Abstract
Objective
Interleukin-17A (IL-17A) genetic polymorphisms are associated with multiple cancer types, including colorectal cancer (CRC). However, no previous study was performed in the Bangladeshi population to evaluate the association. Our study aimed to find the association between two IL-17A variants (rs10484879 C/A and rs3748067 G/A) and susceptibility of CRC.
Methods and Materials
This retrospective case-control study comprised 292 CRC patients and 288 age, sex, and BMI matched healthy volunteers. Genotyping of both variants was done by the tetra-primer ARMS-PCR method, and the results were analyzed by the SPSS software package (version-25.0).
Results
Logistic regression analysis indicated that in case of IL-17A rs10484879 polymorphism, AC and AA genotype carriers showed 2.44- and 3.27-times significantly increased risk for CRC development (OR = 2.44, P = .0008 and OR = 3.27, P = .0133, individually). A significant association was also observed for AC + AA genotype (OR = 2.58, P = .0001). Again, over-dominant and allelic model revealed statistically significant link to CRC risk (OR = 2.13, P = .0035 and OR = 2.22, P = .001). For rs3748067 polymorphism, AG and AA genotype carriers showed 2.30- and 2.45-times enhanced risk for CRC (OR = 2.30, P = .005 and OR = 2.45, P = .031). A statistically significant association was also observed for AG + AA genotype (OR = 2.35, P = .001), over-dominant model (OR = 2.05, P = .014), and allelic model (OR = 2.11, P = .0004).
Conclusion
This study highlights that IL-17A rs10484879 and rs3748067 polymorphisms may be associated with CRC development. However, further functional research with larger samples may reveal more statistically significant outcomes.
Background
Colorectal cancer (CRC) is considered one of the most prevalent carcinomas that mainly affects the areas of the digestive tract. 1 This malignancy is accounted for almost 10% of worldwide occurrences and more than 9% of mortalities by the year 2020. Based on aging projections, population explosion, and human development, the global burden of new cases with CRC is supposed to exceed 3 million within the next 20 years. 2 In 2020, colorectal cancer cases (including anus) were more than 1.9 million, where the death number was about 935000, representing about 1 in ten cancer cases and deaths. In terms of incidence, it ranks about 3rd but 2nd in the case of mortality. 3
CRC risk factors have been found to be heterogeneous across anatomical locations. Smoking, for example, is linked to a higher chance of proximal colon and rectal cancers but not distal colon cancer. 4 Physical exertion is reciprocal to the risk of distal and proximal colon tumor but not tumor of the rectum. 5 The distinguished pathogenic loads, matching microbiological products, and host features may contribute to the heterogeneity of these risk factors.6,7 There are several non-modifiable risk factors such as age, 8 race,2,9 sex, 5 personal history of adenomatous polyps, 10 coronary artery disease,11,12 personal history of inflammatory bowel disease, 13 and modifiable risk factors such as diet,14,15 physical inactivity and sedentary lifestyle,16,17 cigarette smoking,18,19 alcohol consumption,20,21 sedentary lifestyle, 22 overweight and obesity,23-25 chronic inflammatory bowel disease (IBD)26-28 as well as genetic risk factors act an important role in CRC risk, especially when relatives have been diagnosed with the disease early. 29 Moreover, 60-65% of all CRC cases are sporadic, with no family history or inherited genomic abnormalities.2,30
The IL-17A gene, which spans 4252 bp and is found on chromosome 6, location 6p12.2, has three exons and two introns. 31 This gene produces a pro-inflammatory interleukin that is released by activated T lymphocytes and is a major representative of the IL-17 receptor superfamily, which covers six members (IL-17RA-F). The resulting protein has important effects on host immune defense mechanism, immune regulation, and body tissue reconstruction and plays a substantial function in innate immune defense induction. IL-17A is responsible not only for host defense but also for the etiology of various autoimmune diseases and cancers. 32 There are several diseases along with cancers caused by the IL-17A gene, such as multiple sclerosis, 33 psoriasis, 34 Crohn’s disease and ulcerative colitis, 35 systemic lupus erythematosus, 36 lung diseases. 37 It also plays potential role in the formation of stomach cancer,38,39 colorectal cancer,40,41 cervical cancer, 42 lung carcinoma, 43 breast carcinoma, 44 and bladder cancer. 45
