Abstract
Background
The search for rare elements, like mitotic figures, is crucial in pathology. Combining digital pathology with eye-tracking technology allows for the detailed study of how pathologists complete these important tasks.
Objectives
To determine if pathologists have distinct search characteristics in domain- and nondomain-specific tasks.
Design
Six pathologists and six graduate students were recruited as observers. Each observer was given five digital “Where's Waldo?” puzzles and asked to search for the Waldo character as a nondomain-specific task. Each pathologist was then given five images of a breast digital pathology slide to search for a single mitotic figure as a domain-specific task. The observers’ eye gaze data were collected.
Results
Pathologists’ median fixation duration was 244 ms, compared to 300 ms for nonpathologists searching for Waldo (P < .001), and compared to 233 ms for pathologists searching for mitotic figures (P = .003). Pathologists’ median fixation and saccade rates were 3.17/second and 2.77/second, respectively, compared to 2.61/second and 2.47/second for nonpathologists searching for Waldo (P < .001), and compared to 3.34/second and 3.09/second for pathologists searching for mitotic figures (P = .222 and P = .187, respectively). There was no significant difference between the two cohorts in their accuracy in identifying the target of their search.
Conclusions
When searching for rare elements during a nondomain-specific search task, pathologists’ search characteristics were fundamentally different compared to nonpathologists, indicating pathologists can rapidly classify the objects of their fixations without compromising accuracy. Further, pathologists’ search characteristics were fundamentally different between a domain-specific and nondomain-specific rare-element search task.
Introduction
The search for rare elements, such as mitotic figures and tumor cells in lymph nodes, is a critical task in pathology. 1 The enumeration and identification of these elements directly influence the diagnosis, grading, and staging of various cancers. 2 Despite the importance of rare-element search, little is known about how pathologists perform these search tasks. One reason for this is that the analog workflow in pathology makes it difficult to accurately assess how pathologists search for rare elements without relying on a qualitative explanation from the pathologist themselves (eg, think-aloud studies, where pathologists dictate their thought process during their diagnostic assessment).3–5 This qualitative explanation is limited to the conscious efforts made by the pathologist in their search; however, it has been suggested that eye movements during a search task are governed both by conscious and unconscious components.6–8
The evolution from analog to digital pathology allows us to overcome this limitation. Pairing digital pathology with eye-tracking technology offers an opportunity to gain a deeper understanding of how pathologists complete visual search tasks, allowing us to gather quantitative eye gaze data that provides information beyond what can be consciously understood and reported by the pathologist. One such area of interest is pathologists’ diagnostic search for rare elements in digital images.9–12 Specifically, pathologists hone their search skills over years of training and practice, reviewing tens of thousands of slides per year. With experience and repeated exposure to cases, pathologists fine tune and further develop these search skills, which raises the question of whether this training and experience alters their approach to search tasks outside of pathology (ie, nondomain search tasks). The use of nondomain-specific search tasks, such as solving rare-element search puzzles, may be useful in gaining a deeper understanding of how pathologists perform rare-element search.
Since 1987, Handford has been illustrating detailed and crowded scenes of figures, each of which hides a unique character, named “Waldo,” to be found. 13 To increase the difficulty of the puzzles, many scenes also contain several characters designed to look like Waldo, but with minor differences. These characters are intended to trick the reader and are referred to herein as “Waldo confounders”. Together, these puzzles form the famous “Where's Waldo?” picture book series where the objective of the game is to find Waldo in every scene. It is worth noting that this book series has expanded internationally since its creation in the United Kingdom as “Where's Wally?”. As such, the name of the book series slightly differs depending on the region of the world; however, the objective of the puzzles remains the same. 14 For North America specifically, the book series is referred to as “Where's Waldo?” and is what will be used throughout this paper.
The “Where's Waldo?” puzzles have served as a useful general rare-element search task in the field of radiology by offering insight into the search patterns and perceptual skills of radiologists. Specifically, Sahraian et al conducted a study to determine if the accuracy and speed of radiology residents’ diagnostic search of chest x-rays could be improved by practicing “Where's Waldo?” puzzles; however, no such relationship was found. 15 Further, Nodine et al compared radiologists’ performance to that of a lay audience when tasked with searching for Waldo, but found radiologists’ visual training offers no advantage over that of the lay audience. 16 Like radiology, the field of pathology engages one's perceptual search skills in the diagnostic assessment of histopathologic tissue, in which some pathology tasks involve a rare-element search. However, performance on art-based search tasks such as “Where's Waldo?” has yet to be measured in the field of pathology.
The task of “Where's Waldo?” shares several qualities with rare-element search tasks in pathology, one example being the search for mitotic figures. Briefly, a mitotic figure is a cell undergoing division, and the quantification of these figures is used to evaluate the proliferative activity and grade of many types of cancer.17,18 Both the Waldo and mitotic figure search tasks take place within a crowded and dense scene populated with objects similar to the ones being searched for. For pathology specifically, there are many potential mimics of mitotic figures. These include apoptotic bodies, karyorrhectic debris, hyperchromatic cells, inflammatory cells, and tissue artifacts.17,19 These are comparable to Waldo confounders and ultimately make the search task more difficult.
Eye tracking allows for the qualitative and quantitative assessment of visual scan paths, which are the pathway one's eyes follow during visual search.20–23 Specifically, an eye tracker is a device that uses near infrared light to measure where the user is looking on the screen in real time. 24 In the context of eye tracking, there are two types of eye movements: fixations and saccades. A fixation can be defined as the maintenance of one's eye gaze on a single location.25–27 The objects of a fixation are viewed using one's foveated vision, in which visual stimuli are detected by the center of the retina, known as the fovea. 28 The fovea is the region of the eye that can obtain the highest visual acuity, which allows one to see the objects of their fixations clearly, and is represented by a 1° arc length around the point of fixation.29,30 Meanwhile, saccades are rapid eye movements that abruptly change the point of fixation.
