Abstract
Three Treponema sp.—T. medium, T. pedis, and T. phagedenis—are associated with bovine digital dermatitis (BDD). This painful infectious disease has been reported worldwide, causing economic and animal welfare problems. Although detection by real-time PCR (rtPCR) is the method of choice, published assays differ in specificity and sensitivity. We multiplexed target regions of 2 published rtPCR assays and slightly optimized another one to compare their performance. We then used the 2 novel TaqMan rtPCR assays to determine the prevalence of the 3 Treponema sp. in a random sample of hind feet from 137 animals collected at slaughterhouses. An overall prevalence at the animal level of 31.4% was found for T. phagedenis and 10.2% for T. medium, with the latter always being detected together with T. phagedenis. We did not detect T. pedis in our sample set. Typical lesions of BDD were present in 16.2% of animals. Of the animals with BDD lesions, 52.6% were positive for any of the 3 Treponema sp. A significantly lower positive rate for Treponema sp. was found in animals without lesions, underscoring the role of these organisms in the development of BDD. Our study offers insight into the prevalence of BDD-associated Treponema sp. in cattle as well as a tool for routine diagnosis and future control of the disease.
Bovine digital dermatitis (
Clinically, BDD typically is an ulcerative or proliferative lesion on the plantar skin of a hindlimb hoof, proximal to the interdigital cleft between the heel bulbs.15,30 The disease leads to pronounced pain and lameness, which seriously impairs animal welfare. In addition, economic losses are incurred as a result of reduced milk yield, reduced fertility, weight loss, and treatment and monitoring costs.6,8,35 In Switzerland, the prevalence of BDD at the animal level was 29.1% in 2014 3 and fell slightly to 20.7% by 2021. 24 In a 2024 study, a prevalence of 5.4% of BDD stage M2 was found at the cow-level in Swiss herds. 19
The current gold standard for diagnosis of BDD is based on visual clinical classification with stages M0–M4.1.5,13 ELISA testing of bulk tank milk for Treponema antibodies can be used to monitor BDD prevalence at the herd level.23,36 Culture and classical identification of BDD-associated Treponema sp. is cumbersome, expensive, and labor-intensive. Molecular detection is the method of choice, but it is hampered by the lack of genome data 18 and, thus, by a lack of knowledge about specific (virulence) genes that could be targeted to achieve species identification and separation. The difficulty in culturing Treponema sp. is a reason for this deficit. Therefore, many species might still be unknown; data from various studies and GenBank entries indicate the presence of yet uncultivated members of the genus.11,26,33
Various real-time PCR (rtPCR) methods have been developed to improve accuracy in the detection of Treponema sp. involved in BDD, as well as to reduce the workload in the laboratory.2,4,17,38 However, these rtPCR approaches have room for improvement. One method 38 is based on a singleplex approach, in which a separate rtPCR is performed for each treponemal target species and is therefore costly and work-intensive. Problems have been identified 17 with cross-reactions in a triplex assay 2 and a lack of amplification of T. medium and T. pedis in a 4-plex assay. 4 The assay of one study 17 only detected T. phagedenis in 51.7% of biopsies of stages M1–M4.1, but was more sensitive than a modified version of another rtPCR, 38 which resulted in 3.4% fewer T. phagedenis–positive samples.
In view of the high prevalence of BDD in Switzerland, we have modified existing methods to establish a reliable rtPCR for the simultaneous detection of T. phagedenis, T. medium, and T. pedis as a routine detection tool.
Materials and methods
Selection of primers and probes
We evaluated primers and probes of published rtPCR assays in addition to considering reported results. For each of the 3 most recent rtPCR assays,4,17,38 optimal primers and TaqMan probes for corresponding target genes were determined (Primer Express software v.3.0.1; ThermoFisher), assessing all available and compatible reporter and quencher combinations for putative probes. Optimized primers and probes were then checked for specificity and sensitivity in silico using Blast analysis (https://blast.ncbi.nlm.nih.gov) based on all available data on target gene sequences for the 3 Treponema sp. in GenBank (accessed 2025 April 1). The most promising targets were then selected for a novel multiplex rtPCR assay (termed
Target genes and primers/probes used in our study of Treponema sp. in cattle.