There are a few case-control studies till now have been investigated to find out the association of human IL-17A gene polymorphisms (rs10484879 and rs3748067) with the susceptibility of CRC. A Tunisian study has demonstrated that the frequency of the minor allele in rs10484879 was substantially greater in colorectal carcinoma patients than in healthy participants. They also found that heterozygous rs10484879 was linked to an elevated susceptibility to CRC, and the minor allele rs10484879 was linked to a positive family background of this cancer as well as other malignancies. Moreover, they showed that rs3748067 has a positive link with CRC development. 40 Another study examined the role of these polymorphisms in which authors applied information from two population-dependent pilot studies to see whether 69 variants of 13 genes in the cytokine mechanism have any association with colon and rectal tumor risk or not. They found several results with several genes and their respective SNPs. Notably, it was found that the rs10484879 of the IL-17A gene is also associated with colon cancer. 41
Based on the biological, geographical, and ethnic variations, the rate and extent of disease occurrences also vary. The variations may even be seen between the populations from the same geographical area or ethnicity. 46 So, based on previous research around the world, we speculated that rs10484879 and rs3748067 polymorphisms in IL-17A gene might be responsible for CRC development in the Bangladeshi population. To evaluate this hypothesis, the present case-control analysis was performed in our population.
Method and Material
Study Population and Ethical Permission
This retrospective case-control study is comprised of 292 CRC patients and 288 healthy participants. This case-control study was conducted from September 2021 to March 2022. Ethical permission was taken from the Research Ethics Committee, the University of Asia Pacific, Bangladesh [Approval No. UAP/REC/2O21/108(S)] after reviewing the study protocol. Patients with CRC were recruited randomly from the National Institute of Cancer Research and Hospital Dhaka Ahsania Mission Cancer and General Hospital. Controls were chosen following a physical examination by matching age and sex to CRC patients. Data on age, sex, dietary habit, weight, BMI (Body Mass Index), TNM (Tumor Node Metastasis) classification, demographical variables, and lifestyle features were obtained by competent nurses with available expert physicians. Each patient and control subject signed an informed consent document after briefing the purpose of the study. The current pilot investigation was conducted at the Biotechnology Laboratory of the Pharmacy Department, University of Asia Pacific, Bangladesh, according to the Helsinki II declaration. The reporting of this study conforms to STROBE guidelines for case-control studies. 47
Genomic DNA Extraction and Quantification
DNA extraction kit, Axygen® (USA) was used for isolating genomic DNA from whole blood (3 mL) from 292 CRC and 288 healthy participants. The procedure manual that came with the kit was meticulously followed when extracting DNA. The purity of the extracted DNA and their concentrations were measured by Nano Spectrophotometer (Genova Nano, Jenway) at 260/280 nm. The purity of the entire DNA was found to be in the range 1.7-2.0, which was then stored at −80°C.
Preparation of Primers and PCR Working Mix
Primer Sequences and Observed PCR Products.

SNP: Single nucleotide polymorphism; PCR: Polymerase chain reaction; BP: Base-pair.
Validation Process of ARMS–PCR and Subsequent Genotyping
Before starting genotyping, the process of method validation was carried out to choose an appropriate annealing temperature for both genetic variants to obtain the desired fragments of DNA.
48
Primarily, a gradient PCR was run, setting the temperature from 55°C to 65°C by altering the concentration of primers. Based on the outcomes, two annealing temperatures, 63°C and 63.5°C were selected as for amplifying rs10484879 and rs3748067, individually. In terms of the rs10484879 variant, at 63°C temperature, we got 203 bp, 149 bp, and 110 bp DNA fragments (Figure 1), whereas for the rs3748067 variant, at 63.5°C temperature, we got 305 bp, 192 bp, and 168 bp DNA fragments (Figure 2). Next, the products of PCR were examined using gel electrophoresis (1.5% agarose), adding EtBr (ethidium bromide) to confirm the presence of the target genotypes. PCR amplification conditions and the respective amplified fragments for both SNPs (rs10484879 and rs3748067) are described in Table 1. About 20% of the samples were replicated to confirm the genotypes. Observed PCR products for IL-17A gene rs10484879 after 1.5% of agarose gel electrophoresis. Observed PCR products for IL-17A gene rs3748067 after 1.5% of agarose gel electrophoresis.