In this study, we used eye-tracking technology to assess how pathologists search for rare elements in domain-specific and nondomain-specific search tasks. The scan paths of pathologists were compared between a domain-specific and a nondomain-specific search task and were also compared to those of nonpathologist observers for the nondomain-specific search task. This pursuit is driven by an overarching long-term goal of gaining a deeper understanding of how pathologists go from novice observers to highly skilled physicians for visual image search. Specifically in this paper, we took the first steps in addressing this overarching goal by: determining whether pathologists’ scan paths are different from those of nonpathologists, for a nonpathology rare-element search task (“Where's Waldo?”); and determining whether pathologists’ scan paths are different for rare-element search in a nonpathology task (“Where's Waldo?”), compared to a pathology task (mitotic figure search).
Materials and Methods
Observers
Six board-certified pathologists and six graduate students were recruited as observers. The graduate students selected had no formal experience in digital image search. The only exclusion criterion for observers in this study was the use of progressive lenses in the form of glasses or contact lenses, because these negatively affect the precision and accuracy of eye tracking.
Study Design
This study involved two types of tasks undertaken by the observers: the nondomain-specific search task for Waldo and the domain-specific search task for mitotic figures. Images from the “Where's Waldo?” picture book series were selected as the nondomain-specific search task, since this is an activity with which neither observer group would have specific expertise. “Where's Waldo?” was chosen over other “look and find” books in children's literature (eg, I Spy) because the target of the search task (ie, Waldo) remains the same throughout different images in the picture book, similar to how the target of the mitotic figure search task (ie, mitotic figures) remains the same throughout different slides. Conversely, other “look and find” books in children's literature often change the target one is to search for between different images. Therefore, all observers were given a brief introduction to the “Where's Waldo?” search task and subsequently shown a set of five digital images from the picture book series. Observers were informed at the beginning of this study that the goal was to find Waldo in every image, and that each image may or may not contain Waldo. Unknown to the observers was that only three of the five images contained Waldo, which was confirmed by the consensus of three individuals who were not observers in this study. All images were digitally scanned in color and cropped to a square with side length of 1080 pixels. Specifically, images were taken from books one, four, and five of the series corresponding to the “30th Anniversary,” “In Hollywood,” and “The Wonder Book” editions, respectively.31–33 Images were selected to range in difficulty and minimize artifacts introduced by digitization of the printed pages.
Following the five “Where's Waldo?” images, pathologists were then given a set of five images from a digital hematoxylin and eosin–stained breast pathology slide taken from The Cancer Genome Atlas to serve as their domain-specific search task. Each pathology image was cropped to a square with a side length of 1080 pixels and contained 1 mm2 of tissue. Pathologists were informed at the beginning of the study that the goal for this task was to find a single mitotic figure in each image, and that each image may or may not contain a mitotic figure. Specifically, images were selected to contain exactly one mitotic figure or zero mitotic figures. Unknown to the pathologists was that only three of the five pathology images contained a mitotic figure, which was confirmed by the consensus of two board-certified pathologists who were not observers in this study.
The domain-specific and nondomain-specific search tasks of this study were designed to induce uncertainty into the search tasks by including a subset of images that did not include the target figure to be found. The purpose of this was to have the study best mimic a realistic pathologist's rare-element search in a clinical workflow, as a pathologist would not know if there was something to be found in the tissue. As such, it is reasonable to assume that one's search strategy may change when they know there is something to be found in the image. Thus, to preserve the uncertain nature in rare-element search from the clinical workflow, target-absent images were included in both the domain-specific and nondomain-specific search tasks of the study.
For both the domain-specific and nondomain-specific search tasks, observers were given an unlimited amount of time to complete the task and were given verbal feedback on their answers for each image with the opportunity to continue searching or give up if they were incorrect. Observers sat approximately 60 cm from the screen and were not allowed to pan or zoom the images in or out. Ethics approval was not required for this study as all observers are coauthors of this study and all digital pathology slides used in this study are publicly available.
Hardware
All images were displayed on a 27-inch, 1920 × 1080 Dell monitor (model SE2722H, Dell). The Tobii Pro Fusion (Tobii Technologies) was used as the eye-tracking system, operated at 120 Hz and set up using the tripod configuration at the base of the monitor.
Software
The Tobii Pro Eye Tracker Manager (version 2.6.1, Tobii Technologies) and Tobii Pro Lab (version 1.217.49450, Tobii Technologies) applications were used to set up and calibrate the eye tracker. All participants underwent a nine-point calibration and subsequent four-point validation scheme before starting the study. A screen-based project was created in Tobii Pro Lab to sequentially present the images of the study to each participant using the timeline feature of the software. Tobii Pro Lab was also responsible for the data collection from the eye tracker using the Velocity-Threshold Identification Gaze Fixation Filter.
Eye-Tracking Data
The data collected from an eye tracker is a temporal sequence of fixations and saccades, collectively known as a scan path.34,35 A precise definition of scan paths is provided in Supplemental Section A. Scan paths were recorded for each observer throughout each search task to measure the following scan path features: fixation duration, saccade velocity, saccade amplitude, fixation rate, saccade rate, gaze transition entropy, false-negative rate, false-positive rate, directional saccade angle, percent image area covered by foveated vision, cumulative area of fixation overlap, and vector to target arc length. A precise definition of the meaning of each of these features is in Table S1 in Supplemental Section B.
