Abstract
Background
ARID3B (AT-rich interaction domain 3B) has been demonstrated to be associated with the progression and patient prognosis of several human tumors. We conducted the present study to investigate the biological behavior and clinical relevance of ARID3B in gastric cancer (GC).
Methods
Detection of the expression level in GC tissues and cell lines were performed by Western blot and immunohistochemistry. We also retrospectively analyzed the correlation of ARID3B with clinicopathological characteristics and patient prognosis in gastric cancer. The biological functions of ARID3B in GC cells were further explored by transwell migration assays, wound healing assays and cell proliferation assay.
Results
The present study suggested that the expression of ARID3B was significantly lower in GC tissues than in adjacent normal tissues. IHC staining in tissues of 406 GC patients from training and validation sets verified that ARID3B over-expression correlated with clinicopathological features, such as degree of differentiation and clinical stage. Meanwhile, ARID3B was proved to be an independent prognostic factor for GC prognosis. Furthermore, over-expression of ARID3B suppressed proliferation in GC cells according CCK8 assay. We found that over-expression of ARID3B inhibited GC cell migration by transwell assay and wound healing assay. Furthermore, EMT markers were detected in ARID3B over-expression GC cells, which showed that ARID3B may inhibit metastasis of GC cells.
Conclusion
Our results firstly revealed that the expression level of ARID3B was closely correlated with clinicopathological features and may serve as an independent prognostic factor for GC patients. More importantly, ARID3B could suppress GC progression, including cell proliferation, migration and metastasis.
Introduction
Gastric cancer (GC) has the fifth highest incidence and the third highest mortality rate among cancers worldwide. 1 For the early symptoms is not obvious, a large number of GC patients are diagnosed in advanced stage. Some patients have undergone gastrectomy or even combined postoperative adjuvant therapy, yet recurrence and metastasis within 5 years are common. 2 The human stomach carcinogenesis and development contain multiple genetic programming and molecular mechanisms. 3 Therefore, exploring more potential biomarkers for early diagnosis, prognosis and therapeutic target, is imperative to make individualized therapeutic regimens for GC patients.4,5
The AT-Rich interacting domain (ARID) family is a group of DNA-binding proteins that have been studied and found to be important regulators of gene expression and chromatin remodeling. 6 Due to their unique DNA binding domain, ARID family proteins are involved in embryonic development, cell cycle, differentiation and transcriptional regulation.7-10 As an ARID3 subfamily protein, ARID3B shares additional conserved regions that extend beyond the ARID, together with ARID3A and ARID3C.8,11-13 Previous studies showed that deletion of ARID3B could lead to multiple abnormalities and embryonic lethality in mice, which demonstrated that ARID3B plays a vital role in embryonic development.14,15 One possible reason is that ARID3B is an upstream regulator of many cancer stem cell genes, which may be involved in the formation of stem cell phenotypes. 16 In addition to its involvement in cancer development, ARID3B has been linked to cell transformation and tumor progression.17,18 ARID3B have been proved to bind pRB, a key regulation factor of cell cycle, indicating that ARID3B can participate in the regulation of cell cycle. 10 What’s more, ARID3B plays an essential role in p53-mediated apoptosis process by facilitating transcription of relevant critical genes. 19 In addition, ARID3B also can synergize mouse embryonic fibroblasts (MEF) and N-myc oncogene to promote the occurrence of malignant tumors. 18 ARID3B was present in tumors of the salivary gland, esophagus, stomach, intestine, thyroid and cervix. However, the expression of ARID3B was different in kinds of tumor types and changed during tumor progression. 20 For example, ARID3B is thought to be involved in the progression of breast cancer, ovarian cancer and malignant neuroblastoma.18,21-23 However, other studies have shown that overexpression of ARID3B is associated with a better prognosis in patients with colorectal cancer and breast cancer.24,25
So far, the role of ARID3B in the development of GC and its biological behavior are not clear. In the current study, we investigated the expression levels of ARID3B in gastric cancer and the clinical significance in GC patients. We also explored the effects of ARID3B in GC cell lines. Through the above, this study was undertaken to elucidate the functional and prognostic significance of ARID3B in GC.