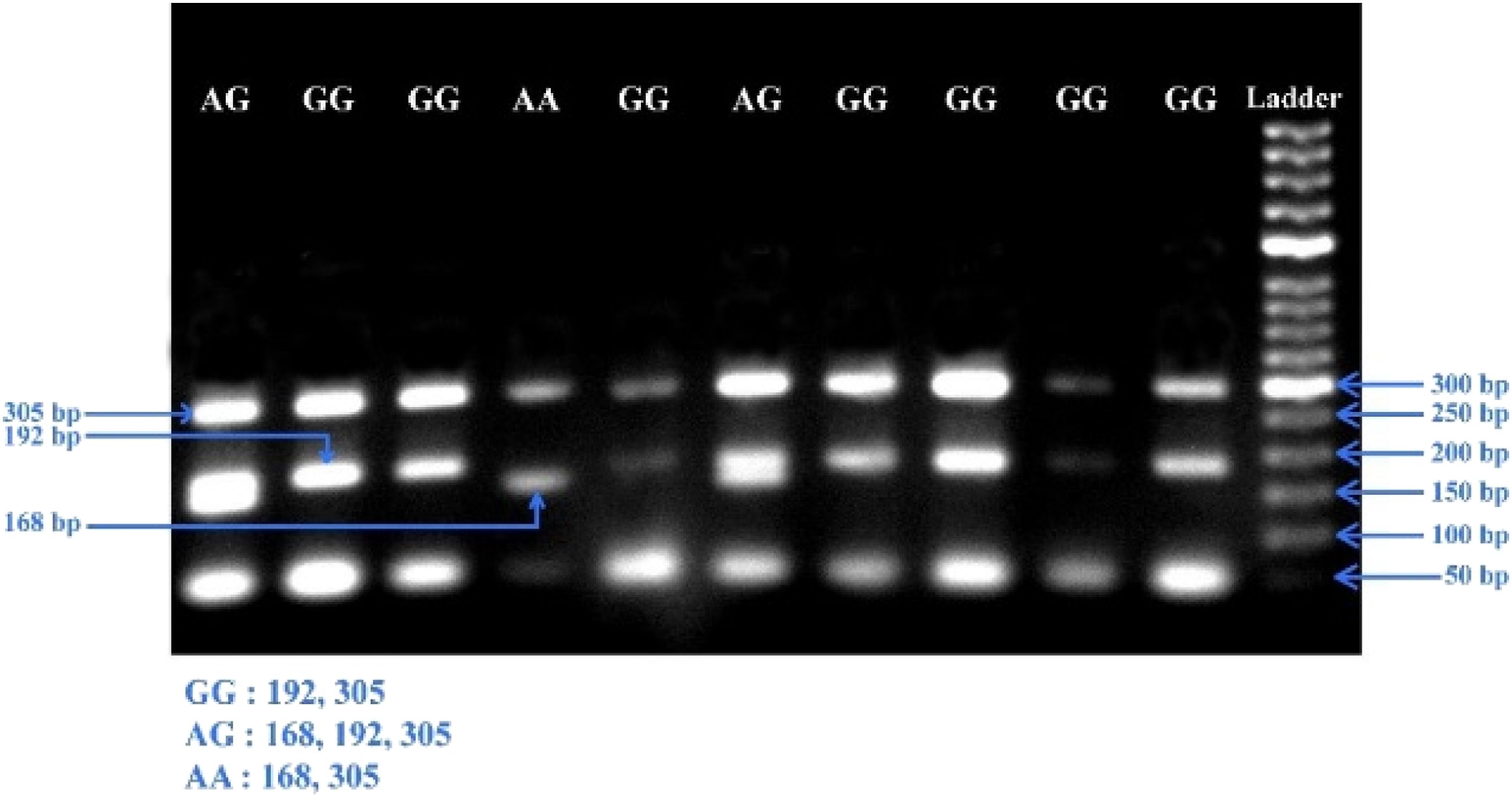
Statistical Analysis
All statistical analyses were carried out applying the statistical software package SPSS (v. 25.0) and MedCalc (v. 20.0). The differences between demographic and clinicopathologic information of cases and volunteers were contrasted by applying χ2-tests and two-sided unpaired t-tests. The association of rs10484879 and rs3748067 with CRC susceptibility was examined using odds ratios (ORs) along with the corresponding 95% confidence intervals (CI). The deviation of genotype frequencies was measured under the Hardy-Weinberg equilibrium (HWE) by applying the χ2-test. Online Sample Size Estimator (OSSE) was used to calculate the statistical power of sample size for this case-control association study. Analysis of the false-positive report probability (FPRP) was utilized to assess the noteworthy relationships of the significant findings.49,50 The FPRP threshold was set at .2, and a prior probability of .1 was provided for a relationship with the investigated genotypes. The P < .05 was regarded statistically significant in this study.
