Abstract
Introduction
Contemporary efforts to predict surgical outcomes focus on the associations between traditional discrete surgical risk factors. We aimed to determine whether natural language processing (NLP) of unstructured operative notes improves the prediction of residual disease in women with advanced epithelial ovarian cancer (EOC) following cytoreductive surgery.
Methods
Electronic Health Records were queried to identify women with advanced EOC including their operative notes. The Term Frequency – Inverse Document Frequency (TF-IDF) score was used to quantify the discrimination capacity of sequences of words (n-grams) regarding the existence of residual disease. We employed the state-of-the-art RoBERTa-based classifier to process unstructured surgical notes. Discrimination was measured using standard performance metrics. An XGBoost model was then trained on the same dataset using both discrete and engineered clinical features along with the probabilities outputted by the RoBERTa classifier.
Results
The cohort consisted of 555 cases of EOC cytoreduction performed by eight surgeons between January 2014 and December 2019. Discrete word clouds weighted by n-gram TF-IDF score difference between R0 and non-R0 resection were identified. The words ‘adherent’ and ‘miliary disease’ best discriminated between the two groups. The RoBERTa model reached high evaluation metrics (AUROC .86; AUPRC .87, precision, recall, and F1 score of .77 and accuracy of .81). Equally, it outperformed models that used discrete clinical and engineered features and outplayed the performance of other state-of-the-art NLP tools. When the probabilities from the RoBERTa classifier were combined with commonly used predictors in the XGBoost model, a marginal improvement in the overall model’s performance was observed (AUROC and AUPRC of .91, with all other metrics the same).
Conclusion/Implications
We applied a sui generis approach to extract information from the abundant textual surgical data and demonstrated how it can be effectively used for classification prediction, outperforming models relying on conventional structured data. State-of-art NLP applications in biomedical texts can improve modern EOC care.
Keywords
Introduction
Contemporary efforts to predict surgical outcomes and postoperative complications usually focus on the associations between traditional surgical risk factors, including age or preoperative albumin.1,2 In addition to risk factors in discrete data fields, we now have access to abundant textual data within digital medical records. In the era of healthcare digitalization, the increasing implementation of Electronic Health Records (EHRs) at UK Hospitals has created valuable data sources for clinical and translational research. 3 Although EHRs hold structured data, a large proportion of clinical notes are in narrative text format. It is estimated that unstructured data accounts for more than 80% of currently available healthcare data. 4 Reading note text and extracting information are resource intensive. Artificial intelligence (AI) has emerged as a potential solution for harnessing these data. More specifically, Natural Language Processing (NLP) is the AI discipline that focuses on extracting information from texts by converting narrative clinical notes into a structured format. The NLP methods have been shown to achieve remarkable results in such tasks using hundreds to thousands of clinical notes. 5 Their implementation has been promoted and accelerated during the COVID era. 6 Nevertheless, clinical research has been heavily affected by the underutilization of unstructured data from EHRs. 7
Amongst the best NLP models employed to date, the Bidirectional Encoder Representations from Transformers (BERT) model was created by Google in 2018. Thanks to its architecture, it can extract information from texts by considering bidirectional contextual information. 8 BERT’s advanced information extraction capacities when combined with the fact that traditional NLP methods such as Word2vec have shown promising results in classification tasks in clinical settings and 9 can lead to the reasonable expectation that a BERT-based classification model would outperform previously used methodologies. Since 2018, several augmentations occurred, with Facebook publishing the RoBERTa language model in 2019, 10 surpassing previously set records. The RoBERTa is a late, robust, unsupervised pre-trained language model that can be used in the context of supervised tasks with outstanding results. 11
Undoubtedly, the abundance of clinical information is locked in clinical narratives. Documentation of EHRs is now developing into standard practice. For instance, surgeons spend significant time documenting and reading, amongst other tasks, narrative descriptions of operative reports and findings. 12 Developing tools to facilitate clinical review of these unstructured data can derive clinically meaningful insights for advanced epithelial ovarian cancer (EOC), a heterogeneous disease. Compared to standard approaches, they can potentiate condensation of results from several tasks and optimize analysis time. One aspiration could be the prediction of no residual disease (R0 resection) following cytoreductive surgery for EOC. Such task of confirming macroscopic clearance remains subjective, 13 to the point that photographic ‘mapping’ has been recommended that allows for an assessment of the surgical effort at primary surgery or provides a baseline for determining the effect of neo-adjuvant chemotherapy at delayed surgery. 14 As a result, most of the quantitative intraoperative assessment tools have mainly focused on their predictive value for suboptimal surgery. 15 To improve modern care, the application of NLP tools could be useful to determine whether processing of unstructured full-text documents improves the ability to forecast outcomes in clinical conditions with significant heterogeneity such as EOC.