For the purpose of this study, it is important to define the following terminology and symbols, which will be used throughout this paper to analyze and interpret scan paths. We define a scan pattern to be a qualitative description of a scan path (eg, left-to-right raster scan path, top-to-bottom raster scan path, sequential fixations on nearby characters in a Waldo image or within a discrete region of tumor in a pathology image). We then define Ftarget as the first fixation on target (eg, Waldo or a mitotic figure), and F−X as the fixation immediately before the observed deviation from a scan pattern, where this deviation leads to Ftarget. The “X” in F−X quantifies the number of fixation points F−X is from Ftarget (eg, if the fixation immediately before the observed deviation from a scan pattern leading to Ftarget is three fixation points before Ftarget, then F−X = F−3). In the event no deviation from the scan pattern was observed, or in cases in which no recognizable scan pattern was seen, the fixation immediately preceding Ftarget was recorded (ie, F−1). For more details in how these data were partitioned, see Table S1, the instructions, and videos 1 and 2 in Supplemental Sections B, C, and D, respectively. With this, vector to target is defined as the vector connecting Ftarget and F−X.
Data Analysis
The Kolmogorov-Smirnov test was used to check for normality, and the Mann-Whitney U test was used for comparative analysis of the measurements listed in Table 1. Bonferroni correction was applied for all statistical tests with α = 0.004.
Group Median Eye Gaze Data and Interquartile Ranges Collected From the Domain-Specific (Mitotic Figure Search) and Nondomain-Specific (Waldo Search) Tasks of the Study.
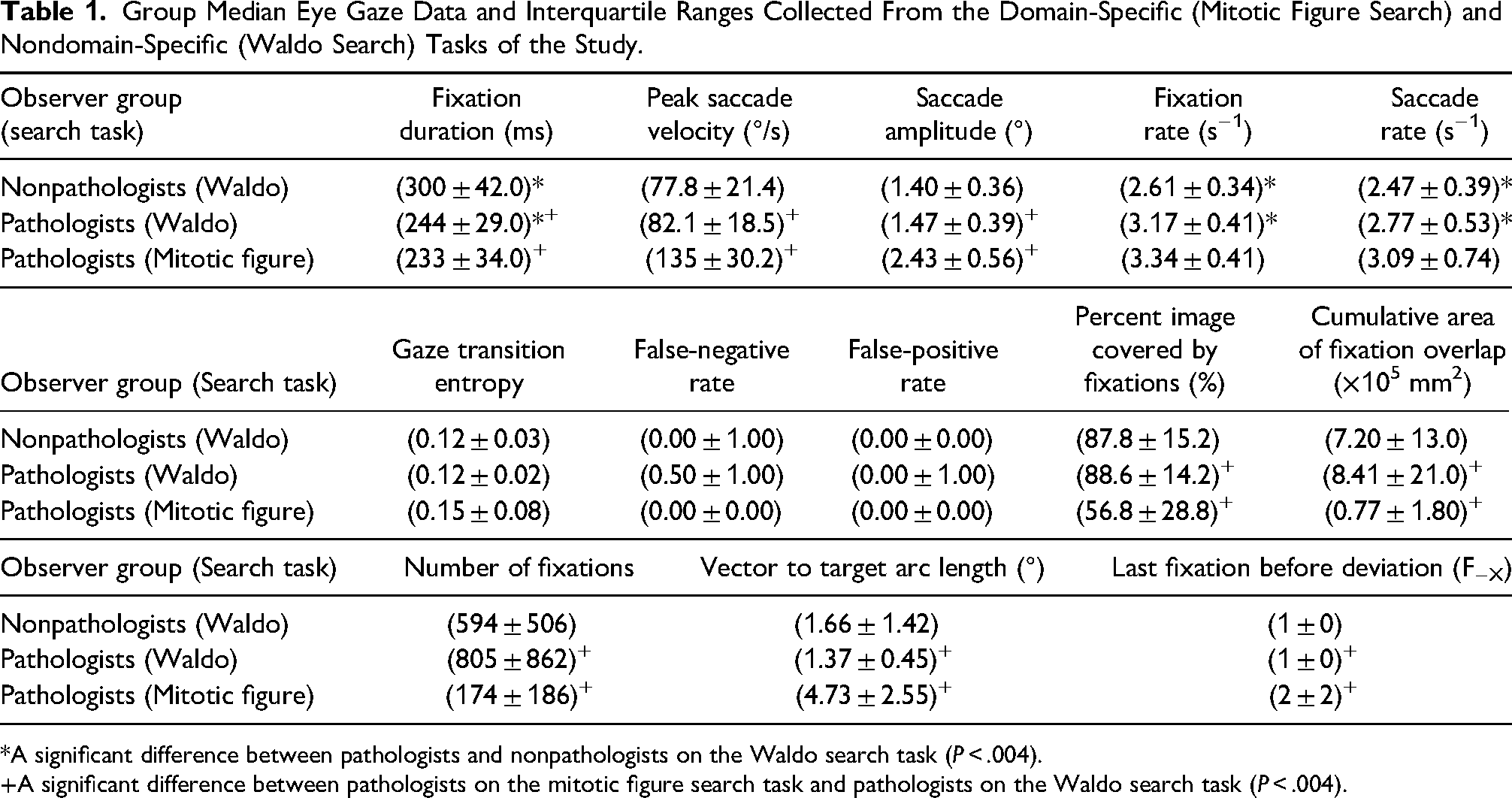
*A significant difference between pathologists and nonpathologists on the Waldo search task (P < .004).
A significant difference between pathologists on the mitotic figure search task and pathologists on the Waldo search task (P < .004).
Results
The Kolmogorov-Smirnov results revealed that the data were predominately not normally distributed. As a result, we used the median and interquartile range to describe the data and the Mann-Whitney U test to test for significance. The group median eye gaze data collected from this study are shown in Table 1.