Materials and Methods
Patients and Tissue Samples
Specimens obtained from 406 GC patients who experienced radical surgery from January 2007 to December 2015 in the Affiliated Tumor Hospital of Nantong University, were retrospectively analyzed in this study. The cohort was split into a training set (n = 104) and a validation set (n = 302). Two patients in the training set had liver metastasis and underwent surgical resection at the same time, which were diagnosed as Stage IV (M1 stage). Duo to the too small sample size to provide meaningful statistical results, we didn’t conduct statistics separately for M stage. The eligibility criteria were as follows: (1) All patients had a clear post-operative pathological diagnosis of gastric cancer; (2) No history of secondary primary tumors; (3) All patients were not treated with neoadjuvant therapy; (4) All enrolled GC patients had detailed clinicopathological data. The cut-off time is the time of the patient’s death, or the follow-up time is December 31, 2020, except for patients who were lost to follow-up. The time interval between the date of surgery and death or the last follow-up visit was considered to be the overall survival (OS). Tumor staging was based on the 8th Edition tumor-node-metastasis (TNM) staging classification system of the American Joint Committee on Cancer. The tissue samples including gastric cancer tissue and the corresponding normal tissue were collected and made into tissue microarray (TMA). This study was approved by the ethics committees of the Affiliated Tumor Hospital of Nantong University (ethics approval number:2023-A 01). We have de-identified all patient details. All procedures involving human participants in this study were in accordance with the Declaration of Helsinki (as revised in 2013).
Immunohistochemical Staining and Evaluation
The gastric cancer tissue chip was taken out and a series of operations were carried out, such as baking, dewaxing, hydration, antigen repair and antibody incubation. Primary anti-ARID3B antibody (ab227870, diluted 1:400; Abcam) was applied for immunohistochemistry (IHC) staining. The tissue chips were reviewed and scored by 2 experienced pathologists in a double-blind method. As mentioned earlier, a semi-quantitative approach was used to determine the score. 26 Five high-power fields of view were randomly selected to count tissue cells. The staining intensity was classified as follows: 0 (negative), 1 (weak), 2 (moderate), and 3 (strong). The combined score was calculated by multiplying the density and intensity of staining (ranging from 0 to 12). During the analysis, scores <6 were considered low expression and scores ≥6 were considered high expression. When the scores were inconsistent, the original wax block would be re-evaluated to determine the final scoring results.
Cell Lines and Cell Culture
Gastric cancer cell lines, including AGS, SGC-7901, MKN-7 and HGC-27, were obtained from Center Laboratory of Affiliated Tumor Hospital of Nantong University. AGS was cultured in McCoy’s 5A medium containing 10% fetal bovine serum (FBS) (Gibco, USA), SGC-7901, MKN-7 and HGC-27 were cultured in RPMI 1640 (HyClone, USA) containing 10% FBS. The culture environment was 5% carbon dioxide, constant temperature 37°C cell incubator. Periodically, the cell medium was replaced with fresh medium and cell expansion was performed.
Western Blot Assay
The cell culture bottle for protein extraction was placed on the ice surface and washed 3 times with PBS. RIPA cell lysate (Beyotime, Shanghai, China) was added for protein extraction. The protein concentration was determined by Nandrop1000 spectrophotometer (Thermo Fisher, USA) and recorded. An equal amount (100 μg) of protein sample was loaded and separated by 10% sodium dodecyl sulfate-polyacrylamide gels (SDS-PAGE) and transferred to polyvinylidene fluoride membranes (PVDF) (Millipore, Billerica, MA, USA). Next, the membranes were fully incubated with antibodies specific for ARID3B (abcam, ab227870, 1:1000), N-cadherin (abcam, ab98952, 1:500), snail (abcam, ab31787, 1:500), twist (abcam, ab175430, 1:500), vimentin (abcam, ab92547, 1:1000) and GAPDH (abcam, ab8245, 1:5000), overnight at 4°C. The next day membranes were washed and then incubated with secondary antibody for 2 h at room temperature. Finally, the proteins were visualized using High-sigECL Western blotting Substrate (Tanon, Shanghai) and exposed to X-ray film for analysis by Image.lab3.0 software.