In this work, we utilized the pre-trained RoBERTa-base language model to predict whether residual disease persists in EOC patients following their cytoreductive surgery. We hypothesized that operative notes contain valuable information associated with surgical outcomes. We aimed to develop an NLP methodology that would address the objectiveness of R0 resection through information hidden in unstructured operative notes.
Methods
Electronic Health Records (EHRs) were queried to identify women with advanced EOC who underwent cytoreductive surgery at St James’s University Hospital, Leeds, from January 2014 to December 2019. The modern EHR dataset included the following clinical features: diagnosis codes (ICD-10 codes), procedure codes (OPCS-4 codes), age at diagnosis, grade, stage, and operative notes with findings. An internally developed advanced EOC clinical database was integrated with the EHR system
16
to provide the availability of discrete and engineered data. Institutional research ethics board approval was obtained through the Leeds Teaching Hospitals Trust (MO20/133 163/18.06.20), and informed written consent was obtained. The study was added to the UMIN/CTR Trial Registry (UMIN000049480). Treatment was pre-operatively planned at the weekly central gynaecological oncology multidisciplinary team (MDT) meeting prior to patient review. The cohort details, hospital setting, indications for surgery, and surgical procedures have been described in our previous studies.13,17 Comprehensive visual assessment of all the areas of the abdomen and pelvis was routinely performed, and no visible residual disease was documented as R0 resection. The analysis took place in three steps: Firstly, words and combinations of words were analyzed based on their frequency and the case they concerned. Following the initial descriptive text analysis, the RoBERTa classifier was employed to predict case outcomes based on operative notes. Lastly, an XGBoost classification model was tasked with predicting the same outcome, this time using tabular discrete data, but also the probabilities that were derived from the RoBERTa classifier of the second step. A flowchart of our approach is shown in Figure 1. Components and the flow of the machine learning pipeline applied in our case.
Textual Descriptive Analysis
For the analysis of the text, word frequencies were calculated, and tables were created using the most common words and n-grams. N-grams are continuous word sequences of words, as they could be found in the text. The length of the n-grams can be as small as one, meaning one word, or as large as the entirety of the text. N-grams are important because they carry contextual information more than simple words do. To find the n-grams that best discriminated between the two cases, we performed an analysis based on the Term Frequency – Inverse Document Frequency (TF-IDF). The TF-IDF is a metric used to quantify n-gram importance in a particular document.
18
The score, as implied by its name, is a function of the number of times the n-gram appears in the document adjusted for the number of times it appears in the rest of the documents, as shown also in equation (1). (i) (ii) (iii) 
For each of the two possible outcomes (R0 resection vs non-R0 resection), we compiled a document consisting of the concatenation of all the individual notes that concerned this outcome. The words inside the documents were reduced to their lemmas, to make the analysis more representative of the real n-gram frequency without accounting for word conjugation. The two resulting documents were inputted into Sklearn’s TfidfVectorizer, which was tasked with assigning scores per n-gram, per document. A high TF-IDF score for an n-gram in a document signifies n-gram importance to this document. The TF-IDF n-gram scores for the documents reporting non-R0 resection were then subtracted from the TF-IDF scores for the documents reporting R0 resection. This way, the higher the absolute difference in scores, the higher the ability of the n-gram to discriminate between the two cases. Positive difference scores show that the n-gram belongs to R0 resection case notes, while negative scores show the opposite.