Pathologists Versus Nonpathologist Observers
For the nondomain-specific rare-element search task (Waldo; top two rows of Table 1), we found that pathologists had a significantly shorter median fixation duration than nonpathologist observers (P < .001). However, the median peak saccade velocity and median saccade amplitude were not statistically significantly different between pathologists and nonpathologist observers (P = .02 and P = .14, respectively). Pathologists’ median fixation rate was greater than that of the nonpathologist observers (P < .001), as was their saccade rate (P < .001). These results are consistent with the visual representation of scan paths between the two groups. For example, Figure 1 compares the scan paths of a nonpathologist observer (L1) and a pathologist observer (P1) when searching the same “Where's Waldo?” image. As shown in the figure, this pathologist's scan path has a higher density of fixations per square area compared to the nonpathologist's search. Also shown in this figure is a lower density scan path in the upper left corner for both observers, which is due to the “Where's Waldo?” image having a text description of that image in that region.
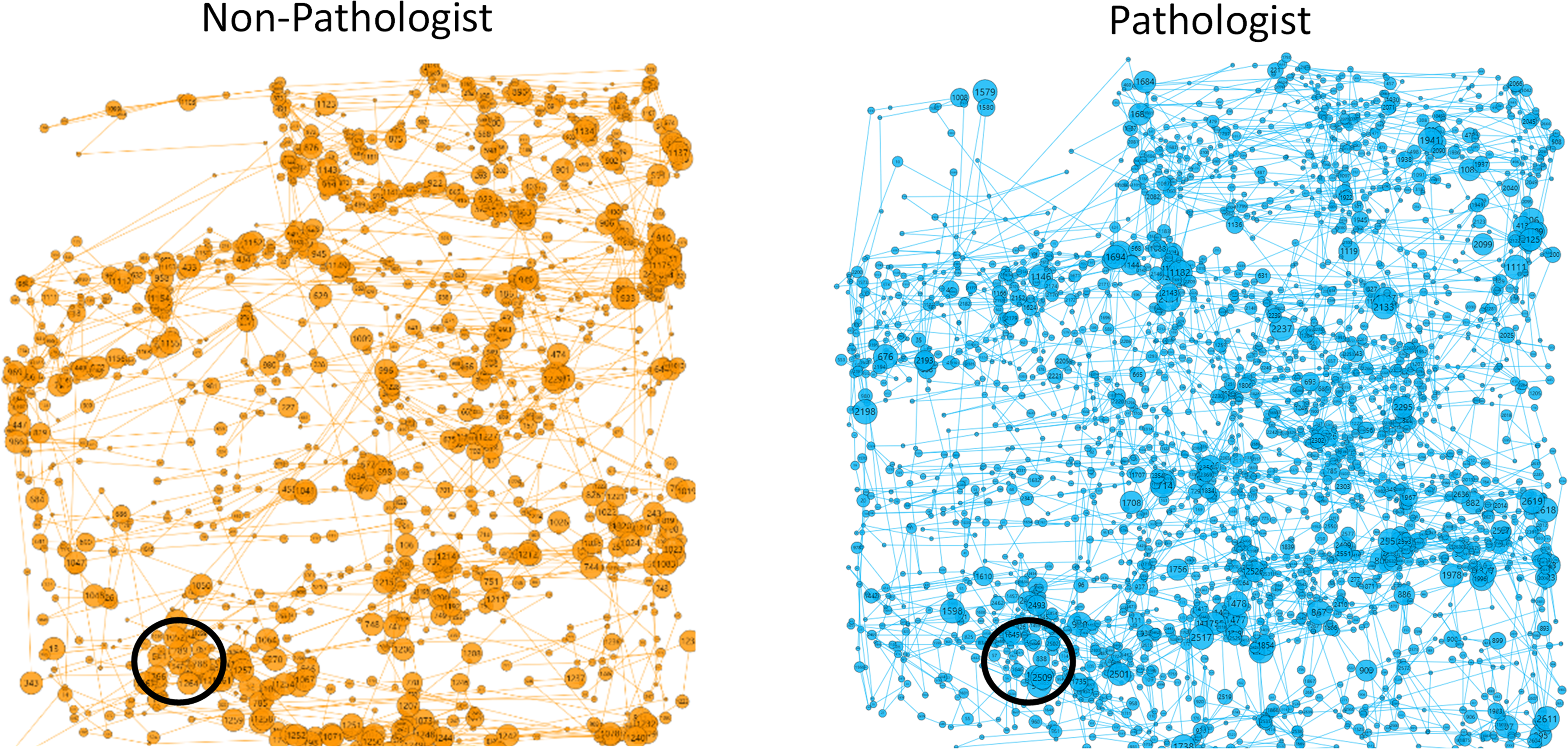
Scan paths for the same “Where's Waldo?” puzzle. Circles represent fixations, radius linearly proportional to fixation duration. Lines represent saccades, length indicating saccade amplitude. Waldo is circled in black. Representative scan path of a nonpathologist observer, L1 (left, orange). Representative scan path of a pathologist observer, P1 (right, blue). The total size of each square image as shown on the screen was 13 cm × 13 cm.
Median gaze transition entropy was equivalent among the pathologists and nonpathologist observers at a value of 0.12 (P = .24). There was also no significant difference for both the false-negative rate and false-positive rate between observer groups (P = .48 and P = .40, respectively). To assess the geometric persistence in scan path for pathologists versus nonpathologist observers, their directional saccade angle (θd) values were computed and are compared in Figure 2. For reference, θd ≈ 0° and θd ≈ 360°represents two consecutive saccades moving in the same direction (ie, persistent), whereas θd ≈ 180° represents two consecutive saccades moving in opposite directions (ie, antipersistent). A specific example of the set of θd for the same Waldo image of a single nonpathologist observer (L1) and pathologist observer (P1) are provided in Supplemental Section E. The corresponding time lapsed videos of L1 and P1's scan paths are provided as videos 3 and 4, respectively, in Supplemental Section D for reference.
