Figure 3B and D).

The effects of microRNA (miRNA) intervention in H9c2. Ten miRNA mimics and 5 miRNA inhibitors were transfected in H9c2 followed by 15 hours of 0.2% oxygen hypoxia and 2 hours of reoxygenization (I/R). Increase in WST activity and/or decrease lactate dehydrogenase (LDH) release indicate the protection achieved by miRNA interventions. A and C, WST activity and LDH release following mimics transfection. Five of 10 miRNA transfected mimics were protective. B and D, WST and LDH following inhibitor transfection. *P < .05, **P < .01, and ***P < .001 vs mimic or inhibitor control by 2-tail unpaired t test. Each transfection was conducted 3 times in triplicate.
MicroRNA Overexpression Protective of NRVM in I/R
Parallel experiments were conducted using NRVM. Preliminary work indicated that 6 hours of hypoxia followed by 2 hours of reperfusion induced significant cell injury as determined by assessments of cell morphology, cell viability, and apoptosis as well as measurement of WST and LDH. The transfection efficiency was about 90% (Supplemental Figure 3). Among the 5 protective miRNA mimics tested in H9c2 experiments, miRNA-221, -150, and -206 were similarly protective in NRVM as indicated by decreased LDH release (P < .05 vs I/R-mimic control, Figure 4A). The increase in WST activity was detected with miRNA-221 mimics (Figure 4C). Conversely, the corresponding miRNA inhibitors had partial adverse effects (Figure 4B and D).
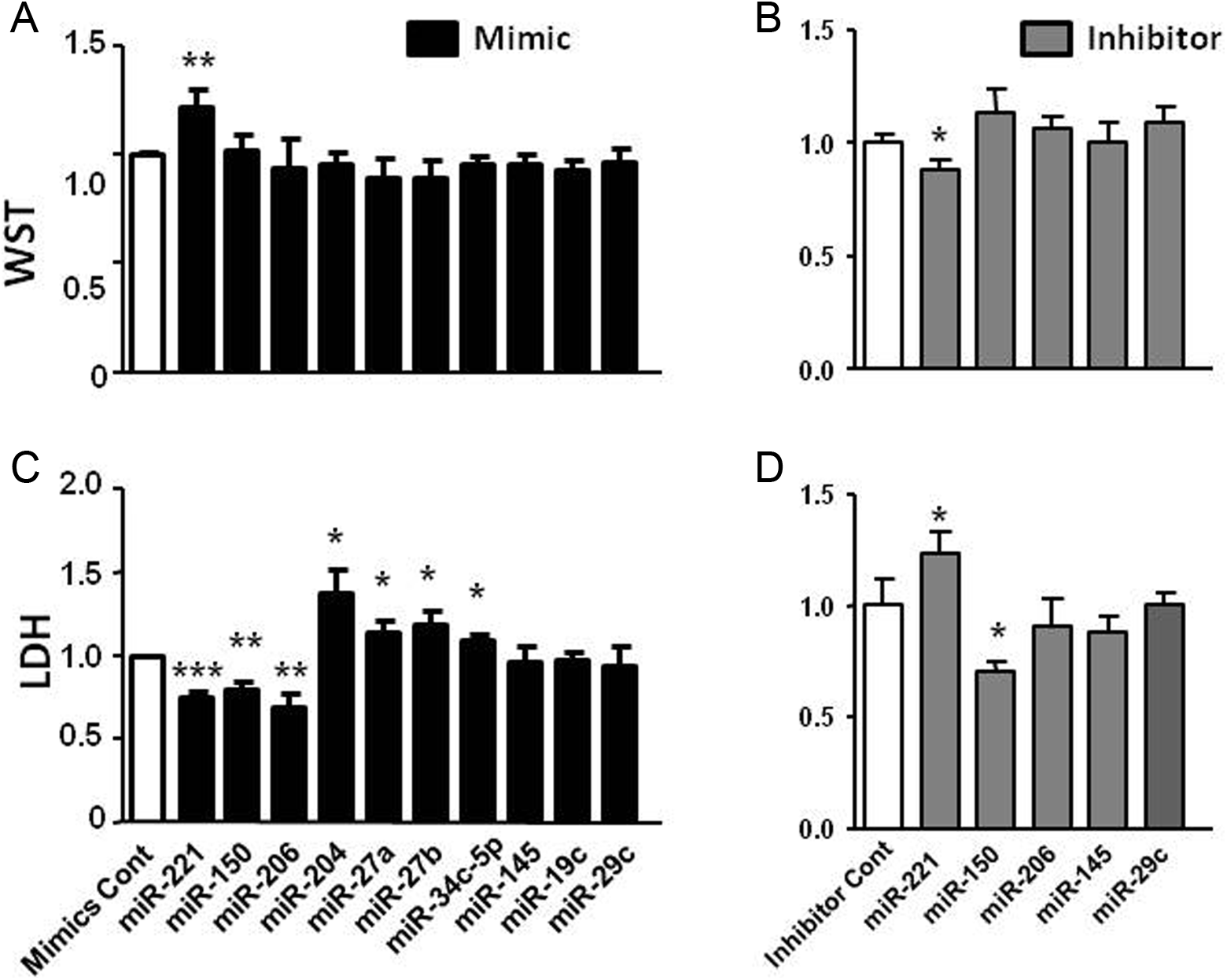
