Abstract
In recent years, the relationship between microbes and tumors has led to a new wave of scholarly pursuits. Due to the growing awareness of the importance of microbiota, including those within tumors, for cancer onset, progression, metastasis, and treatment, researchers have come to understand that microbiota and the tumor microenvironment together form a dynamic and complex ecosystem. Liquid biopsy technology, a non-invasive and easily repeatable method for sample collection, combined with emerging multi-omics techniques, allows for a more comprehensive and in-depth exploration of microbial signals and characteristics in bodily fluids. Microbial biomarkers hold immense potential in the early diagnosis, treatment stratification, and prognosis prediction of cancer. In this review, we describe the significant potential of microbial biomarkers in liquid biopsy for clinical applications in cancer, including early diagnosis, predicting treatment responses, and prognosis. Moreover, we discuss current limitations and potential solutions related to microbial biomarkers. This review aims to provide an overview and future directions of microbial biomarkers in liquid biopsy for cancer clinical practice.
Introduction
Recently, the significance of the microbiota in cancer has gained formal recognition, revealing substantial potential in cancer prevention, diagnosis, and treatment. 1 There are 10 to 100 trillion symbiotic microbial cells in each person’s microbiota, and the microbiota is mainly composed of bacteria, archaea, fungi, viruses, and protists, whose genes make up the human microbiome. 2 Microbes are present in various parts of the human body, including the gut, oral cavity, and lungs, and recent discoveries have also revealed the presence of intratumoral microbes. 3 Consequently, the human microbiome constitutes a dynamic and complex ecosystem through co-evolution with these microbes. The microbiota is believed to exert influence on the development, 4 metastasis, 5 and treatment response of various cancers, including colorectal, liver, lung, and brain tumors.6-11 Current research has discovered that certain microbes can impact cancer through multiple mechanisms, which include influencing immune responses,12-15 causing DNA damage, 16 participating in inflammatory reactions, 17 and affecting cell proliferation. 18 Importantly, a discernible disparity in microbiota constitution and abundance from oral and fecal samples has been identified between cancer patients and healthy individuals, 19 suggesting that microbes could potentially play a role in predicting and distinguishing cancer. Emerging tumor specificity and clinical relevance of microbiota has gained more importance in the exploration of diagnostic and prognostic biomarkers and therapeutic targets, hence, the role of microbial markers in cancer is gradually gaining recognition. 20 Microbes produce a variety of substances, including DNA, RNA, metabolites, and extracellular vesicles, and these substances, along with the microbes themselves, can enter bodily fluids such as blood, saliva, gastric juice, and intestinal fluid through systemic circulation. Consequently, they can be detected by liquid biopsy technology, a method that captures microbial signals in these fluids.
Liquid biopsy, which analyzes biomarkers released in body fluids, has wide-ranging applications in cancer screening, patient stratification, and monitoring due to its relatively low invasiveness and the capacity for repeated testing over time. 21 Through the detection of genetic and epigenetic genomic alterations, the evolution of the genome/acquired drug resistance can be monitored, enabling early diagnosis and disease progression monitoring.21,22 In the dynamic realm of oncology, liquid biopsies have emerged as pivotal tools, revolutionizing our approaches to cancer detection and management. However, a lacuna persists in the articulation of the evolving role of microbial biomarkers within this domain. The intricate interplay between microbial entities and cancer—a confluence of diagnostics, therapeutics, and prognostics—remains an area for exploration, promising to imbue liquid biopsy research with newfound depth and breadth.
In this review, the roles of various microbial biomarkers in cancer diagnosis, treatment response prediction, and prognosis are summarized. Moreover, the current limitations and challenges of microbial biomarker analysis in liquid biopsy and potential solutions are presented. We aim to review and forecast the application of microbial biomarkers in liquid biopsy for clinical cancer practice, with the expectation of improving patient outcomes.
The Role of Microbiota in Carcinogenic Mechanism and as Biomarker
Carcinogenesis Mechanisms Involving Specific Microbiota.