ARID3B Knockdown and Over-Expression
In order to knock-down ARID3B in GC cells, short hairpin RNAs (shRNA) targeting the ARID3B transcript were obtained from Genechem. Sequences were (5’-3’): AGTAGCATTGGGAGCATTA and TAATGCTCCCAATGCTACTCG. For ARID3B over-expression in GC cells, full-length human ARID3B cDNA (NM_006465) was constructed into a lentiviral expression vector, and empty vector was used as the control. Lentiviral transfection was performed according to the protocol of the manufacturer. The cells prepared for transfection with lentivirus were counted and inoculated into 24-well plates with a density of 30 000 cells per well. On the second day, lentivirus (MOI: 50) was added and cell proteins were extracted 72 to 96 s later to detect infection effect.
Transwell Migration Assays
GC Cell migration were assessed by performing the Transwell assay as previously described. 27 Transwell assays were performed by boyden chambers with a polycarbonate nucleopore membrane. Cells were suspended in 200 μL of serum-free RPMI 1640 medium and seeded into the upper chamber at a density of 10 × 104 cells for migration assays. The lower chamber was filled with 500 μL of medium containing 10% FBS. Cells were incubated in a 37°C cell incubator for 24 h. Cells were fixed in methanol and stained with crystal violet. Cell counting was performed using a phase contrast microscope (Olympus, Germany) in 5 random areas of the invaded Matrigel layer.
Wound Healing Assay
A wound healing assay was conducted to further investigate tumor cell migration. The cells were seeded at 80% to 90% confluence in 6-well plates. The cell monolayer was scratched in a straight line with 10 μL pipette tips drawn across the center of each well. After 48 h, the wounds were observed under a microscope and taken photographs at 0, 24 and 48 h. The wound area ratio at 48 h would be compared with 0 h.
Cell Proliferation Assay
GC cells were inoculated in 96-well plates and cultured for 0, 24, 48 and 72 hours. Cell viability was assessed using the Cell Counting Kit-8 (CCK-8) protocol (Beyotime, Shanghai, China). Because the cell culture time was long, the circle around the 96-well plate was most likely to evaporate, so the circle around the 96-well plate was abandoned and the same amount of PBS or culture medium was added. After adding 10 μl of CCK-8 reagent and incubating for 2 hours protected from light, OD 450 was measured by enzyme label instrument (Thermo Fisher,USA). Note that there are no bubbles in each hole before using the enzyme marker.
Database Analysis
Kaplan-Meier Plotter for the OS data of ARID3B in GC patients was obtained from the Human Protein Atlas (https://www.proteinatlas.org/). The Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway was applied to enrich related genes and identify the key biological pathways, by using the clusterProfiler, enrichment, and ggplot2 R packages to enrich related genes.
