Abstract
Introduction
Ovarian cancer (OC) remains one of the most lethal gynecological malignancies, primarily due to challenges in early detection and the consequent poor prognosis. Genetic predisposition plays a critical role in OC development, with the Glutathione Peroxidase 1 (GPX1) gene receiving increasing attention. The GPX1 gene polymorphism rs1050450 has been implicated in various cancers, potentially through its impact on oxidative stress mechanisms.
Objective
This study aimed to investigate the association between the GPX1 (rs1050450) polymorphism and the risk of developing OC in a Turkish population.
Methods
A retrospective case-control study was conducted involving 90 women diagnosed with OC and 90 healthy controls. Genotyping of the GPX1 (rs1050450) polymorphism was performed using real-time PCR (RT-PCR). Statistical analyses were conducted using the SPSS software, with chi-square and t-tests applied where appropriate.
Results
The CC genotype of the GPX1 (rs1050450) polymorphism was significantly associated with a reduced risk of OC (P = 0.002; OR = 0.304; 95% CI = 0.161-0.577), whereas the TT genotype was linked to an increased risk, demonstrating a threefold elevation in susceptibility (P = 0.036; OR = 3.308; 95% CI = 1.024-10.682). Additionally, the T allele was associated with an approximately threefold increased risk of developing OC (P = 0.0002).
Conclusion
These findings suggest that the GPX1 (rs1050450) polymorphism may play a significant role in OC susceptibility, with the CC genotype offering potential protective effects and the TT genotype indicating increased risk. This genetic variant may serve as a useful marker for assessing OC risk; however, further studies involving larger and more diverse populations are needed to validate these results.
Plain Language Summary
Ovarian cancer (OC) is one of the deadliest gynecological cancers, largely due to its late diagnosis and poor prognosis. Several factors contribute to the development of OC, including genetic susceptibility. Glutathione Peroxidase 1 (GPX1) is an important antioxidant enzyme that protects cells from oxidative stress, which can damage DNA and contribute to cancer development. However, the role of GPX1 in ovarian cancer remains unclear, as it may either suppress or promote tumor growth depending on the context. A specific genetic variation in the GPX1 gene, known as rs1050450, has been linked to the risk of several types of cancer. In this study, we investigated whether this genetic variant is associated with an increased or decreased risk of ovarian cancer in Turkish women. Our findings suggest that women who carry the CC genotype of the GPX1 gene may have a lower risk of developing ovarian cancer, while those with the TT genotype may face a higher risk. Understanding the impact of genetic factors like GPX1 on ovarian cancer could help improve early diagnosis and guide personalized treatment strategies. However, additional research involving larger and more diverse populations is necessary to confirm these results and explore their clinical relevance.
Keywords
Introduction and Purpose
Ovarian cancer (OC) is one of the most lethal gynecologic malignancies, 1 characterized by late-stage diagnosis and poor prognosis. 2 Despite advances in treatment, the overall survival rate for OC remains low due to the high recurrence rate and resistance to chemotherapy. 3 Understanding the genetic factors involved in OC can provide insights into its pathogenesis and lead to the development of targeted therapies.4,5
Glutathione Peroxidase 1 (GPX1) is an antioxidant enzyme that plays a crucial role in protecting cells from oxidative damage by reducing hydrogen peroxide to water. The GPX1 gene, located on chromosome 3p21.3, encodes this enzyme. 6 Polymorphisms in the GPX1 gene have been implicated in various cancers. 7 However, the debate regarding the link between GPX1 variations and cancer vulnerability persists across research studies. It is essential to highlight that higher levels of GPX1 have been consistently reported in different cancer types. 8 Conversely, independent research has revealed a relationship between decreasing circulating GPX1 levels and an elevated susceptibility to cancer, 9 Thus, its function can be contradictory, operating as both a promoter and a suppressor of tumors, dependent on the cancer type. GPX1 can regulate cell proliferation, cell apoptosis, cell migration, and the immunological response to cancer; Additionally, it is involved in the response to chemotherapy and radiotherapy. 10 The most studied polymorphism, Pro198Leu (rs1050450), results in an amino acid change from proline (CCC) to leucine (CTC) that affects the enzyme’s activity and potentially influences cancer risk. 11
Several studies recently established a bond between the existence of GPX1 (rs1050450) gene polymorphisms and the development of malignancies, such as breast, 12 prostate, 13 bladder, 14 lungs, 14 leukemia, 15 and colorectal cancers. 16 However, data on its role in OC are limited and inconclusive. Therefore, further research is required to clarify the connection between GPX1 polymorphisms and OC risk.