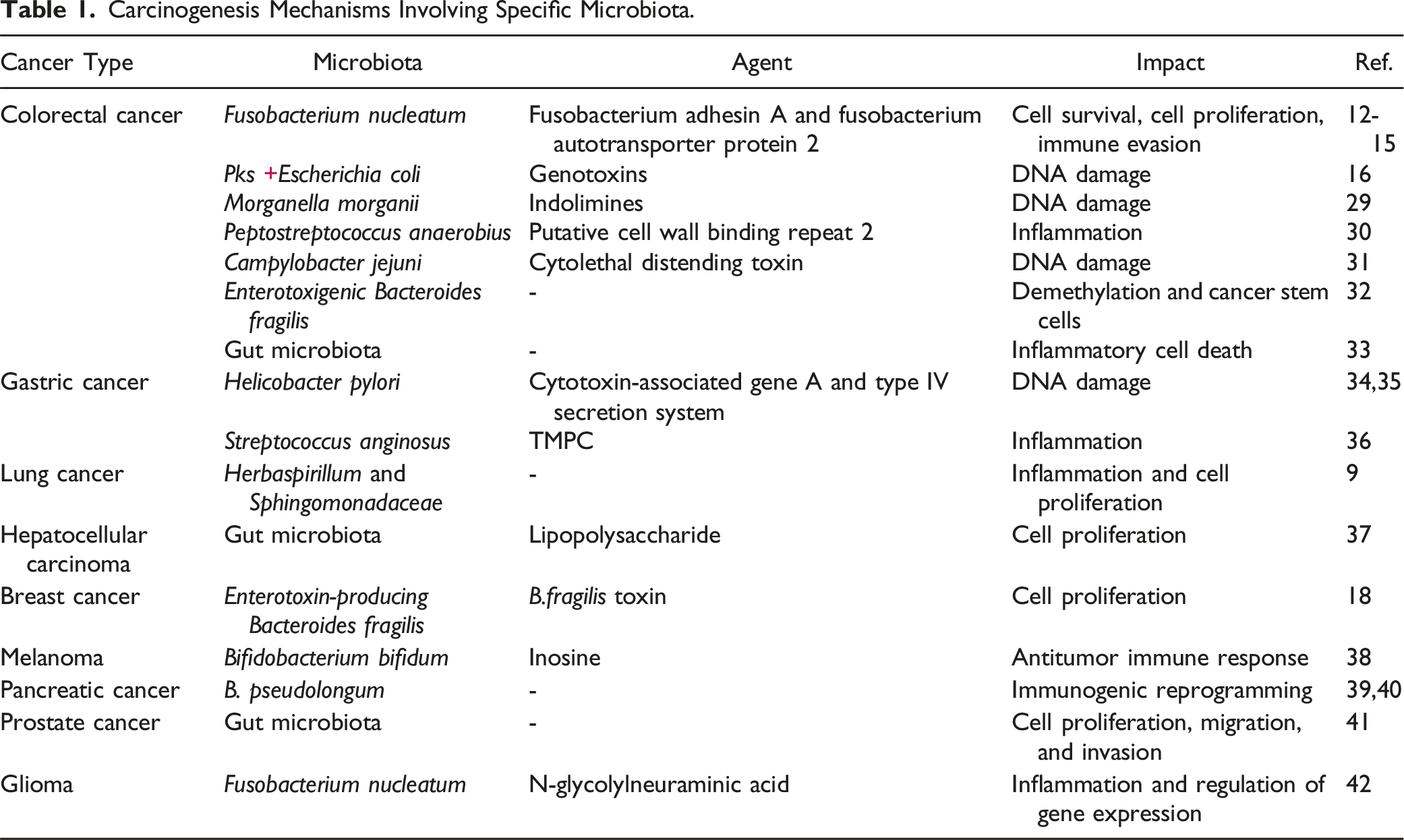
First, microbial characteristics in body fluids have been identified as predictors of cancer types, capable of distinguishing between healthy and cancerous tissues, indicating significant potential for microbial biomarkers in early cancer screening and diagnosis.
27
Second, characterizing microbial community profiles at different stages of cancer progression and in patients with varying treatment responses can aid in therapeutic stratification and in evaluating treatment responsiveness, particularly in the realm of cancer immunotherapy. Furthermore, variations in microbial diversity and composition within different tumors are indicative in survival assessments, with microbiome analysis providing prognostic insights.