The effects of microRNA (miRNA) intervention in neonatal rat ventricular myocyte (NRVM). Ten miRNA mimics and 5 of effective miRNA inhibitors were transfected in NRVM followed by 6 hours of 0.2% oxygen hypoxia and 2 hours of reoxygenization (I/R). Increased WST activity and/or decreased lactate dehydrogenase (LDH) release indicate the protection by miRNA interventions. A and C, WST activity and LDH release following mimics transfection. Three of 10 transfected miRNA mimics were protective. B and D, WST and LDH following inhibitor transfection. *P < .05, **P < .01, and ***P < .001 versus mimic or inhibitor control by 2-tail unpaired t test. Each transfection was repeated 3 times in triplicate.
Ischemia/Reperfusion Dysregulated miRNAs in H9c2 and NRVM
Endogenous expression of the 5 miRNAs of interest was measured in H9c2 and NRVM in I/R. When normalized to U6B and normoxia using ▵▵CT method, miRNA-206, -29c, and -221 were significantly downregulated by I/R in H9c2 (P < .05 vs normoxia, Figure 5A), whereas expression of miRNA-150 and -145 was unchanged. The overall pattern of miRNA downregulation was similar in NRVM with miRNA-145 significantly downregulated as well (P < .05 vs normoxia, Figure 5B). In order to show the expression level of individual miRNAs relative to each other, the expression level was normalized to U6B only using ▵CT method, a well-accepted housekeeping small nucleotide. The expression of miRNA-150 was extremely low in both cell types and that of miRNA-206 was most abundant in H9c2 and miRNA-145 was most abundant in NRVM.
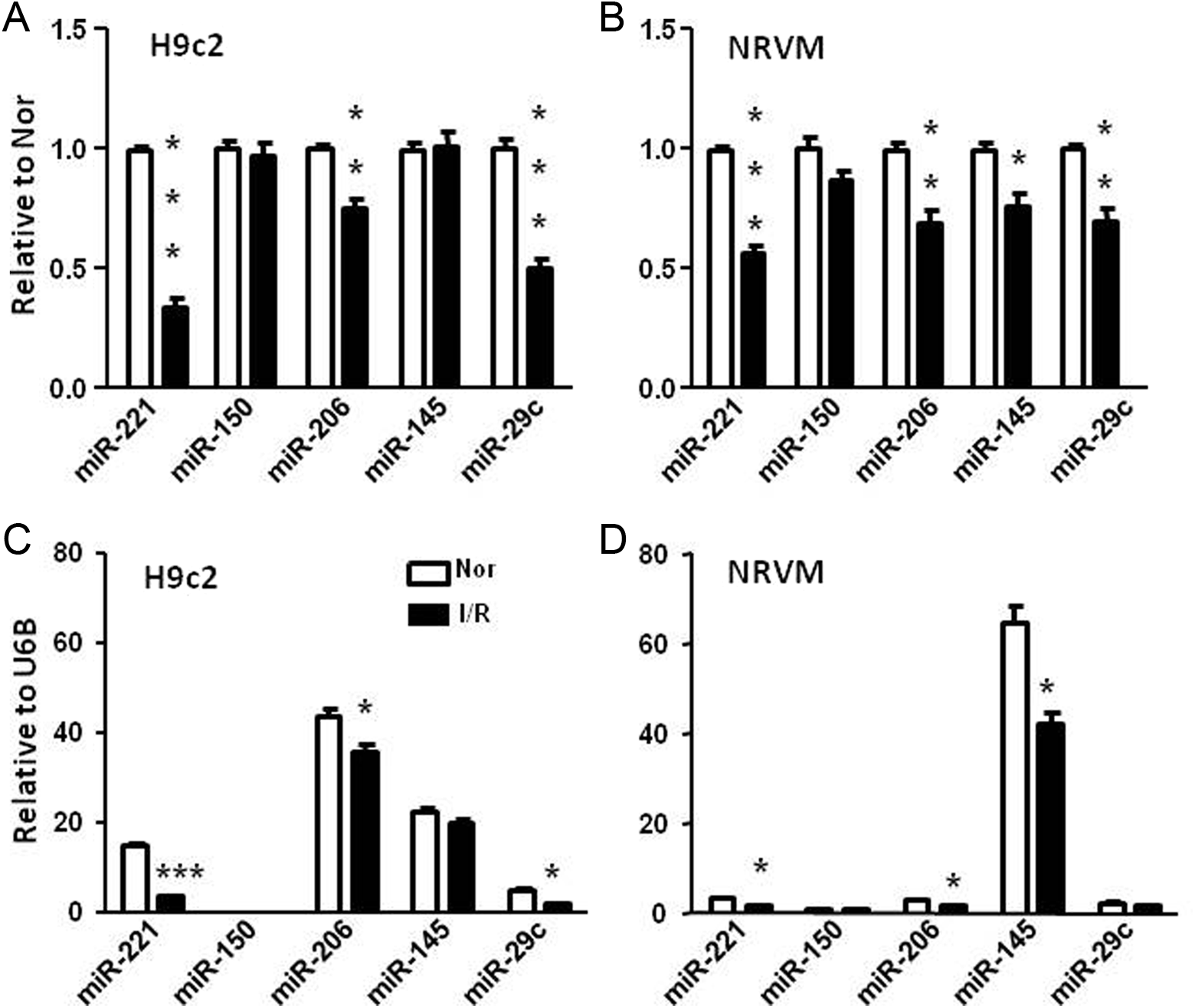