This study aims to investigate the association between GPX1 Pro198Leu polymorphism and the risk of ovarian cancer in a population of Turkish women. We hypothesize that the Pro198Leu polymorphism is significantly associated with OC susceptibility and could serve as a potential biomarker for early detection and personalized treatment strategies.
Understanding the genetic factors contributing to OC can improve risk assessment, early detection, and the development of targeted therapies. This study’s findings could contribute to the growing body of knowledge about the genetic underpinnings of OC and support the potential use of GPX1 polymorphisms as biomarkers in clinical practice.
Materials and Methods
Study Population
This retrospective, hospital-based case-control study was conducted at Yeditepe University Hospital in Istanbul, Turkey. The study included 90 women with histologically confirmed ovarian cancer (OC), all of whom met the following inclusion criteria: Turkish ancestry, age 18 years or older, and no previous history of malignancy. The control group consisted of 90 healthy women attending routine health screenings at the same institution during the study period. Controls were matched geographically to the cases and confirmed to be cancer-free through clinical evaluation.
Sample size calculations were performed using Slovin’s, Krejcie’s, and Morgan’s methods to ensure adequate statistical power. To minimize selection bias, cases and controls were recruited from the same geographic region and medical facility. The reporting of this study conforms to STROBE guidelines. 17
All participants completed a structured questionnaire to provide demographic information, cancer history, menopausal status, and reproductive history. Additionally, 5 mL of venous blood was collected from each participant in EDTA tubes. Samples were stored at 4°C and processed for DNA extraction within 3 to 7 days.
Verbal informed consent was obtained from all participants prior to enrollment, and permission was granted for the use of their data and biological samples. The study protocol was approved by the Clinical Research Ethics Committee (KAEK) of Yeditepe University (Istanbul, Turkey) under application number 2387. Ethical approval was granted on April 13, 2022 (KAEK Decision No: 1592). The committee was chaired by Prof. Dr Turgay Çelik, with Prof Dr Turgay İsbir serving as the project coordinator and Prof Dr Rukset Attar as the responsible investigator.
Genomic DNA Isolation and Purification
Peripheral blood samples were collected in EDTA tubes and stored at +4°C until processing. Genomic DNA was extracted using the iPrep PureLink gDNA Blood Isolation Reagent and the iPrep PureLink DNA Isolation Robot (Invitrogen, Thermo Fisher Scientific Inc.). The concentration and purity of the extracted DNA were assessed with a NanoDrop 2000 spectrophotometer (Thermo Fisher Scientific Inc.), and samples with OD260/OD280 ratios between 1.7 and 1.9 were considered suitable for analysis.
Genotyping of GPX1 (rs1050450) Polymorphism
Genotyping of the GPX1 (rs1050450) polymorphism was performed using a TaqMan® SNP Genotyping Assay (Applied Biosystems) on a 7500 Real-Time PCR system. The assay included sequence-specific primers (forward: AGGAGGGGCGCCCTAGGCACAGCTG; reverse: GCCCTTGAGACAGCAGGGCTTCGAT) and dual-labeled TaqMan® MGB probes (VIC® dye for allele C and FAM™ dye for allele T) to detect allelic variants.
A Composition of Real-Time PCR Reaction Mixtures.

Statistical Analysis
All statistical analyses were performed using SPSS version 28 (IBM Corp., Armonk, NY, USA). Descriptive statistics were calculated for demographic and clinical variables. Differences in genotype frequencies between the OC and control groups were evaluated using Pearson’s chi-square test. A P-value of less than 0.05 was considered statistically significant.
For continuous variables, such as age, independent t-tests were used to compare group means, following verification of normality and homogeneity of variance with Levene’s test. Logistic regression analysis was employed to assess the association between GPX1 (rs1050450) genotypes and ovarian cancer risk, with adjustments for potential confounders.