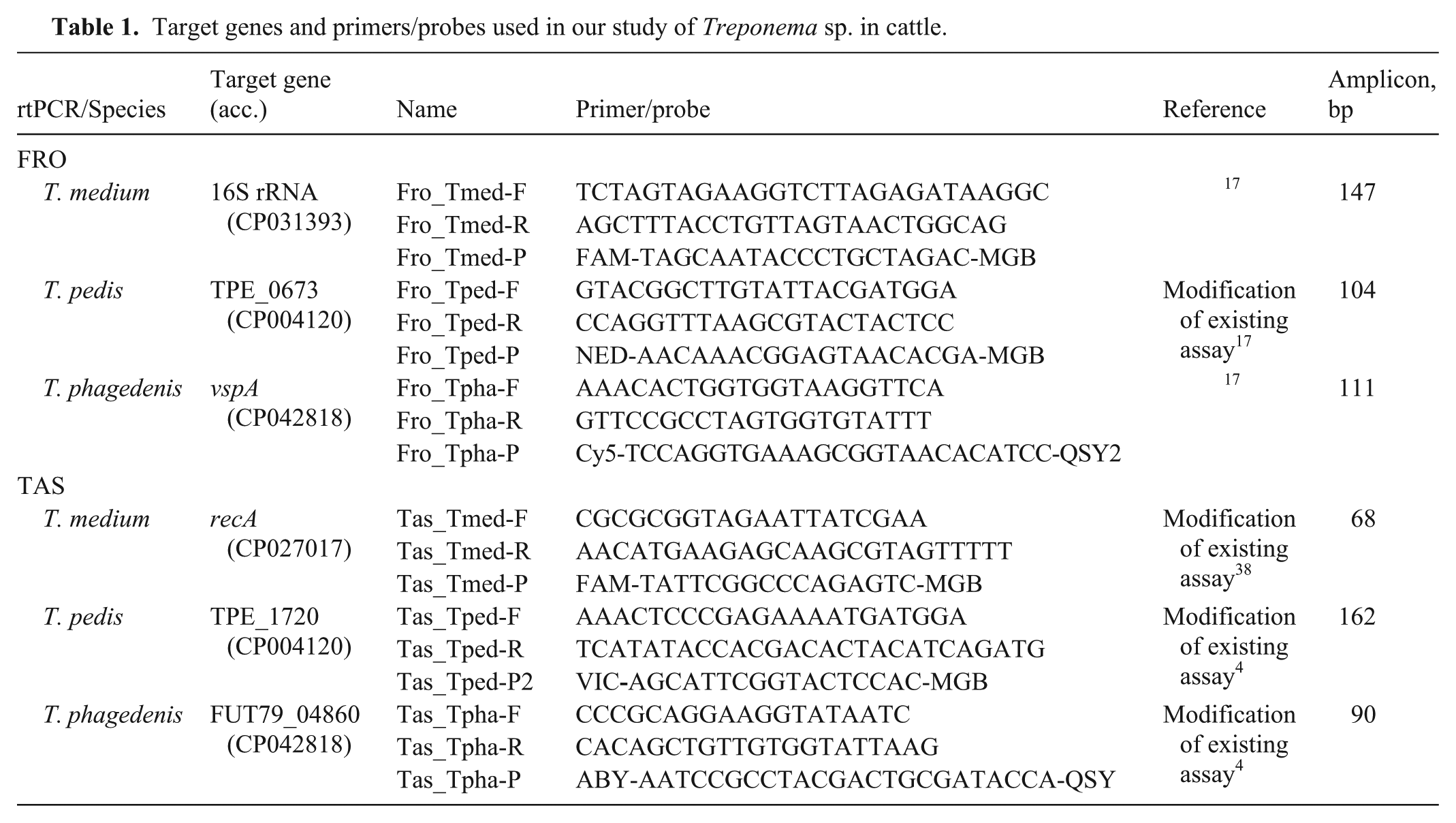
Real-time PCR assays
The rtPCR was carried out in a total volume of 25 μL containing master mix (1× TaqPath BactoPure microbial detection master mix; ThermoFisher), 150 nM primers, 250 nM TaqMan probes, 1 μL of VetMAX Xeno internal positive control-VIC assay (ThermoFisher), and 2.5 µL of template DNA. Alternatively, the VetMAX Xeno internal positive control-LIZ assay (ThermoFisher) was used if necessary for fluorescence compatibility. The rtPCR was performed in a real-time PCR system (QuantStudio 5; ThermoFisher). The run program consisted of 30 s at 60°C, 2 min at 95°C, followed by 40 cycles with 10 s at 95°C, and 60 s at 60°C, with a post read for 30 s at 60°C, according to the manufacturer’s user guide. Software (Design & analysis software v.2.6.0; ThermoFisher) was used for the rtPCR run setup and analysis.