Dysregulation of microRNAs (miRNAs) induced by I/R in H9c2 and neonatal rat ventricular myocyte (NRVM). All miRNA expressions were normalized to normoxia control (Nor) of H9c2 (A) and NRVM (B) and normalized to U6B expression (C) and (D). *P < .05, **P < .01, and ***P < .001 versus Nor by 2-tail unpaired t test. White bars indicate normoxia control and black bars indicate ischemia/reperfusion (I/R)-control. N = 3 in triplicate.
Assessment of RISC-Loaded miRNA Upregulation Following Mimic Transfection
MicroRNA changes following miRNA transfection were detected by stem-loop qPCR in cell extract. Notably, increases in miRNA induced by 25 nmol/L mimic transfection were 1000-fold or more (Figure 6A). Varying concentrations of mimic transfection allowed desirable fold changes to be achieved (Figure 6B). However, biological impact is determined by the regulations of functional RISC-loaded miRNAs. Therefore, Ago-2 immunoprecipitation (IP) was undertaken to measure RISC-loaded miRNAs. Figure 6D shows the Western blot results. Ago-2 was highly enriched in Ago-2–specific antibody pull down. In contrast, Ago-2 was barely detected in IgG nonspecific pull down. These results indicated that Ago-2 pull down was specific (Figure 6D). Following miRNA transfection, RISC-loaded miRNAs were upregulated by 20- to100-fold (Figure 6C) for most miRNAs. The increase in miRNA-150 mimic transfection was as high as 105- and 103-fold in total cell extract and Ago-2 IP, respectively. This was not surprising, since extremely low baseline expression was expected to associate with many-fold elevation of expression upon transfection.