28
In Figure 1, we have summarized the applications of microbial biomarkers in cancer. Liquid biopsy techniques enhance the feasibility of utilizing cancer microbial biomarkers as non-invasive, convenient, repeatable, and early-sensitive diagnostic targets. Table 2 presents current research advancements in the application of microbial biomarkers in cancer diagnosis, treatment, and prognosis. Microbiota plays a crucial role in the iterative process of cancer management, encompassing several key areas: (1) Diagnosis: the presence of microbial biomarkers in body fluids demonstrates significant potential for early cancer diagnosis. (2) Monitoring: These biomarkers can track cancer metastasis and accurately pinpoint the primary disease sites. (3) Therapeutic evaluation: dynamic changes in microbial abundance, as indicated by microbial biomarkers, can preemptively assess treatment responsiveness. (4) Prognosis: variations in the relative abundance of microbes can distinguish between low-risk and high-risk cancer scenarios. A Summary of Research on Different Kinds of Microbial Biomarkers in Cancer Diagnosis, Therapy, and Prognosis.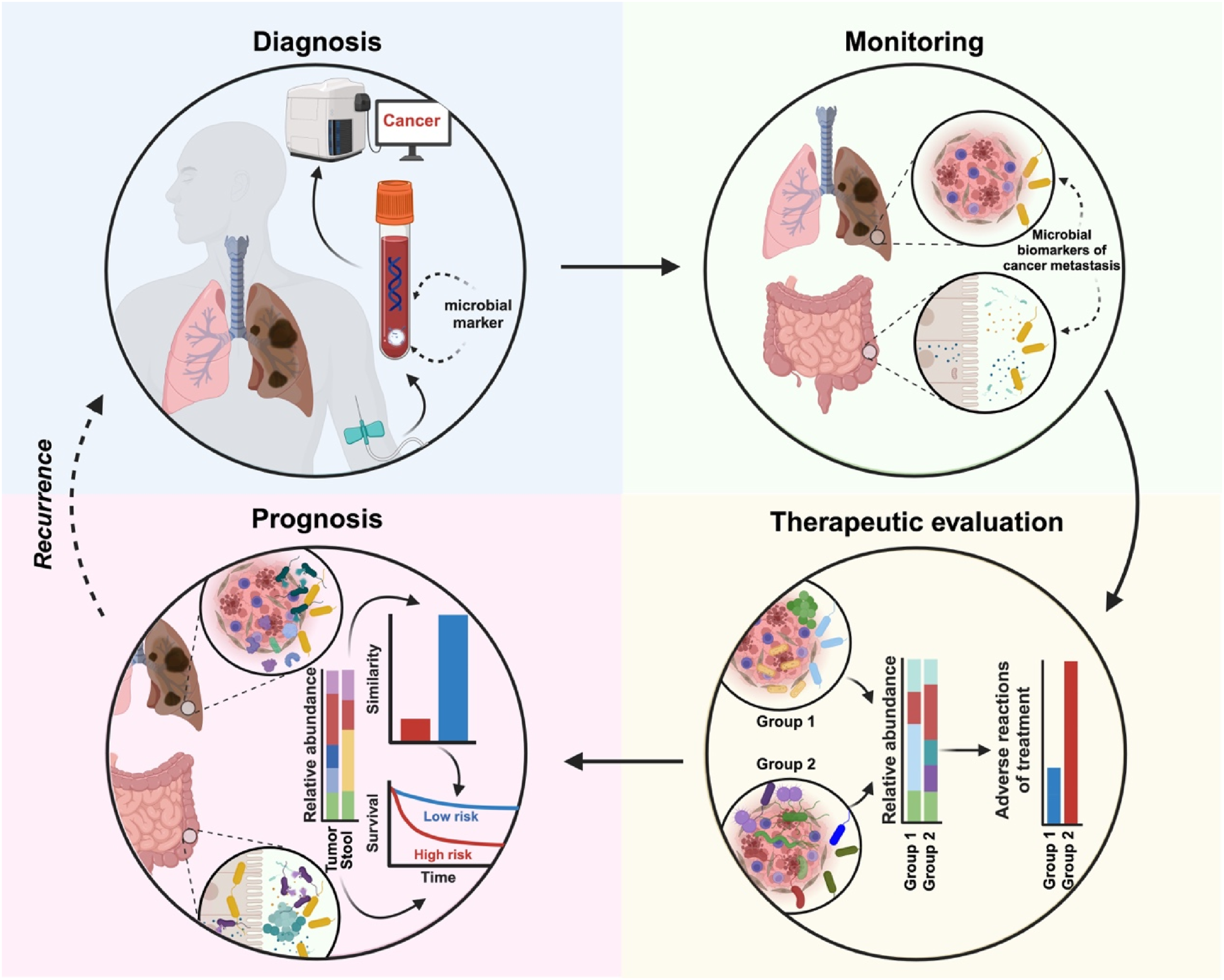

Microbial Biomarkers as Promising Innovations for Cancer Diagnosis
As early detection of cancer as possible is crucial to reducing tumor-associated mortality and morbidity. With the advent of precise detection techniques, researchers elucidate that some niches (eg, blood, urine, saliva and bronchoalveolar lavage fluid) once considered sterile have been quantified low-biomass microbes, and their predictive value has been increasingly unearthed. Thus, liquid biopsy, which identifies microbial biomarkers, could revolutionize early cancer diagnosis, as well as benefit precision oncology in addressing tumors at their curable stage. 22
Microbial Biomarkers Derived from the Blood
Blood has traditionally been regarded as a “sterile” environment, devoid of bacterial growth in non-communicable diseases. However, recent advancements in molecular techniques and bioinformatics, such as the sequencing of the 16S ribosomal RNA gene, have led to the identification and validation of a blood bacterial microbiome even in non-infectious disease states. Given economic considerations, an increasing number of researchers are utilizing this 16S rRNA technology to demonstrate associations between blood bacterial DNA signatures and various cancer types. For instance, Dong et al employed 16S rRNA sequencing targeting the V1-V2 region to reveal the presence of bloodborne microbes (e.g., Acinetobacter, Bacteroides, Sphingomonas, Comamonas, Haemophilus parainfluenzae, and Pseudomonas stutzeri), enabling the differentiation between gastric cancer patients and healthy individuals. 