Natural Language Classification With RoBERTa
We utilized the pre-trained RoBERTa-base language model to extract information from the unstructured surgical notes through transfer learning. Transfer learning is the process of re-training part of a pre-trained model on specific data to fine-tune its performance for a specific task. The initial training often uses vast datasets that hold most of the information relevant to the task at hand. Re-training allows for finer details to be captured by the model. The main advantage of transfer learning is that the resulting model can reach high performance without needing to use large amounts of data. The RoBERTa-base language model is pre-trained on a large corpus of English data using the BERT-base architecture and has 125 million parameters.
10
The surgical data was used to train and test the model at a ratio of 4:1. The model was trained for 40 epochs. Discrimination was measured using the most common performance metrics for classification tasks, namely, with: (i) Accuracy = (ii) Precision = (iii) Recall = (iv) F1-score = 2* (v) Area under the receiver operating characteristic curve (AUC) (vi) Area under the precision-recall curve (AUPRC)
Understanding how the RoBERTa reached the conclusions is essential in evaluating its performance. For such an extremely complicated model, tracing the individual impact of the text tokens on the final prediction is a task requiring advanced decompositional methods. In this effort, we employed the transformers-interpret Python library 19 to explain and visualize the factors that contributed to the model’s prediction accuracy. In turn, the library employs the Captum model of interpretability and understanding library. 20 Using integrated gradients, the library evaluates the contribution of each input feature to the model output of the model. The net result is an attribution score for each token; that is positive when the token contributes towards class prediction and negative in the reverse scenario.
As a final step, we employed a surrogate model in order to augment the explainability effort. In this context, a surrogate model denotes a simpler model than the powerful original one (in our case RoBERTa), whose outputs are interpretable. The surrogate model trains on the outputs of the original and, through its interpretable coefficients, offers a way to access the decision process of the complex, original model. The surrogate model used was a simple logistic regression. The dependent variable was the RoBERTa predictions in the form of binary integer values, created by setting a threshold of .5 on the original probabilities, while the independent variables were the TF-IDF sentence vectors created through the method of TF-IDF vectorization. In this way, the aim was 2-fold: The surrogate model would serve both as an explainer and as a conceptual link between the RoBERTa outputs and the TF-IDF scores created in the descriptive analysis.
XGBoost Classification Model
Subsequently, we trained an XGBoost model 21 to predict R0 resection using a combination of structured and unstructured data sources. The independent variables included the Aletti surgical complexity score (SCS), the size of the largest bulk of the disease in centimeters, the age of the patient, the Pre-Surgery CA125, the Intraoperative Mapping of ovarian cancer (IMO) score, the operative time in minutes, the estimated blood loss (EBL), the pre-treatment CA125, the tumour grade encoded as a binary variable, the Peritoneal Carcinomatosis Index (PCI), the timing of surgery (encoded as a binary variable where primary debulking equaled 0 and interval debulking surgery equaled 1), the ANAFI score and the probabilities that the RoBERTa classifier outputted when solely tasked to predict R0 resection (real number in the interval of 0 to 1). The PCI and IMO scores were calculated at the beginning of surgery to describe the intraoperative location of the disease.22,23 The Aletti SCS was assigned to describe the surgical effort. 