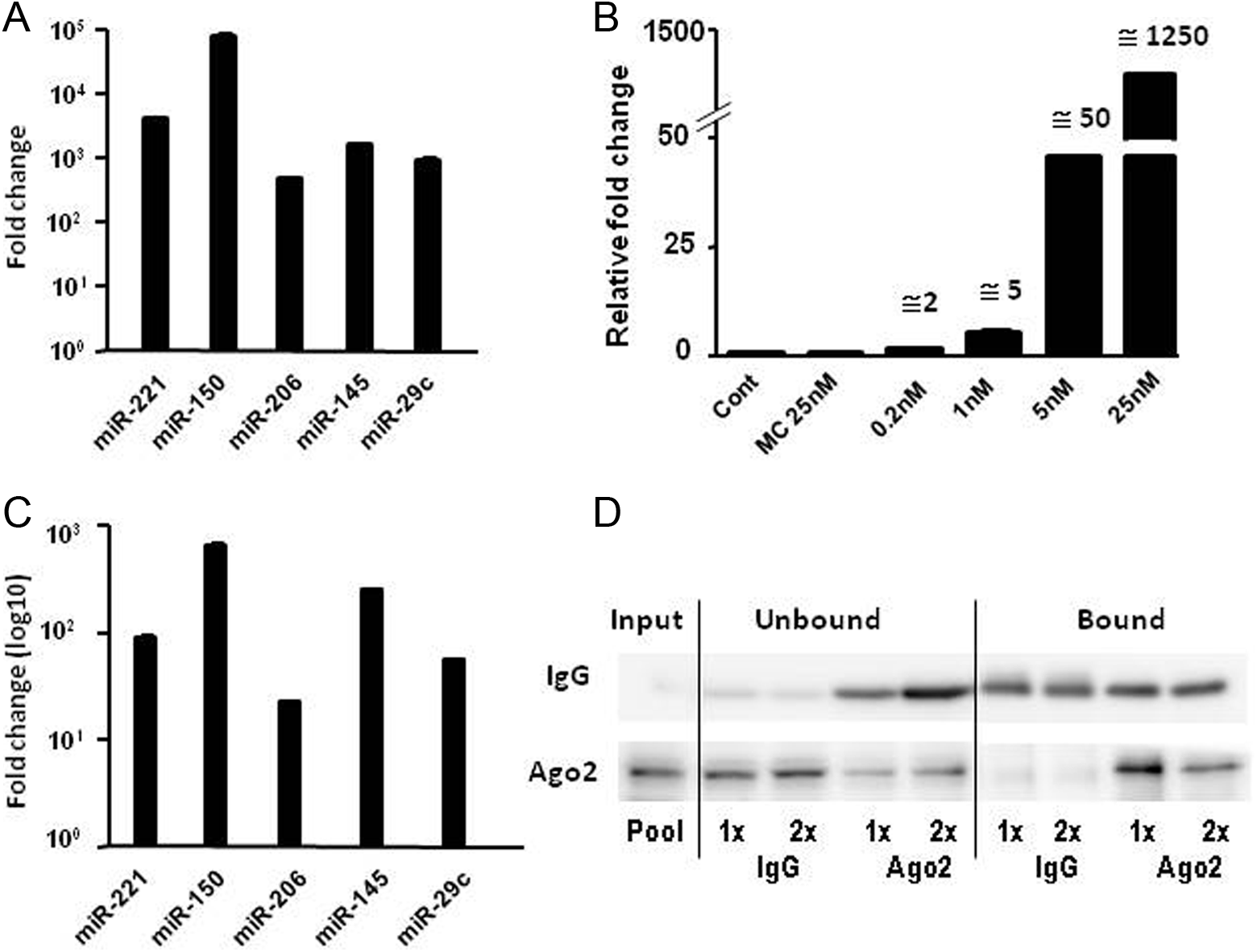
Upregulations of microRNA (miRNA) induced by miRNA mimic transfection in H9c2. A, Upregulations of miRNA after 25 nmol/L mimic transfection in total cell extract. B, Upregulation of miRNA induced by different concentrations of miRNA mimic transfection in total cell extract. C, Upregulation of miRNA after 25 nmol/L mimic transfection in Ago-2 immunoprecipitation (IP). D, Western blot showing specific Argonaute-2 (Ago-2) pull-down with IP. Immunoglobulin G (IgG) antibody was used as nonspecific pull-down control in IP.
MicroRNA-221 Mimic Transfection Reduced Hypoxia/Reoxygenation Induced-Apoptosis and Autophagy in H9c2 Cells
MicroRNA-221 was selected for exploration of possible protective mechanisms as a proof of principle. As shown in Figure 7A, I/R-induced cell death and increase in caspase 3/7 activity were significantly reversed by miR-221 mimic. LC3 I/II was measured as an indicator of autophagy. LC3-II was greatly reduced by miR-221 mimic transfection, Figure 7B.