43 While the 16S rRNA technique has facilitated the identification of novel microbial biomarkers, it is limited to genus-level information, rendering it insufficient for tracing microbiota at the species and subspecies levels. To overcome these limitations, whole-genome sequencing (WGS) has been employed, which covers the entire genome and allows for species and subspecies identification. This advancement has propelled circulating microbial DNA (cmDNA) as a promising marker. For example, Xiao et al. demonstrated that alterations in the cmDNA of 28 bacterial species (e.g., Eubacterium rectale, Bifidobacterium adolescentis, Ruminococcus torques, Roseburia intestinalis, and Propionibacterium freudenreichii) in blood could distinguish colorectal cancer(CRC) patients from healthy controls, with an AUC of 0.971. 44 Their random forest model further revealed that these cmDNA characteristics have the potential to serve as a non-invasive biomarker for CRC. Moreover, another study employing WGS also identified bacterial cmDNA and developed a random forest model to detect early-stage lung cancer, achieving a sensitivity of 86.5% in stage I lung cancer. 45 This underscores the potential of cmDNA in early cancer diagnosis. Furthermore, a cmDNA marker-based model, including species such as Pseudomonas, Streptococcus, Staphylococcus, Bifidobacterium, and Trabulsiella, achieved an AUC of 0.879 in distinguishing hepatocellular carcinoma from control groups. 46 Despite RNA’s susceptibility to degradation, RNA sequencing can detect microbial characteristics more effectually than DNA sequencing, which is because microbes, including bacteria, possess a higher RNA-to-DNA ratio after transcription, making RNA more detectable by RNA sequencing compared to DNA sequencing. Thus, microbe-derived RNA is also a potential cancer biomarker. For example, a machine-learning model based on bacterial RNA and human features achieved an AUC of 0.9 in identifying the primary location of various cancers, with an improvement of 0.08 over models using only human features. 47 In addition to bacterial DNA and RNA, other microbial signatures, such as bacterial-associated metabolites and antibodies, could also complement traditional biomarkers (e.g., CEA and CA19-9), enhancing the performance of early cancer diagnosis.48,49 Beyond bacteria, the identification of fungi has enabled models that, after batch correction for mycobiome species, can strongly discriminate 32 cancer types from healthy individuals, demonstrating the diagnostic significance of fungi as biomarkers. 50 In summary, the single modality of microbial biomarkers derived from blood holds significant potential as an innovative tool for cancer diagnosis.
Another form of microbial signal, microbial extracellular vesicles (EVs), differs from the aforementioned biomarkers as they encapsulate a variety of soluble products,
67
such as metabolic byproducts, lipopolysaccharides, quorum sensing peptides, nucleic acids, proteins, and membrane vesicles (Figure 2). The interaction between these EVs and the human body is believed to be intricately linked to cancer mechanisms.68-70 Researchers are now exploring whether microbial EVs in the bloodstream could contribute to early cancer detection through the application of omics technologies. For example, Kim et al conducted a metagenomic analysis of serum-derived microbial EVs to develop a model that distinguishes ovarian cancer from benign ovarian tumors, highlighting the significant potential of microbial EVs in early diagnosis.