24 The ANAFI score is an AI-derived novel intraoperative score that assigns specific weights to the EOC dissemination patterns (ANAtomic FIngerprints). 25 It appears to be more predictive of R0 resection than the entire PCI and IMO scores whilst it retains its prognostic power. Most of these discrete and engineered data predictors have been interrogated in our previous studies.13,17,25–27
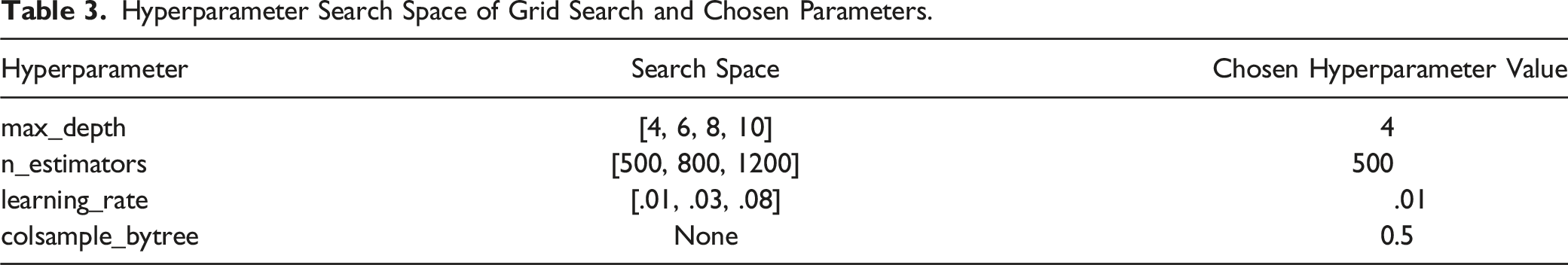
The hyperparameters of the XGBoost model were selected by using an exhaustive grid hyperparameter search. The grid search also implemented cross-validation. The hyperparameter grid is shown in Table 3. The feature importance was determined using the Shapley additive explanations (SHAP) framework to interpret the model’s predictions based on the Shapley values. 28
Results
Cohort Statistics.

Textual Descriptive Analysis
Top 10 n-Grams With the Highest TF-IDF Difference Scores per Case Outcome. The Two Top-Level Header Columns Indicate the Case According to Which the Score Difference was Sorted. R0 n-Grams had High Positive Score Difference While Non-R0 n-Grams had High Negative Score Difference.


N-gram word clouds for findings notes where residual disease is non-zero (left) and zero (right).
The words ‘normal’ and ‘miliary’ best discriminated between both groups. For non-R0 resection prediction, these included n-grams related to the EOC dissemination, such as ‘omental cake.’ The appearance of the cancer was best described by the predictive n-gram ‘miliary disease.’ The average word count was 320 ± 98 vs 292 ± 105 in case notes with non-R0 vs R0 resection, respectively. The average stop word count was 11.22 ± 6.27 vs 9.86 ± 5.89 in case notes with non-R0 vs R0 resection, respectively.
Natural Language Classification With RoBERTa
The model reached high evaluation metrics (area under ROC .86; area under precision-recall curve .87, precision, recall, and F1 score of .77 and accuracy of .81 [Figures 3–5]), surpassing even specialized BERT and DistilBERT models tested (BioBERT
29
: R .6, P 0.84, F1 .7, ACC .79, AUROC .84, AUPRC .84, ClinicalBERT
30
: R .68, P 0.72, F1 .7, ACC .76, AUROC .82, AUPRC .82 and BioClinicalBERT
31
: R .64, P 0.76, F1 .69, ACC .77, AUROC .81, AUPRC .79). The true positives, true negatives, false positives and false negatives were 35, 56, 10 and 10, respectively. Receiver operating characteristic curve and area under the curve for the RoBERTa classifier. Precision-recall curve and area under the curve for the RoBERTa classifier. Confusion matrix for the RoBERTa classifier.


The explanations of the model’s predictions were visualized in a highlighted text plot, where negatively contributing tokens are coloured red, and positively contributing tokens are coloured green, whereas the colour intensity is translated to an attribution score strength (Figure 6). Explainability of the RoBERTa inference on textual data. The green highlighting indicates the section of text that contributed positively to the classification of the note as belonging to the assigned class, while the red highlighting indicates the opposite. Examples are text instances correctly classified as describing cases where (A) residual disease persisted and (B) no residual disease persisted after surgery.