Results
Distribution of Demographics in Participants
Demographic and Clinicopathological Characteristics of Colorectal Cancer Patients and Controls.

BMI: Body mass index; TNM: Tumor node metastasis; CRC: Colorectal cancer; SD: Standard deviation.
Genotype Frequencies of Participants
Distribution of Genotype Data for rs10484879 and rs3748067.
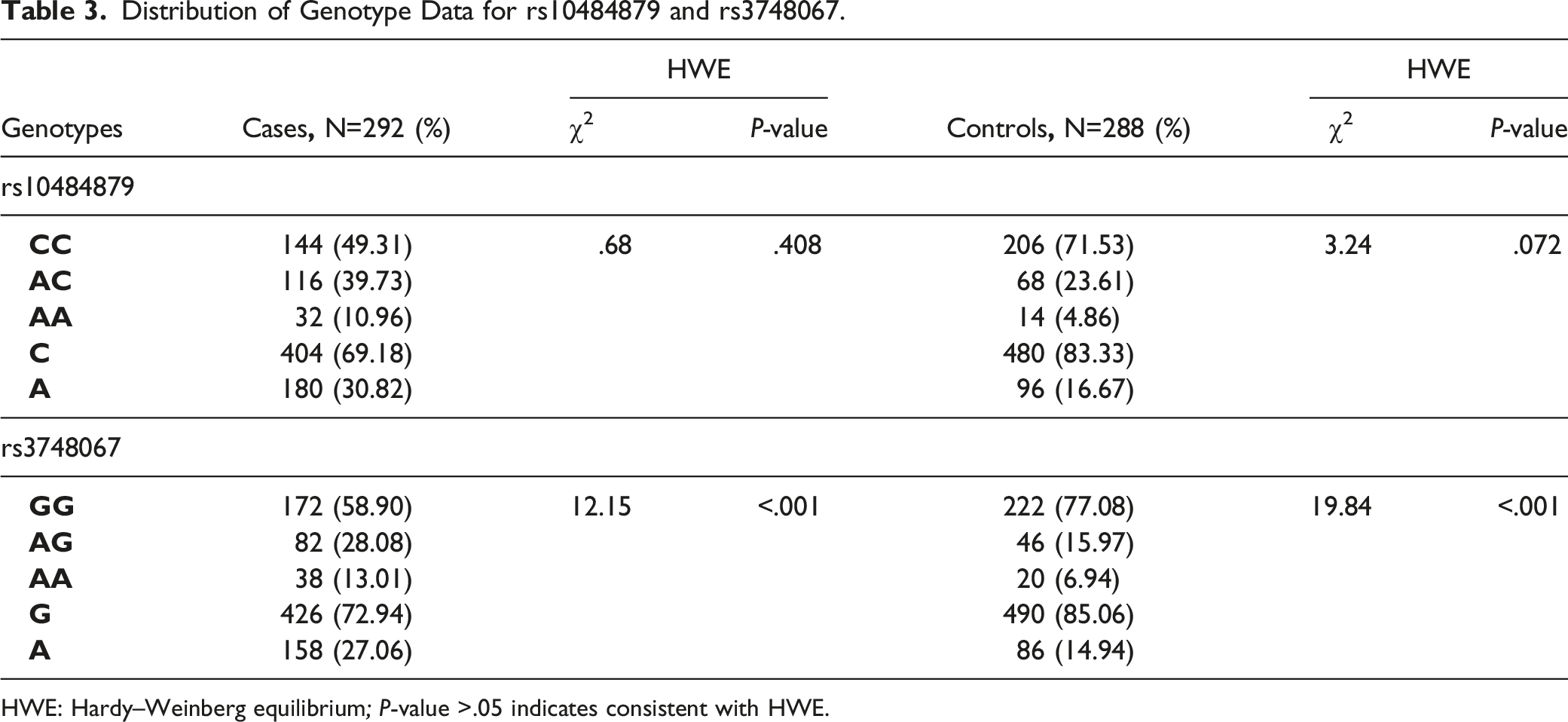
HWE: Hardy–Weinberg equilibrium; P-value >.05 indicates consistent with HWE.