Overexpression of microRNA-221 (miRNA-221) reduced ischemia/reperfusion (I/R)-induced apoptosis and autophagy in H9c2. A, Cell morphology, cell number count, and caspase-3/7 activity of normoxia (Nor), I/R-mimic control (MC-I/R), and I/R-miRNA-221(miR-221-I/R). B, Western blot of LC3. ***P < .001 versus normoxia control and

Quantitative polymerase chain reaction (qPCR) and Western blot measurements of miRNA-221 target genes in the regulation of apoptosis and autophagy in H9c2. A, Messenger RNA (mRNA) and (B) and (C) protein expressions in normoxia (Nor), ischemia/reperfusion (I/R)-mimic control (MC-I/R), and I/R-miRNA-221(miR-221-I/R). *P < .05, **P < .01, and ***P < .001 versus normoxia control,
MicroRNA-221 Mimic Transfection DownRegulated Multiple Apoptosis- and Autophagy-Related Targets at mRNA and Protein Level in H9c2 Cells
Using miRNA target prediction (http://www.mirbase.org linked to microrna.org, mirdb.org, Targetscan.org), a group of candidate target genes known to regulate apoptosis and autophagy was identified and selected for measurement of mRNA and protein expression (Figure 8). Intrinsic apoptosis pathway Bcl-2 family members Bak1, Bim, Bmf and Puma, PDCD10, and autophagy-regulated genes Ddit4, Tp53inp1, and p27 were upregulated by I/R. Increased mRNA expression of Bak1, Puma, Ddit4, Tp53inp1, and p27 was reversed by miRNA-221 overexpression (Figure 8A). The protein expressions of Bim, Bmf, Ddit4, Tp53inp1, and p27 were reduced. Among them, I/R-induced increases in Ddit4 and Tp53inp1 were reversed (Figure 8B and C). These data reflect the fact that a single miRNA may target multiple genes in the same signaling pathway and induce synergetic effects. These gene targets need to be validated in future studies. Especially in view of the multiple mechanisms involved in I/R injury, for example, Ca2+ overload, mitochondrial adenosine triphosphate production, protein kinases, protein ubiquitination, and so on. Additional predicted miRNA-221 gene targets are listed in Supplemental Table 1.
Discussion
We have reported an approach to screening for and selecting potential miRNA therapeutic targets in I/R injury. NaHS and UCN2 pretreatments are cardioprotective in I/R. MicroRNA array analyses indicated that these protective treatments induced similar changes in overall miRNA profiles. Approximately 20 miRNAs were commonly upregulated and 20 miRNAs downregulated by 2-fold or more. Ten of 20 upregulated miRNAs with antiapoptotic and antiautophagy potentials were selected and tested by miRNA mimic intervention. Using a miRNA functional screening approach, we demonstrated that mimics of miRNA-221, -150, and -206 are protective in both H9c2 and NRVM (and miRNA-145, 29c in H9c2 only) in in vitro models of I/R. Upregulation of RISC-loaded miRNA was confirmed by Ago-2–specific IP. Further examination of miRNA-221 overexpression demonstrated that multiple genes regulating apoptosis and autophagy were downregulated.
To reverse the disease-related change is the mainstay strategy for assessing new candidate therapies. Restoring I/R-induced miRNA changes is a common strategy for discovery of therapeutic miRNA targets for cardioprotection. 24,25 However, changes in miRNAs in disease states may be neutral, deleterious, or beneficial with respect to end-organ effects. Hence restoring disease-induced dysregulation of individual miRNAs may not always have the hoped for therapeutic effect and it may leave many potential therapeutic targets undetected. As shown in this study, miRNA-150 and -145 were not decreased by I/R. However, upregulation of these miRNAs had protective effects. These data support the necessity to explore supplementary approaches to target selection. We have selected miRNAs that change concurrently with benefit from cardioprotective interventions and this approach has yielded miRNAs with promising subsequent results on functional screening in vitro.