Statistical Analysis
Statistical analysis and graphic plotting were done with SPSS 26.0 software (IBM Corp., USA), GraphPad Prism version 8.0.1 (La Jolla, USA) and R programming language version 3.2.2 (R Foundation for Statistical Computing, Vienna, Austria). The Fisher’s exact test, chi-square test and independent t-test were used for comparisons among groups. The survival curves were calculated with Kaplan-Meier method and log-rank test. Hazard ratios (HR) and 95% confidence intervals (95% CI) for clinicopathological characteristics in univariate and multivariate analysis were analyzed by Cox proportional hazards regression. Nomogram models were analyzed by the “rms” package of R programming language. The prognostic accuracy of the prognostic models was demonstrated by Harell concordance index (C-index). Each experiment was repeated 3 times. All statistical analyses were performed using a two-tailed test and P-value <.05 was considered statistically significant. The reporting of this study conforms to the Strengthening the Reporting of Observational Studies in Epidemiology (STROBE) guidelines. 28
Results
ARID3B is Down-Regulated in Human Gastric Cancer
IHC was performed to detect ARID3B expression in a GC tissue array containing tumor tissue from GC patients and adjacent normal tissue. As shown in Figure 1(A), ARID3B protein was mainly distributed in the nucleus. Meanwhile, the results of IHC showed that ARID3B expression was significantly higher in adjacent normal tissues compared with primary GC (Figure 1(A)and (B)). Besides, when subgroup analyses were conducted by differentiation and TNM stage, the IHC results showed that the expression of ARID3B were higher in well differentiation than poor differentiation (Figures 1(C)-(E)). The same results were observed in the TNM stage I than stage II and III. The above results indicated that ARID3B may inhibit the progression of GC (Figures 1(F)-(H)). ARID3B is down-regulated in human gastric cancer in the GC tissue array by IHC. (A, B) ARID3B protein expression was mainly distributed in the nucleus and was significantly higher in adjacent normal tissues compared with primary GC. (C, D, E) ARID3B protein expression in well differentiation, moderate differentiation and poor differentiation. (F, G, H) ARID3B protein expression in TNM stage I, stage II and stage III. Original magnification:×4, scale bar: 500 μm; ×20, scale bar: 100 μm.
The Expression of ARID3B is Associated With Clinicopathological Features of Gastric Cancer
Relationship Between the Expression of ARID3B and Clinicopathological Features of GC Patients.

High ARID3B Expression Indicated Better Prognosis in Patients With Gastric Cancer