As shown in the figure, it is easy to discern the fact that there is a correlation between word contribution to the prediction and TF-IDF score difference. This makes sense, as n-grams with high TF-IDF score difference tend to discriminate better between the two cases. However, it is equally important to see that not all words that have high prediction contribution score appear as entries in Table 2. That is due to the fact that since RoBERTa is able to capture contextual meaning spanning several words that could also be non-sequential, it is possible that local information that was not apparent through simple TF-IDF analysis was now deemed as important to the prediction.
The results of the surrogate logistic regression model employed further reinforce the results acquired from the RoBERTa model. Specifically, n-grams that reached high TF-IDF difference scores (Table 2) appeared as top coefficients in the logistic regression, in either direction (Figure 7). Top 10 n-grams with the lowest (green) and highest (red) coefficients of the logistic regression model. The negative sign denotes non-existence of residual disease and vice versa.
XGBoost Classification Model
Hyperparameter Search Space of Grid Search and Chosen Parameters.

Explainability plots for the XGBoost classification model. The beeswarm feature impact plot (left) visualizes the relationship between direction of model prediction and value of feature. The bar feature impact plot (right) shows the absolute impact of each feature on model prediction.
Discussion
In this proof-of-principle study, we demonstrated the capability of the RoBERTa classifier to extract and process information from unstructured operative note formats that can enable important clinical tasks, such as R0 resection prediction following EOC surgical cytoreduction. We showcased how EHRs can be a helpful data source for supporting surgeons’ activities by automated data coding for quality assessment while reducing the burden of chart review. As an estimated 70% of clinicians report EHR-related, specialty-specific burnout, 32 this information may guide healthcare organizations on how to remediate burnout amongst their staff. Equally, we surmise this effort can help establish interoperability standards of surgical narration to ensure objectivity when it comes to reporting residual disease. Working with EHR data is relatively challenging due to data heterogeneity. Being able to quickly retrieve important information stored in surgical narratives carries the potential to improve understanding of patient journeys and identify subgroups of patients for research purposes. For those reasons, the design and application of a system that could offer the NLP AI-derived insights directly to the surgeon in real-time would be extremely beneficial. The system could offer objective feedback on written notes. A study on the effects of such a system should be investigated.
The driving motivation behind this effort was to explore the potential of using the RoBERTa algorithm in the EOC domain. This transformer architecture has been recently used to extract adverse drug events from biomedical text to monitor drug safety. 33 Barber et al initially developed an NLP-augmented algorithm that improved the ability to predict postoperative complications and hospital readmissions among women with EOC undergoing surgical cytoreduction. 34 They compiled discrete data with different types of NLP features from unstructured clinical notes and sequentially employed machine learning to build new sets of features. Herein, we purely used a novel NLP tool that recognizes the specific local textual context, thus enabling a recommendation concerning the prediction of residual disease. Pre-processing steps contributed to the rather high AUROC of .86, which shows how surgeons tend to capture more of the predictive information in their words. This ‘hunch’ critically layered upon situational awareness, and human factors have been addressed in our previous study. 13 The model specificity was higher than its sensitivity, which is critical, should this be used as a cancer screening tool for quality control. Reports of surgical findings are less restrictive in vocabulary than other EHRs, but their efficiency at scale has never been previously examined. They do not usually contain highly complex sentence structures, so they are not incorrectly abstracted as a result. By avoiding the model to make assumptions, this advantage would potentially explain the high-performance accuracy.