51
Their model, which incorporated serum CA-125 levels, age, and the relative abundance of Acinetobacter, achieved an AUC of 0.898. Similarly, Uemura et al. identified a BTS index based on microbial EVs (e.g., Bacteroidia, TM7-1, and Sphingomonadales) as an early screening tool for renal cell carcinoma, utilizing 16S rRNA metagenomic analysis of serum.
52
Their integrated model of microbial DNA features achieved an AUC of 0.88. Moreover, another study employing next-generation sequencing of microbial EVs in blood developed a diagnostic algorithm for breast cancer, achieving a sensitivity of 0.964, specificity of 1.000, and overall accuracy of 0.996 through the application of machine learning.
53
However, genomics alone cannot fully explain the cellular processes that occur post-transcription, such as translation and modification, leading to a loss of critical signals from bacterial EVs, particularly those involving metabolites and proteins. This limitation hinders our comprehensive understanding of the interaction between EVs and the human body. To address this issue, a further study using fecal samples combined metagenomics with metabolomics to investigate the relationship between microbial EVs and metabolomic profiles in patients with CRC compared to healthy controls, employing binary logistic regression to assess diagnostic performance.
71
Metabolomic profiling revealed alterations in the levels of seven amino acids, four carboxylic acids, and four fatty acids in the CRC group. Similarly, another integrative study utilizing (meta)genomics and proteomics conducted a comprehensive analysis of proteins within EVs from fecal samples of patients with solid tumors and healthy individuals.
72
Next-generation sequencing and machine learning were employed to evaluate microbial EVs as biomarkers for cancer diagnosis and treatment. In summary, the integration of multi-omics approaches enables the detection of multi-dimensional microbial signals within EVs, thereby enhancing diagnostic performance, although the two aforementioned multi-omics studies were conducted using fecal samples. Similarly, integrated microbial biomarkers from blood have the potential to enhance the performance of cancer detection and diagnosis, marking a critical direction for future advancements in liquid biopsy. Microbiota releases a wide array of substances in the form of extracellular vesicles, such as metabolites, lipopolysaccharides, quorum-sensing peptides, nucleic acids, proteins, and membrane vesicles. These substances interact with both the host and other microbial communities, circulating through body fluids to reach distant organs.
Microbial Biomarkers Derived from the Other Body Fluids
Beyond blood, liquid biopsy can detect microbial signatures in various bodily fluids, thereby enhancing the detection of different cancers. For instance, the vital advantage of saliva is its applicability in both primary and secondary healthcare settings, with sample collection being less dependent on the operator. Similar to blood, 16S rRNA technology also enables the tracking of microbial genera DNA in saliva. Notably, Sun et al utilized this technology to identify 25 microbial genera in saliva (e.g., Veillonella, Prevotella, Leptotrichia, Rothia, Capnocytophaga, Aggregatibacter, Campylobacter, Megasphaera, Tannerella, and Granulicatella) as promising new biomarkers for detecting gastric cancer, achieving a sensitivity of 0.9754. Additionally, Flemer et al also used 16S rRNA sequencing to establish oral microbial taxa, including Streptococcus and Prevotella spp., which demonstrated strong performance in distinguishing CRC or adenoma from healthy individuals, with a sensitivity of 53% (CRC)/67% (polyps) and specificity of 96%. 55 By combining the salivary model with fecal data, its sensitivity increased to 76% (CRC)/88% (polyps). In addition to microbial DNA in saliva, RNA signatures also show potential as new biomarkers. For example, a tool named CancerDetect for Oral & Throat cancer™ achieved an AUC of 96% in detecting oral squamous cell carcinoma and oropharyngeal squamous cell carcinoma. 56
Beyond saliva, gastric fluid and bronchoalveolar lavage fluid are also components of liquid biopsy. Metagenomic analyses of microbial EVs in gastric fluid revealed that the abundance of Streptococcus, Gemellaceae, Oribacterium, and TM7-3 increased in gastric cancer patients compared to healthy individuals. 57 Another study using 16S rDNA showed that the abundance and diversity of bacteria in bronchoalveolar lavage fluid could help assess the severity of non-small cell lung cancer. 58
Moreover, considering that microbial abundance and Shannon diversity in urine do not significantly differ between urothelial tumors, urine could serve as a diagnostic biomarker for bladder cancer. 59 Interestingly, a classifier constructed using urine-enriched bacterial genera was able to distinguish between obstructive urinary tumors and healthy patients with an accuracy of 97.96%. 60
Consistent with the above, future studies should focus on integrating multiple sample types and various microbial categories to explore ways to improve the stability and robustness of diagnostic models, while also broadening the applicability of biomarkers across different tumors and populations.