To further identify the influence of ARID3B expression status on the OS of GC patients, we first applied Kaplan-Meier survival analyses. As shown in Figures 2(A)-(C), patients with high ARID3B expression survived longer in both the training (P < .001) and validation sets (P < .001), which were consistent with the result from the Human Protein Atlas (https://www.proteinatlas.org/) (P = .047). Interestingly, in the group of III-IV clinical stage from the 2 sets, we found that low ARID3B expression indicated poor prognosis (training set: P = .007; validation set: P < .001) (Figure 2(E), (G)), while no significant difference in prognosis was observed in I-II clinical stage (training set: P = .120; validation set: P = .200) (Figure 2(D), (F)). When stratified by differentiation, patients with high ARID3B expression had longer overall survival both in well (P = .013) and poorly differentiated groups (P = .037) in the training set (Figure 2(H), (I)). While in the validation set, high ARID3B expression predicated better prognosis in poorly differentiated group (P < .001), but no in well differentiated group (P = .100) (Figure 2(J), (K)). High ARID3B expression indicated better prognosis in patients with gastric cancer according to the Kaplan–Meier survival curves. (A) Patients with high ARID3B expression had significantly better survival from the database result of the Human Protein Atlas. (B-C) In the training set (B) and validation set (C), patients with high ARID3B expression had significantly longer overall survival. (D-G) Stratified by clinical stage, no significant difference in prognosis was observed in I-II clinical stage (training set: D; validation set: F). Low ARID3B expression indicated poor prognosis (training set: E; validation set: G) in the group of III-IV clinical stage. (H-I) Stratified by differentiation, patients with high ARID3B expression had longer overall survival both in well (H) and poorly differentiated groups (I) in the training set. (J-K) High ARID3B expression predicated better prognosis in poorly differentiated group (J), but no in well differentiated group (K) in the validation set. P < .05 means statistically significant.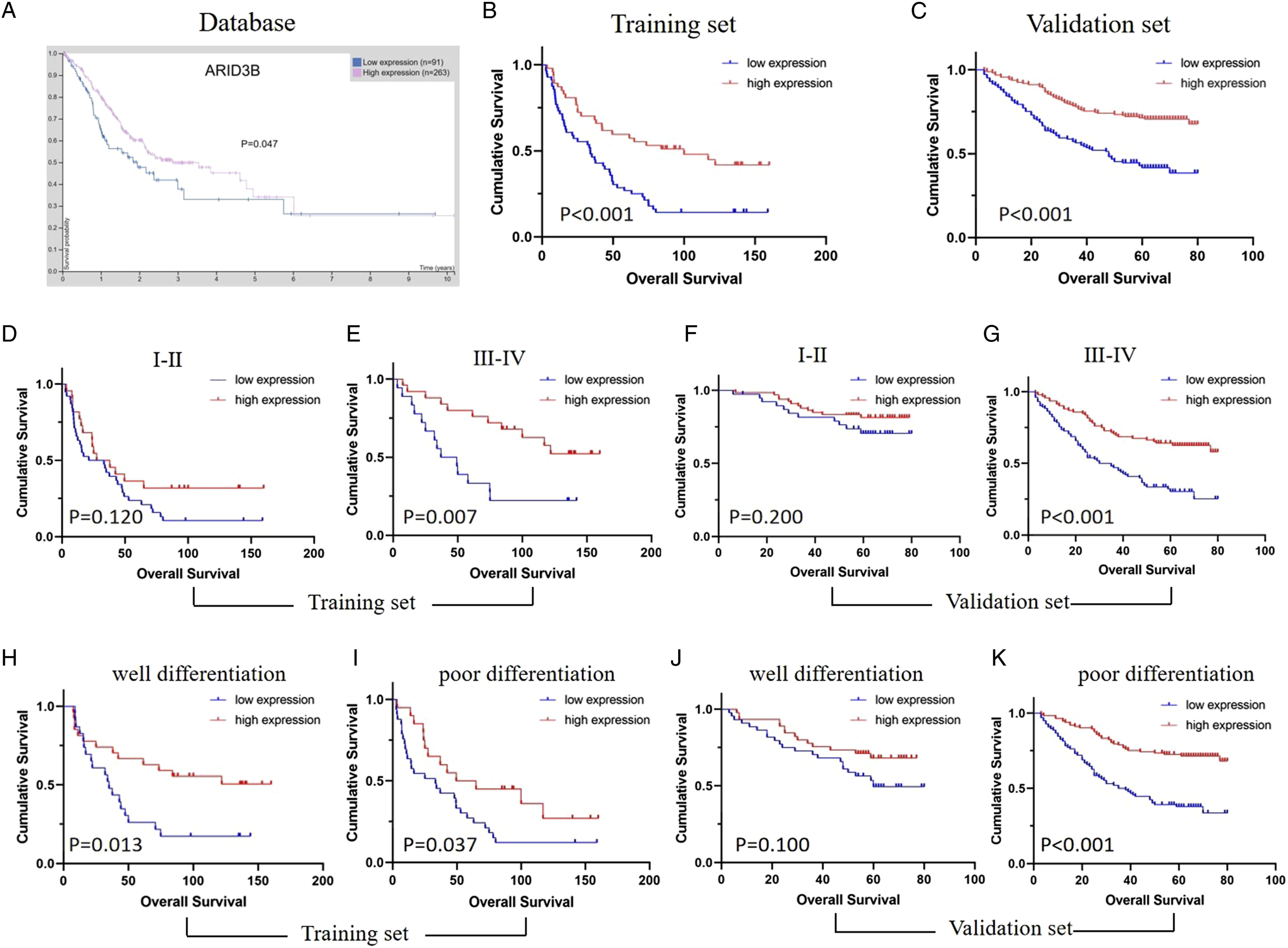
Univariate and Multivariate Cox Regression Analyses for Overall Survival of Gastric Cancer.

Moreover, to visualize the prognostic application of ARID3B expression, prognostic nomograms were conducted to predict the 3- and 5-year survival of the GC patients based on ARID3B expression and other well-known prognosticators from the Cox regression analysis. The nomograms showed improved prognostic accuracy for GC when integrating ARID3B expression level both in the training set (C-index: .669) and validation set (C-index: .716) (Figure 3(A), (B)). Nomograms for predicting the 3- and 5-year survival of the GC patients. (A-B) Nomograms predict the 3- and 5-year survival of the GC patients based on ARID3B expression and other well-known prognosticators from the Cox regression analysis in the training set (A) and validation set (B).