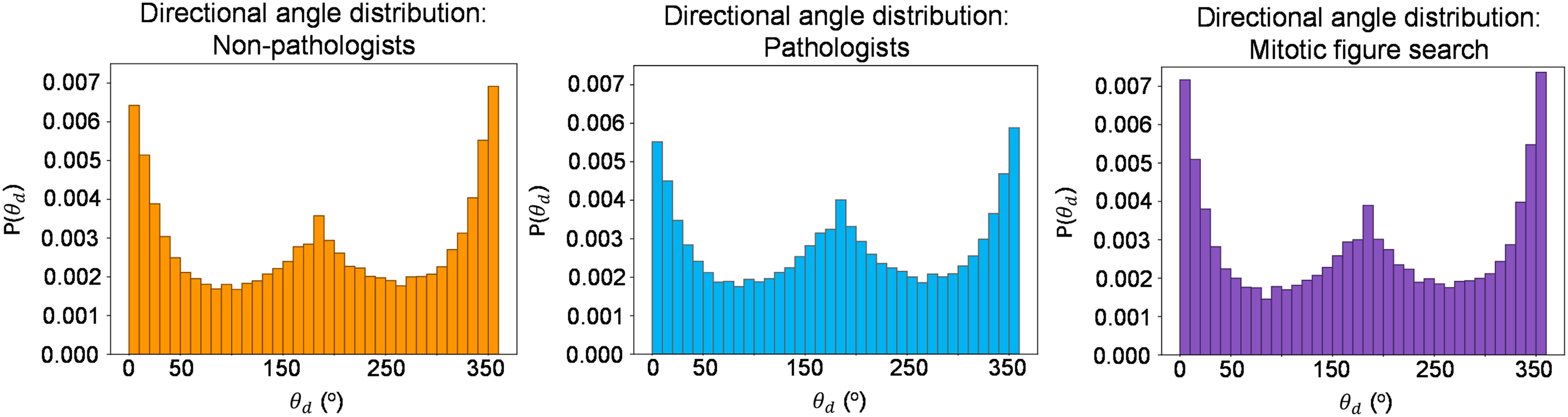
Nonpathologists’ collective directional angles searching for Waldo (left, orange). Pathologists’ collective directional angles searching for Waldo (middle, blue). Pathologists’ collective directional angles searching for mitotic figures (right, purple).
Domain-Specific Versus Nondomain-Specific Search Tasks
The bottom two rows of Table 1 compare how pathologists search for rare elements in a pathology specific search task (mitotic figure) compared to a nonpathology specific search task (Waldo). Compared to pathologists’ scan paths on the Waldo task, the mitotic figure task was associated with a greater saccade amplitude (P < .001), less area covered by foveated vision (P < .001), lower fixation duration (P < .001), and higher saccade velocity (P < .001). There was no statistically significant difference in fixation rate nor saccade rate.
Consistent with these results, Figure 3 compares the scan paths of a single pathologist (P5) for one Waldo image and one pathology image. The domain-specific scan path for this pathologist, shown on the right, is visually sparser than their nondomain-specific scan path shown on the left, signifying that this pathologist covered less square area with their foveated vison for the domain-specific search task. Further, the fixation duration is shorter and the saccade amplitude is longer on average for this pathologist's domain-specific search compared to the nondomain-specific search.
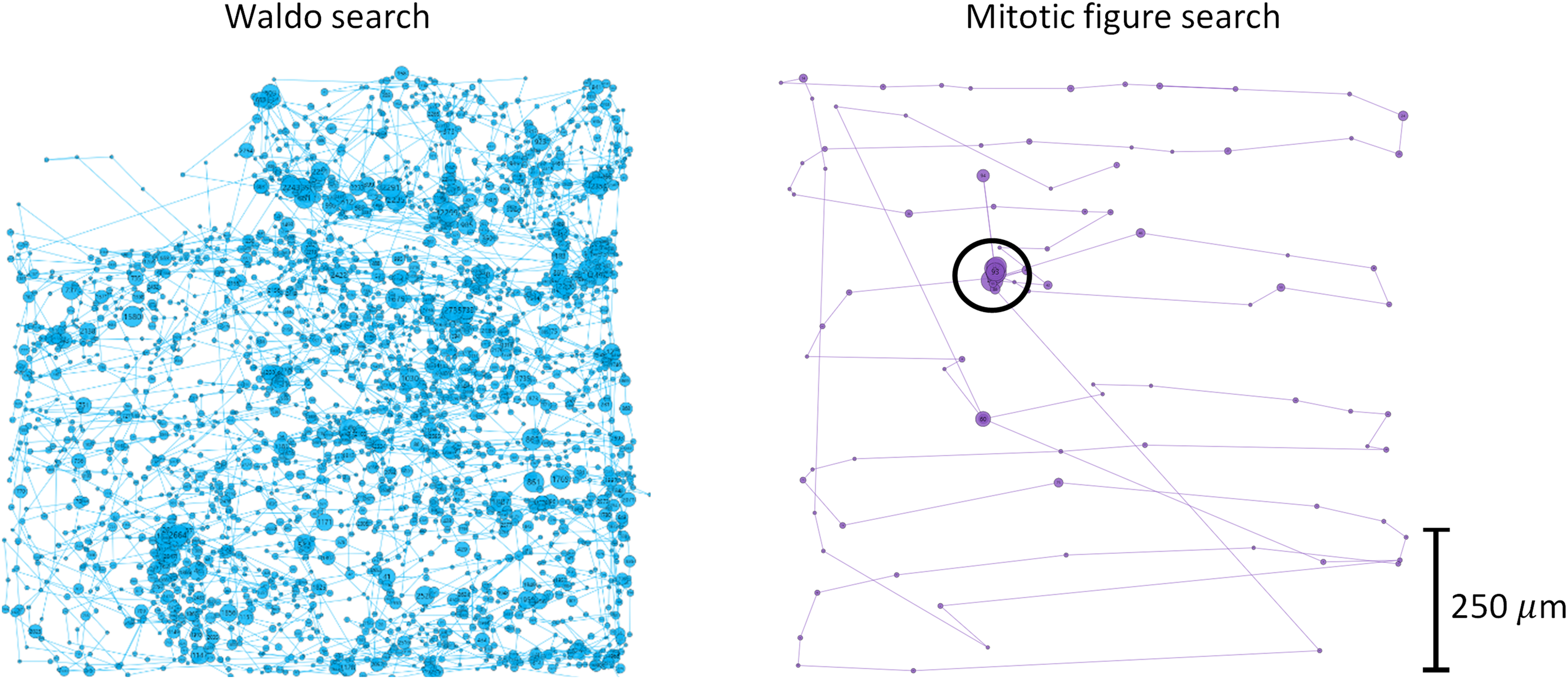
Scan paths from P5 observer for the nondomain search task (left, blue) and domain search task (right, purple). Circles represent fixations, radius linearly proportional to fixation duration. Lines represent saccades, length linearly proportional to saccade amplitude. Target of search is circled in black on the right. No target was present in the Waldo search on the left in this example. The scale bar on the right shows the physical tissue size of the digital breast pathology slide. The total size of each square image as shown on the screen was 13 cm × 13 cm.