Association of rs10484879 and rs3748067 With CRC
Association of rs10484879 and rs3748067 Polymorphisms with Colorectal Cancer.
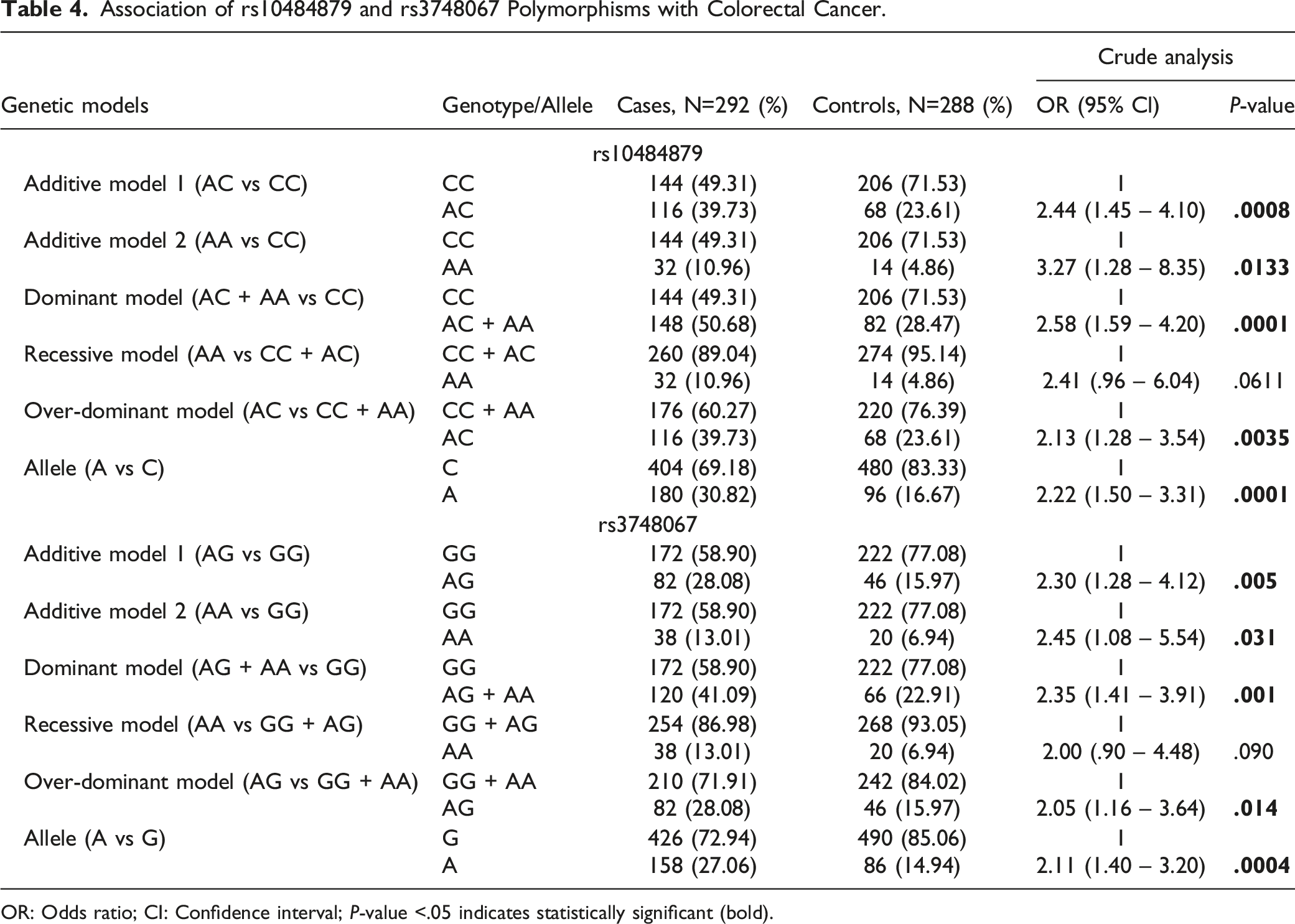
OR: Odds ratio; CI: Confidence interval; P-value <.05 indicates statistically significant (bold).