Microbial Biomarkers as Promising Innovations for Cancer Therapy
Cancer treatment methods, including chemotherapy, radiotherapy, and immunotherapy, are continually evolving, yet only a subset of patients respond to these therapies. Biomarkers that predict treatment response can assist in stratifying patients for clinical treatment, guiding the escalation or de-escalation of adjuvant therapies, increasing the likelihood of cure, and minimizing unnecessary treatment toxicity. Numerous studies have revealed the role of microbial biomarkers in predicting tumor treatment responses, including immunotherapy response in gastric cancer, 73 melanoma, 74 and hepatocellular carcinoma, 75 as well as predicting chemoradiotherapy outcomes in rectal cancer.76,77 However, these studies were based on fecal samples, which may not be strictly classified as liquid biopsy techniques.
In the field of microbial liquid biopsy, researchers have begun exploring microbial biomarkers in body fluids that could predict cancer chemotherapy response. For example, Zhou et al integrated blood microbiome data and tumor RNA sequencing in non-small cell lung cancer patients, demonstrating that cmDNA could predict patient response to chemotherapy. 61 Their model identified that patients with high scores might benefit more from five specific drugs: GSK1904529 A_1093, KRAS (G12 C) Inhibitor-12_1855, Crizotinib_1083, Gefitinib_1010, and Dabrafenib_1373. Beyond chemotherapy, potential microbial biomarkers for predicting immunotherapy response have also been revealed through liquid biopsy techniques. For instance, serum microbial EVs have been shown to predict the efficacy of ICI treatment. 52 Additionally, Wu et al. were the first to combine serum and fecal samples for a joint genomic and metabolomic analysis to differentiate hepatocellular carcinoma patients who respond to immunotherapy, achieving an AUC of 0.796. 62 Interestingly, they suggested that a classifier based on serum outperformed one based on the gut microbiota.
Based on the aforementioned information, we hypothesize that combining microbial biomarkers from both feces and serum samples (or even other samples) could improve the prediction of cancer treatment outcomes. Moreover, the synergistic predictive performance of fungal and bacterial biomarkers might surpass that of single-microbe predictions. 78 Additionally, since intratumoral microbes are closely related to the tumor microenvironment, 28 further research is needed to explore how microbial features in body fluids can be used to track intratumoral microbes, thereby predicting treatment response.
Microbial Biomarkers as Promising Innovations for Cancer Prognosis
Beyond diagnosis and therapy, microbial prognostic markers also hold significant clinical importance. Prognosis can be categorized into two types: survival prediction and risk prediction for recurrence or metastasis. The former is used to assess a patient’s overall prognosis, enabling more precise treatment stratification and facilitating shared decision-making among doctors, patients, and their families. Given that many cancers are prone to post-surgical recurrence, leading to persistently high mortality rates, the latter is primarily used to predict the likelihood of post-surgical recurrence or metastasis, which drives further preventive treatment or helps avoid unnecessary overtreatment and the associated side effects.
Similar to diagnostic and therapeutic applications, microbial DNA derived from blood can be used to predict cancer patient survival. For instance, Messaritakis et al used pathologically complete response (PCR) to analyze the peripheral blood of CRC patients, demonstrating a correlation between microbial DNA fragments in the blood and patient survival. 63 They found that genomic DNA encoding 16S rRNA, the glutamine synthase gene of Bacteroides fragilis, the β-galactosidase gene of Escherichia coli, and 5.8S rRNA of Candida albicans were significantly associated with shorter survival. Additionally, a prognostic model for non-small cell lung cancer, based on circulating microbial signatures, included 14 microbes (e.g., Candidatus Babela, Methanotorris, Anaeromusa, Hirschia, Parascardovia, and Agreia were associated with poor prognosis, while Kozakia, Andromedalikevirus, Natronococcus, Demequina, Desulfuromonas, Blastococcus, Anaerobacillus, and Ewingella were associated with better prognosis). 61 This model achieved AUCs of 0.890, 0.920, and 0.878 for predicting 1-year, 3-year, and 5-year survival, respectively.