There was no statistically significant difference in median gaze transition entropy between the two groups (P = .10), nor was there a statistically significant difference in their false-negative rate and false-positive rate (P = .02, P = .81, respectively). Figure 2 above compares the geometric persistence of pathologists’ search for Waldo to their search for mitotic figures.
We examined the vector to target arc length to gain an understanding of observers’ abilities to detect the target at different distances away from a current fixation point. Figure 4 is a visual representation of each vector to target measurement for every instance where an observer fixated on the target for the first time. The data were then partitioned into instances where the observer deviated from their scan pattern (depicted as the green arrows in Figure 4) and where the observer did not deviate from their scan pattern (depicted as the black/gray arrows in Figure 4). For pathologists’ search for Waldo, only one vector to target measurement was from a deviation in scan pattern (Figure 4, left). Conversely, every vector to target measurement came from a deviation in scan pattern with pathologists’ search for mitotic figures (Figure 4, right). As a result, the difference in vector to target arc length between these two search tasks is statistically significant (P < .001), and the median F−X value for pathologists’ search for Waldo and for mitotic figures was F−1 and F−2, respectively (P = .003).
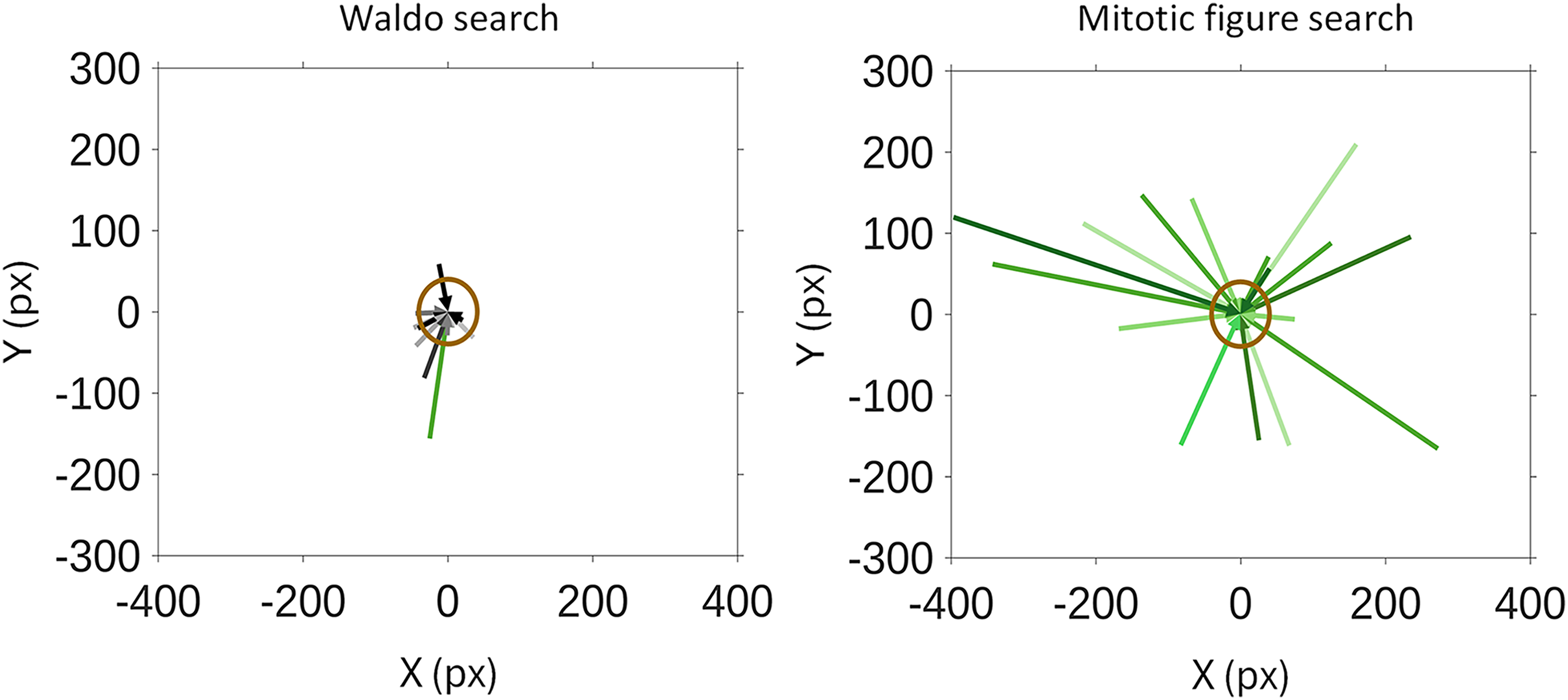
Vector to target measurements for each pathologist observer in their search for Waldo (left) and mitotic figures (right). Green arrows represent vector to target measurements when a deviation from scan pattern was observed. Black/gray arrows represent vector to target measurements where no deviation in scan pattern was observed. The target of each image in both search tasks was normalized to the centers of the figures at (0,0) pixels. A 1° arc length around the target of the image is shown as a brown circle, which represents the amount of image visible in one's foveated vision when fixating on the target.