Statistical methods were selected based on the nature of the data and the study objectives. Results are reported as odds ratios (OR) with 95% confidence intervals (CI) for all associations.
Results
Study Group and Recruitment
Participants were recruited from the Gynecology Clinic of Yeditepe University Hospital. The patient group consisted of 90 women with histologically confirmed ovarian cancer, selectively recruited based on predefined criteria (Turkish heritage, age ≥18 years, geographic origin within Turkey, and no prior malignancies).in addition to 90 Controls were selected from women visiting the hospital for routine gynecologic evaluations as part of their regular checkups. Both groups included individuals of Turkish heritage (Caucasian) aged 18 years and older. Patients were rigorously selected to ensure diagnostic confirmation and demographic homogeneity, while controls represented a convenience sample from the same clinical setting. All eligible participants meeting inclusion criteria (cases: confirmed ovarian cancer; controls: cancer-free status) were retained in the final analysis, with no exclusions due to strict adherence to eligibility requirements (Figure 1). Flowchart of Participant’s Enrollment.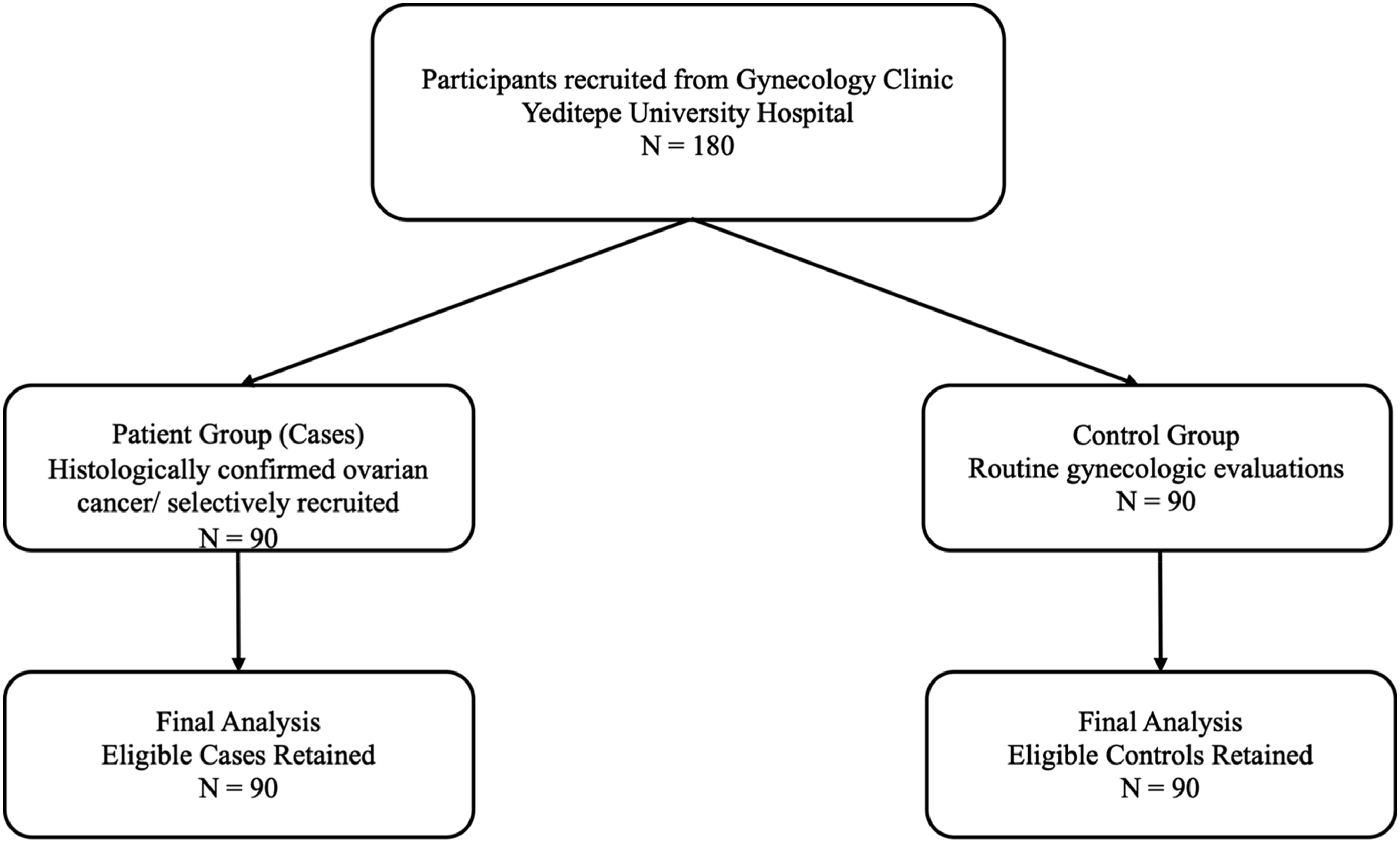