ARID3B Suppresses Gastric Cancer Cell Proliferation, Migration and Metastasis in Vitro
We conducted KEGG pathway analysis to identify the possible role of ARID3B playing in GC. As shown in Figure 4, ARID3B may regulate the pathway of homologous recombination, mismatch repair, DNA replication, base excision repair, linolenic acid metabolism and lysine regulation. Analysis of biological processes showed that ARID3B was primarily involved in DNA replication and cell cycle regulation. Cellular component showed that ARID3B mainly related to the formation of cytoskeleton. Molecular function suggested that ARID3B functioned in phosphokinase activity, protein deacetylation, ubiquitination, p53 binding and transcription factor (Figure 5). The possible functions of ARID3B playing in GC by database analysis. ARID3B may regulate the pathway of homologous recombination, mismatch repair, DNA replication, base excision repair, linolenic acid metabolism and lysine regulation. P < .05 means statistically significant. The possible roles of ARID3B playing in GC by KEGG pathway analysis. (A-B) Biological process analysis indicated that ARID3B was mainly involved in DNA replication and cell cycle regulation. (C) Cellular component showed that ARID3B mainly related to the formation of cytoskeleton. (D) Molecular function suggested that ARID3B functioned in phosphokinase activity, protein deacetylation, ubiquitination, p53 binding and transcription factor. P < .05 means statistically significant.

To explore the potential role of ARID3B in the development of GC, the applicable cell models were established. WB was conducted to assess the expression levels of ARID3B in 4 GC cell lines (AGS, SGC-7901, MKN-7 and HGC-27). ARID3B expression levels were significantly higher in AGS than in other cell lines (Figure 6(A)). ARID3B was knocked-down in AGS cells or over-expressed in SGC-7901 cells, which were validated by WB (Figure 6(B)). ARID3B suppresses gastric cancer cell proliferation, migration in vitro. (A) The expression levels of ARID3B in 4 GC cell lines (AGS, SGC-7901, MKN-7 and HGC-27) by WB. (B) ARID3B was knocked-down in AGS cells and over-expressed in SGC-7901 cells, which were verified by WB. (C) ARID3B knockdown markedly promoted the proliferation of AGS cells by CCK8 assay. (D) ARID3B over-expression impeded the growth of SGC-7901 cells by CCK8 assay. (E) Transwell assays showed that the over-expression of ARID3B inhibited the migration of SGC-7901 cells and ARID3B knockdown promoted the migration of AGS cells. (F) The number of migratory cells were counted by Image J, Original magnification:×10, scale bar: 100 μm. Data are presented as the mean ± SD of 3 independent experiments. *P < .05, **P < .01, ***P < .001, ****P < .0001.
We then assessed the role of ARID3B in cell proliferation by CCK8 assay. The study clearly demonstrated that ARID3B knockdown significantly promoted the proliferation of AGS cells, while ARID3B over-expression impeded the growth of SGC-7901 cells (Figure 6(C), (D)). Next, the results of transwell assays showed that the over-expression of ARID3B inhibited the migration of SGC-7901 cells and ARID3B knockdown promoted the migration of AGS cells (Figure 6(E)). The numbers of migratory cells were counted and showed statistically significant both in the 2 groups (Figure 6(F)). Similarly, in the Wound healing assay, the relative healing of wounded areas in the over-expression of ARID3B groups were reduced compared to the control groups (Figure 7(A)). Conversely, in the ARID3B knockdown groups, the relative healing of wounded areas were increased compared the control groups (Figure 7(B)). Epithelial-mesenchymal transition (EMT) has been well known as an important process in the metastasis of GC cells.