In the case of rs3748067, it was found that AG genotype carrier showed 2.30-times significantly elevated association with CRC (AG vs. GG: OR = 2.44, P = .005). Again, AA genotype depicted 2.45-times enhanced risk, and the association was significant (AA vs. GG: OR = 2.45, P = .031). The dominant model showed 2.35-folds more risk for CRC development in terms of GG genotype, and the association was also statistically significant (AG + AA vs. GG: OR = 2.35, P = .001). In the recessive model, AA genotype showed 2.00-folds enhanced risk for CRC development, and the association was non-significant with respect to the GG + AG genotype (AA vs. GG + AG: OR = 2.00, P = .090). The over-dominant model revealed 2.05-times elevated risk for CRC development, and the association was statistically significant (AG vs. GG + AA: OR = 2.05, P = .014). In the allelic model, A allele showed 2.22-times enhanced risk for CRC development, and the risk was significantly associated with CRC (A vs. G: OR = 2.11, P = .0004).
False-Positive Report Probability Values for the Association Between rs10484879 and rs3748067 with Colorectal Cancer Risk.

aStatistical power was calculated using the number of observations in the subgroup and the OR and P values in this table.
The results in false-positive report probability analysis were in bold if the prior probability <.2.
Discussion
Previous investigations have analyzed various polymorphisms in multiple genes that may be associated with CRC susceptibility. The IL-17A genetic polymorphisms have also been evaluated in different studies to correlate with CRC development. Keeping the possibility of different outcomes due to population or regional variation, we conducted this case-control study in the Bangladeshi population to examine the susceptibility of CRC by two common IL-17A genetic polymorphisms, including rs10484879 and rs3748067.
SNP rs10484879 in the IL-17A gene is located on the 3’-untranslated region (UTR) in chromosome 6 position 52187159. IL-17A gene rs10484879 has been extensively studied due to its potential association with different diseases in different ethnic populations. Previous studies with rs10484879 have been performed in various diseases, such as vulvovaginal candidiasis, 51 hepatitis B virus infection, 52 psoriasis, 53 periodontal disease and peri-implantitis. 54 The connection between IL-17A rs10484879 polymorphism and CRC susceptibility was studied for the first time in 2013. 41 Bondurant and colleagues conducted a case-control study in Northern California and Utah to look at the effects of IL-17A gene rs10484879 polymorphism on the risk of colon tumor (total subjects = 3,511) and rectal cancer (total subjects = 1,713) utilizing the information from two population-dependent pilot studies. In that study, particularly for rs10484879, GT + TT combined genotype carrier was at .76-times more susceptibility for the progression of colon tumor than GG genotype carrier and it was statistically significant (ORs = .76; P-value .011). Then in 2018, Bedoui and colleagues 40 further carried a case-control study on the Tunisian population to correlate the IL-17A gene rs10484879 polymorphism with CRC. The study included 293 CRC patients and 268 healthy controls who were recruited, matching age, sex, and BMI. The researchers demonstrated that the minor allele frequency in rs10484879 in CRC patients was considerably greater than in control subjects. Moreover, the minor allele of rs10484879 was associated with a previous family background of CRC and related malignancies (P = .002), CRC classification (P = .044), chemo reaction (P = .001), and therapy (P = .038). The study reported that, CT genotype and TT genotype carrier had 2.63- and 1.27-times more risk for CRC (CT vs. CC: OR = 2.63; P < .001 and TT vs. CC: OR = 1.27; P < .001). Again, dominant association model (OR = 2.43; P < .001) showed significantly elevated risk for CRC. In the present case-control study, our analysis also revealed that in case of rs10484879, additive model 1 (P = .0008), additive model 2 (P = .0133), dominant model (P = .0001), over-dominant model (P = .0035), and allelic model (P = .0001) showed the enhanced association for the development of CRC and all of the genetic models are statistically significant.