Evaluation, Assessment, and Interpretation of Real-Time PCR Findings
Our study used the 7500 Fast-Real Time PCR Instrument to assess allelic discrimination automatically. The instrument’s software translated the fluorescence irradiation readings using the dyes found in the probes. The FAM dye displayed a blue coloring, whereas the VIC dye had a green tint. The FAM and VIC dyes were compared to the ROX dye as a reference color. However, it is essential to note that discrimination between alleles could not appear for particular samples. In such cases, the Thermo Fisher Cloud platform was utilized to determine the undetermined samples. Our experiment was conducted multiple times for patients and control samples. Figure 2 shows some of the analyses of allelic discrimination through the examination and interpretation of radiance curves. Allelic Discrimination Plot of GPX1 Genotypes.
Demographic Features Analysis
Demographic data analysis (shown in Figure 3) revealed a highly significant P-value (0.0002) for the distribution of alcohol. The OC group and the control group showed no statistically significant differences in the mean age (P = 0.582) or cancer history (P = 0.412). The OC group exhibited a greater prevalence of postmenopausal women (79.5%) than the control group (17.4%). Moreover, the control group conveyed a greater percentage of individuals with one or fewer births (56.5%) in contrast to the OC group (27.3%), as well as a higher proportion of participants with one or fewer pregnancies (56.5%) compared to the OC group (22.7%). The data indicate that higher parity and pregnancy rates may provide a preventive benefit against ovarian cancer. Clinical Features and Histopathological Parameters for OC Patients and Control group. *(S) = Significantly Different (P < 0.05), NS = Non-Significant (P > 0.05).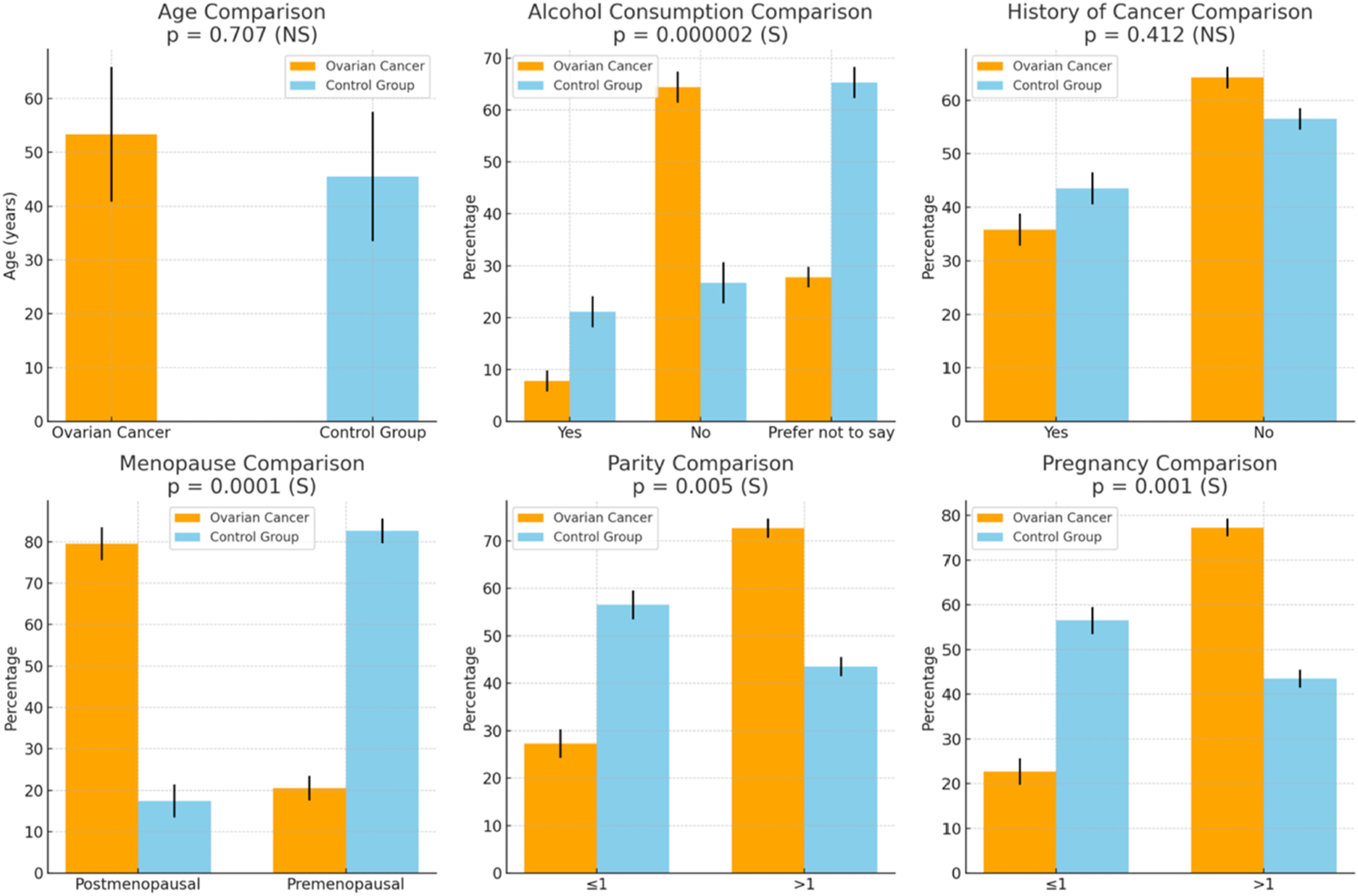
Comprehensive Overview: Clinical Features and Histopathological Parameters of Ovarian Cancer Patients.
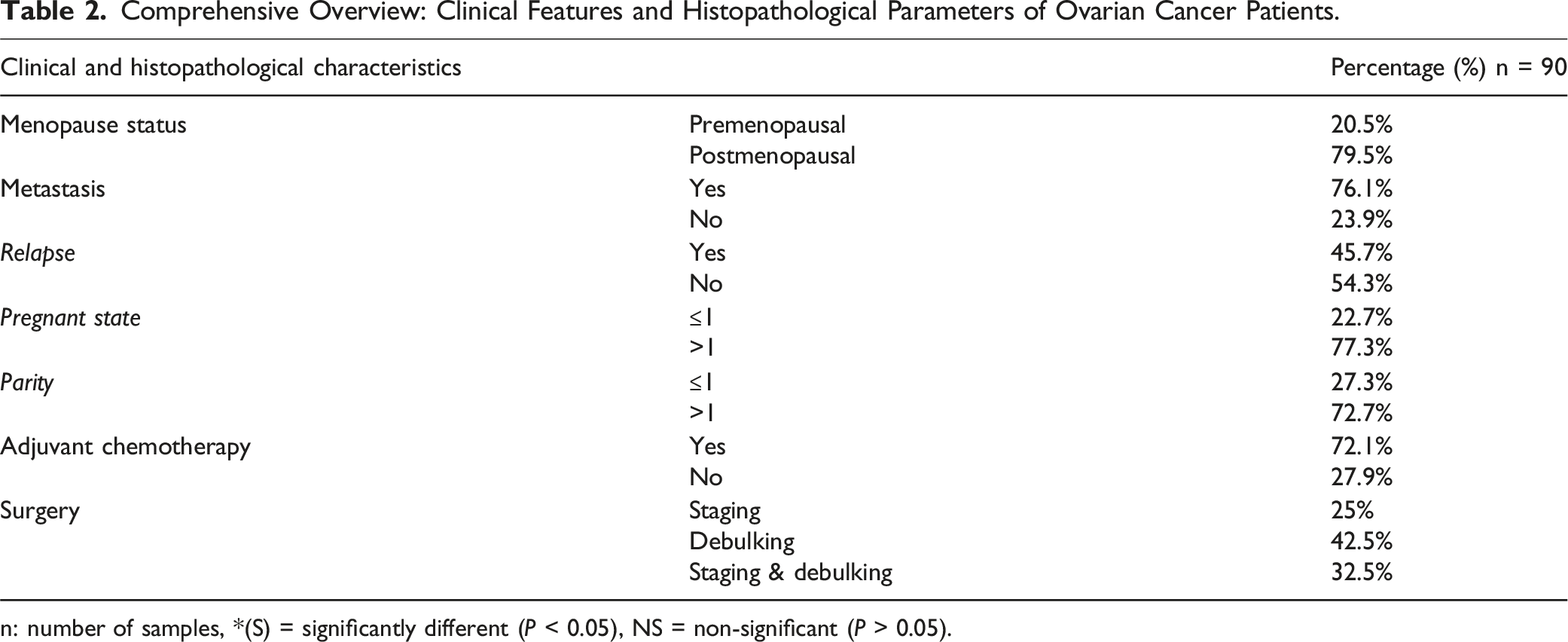
n: number of samples, *(S) = significantly different (P < 0.05), NS = non-significant (P > 0.05).