29
Next we explored the association between ARID3B and the EMT markers. Compared with the control cells, ARID3B-overexpression SGC-7901 cells expressed lower level of N-cadherin, snail, twist and vimentin, while ARID3B knockdown AGS cells expressed higher at protein level by WB (Figure 7(C)). Based on these above results, ARID3B could inhibit the proliferation, migration and metastasis of GC cells. ARID3B suppresses gastric cancer cell migration and metastasis in vitro. (A) Wound healing assay showed that the relative healing of wounded areas in the over-expression of ARID3B groups were reduced compared with those in the control groups in 48 h. The percentage of wound area closure was analyzed by Image J. (B) The relative healing of wounded areas in the ARID3B knockdown groups were increased compared with those in the control groups in 48 h. The percentage of wound area closure was analyzed by Image J. (C) Compared with the control cells, ARID3B over-expression SGC-7901 cells expressed lower level of N-cadherin, snail, twist and vimentin, while ARID3B knockdown AGS cells expressed higher at protein level by WB. Original magnification:×10, scale bar 500 μm. Data are presented as the mean ± SD of 3 independent experiments. *P < .05, **P < .01, ***P < .001, ****P < .0001.
Discussion
The functions of ARID3B have been reported to be varied among various pathological types of cancer. However, the role of ARID3B in the development of gastric cancer remains unclear. This study firstly demonstrated for its key function in GC through a combination of clinicopathological data analysis and functional testing. The results showed that high ARID3B expression could inhibit the proliferation, migration, invasion, metastasis of GC cells, and is apparently associated with better prognosis of GC patients. First, we found that the expression of ARID3B was higher in GC tissues than adjacent normal gastric tissues. Second, IHC staining in tissues of 406 GC patients from training and validation sets verified that ARID3B over-expression correlated with clinicopathological features, such as degree of differentiation and clinical stage. Moreover, survival analyses showed that the down-regulated ARID3B predicted a poor prognosis of GC patients. Meanwhile, ARID3B was proved to be an independent protective factor for GC prognosis using COX regression analysis and prognostic nomograms. Furthermore, over-expression of ARID3B suppressed proliferation in GC cells according CCK8 assay. Through transwell assays and wound healing assay, we found that forced expression of ARID3B inhibit the migration and invasion of GC cells. Furthermore, EMT markers were detected in ARID3B over-expression GC cells, which showed that ARID3B may repress metastasis of GC cells. These results clearly indicated that ARID3B may be a key inhibitor in gastric cancer progression and may serve as a prognostic indicator for GC patients.
The ARID protein family is a group of transcriptional regulators which contain a distinctive DNA-binding domain. 6 Proteins of this family have been involved in various biological processes, such as cell cycle and differentiation.7-10 ARID3B, as a important member of the ARID3 subfamily, have been proved to play a key role in tumor initiation and progression in various cancers.17,18 According to the bioinformatics analysis in GC, we supposed that ARID3B may be mainly involved in DNA replication and cell cycle regulation. Mounting studies have also confirmed that ARID3B play important functions in the apoptosis which related to the cell cycle regulation. Pratama et al. identified that ARID3B play a differential role in p53-mediated apoptosis. Possible mechanism was that ARID3B can binding to the p53 promoter, facilitate transcription of p53 and its target genes and induce the function of promote apoptosis. 19 Previous study reported that in wild-type p53 ovarian cancer cell lines, over-expression of ARID3B activates transcription of genes related to the apoptotic pathway, thereby promoting apoptosis in ovarian cancer cells. 21 In the TNF-related apoptosis, over-expression ARID3B could up-regulate the pro-apoptotic BIM, Tumor Necrosis Factor alpha (TNFα) and TNF-related apoptosis inducing ligand (TRAIL) which resulting in cell death. 21 However, when interacting with Mycn during mouse neural crest differentiation, Mycn promote cell cycle progression and ARID3B prevent cell death. 30 ARID3B has contradictory effects on apoptosis, which may depend on the different target genes and cancer cell lines.