IL-17A gene rs3748067 polymorphism is located on the 3’-untranslated region (UTR) in chromosome 6 position 52190541. This SNP has been extensively studied due to its potential association with different diseases in different ethnic populations. Previous studies with r3748067 in various diseases are asthma, 55 breast cancer, 44 cervical cancer,56,57 CRC,40,58 congestive heart failure, 59 coronary artery disease,60,61 gastric cancer,30,62 gastrointestinal diseases, 63 rheumatoid arthritis, 64 laryngeal cancer, 65 lung cancer, 66 otitis, 67 pneumoconiosis, 68 psoriasis, 69 tuberculosis,70-72 ulcerative colitis, 73 and viral myocarditis. 74
The connection between IL-17A rs3748067 genetic variant and susceptibility of CRC has been studied for the first time in 2018. 40 The study by Bedoui and colleagues evaluated the influences of IL-17A gene variants on the susceptibility of CRC by a pilot study on the Tunisian population. The study included 293 CRC patients and 268 healthy controls and examined the link of CRC susceptibility with six IL-17A polymorphisms (rs10484879, rs3819024, rs3748067, rs3819025, rs7747909, and rs2275913) in Tunisians. They showed that AG and AA genotype carrier was at .56-times and 2.42-times significant risk for the development of CRC (AG vs. GG: OR = .56; P = .003 and AA vs. GG: OR =2.42; P = .003). In terms of the dominant model and recessive model, the AG + AA genotype showed a significantly .67-times decreased risk for CRC susceptibility (AG + AA vs. GG: OR = .67; P = .04). In the recessive model, AA genotype carrier depicted 2.83-folds significantly elevated risk for CRC (AA vs. GG + AG: OR = 2.83; P = .04). However, no further studies evaluated the role of this polymorphism in other populations. To evaluate the effect of rs3748067 on CRC, we performed this study in our population. Our case-control study revealed that rs3748067 is associated with significantly increased CRC susceptibility under the additive model 1 (P = .0008), additive model 2 (P = .031), dominant model (P = .001), over-dominant model (P = .014), and allelic model (P = .0004) in the studied population.
It would be noteworthy to mention that we have performed this association study in Bangladesh for the first time. Before this, IL-17A rs10484879 had been studied in two countries, and rs3748067 had been studied in only one country. Notably, our results are compatible with the previously performed investigations, but further genotyping research in different ethnic populations is needed to confirm the link between this SNP and CRC susceptibility.
At the same time, there were some limitations in our study, which should be resolved in the upcoming studies. The number of participants involved in the study was comparatively low, which we thought was not large enough to represent the actual scenario of the country. Again, we could not find all the associated data of the patients, which may affect the risk. Moreover, we have selected only a known SNP from the public database. Furthermore, the interaction of this polymorphism with other modifiable risk factors has not been addressed. As a result, further studies need to be done to confirm these findings.
Conclusion
To summarize, our study highlights that IL-17A rs10484879 and rs3748067 polymorphisms may be associated with colorectal cancer development. However, further functional research with larger samples may reveal more statistically significant outcomes.
Footnotes
Acknowledgments
The authors would like to acknowledge the Department of Pharmacy, University of Asia Pacific for their full support during this study.
Author Contributions
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This research received a partial grant from the Department of Pharmacy, University of Asia Pacific.
Ethical Approval
Ethical permission was taken from the National Institute of Cancer Research and Hospital (NICRH), and Dhaka Ahsania Mission Cancer and General Hospital.
Data Availability
All data related to the manuscript are provided in the manuscript main file, figure, and tables. The corresponding author will provide additional information on a valid request if required.