The Proportional Breakdowns of Stages and Cell Types Within the Ovarian Cancer’s Patients.
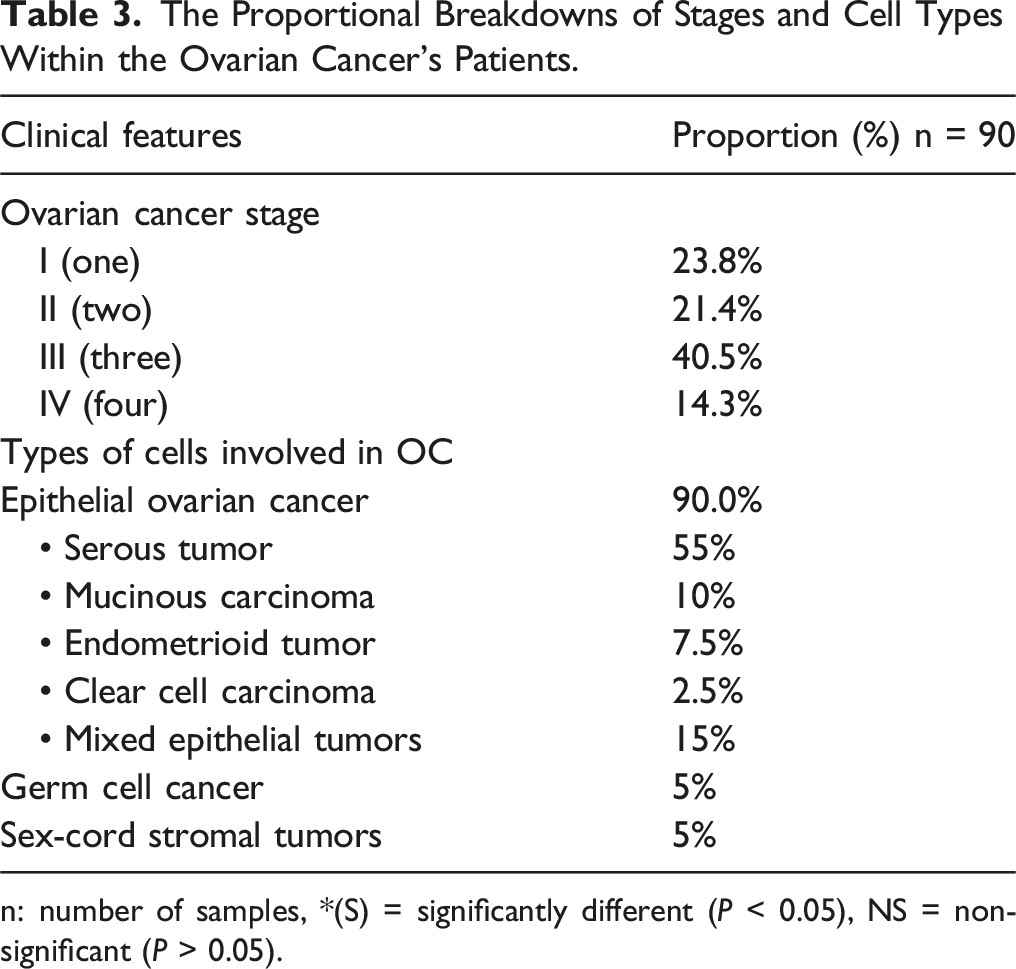
n: number of samples, *(S) = significantly different (P < 0.05), NS = non-significant (P > 0.05).