Discussion
This is the first study to assess the generalizability of pathologists’ scan paths in searching for rare elements outside their domain (“Where's Waldo?”) compared to those of nonpathologist observers, and with comparison to their scan paths for a domain-specific rare-element search task (mitotic figures).
This study showed that for nondomain-specific search tasks, pathologists have higher saccade rates and fixation rates, with shorter fixation durations compared to nonpathologists. This indicates that pathologists saccade more frequently than nonpathologists as they search. Further, the short fixation durations suggest that pathologists classify the objects of their fixations faster than nonpathologists with no decrease in accuracy (indicated by the similar false-negative and false-positive rates between groups). This raises the possibility that the training and experience a pathologist receives for diagnostic search of rare elements leads to a generalizable ability to rapidly classify the objects of their fixations without compromising accuracy.
Past literature for eye tracking in digital pathology supports the observed differences between pathologists and nonpathologist observers in the current study. Specifically, several studies reported that pathologists with greater experience had shorter fixation durations and longer saccade amplitudes for rare-element search tasks in pathology, when compared to pathologists with less experience.3,36–42 These results suggest that pathologists become more efficient in their search through experience, allowing them to search faster without compromising their diagnostic accuracy. The present study extends these observations to a nondomain-specific task.3,36–42 With the support of existing literature, this study demonstrates that pathologists’ eye movement characteristics associated with rare-element search tasks translate to search tasks outside pathology.
These observed differences in pathologists’ scan paths were further amplified when assessing them on a task within their domain. Specifically, pathologists had statistically significantly shorter fixation durations with faster saccade velocity and larger saccade amplitude when searching for mitotic figures compared to their search for Waldo. Despite this, there were no statistically significant differences in the two groups’ false-negative and false-positive rates, implying that pathologists can classify the objects of their fixations more rapidly within their domain than outside their domain without compromising accuracy. This is supported by the learning theory in psychology which states that one's performance relies on the similarity of the training and testing situations. 43 As pathologists gain rare-element search experience primarily through pathology-related search tasks, this explains pathologists’ superior performance in searching for mitotic figures versus Waldo.
A qualitative analysis of θd for pathologists and nonpathologist observers for the nondomain search task (Figure 2) revealed that both groups had directional peaks around 0° and 360°, signifying that they predominantly searched with sequential saccades in a consistent direction. Additionally, both pathologists and nonpathologist observers had a directional peak around 180°, implying both groups changed the directionality of their saccades with similar consistency. These results indicate that the directional eye movements of pathologists are very similar to those of nonpathologists when given the same search task; however, in relation to the observations made above, pathologists perform these movements faster than nonpathologists, without compromising the accuracy of their search. A similar observation was made comparing θd for pathologists’ nondomain-specific and domain-specific rare-element search (Figure 2).
The differences in visual scan path between a pathologist and a nonpathologist observer in this study are shown in the pair of scan paths illustrated in Figure 1 (also see time-lapsed videos of both scan paths provided as videos 3 and 4 in Supplemental Section D). Qualitatively, the nonpathologist observer's scan path is a loose search of the image with no particular pattern. Conversely, the pathologist applies a methodical approach to their search as their scan path immediately undergoes a series of vertical and horizontal raster scans across the image for the entire duration of the search. Despite searching differently however, both groups had an equivalent median gaze transition entropy, signifying a low complexity and high predictability in scan path for the “Where's Waldo?” search task.
With respect to pathologists’ domain-specific and nondomain-specific search tasks, there was a substantial difference in the amount of the image covered by foveated vision. Pathologists needed to cover about 89% of the image with their foveated vision to find Waldo, whereas they only needed to cover about 57% of the image to find the mitotic figure with much shorter fixation durations and longer saccades. Additionally, pathologists had a more complex and less predictable scan path when searching for mitotic figures as shown by the increase of 0.03 in gaze transition entropy from the Waldo search (Table 1); however, this difference was not statistically significant.