Synthetic double-stranded gene fragments containing all 3 amplified target sequences separated by “AATAA” were designed for each multiplex rtPCR assay (GenScript; Suppl. Table 1 ).
Analytical performance
The assays were tested with genomic DNA isolated from type strains and field strains of T. medium, T. pedis, and T. phagedenis, as well as the synthetic positive controls. Dilution series containing 2–2 × 106 DNA copies/µL were used to determine efficiency, linearity, repeatability, and limit of detection (LOD) in triplicates.
Hoof sample collection and DNA extraction
Convenience samples from 20 cattle collected in a 2023 study 29 and 117 additional samples obtained in 2025 May–Sep were included in our study ( Suppl. Table 2 ). Hind feet were collected at 2 abattoirs in the canton of Bern and transported to the laboratory at ambient temperature within 6 h and processed within 2 h of arrival. Hoofs were cleaned of dirt and visually inspected for lesions. If lesions were present, they were classified M1 to M4.1. Absence of lesions was classified as M0. A punch biopsy was then taken at the heel bulb, where BDD lesions are normally observed. The tissue was pierced with a 6-mm punch biopsy (KAI Europe) and cut with a scalpel into 3 smaller pieces on a sterile petri dish. These were placed in an Eppendorf tube containing 800 µL of lysis buffer (4 M guanidine thiocyanate, 0.01 M Tris-HCl, 1% beta-mercaptoethanol) and stored overnight at 4°C. DNA was then extracted from 500 µL of lysate by mixing with magnetic beads (MagneSil RED; Promega) and processing on a semi-automated extraction robot (KingFisher DuoPrime; ThermoFisher). 40 An internal positive control (IPC; VetMAX Xeno internal positive control DNA, 20,000 copies, ThermoFisher) was added to each sample before extraction to serve as an extraction and rtPCR control. 27 DNA was eluted in 80 µL of pyrogen-free water and stored at −20°C until further use.
Statistical analysis
The Fisher exact test with a value of significance p ≤ 0.05 was used for statistical analysis.
Results
Performance of FRO and TAS rtPCR assays
First, we tested the 2 rtPCR assays (FRO and TAS) in the multiplex rtPCR format as well as in individual singleplex reactions, with the corresponding synthetic controls providing equimolar target sequences for each amplification. Although Ct values for initial dilution series of T. phagedenis and T. medium controls were consistent between the 2 rtPCR assays, signals for T. pedis were always later, with up to ~10× loss of sensitivity. This was more distinct in the TAS rtPCR. We therefore changed the originally identical fluorescence of FRO and TAS probes (Table 1), which slightly improved the performance of the TAS rtPCR.