Association Between GPX1 Genotypes and Ovarian Cancer Risk
The Genotypes Frequencies of GPX1 Gene in All Study Groups.

Logistic Regression Analysis of GPX1 Genotypes.

(B, B coefficient; S.E, standard error; OR, odd ratio; CI, confidence interval).
Discussion
Gynecological malignancies, involving ovarian, cervical, and uterine cancers of the female reproductive system, represent a considerable worldwide health burden. 18 Among these, ovarian cancer appears to be the most fatal, contributing to a significant proportion of gynecological cancer-related deaths globally. 19 In 2020, ovarian cancer was responsible for nearly 207 252 deaths globally, underscoring the essential need for improved understanding and treatment techniques.20-22 Within ovarian malignancies, epithelial ovarian cancers (EOCs) are most prevalent, constituting about 90% of cases.23,24 Often detected at an advanced stage, High-Grade Serous Ovarian Cancer (HGSOC), a prominent subtype of EOC, 25 complicates therapeutic efforts. In our study, which covered 90 patients with ovarian cancer, 90% were diagnosed with EOCs, and HGSOC accounted for 55% of these cases, underscoring its higher incidence among ovarian cancer subtypes.
Age, 26 menopausal status, 27 and family history 28 are well-established risk factors for ovarian cancer. Our study reaffirmed the significance of these factors, with the average age of patients being 53.31 years and a higher prevalence of the disease among postmenopausal women (79.5%). Additionally, 35.8% of patients reported a family history of cancer, supporting the notion that genetic predisposition plays a critical role in ovarian cancer development.
However, a particularly intriguing aspect of our study is the exploration of the GPX1 gene and its Pro198Leu polymorphism (rs1050450), which has been implicated in cancer susceptibility across various types. 9 GPX1, a member of the glutathione peroxidase family, is a crucial antioxidant enzyme responsible for detoxifying hydrogen peroxide and protecting cells from oxidative damage. 29 The Pro198Leu polymorphism, specifically, has attracted attention due to its potential role in influencing cancer risk. 9
In light of this, our research has investigated the potential function of the GPX1 (rs1050450) polymorphism in the susceptibility to ovarian cancer, and it is noteworthy that the research we conducted represents the first attempt to investigate within the Turkish population.
The results revealed a significant association between the GPX1 gene and ovarian cancer risk, with a P-value of 0.001. Notably, individuals with the CC genotype of GPX1 exhibited a reduced risk of ovarian cancer, with an odds ratio (OR) of 0.304 (95% CI 0.161-0.577). On the other hand, the TT genotype was associated with a higher risk of ovarian cancer, with an OR of 3.308 (95% CI 1.024-10.682) and a statistically significant P-value of 0.036. Furthermore, the C allele was associated with a protective effect against ovarian cancer, while the T allele significantly increased susceptibility, with an OR of 3.286, indicating that individuals with the T allele are nearly three times more likely to develop ovarian cancer compared to those without the T allele.
The association between the GPX1 polymorphism (rs1050450) and cancer risk has been extensively investigated across multiple malignancies, with prior studies demonstrating consistent trends in its role as a genetic modifier of disease susceptibility,30-32 Our findings align with this established body of evidence, further underscoring the polymorphism’s relevance beyond ovarian cancer. For instance, the TT genotype of GPX1 (rs1050450) has been robustly linked to elevated risk of bladder cancer, 11 as evidenced by a case-control study within a Turkish cohort. Kucukgergin et al reported a significant association between the TT genotype and bladder cancer risk (OR = 1.67; 95% CI = 1.17-2.40; P = 0.005), highlighting its potential as a biomarker for urothelial carcinogenesis. 33 Similarly, Chen et al demonstrated a significant association between the GPX1 polymorphism and lung cancer particularly non-small cell lung cancer (NSCL), 31 These observations suggest that the GPX1 (rs1050450) polymorphism may influence susceptibility to multiple cancer types, not just ovarian cancer.