Another important function of ARID3B is its association with cancer stem cells (CSC). ARID3B was first reported to be expressed at high levels in embryonic (ES) and induced pluripotent stem cells in ovarian cancer.30,31 Further study revealed that ARID3B over-expression is associated with elevated expression of many stem cell markers (LGR5, Oct4 and Nanog), which could increases CSC production.31,32 What’s more, ARID3B can also up-regulate metastasis associated genes of the Wnt signaling pathway (SFRP1 and WISP1) in ovarian cancer.33,34 Furthermore, as a target of miR-125a, ARID3B promotes migration of breast cancer cells and invasion of ovarian cancer cells and tends to function as an oncogene. 22 However, in gastric cancer, previous study examined 42 gastric cancer tissues and 10 normal stomach sections, of which 34 GC tissues exhibited decreased ARID3B staining intensity. 20 Consistent with the above result, we found that ARID3B suppresses gastric cancer cell proliferation, migration, invasion and metastasis in vitro. Besides, patients databases of the German population-based case-control DACHS study were analyzed and showed that overexpression of ARID3B is associated with longer disease-specific survival time of obese colorectal cancer patients. 24 Another study using the TCGA database demonstrate that low mRNA expression of ARID3B were interrelated with worse OS in each molecular subtypes of breast cancer patients. 25 Similar results were observed in this study that down-regulated ARID3B predict a poor survival of GC patients and ARID3B can act as an independent protective factor for GC prognosis. The contradictory results of ARID3B function may due to the different target genes, tissue micro-environment and cancer types. Other reasons may include inter-cohort variation, diverse experimental conditions and different cell models. The limitation of our study is that required sample size is not calculated and justified for patients and tissue samples. However, all eligible cases in the time interval were included, which is enough to support our conclusion. What’s more, there is no functional experiment to explore the mechanism of ARID3B and gastric cancer, that is what we need to do next. Therefore, validation of our results by more studies is crucial for further understanding the role of ARID3B in cancer.
Conclusions
In general, our finding firstly revealed that the expression level of ARID3B is closely related to the clinicopathological features and may serve as an independent prognostic factor in GC patients. In addition, we explored the effects of ARID3B on GC progression, including the suppression of cell proliferation, migration and metastasis. The above suggests that ARID3B may be a potential prognostic biomarker and a therapeutic target for patients with GC. However, further studies are need to clarify the specific molecular mechanisms of ARID3B involved in the development of GC.
Supplemental Material
Supplemental Material - Over-Expression of ARID3B Suppresses Tumor Progression and Predicts Better Prognosis in Patients With Gastric Cancer
Supplemental Material for Over-Expression of ARID3B Suppresses Tumor Progression and Predicts Better Prognosis in Patients With Gastric Cancer by Xunlei Zhang, Xinyue Qiu, Wenjing Zhao, Li Song, Xingsong Zhang, Lei Yang, and Min Tao in Cancer Control.
Footnotes
Acknowledgments
Not applicable.
Author Contributions
ZXL and TM conceived and designed the experiments; ZXL, QXY and ZWJ performed the experiments; SL and YL performed the statistical analysis; ZXS and ZXL collected clinical samples and corresponding clinical data; ZXL and QXY wrote the manuscript; TM and YL supported the experiments; All authors read and approved the final manuscript. Xunlei Zhang, and Xinyue Qiu are contributed equally to this work.
Author’s Note
Part of data generated or analyzed during this study are included in this article. The other datasets used or analyzed during the current study are available from the corresponding author on reasonable request.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The work was supported by grants from Natural Science Foundation of Jiangsu Province (BK20191208), the General Project of Nantong Health Committee (MA2020009), Nantong Science and Technology Foundation grant (JCZ20204), The Science and Technology Plan Project of Suzhou (SLT201913), Beijing Xisike Clinical Oncology Research Foundation (Y-XD2019-227), Horizontal Research Foundation of Soochow University(P142900221).
Ethical Approval
This study was approved by the Ethics Committee of the Institutional Ethical Review Board of the Affiliated Tumor Hospital of Nantong University (ethics approval number:2023-A 01).
Supplemental Material
Supplemental material for this article is available online.
Appendix
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