To establish a consistent and reproducible model is very important to the success of research work. Ischemia duration is correlated to the degree of injury. Different preparations, namely isolated heart, cultured cells, and in vivo animal models, respond to ischemia differently as do varying species of animals or cell types. Therefore, there is no fit-for-all protocol. In our study, protocols were chosen according to functional recovery in the isolated heart setting or cell death in vitro. In the isolated rat heart, 20-minutes global ischemia followed by reperfusion resulted in ∼40% ± 10% functional recovery compared with baseline. In H9c2 and NRVM cells, 15 or 6 hours of hypoxia and reoxygenation, respectively, caused ∼40% ± 20% cell number loss compared with normoxia.
How actions of cardioprotective drugs may be mediated by miRNA regulation is a promising field for research, but little is known. A good example is that propranolol, an early β-blocker (a class of drugs well proven in treatment of MI and I/R injury). It was reported that the decrease in miRNA-1 caused by propranolol is associated with cardioprotection in MI. 26 Chronic administration of propranolol reverses MI-induced dysregulation of multiple miRNAs. 27 NaHS and UCN2 are well confirmed as cardioprotective in I/R. This protective effect is mediated through the activation of cell survival signaling pathways of mTORC2 (mammalian target of Rapamycin complex 2)/Akt (v-akt murine thymoma viral oncogene homolog 1) and MAPKs (mitogen-activated protein kinases) of ERK1/2 (mitogen-activated protein kinase 1) and p38 (mitogen-activated protein kinase 14), respectively. 22,28,29 To our surprise, these 2 apparently disparate treatments induced very similar patterns of miRNA expression profiling as assessed by miRNA array. Therefore, it seems reasonable to look for potential therapeutic miRNA targets through miRNA changes appearing in cardioprotection. We accept that screening guided by changes induced by both pre- and posttreatment strategies are relevant. Whether these targets can be developed to useful therapeutic treatment will be determined by further studies. We are aware that posttreatment (drugs given after reperfusion) may seem more clinically relevant although pretreatment during the “door-to-needle” phase of PCI for acute MI is highly feasible. In our NaHS and UCN2 studies, pretreatment was used for cardioprotection. From functional screening experiments, 5 of 10 miRNAs were shown to have protective effects in H9c2 against I/R injury. Among them, miRNA-221, -206, and -29c were downregulated during I/R; miRNA-150 and 145 were unchanged. Three of 5 miRNAs had similar protective effects in NRVM, for example, miRNA-221, -206, and -150.
Downregulation of miRNAs appears to be more common than upregulation during ischemia and I/R (Supplemental Figure 1). Therefore, an increase in the specific miRNA in I/R may be a beneficial therapeutic approach if this “cue” is observed with administration of an effective cardioprotective agent in I/R injury. Synthetic miRNA mimics are hairpin structured oligonucleotides. One strand is identical to the miRNA of interest as the “guide strand.” The other, or “passenger” strand, is partially complementary and usually modified to increase stability and uptake. The use of miRNA mimics is a less studied area compared to the use of miRNA inhibitors. A few studies have investigated the transfection of miRNA mimics into NRVM and H9c2 across a wide range of concentrations (10 to 100 nmol/L) but the transfection-induced miRNA changes have not been reported. 14,30 –32 In this study, mimics were transfected at 25 nmol/L. With 90% or above transfection efficiency, increases in specific miRNAs were extreme high at 103- to 105-fold. With stepped concentrations of 0.2, 1, 5, and 25 nmol/L mimic transfection, increases in specific miRNA could be well controlled at 2-, 5-, 50-, and 1250-fold, respectively. In our preliminary study, 25 nmol/L mimics were necessary to achieve protective effects. We assessed the corresponding upregulation of RISC-loaded miRNA. In a previous report, 60 nmol/L mimic transfection induced upregulation of the specific miRNA by 1176-fold, with concurrent RISC-loaded miRNA increase of only 92-fold. 33 According to our results from IP of Ago-2, increases in the RISC-loaded miRNA were between 20-fold and 100-fold, in accord with the previously reported results. 33 One exception was miRNA-150, which induced a close to 1000-fold increase reflecting extremely low baseline expression levels.