These findings have implications that extend beyond epidemiological associations, as the GPX1 polymorphism may also influence cancer cells’ biological behavior. 34 GPX1 has been shown to be overexpressed in several malignancies, including ovarian serous cystadenocarcinoma, endometrial cancer, and melanoma, indicating its potential role in promoting tumor growth and progression. 34 However, the function of GPX1 may vary depending on the cancer type. For instance, Cullen et al observed a progressive decrease in GPX1 levels from normal pancreatic cells to chronic pancreatitis and eventually to pancreatic cancer, suggesting that GPX1 may act as a tumor suppressor in certain contexts. 35
Functional enrichment analyses have highlighted GPX1’s involvement in several critical biological processes, including the regulation of reactive oxygen species (ROS), iron metabolism (ferroptosis), cell growth signaling, glutathione metabolism, and p53-mediated metabolic pathways. The diverse roles of GPX1 underscore its complexity and the need for further research to fully understand its impact on cancer biology. 35
Despite the valuable insights provided by this study, several limitations should be acknowledged. First, the relatively small sample size of 180 participants (90 cases and 90 controls) may limit the statistical power to detect weaker associations and reduce the precision of the estimated effects. Additionally, the retrospective design of the study may introduce recall bias and restrict the ability to establish causality. Furthermore, while potential confounding factors such as age and reproductive history were considered, other unmeasured confounders—such as environmental exposures and dietary habits—may have influenced the results. Finally, the study focused exclusively on a single polymorphism (GPX1 rs1050450), and did not assess other genetic variations or gene-environment interactions that could contribute to ovarian cancer risk. Therefore, future research involving larger sample sizes from diverse populations and conducted across different research settings is necessary to validate these findings and further investigate the underlying mechanisms.
Conclusion
Our findings highlight the critical importance of exploring potential applications for the diagnosis, treatment, and management of ovarian cancer within Turkish populations. By finding specific genetic markers, this research provides the framework for enhancing early detection, guiding treatment procedures, informing treatment protocols, and facilitating more personalized management strategies for ovarian cancer patients in this demographic area. Our analysis reveals a significant association between the GPX1 Pro198Leu polymorphism and ovarian cancer risk, with the TT genotype connected to heightened susceptibility and the CC genotype displaying a protective effect. These findings point out the critical significance of genetic variables in ovarian cancer and provide a foundation for further study on the relevance of oxidative stress to cancer development. This study underlines the critical need for larger, broader cohort studies to validate these results and expand our understanding of ovarian cancer pathophysiology and etiology. By converting our findings into practical therapeutic insights, we strive to bridge the gap between research and clinical application, ultimately enhancing care and support for individuals affected by ovarian cancer.
Footnotes
Acknowledgments
I am deeply grateful to everyone who contributed to the success of this project. First and foremost, I extend my sincere thanks to my co-authors for their invaluable insights, teamwork, and dedication throughout the research process. Their expertise, ideas, and enthusiasm have greatly enriched the quality and impact of this work. I also wish to express my appreciation to Yeditepe University for providing the essential resources, facilities, and support that enabled this research. The university’s infrastructure and academic environment were instrumental in shaping and executing this study. I am particularly thankful to those who offered guidance, insights, and constructive feedback during the preparation and revision of this manuscript. Their thoughtful suggestions have greatly strengthened this research. Additionally, I acknowledge the support of artificial intelligence tools used to refine and enhance the writing at various stages of this project.
Statements and Declarations
Author Contributions
Rukset Attar was responsible for the recruitment of study participants, clinical investigations, and data collection. Assoc. Prof. Seda and Selvi Duman Bakırezer contributed to the research’s design and implementation. Sara Yaser Barham played a key role in drafting and revising the manuscript, ensuring clarity and coherence in the presentation of findings. Baker was involved in the research’s design and execution, conducting experiments and performing statistical analyses. All authors have reviewed and approved the final version of the manuscript.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
Conflicting Interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Data Availability Statement
The data generated in the present study may be requested from the corresponding author or Assoc. Prof. Seda Yılmaz.