Pathologists covered significantly less of the image with their foveated vision and had significantly fewer fixations in their search for mitotic figures compared to their search for Waldo; this implies that pathologists in this study were using a mechanism other than foveated vision to effectively detect the presence of mitotic figures. One mechanism to explain this may be the use of peripheral vision. To investigate this, we measured the vector to target arc length for each observer on each search task. As shown in Table 1 and Figure 4, the median vector to target arc length is significantly larger for pathologists searching for mitotic figures than their search for Waldo. Specifically, pathologists had a median vector to target arc length of about 4.7° when searching for mitotic figures, which is well beyond the range of foveated vision (1°) and borders between parafoveal and peripheral vision.29,30 Additionally, all but one vector to target measurement for pathologists’ search for Waldo did not deviate from a scan pattern, whereas every vector to target measurement came from a deviation in scan pattern for pathologists’ mitotic figure search. Conversely, no significant difference was observed in the vector to target arc length between pathologists and nonpathologist observers for the nondomain-specific search task (median vector to target arc length of about 1.4° and 1.6°, respectively), implying that the pathologists in this study have no inherent ability to deviate from their scan pattern toward a nondomain-specific target outside their foveated vision. These results lead to a speculation that pathologists, through experience, may be able to detect mitotic figures in their peripheral vision, leading to more efficient eye movements and coverage of the image. This is a surprising finding, given that nonfoveal vision is less than 20/20, and potentially as poor as 20/400.44,45 This speculation therefore requires further testing in a subsequent study specifically designed to test the effect of peripheral vision in rare-element detection in pathology. There is extensive literature that exists to support the idea that peripheral vision is good for detecting motion and motion changes46–50; however, there is considerably less literature concerning peripheral vision and object recognition.51–53 The results of this study demonstrate a need to further explore the effect of peripheral vison in object detection and the corresponding effect on rare-element search tasks in pathology.
Together, the results of this study imply that some components of pathologists’ domain-specific scan paths may translate to nondomain-specific tasks. These components included elements corresponding to the mechanics of how the eye moves (eg, fixation duration, fixation rate, and saccade rate). However, elements corresponding to the overall scan path (eg, percent image area covered by foveated vision, cumulative area of fixation overlap, gaze transition entropy, directional saccade angle, and vector to target arc length) did not differ between pathologists and nonpathologists.
There are several limitations to this study. The exclusion criteria against the use of progressive lenses for the observers of this study posed a risk to bias the results toward a younger population of pathologist observers, since progressive lenses are typically worn by individuals 40 years of age or older. 54 To overcome this limitation, it is possible for a future study to provide single vision glasses with the observer's prescribed refractive correction if they use progressive lenses in their day-to-day life; however, such an approach would often require the observer to undergo a neural adaptation period for adjusting to the new glasses. Further, the observers did not undergo a comprehensive eye exam as part of this study, which could have revealed an underlying eye condition that may have affected one's foveated vision (eg, amblyopia, anomalous retinal correspondence). It is also important to note that the results from the domain-specific to nondomain-specific comparison in this study may be affected by the magnification level of the digital images, as the characters in the Waldo images were not the same size as the cells in the pathology images. Additionally, these results may be affected by the difficulty of directly comparing mitotic figures to Waldo images due to the relative difficulty of these tasks likely not being the same. To overcome these limitations, it is possible for a future study to design a nondomain-specific rare-element search task such that the figures in the image are of similar size to those in the domain-specific rare-element search task. Further, this search task can be designed to have similar contrast to what is seen in pathology slides to make the relative difficulty of the two search tasks more equal. Overall, although these are necessary limitations when conducting a study comparing a search on pathology images to nonpathology images, they lead to caution in interpreting the results of these comparisons.
Conclusion
This study demonstrates that pathologists’ scan paths are different from those of nonpathologists for a nondomain-specific rare-element search task (ie, searching for rare elements in nonpathology images), and different from their scan paths in search for rare elements in a domain-specific task (ie, searching for rare elements in pathology images). These results suggest that, in part, pathologists’ training and experience in diagnostic rare-element search translates to tasks outside of pathology, suggesting there are fundamental differences in how pathologists search for rare elements. Understanding the nature of these differences yields important insights: scan paths are particular to both the observer group and the type of rare-element search task; and, compared to nonpathologists, pathologists have the ability to more rapidly classify the targets of their fixations without compromising the accuracy of their search. These insights assist with identifying commonalities in the visual search characteristics of experienced pathologists. This has potential implications to refine and develop training resources for trainee pathologists by identifying the qualities of experienced pathologists’ visual scan paths that lead to efficient rare-element search. Identifying these qualities would then allow for the targeted development of these efficient visual search skills in trainee pathologists through eye gaze–guided training resources.
Supplemental Material
sj-pdf-1-ijs-10.1177_10668969241294239 - Supplemental material for Identification of Distinct Visual Scan Paths for Pathologists in Rare-Element Search Tasks
Supplemental material, sj-pdf-1-ijs-10.1177_10668969241294239 for Identification of Distinct Visual Scan Paths for Pathologists in Rare-Element Search Tasks by Alana Lopes, Sean Rasmussen, Ryan Au, Vignesh Chakravarthy, Tricia Chinnery, Jaryd Christie, Bojana Djordjevic, Jose A. Gomez, Natalie Grindrod, Robert Policelli, Anurag Sharma, Christopher Tran, Joanna C. Walsh, Bret Wehrli, Aaron D. Ward and Matthew J. Cecchini in International Journal of Surgical Pathology
Footnotes
Additional Information:
Please be aware that a preliminary abstract of this manuscript was presented at the 2024 Pathology Informatics Summit in Ypsilanti, Michigan, United States of America on May 22, 2024.
Coauthor Sean Rasmussen has moved to a new institution after completing the research. Their new affiliations are:
Department of Pathology and Laboratory Medicine, Queen Elizabeth II Health Sciences Centre, Halifax, NS B3H 3A7, Canada
Department of Pathology, Dalhousie University, Halifax, NS B3H 4R2, Canada
Data Availability Statement
The information related to the data analysis as part of this study is available upon request.
Declaration of Conflicting Interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval and Informed Consent Statements
Not applicable, because all observers are coauthors of this study and all digital pathology slides used in this study are publicly available.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
Supplemental Material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