Regarding the risk of recurrence or metastasis, cmDNA in blood, tracked using WGS, also offers potential prognostic value. For example, Chen et al developed a cmDNA model that accurately predicted post-surgical recurrence (sensitivity of 72.7%, specificity of 84.6%, AUC of 88.1%), indicating its potential as a prognostic marker. 45 Additionally, Ma et al 64 analyzed plasma from lung cancer patients using WGS and elucidated the potential of intratumoral microbial signatures and microbial cmDNA as biomarkers for lung cancer recurrence. They constructed a random forest model based on seven genera (Klebsiella, Massilia, Microbulbifer, Cutibacterium, Comamonas, Staphylococcus, and Hydrogenophilus) with an AUC of 0.857 for recurrence prediction. Beyond blood, the microbial composition of bronchoalveolar lavage fluid has also been linked to cancer recurrence, as evidenced by a study using 16S gene sequencing, which showed that microbial DNA predicted early-stage non-small cell lung cancer recurrence with an AUC of 0.77. 65 Other microbial features, such as RNA, can also enhance cancer prognosis prediction and clinical management. For instance, Hurst et al. integrated genomic and transcriptomic analyses of urine to identify four bacteria (Porphyromonas sp. nov., Varibaculum sp. nov., Peptoniphilus sp. nov., and Fenollaria sp. nov.) capable of predicting prostate cancer metastasis, demonstrating the potential of anaerobic bacteria in metastasis prediction. 66
Previous research has primarily focused on the microbiota outside the tumor; however, recent studies have discovered that intratumoral microbiota is also associated with prognosis, potentially emerging as a new class of prognostic markers. 79 Future studies should leverage multi-omics technologies to profile microbial signals in various bodily fluids, including intratumoral microbiota, to identify additional microbial prognostic markers.
Discussion
Undoubtedly, microbial biomarkers have revolutionized the technologies and methods of precision oncology. This review describes the roles of various microbial biomarkers in the early diagnosis, prediction of treatment responses, and prognosis of cancer patients. By reviewing and forecasting the current status and challenges of liquid biopsy in cancer, this review aims to promote the clinical application of microbial biomarkers and provide potential perspectives for the precise management of cancer. As a non-invasive and repeatable method, liquid biopsy helps avoid invasive examinations or overtreatment of patients, which could exacerbate their conditions and impose unnecessary burdens on the healthcare system. However, due to certain technical and research design limitations, several issues need further discussion and resolution.
Technological Concern
With the advent of 16S rRNA sequencing, this low-cost, short turnaround technique has made it possible to detect microbial features and signals in body fluids. However, it cannot fully characterize bacterial species or reveal the complete genetic content. 80 To address this, researchers can use WGS and NGS technologies to comprehensively characterize bacterial genes, which also makes the study of fungal biomarkers feasible, despite the increased cost and time associated with these new technologies. Additionally, since RNA is more effectively detected than DNA at the same depth, 47 RNA sequencing and transcriptomics can better capture dynamic labels in body fluids and facilitate the study of viral biomarkers, compared to tracking DNA alone. Recently, multi-omics technologies have been increasingly applied in liquid biopsy to identify more accurate cancer biomarkers. 81 Therefore, future studies should also consider incorporating microbe-related metabolomics and proteomics.
Data Analysis Related Concern
Apart from technology, other limitations in biomarker mining are also related to the accuracy of data processing. Gihawi et al. recently unearthed a significant oversight in a large-scale microbial study, revealing instances where human sequences were erroneously classified as bacterial due to data analysis errors. 82 This can be primarily attributed to two factors: (1) computational errors have resulted in numerous human sequences being incorrectly identified as false-positive bacterial sequences; (2) during the raw data transformation, an artificial signature was generated, which ultimately rendered the microbiome-based cancer identification classifier completely ineffective. This discovery underscores the need for meticulous computational methods and data processing in large-scale microbial biomarker research. Currently, machine learning-based computation and data processing methods have been employed to enhance the robustness and accuracy of microbial biomarker classification. For instance, MetaBoot, a machine learning framework that integrates mRMR (minimal redundancy maximal relevance) and bootstrapping in a top-down strategy, can identify non-redundant and robust microbial genome biomarker features. 83 Furthermore, A multivariate feature analysis-based machine learning model has also been applied to metabolomics biomarker mining. 84 Beyond machine learning, deep learning, which possesses end-to-end automated learning capabilities, has garnered significant attention in microbial research. A deep learning method based on Graph Embedding Deep Feedforward Network can be used for embedding network prior knowledge into deep feedforward neural networks to guide microbial feature selection. 85 Besides, the MEGMA-AggMapNet-GFI metagenomic deep learning pipeline can robustly identify research-grade and reproducible disease microbial biomarkers. 86 In summary, the integration of personalized machine learning or deep learning with advanced derivative tools enhances the accuracy, robustness, and non-redundancy of biomarker screening, holds promise for advancing the implementation of “precision” medicine and promoting the development of biomarkers for diagnosis, prognosis, and therapeutic prediction targets.