Our data indicated that upregulation of miRNAs-221, -150, and -206 are protective in an in vitro model of I/R injury. Although the roles of these 3 miRNAs in the heart are poorly understood, some previous findings support their therapeutic potential. A miRNA cocktail of miRNA-221, -21, and -24 improves cardiac function following MI. 34 Upregulation of miRNA-221 in cancer cells has antiapoptotic effects. 35 Upregulation of miRNA-221 (-222 cluster) has proproliferative, promigration, and antiapoptotic effects in vascular smooth muscle cells. 36 MicroRNA-150 has been reported to be downregulated in the failing heart, 24 may play a protective role in the heart through the regulation of inflammation. 37 Low levels of miRNA-150 may be associated with cardiac rupture after MI 38 and lower survival. 39 Not much is known about the possible role(s) of miRNA-206 in the heart. It is believed to be a muscle enriched miRNA and might play a role in cardiac hypertrophy. 40 Overexpression may reduce the effects of inflammatory cytokines. 41 These results point to plausible cardioprotective mechanisms for these miRNAs. From target prediction, multiple genes with regulatory effects upon apoptotic and autophagy pathways are listed. As a proof-of-principle study, we investigated the protective mechanisms of miRNA-221. Overexpression of microRNA-221 did decrease I/R-induced apoptosis as indicated by lower caspase-3/7 activity. It inhibited I/R-induced autophagy as indicated by decreased LC3-II. Using target prediction and qPCR assessment, we demonstrated that miRNA-221 downregulated expression of BAK1, Puma, Ddit4, Tp53inp1, and p27 through which I/R-induced apoptosis and autophagy might be inhibited. This change was further confirmed by Western blots demonstrating downregulation of the proautophagy targets Ddit4, Tp53inp1, and p27. Changes in proapoptotic gene expression were not fully paralleled by changes in protein. BAK1 and Puma were downregulated at mRNA level vs Bim and Bmf, which were downregulated at protein level. This remains consistent with miRNA regulation of protein expression through accelerated mRNA degradation and impaired translation. 42
Limitations of our study include the cell types investigated. Both H9c2, a rat cardiac myoblast, and NRVM are well characterized, low cost, and easy to maintain, therefore they are frequently used cell models in cardiac research including miRNA studies. 43 –45 However, one is not a cardiac myocyte and the other neonatal rather than adult.
In conclusion, the strategy of selecting miRNA target through upregulations of miRNAs paralleling cardioprotection induced by 2 biological agents, coupled to functional screening, successfully identified 3 miRNAs (-221, -150, and -206) of 10 to be cardioprotective in both H9c2 and NRVM models of I/R. To further validate the feasibility of this approach, miRNA-221 was selected as a proof-of-principle study. Predicted apoptosis and autophagy-related miRNA-221 targets were measured by qPCR and Western blot. MicroRNA-221 significantly reduced the mRNA and protein expressions of Ddit4, Tp53inp1, and p27, but differently reduced BAK1 and Puma at mRNA level but Bim and Bmf at protein level. This report demonstrates that mimicking miRNA changes caused by cardioprotective agents in combination with functional screening enables investigators to efficiently identify novel miRNAs with therapeutic potential in cardiac I/R.
Footnotes
Author Contribution
Yue Zhou, and Qiying Chen contributed equally to this work. Zhou, Y contributed to design, contributed to acquisition, analysis, and interpretation; gave final approval; and agreed to be accountable for all aspects of work ensuring integrity and accuracy; Chen, Q contributed to acquisition, analysis, and interpretation; gave final approval; and agreed to be accountable for all aspects of work ensuring integrity and accuracy; Sheng Lew, K contributed to acquisition and gave final approval; Mark Richards, A critically revised the manuscript and gave final approval; Wang, P contributed to conception and design; contributed to interpretation; drafted the manuscript; critically revised the manuscript; gave final approval; and agreed to be accountable for all aspects of work ensuring integrity and accuracy.
Author’s Note
This work was done at Cardiovascular Research Institute; Department of Medicine, Yong Loo Lin School of Medicine, National University of Singapore, Singapore.
Acknowledgments
We thank Dr. Leelee Wong for the assistance in miRNA array data analysis (Cardiovascular Research Institute, NUS, Singapore). We thank Drs Armugam Arunmozhiarasi and Jeyaseelan Kandiah for the assistance in miRNA array experiment (Department of Biochemistry, NUS, Singapore). This work was supported by Cardiovascular Research Institute Start-up Fund (National University of Singapore, Singapore) and National Medical Research Council Center Grant (Ministry of Health, Singapore).
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by Cardiovascular Research Institute Start-up Fund (National University of Singapore, Singapore) and National Medical Research Council Center Grant (Ministry of Health, Singapore).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