More importantly, we demonstrated a distinct pattern of word differential expression between R0 resection and non-R0 resection operative notes from 555 surgical events. While survival is the ultimate treatment outcome, prediction of residual disease is a key issue in the advanced EOC trajectory. This disease quantification can valuably complement our previous work using AI to predict EOC-specific surgical outcomes13,17,25–27 and validate the paradigm shift towards complete clearance to improve the survival outcomes of these patients.13,35 The use of language in medicine is often underestimated not that all Gynaecologic Oncology Surgeons speak the same language. 36 Historically, the quantification of both peri-operative disease burden and post-operative residual disease in advanced EOC was subject to significant intra- and inter-observer variability, particularly in the case of miliary peritoneal disease. While addressing the need to improve standardization and reproducibility of surgical outcomes, we made some interesting observations. Despite several words or n-grams being commonly shared between examined surgical outcomes, several descriptive words were found to be predictive of residual disease. For instance, the words ‘stuck’ and ‘adherent’ tend to describe a more complex and morbid surgery; dissemination leading to residual disease was best described by ‘(small volume) miliary disease’ or ‘miliary in all peritoneal surfaces’. ‘Excellent response to chemo’ was clearly an obvious indication to achieve R0 resection. Not surprisingly, words demonstrating hepatobiliary involvement were referring to those patients who had had macroscopically complete resection of all visible tumours. 37 Notably, the ‘completion of cytoreduction’ (CC) scoring system was developed to evaluate the extent of resection for peritoneal malignancies. 22 We clearly showed that the word ‘miliary’, if quantified, rather refers to CC1 (residual disease nodules up to 2.5 mm in size) of the perhaps outdated Sugarbaker classification (PCI). 22 We provide valid language evidence that the CC score is more likely to give a convincing and reproducible description of residual disease in EOC. In addition, the subtle performance superiority of textual data when compared with discrete surgical data can be also invoked. Going forward, data integration between structured and unstructured formats can promote innovative thinking to perfect the prediction of surgical outcomes, offer important indications for the treatment of patients and contribute to policies and clinical guidelines with the goal of reducing the future risk of unnecessary morbidity and mortality. 38
The challenges of Machine Intelligence in healthcare have been consistently addressed. 39 Transfer learning requires a close collaboration between clinicians and computer scientists. Until that happens, the inherent resilience in these tools will delay their widest adaptation. We anticipate the portability of our RoBERTa algorithm across similar practice settings by conducting original studies albeit we acknowledge the heterogeneous nature of the clinical language. Historically, linguistic models are evaluated by perplexity, that is, the probability of predicting the word in its context. Our study used retrospective data from a single institution. On that note, our data size was small-to-moderate, which entertains the general wisdom that the fewer data feeds the model, the higher the perplexity can get. Our important observations were made in a tertiary referral center; hence, they might not be generally applicable. We reiterate our strong preference for explainable NLP methods, 40 which has been showcased in this work. Understanding the features that drive a model prediction can potentially support decision-making in the healthcare domain. As NLP is moving to deep learning, it is becoming increasingly challenging for these complex non-linear data transformations to satisfy transparency. 41
The latest hype from the technological advancements in large language models has been embraced with some cautious excitement. Undoubtedly, AI-based chatbots engage in a capacity to understand multiple languages and possess knowledge of various topics. They can generate fabricated information in healthcare settings word by word. 42 In ovarian cancer research, most efforts focus on addressing the disease heterogeneity. 43 It is likely that this heterogeneity contains ‘special grammars’ that cannot be distilled from simply vast amounts of pre-trained textual data resources. Our work highlights the need for a bespoke, proprietary ovarian cancer-specific natural language that can pay attention to detail and learn beyond human knowledge.
Conclusion
We applied a sui generis approach to extract the information from the abundant textual surgical data through the use of an NLP model utilizing transfer learning and demonstrated how such tasks can be effectively modeled for the classification of prediction important surgical tasks, such as R0 resection following advanced EOC surgical cytoreduction. State-of-art NLP applications in biomedical texts can improve modern EOC care.
Footnotes
Acknowledgments
We wish to thank all the staff caring for our ovarian cancer patients at LTHT.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