Contamination Removal Concern
In addition to minimizing computational errors, the utilization of filters or contamination removal procedures is crucial for investigating biomarkers in low-biomass samples. For instance, when Nejman et al observed that some of the 9190 bacterial species detected in various tumor or normal tissues might represent sample contamination, they employed a stringent set of six filters in 5R 16S rDNA sequencing to control potential contaminants. Filter 1 removed species found in control samples, while Filters 2 to 4 controlled for batch effects, including those introduced during DNA extraction and PCR amplification. Filters 5 and 6 addressed potential contamination before sample processing. 87 Similarly, Qiao et al utilized a contamination-removal procedure known as the RIDE checklist to filter out contaminants from low-biomass samples. Filter 1 eliminated the impact of primer sequence variants containing host-derived mitochondria or chloroplasts in taxonomic annotation. Filters 2 to 4 managed three different batch effects, including those from sequencing library batches, DNA extraction batches, and PCR amplification batches. Filters 5 and 6 were used to remove single amplicon sequence variants and to retain amplicon sequence variants present in at least two samples, respectively. 88 Eliminating multi-layered contamination is essential to offering microbiota profiling, which enables the effective identification of biomarkers.
Sample Sizes and Validation Related Concern
Moreover, many studies have limitations in their test sets, such as small sample sizes and the lack of inclusion of diverse populations, necessitating further validation in external cohorts to increase credibility. However, many studies lack validation cohorts, which may weaken their findings due to potential confounding factors such as age, tumor stage, smoking, and alcohol consumption.55,61,64 Therefore, future studies should aim to include larger initial sample sizes and diverse populations as validation cohorts. Additionally, one study attempted to overcome the limitations of validation cohorts by separating the test set from the training set from the outset and implementing strict internal validation. 51 This compromise may become a viable solution for many researchers.
Consistent Performance Related Concern
Although microbial biomarkers hold significant potential for cancer detection, their clinical application has been limited by the challenge of achieving consistent diagnostic performance across different tumors and populations. To address these challenges, multi-kingdom biomarkers may offer the stability needed across diverse clinical contexts. Firstly, multi-omics approaches enable the detection of multi-dimensional microbial signals to enhance diagnostic performance in liquid biopsy.71,72 Secondly, multi-sample approaches, such as combining serum and fecal samples, offer greater accuracy compared to single-sample or traditional clinical methods. 48 Finally, multi-kingdom microbiota analyses, which include archaea, bacteria, fungi, and viruses, can not only enhance diagnostic accuracy 50 but also ensure significant consistency across different population cohorts. 89 These strategies hold the potential for accelerating the clinical translation of microbial biomarkers from different body fluids and providing more options for early cancer screening, which enables clinical interventions at an earlier, more treatable stage of cancer.
Conclusion
In conclusion, microbiota plays a significant role in the onset, progression, and metastasis of cancer, contributing to a complex and multi-level landscape including bacteria, fungi, viruses, immune cells, and cancer cells. While the relationship between gut microbiota and cancer is increasingly elucidated, the interactions between extra-intestinal microbiota and cancers at different anatomical sites remain unclear. Although single-omics technologies effectively process large amounts of microbial data, they fall short of exploring the upstream and downstream mechanisms that link microbial biomarkers to cancer phenotypes. The future of research on tumor microbial biomarkers lies in the application of multi-omics approaches to identify more effective biomarkers and to uncover the true role of microbiota in cancer progression. Despite some unresolved mysteries, it is undeniable that microbial biomarkers hold immense potential for clinical translation. As clinical application and practice continue to advance, microbial biomarkers are poised to provide new perspectives and strategies for the diagnosis, treatment, and prognosis of cancer, ultimately improving patient outcomes.
Footnotes
Acknowledgments
We thank Lu Zhang, Shilan Chen, and Xiaoxi Cai for their support.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Ethical Statement
Data Availability Statement
Data sharing is not applicable to this article as no datasets were generated or analyzed during the current study.