We then tested extended dilution series from 5–5 × 106 copies/reaction of synthetic controls. Both the FRO and the TAS multiplex rtPCR results were linear over 7 logs (
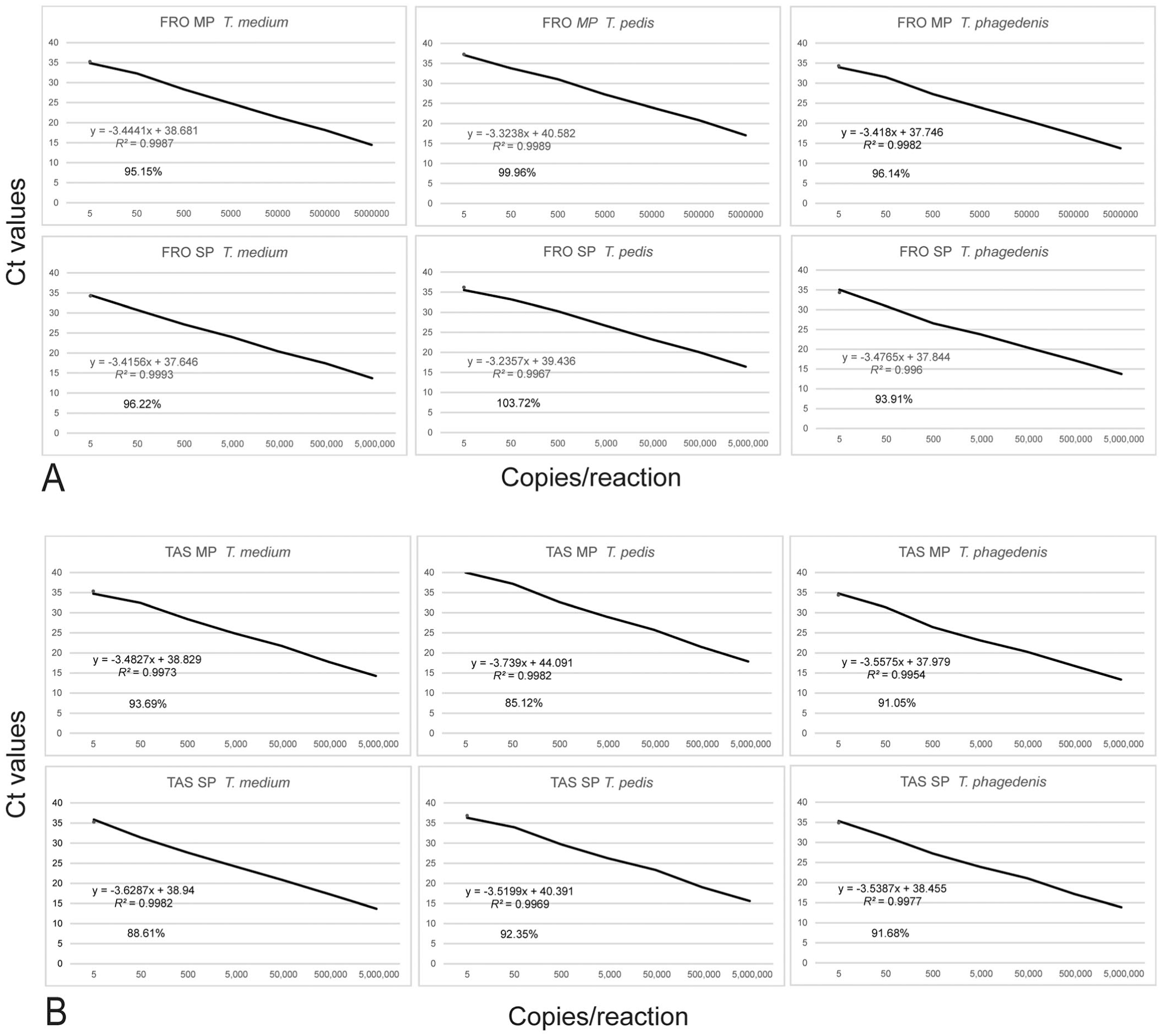
Comparison of
Prevalence of Treponema sp. in cattle
We used the FRO and TAS rtPCR assays for a study of the prevalence of BDD-associated Treponema sp. in Swiss slaughter cattle (Suppl. Table 2). Of the 137 animals tested, 40 (29.2%) were positive for T. phagedenis in both rtPCR assays, 1 was positive only in the FRO assay, and 2 were positive only in the TAS assay, resulting in an overall prevalence of 31.4%. Regarding T. medium, 13 animals were positive in both rtPCR assays, whereas the TAS assay detected an additional T. medium–positive animal, resulting in a T. medium prevalence of 10.2%. The 14 T. medium–positive animals were all positive for T. phagedenis. The TAS rtPCR assay detected T. pedis in 8 of 137 (5.8%) animals. However, the high Ct values (>37) indicated low amounts of target DNA or low specificity. We were able to sequence the 162-bp PCR product amplified from 2 animals with slightly lower Ct values (8 and 9; Suppl. Table 2). The 2 sequences were identical and were the correct size; however, the sequence only had 86% similarity to the target gene, including 2 mismatches in the probe. When performing Blastx, the best hit was 96% to the hypothetical protein of a Treponemaceae bacterium (GenBank MEL5719964) identified in the hoof lesion of an elk. We therefore conclude that the signals obtained for T. pedis in the TAS rtPCR assay resulted from nonspecific amplification of a gene homolog from an as-yet unidentified Treponema sp. in the absence of the specific target sequence.
The prevalence, as assessed by visual inspection, was 19 of 117 (16.2%) in animals for which such data were available (Suppl. Table 2). Fourteen feet from 9 animals had M1 lesions, 4 feet from 3 animals had M2, 4 feet from 4 animals had M3, 9 feet from 6 animals had M4 scores, and 1 foot from 1 animal had M4.1. Prevalence of Treponema sp. in animals that had limbs with typical lesions was 10 of 19 (52.6%), but was 26 of 98 (26.5%) in animals that had limbs without lesions, which was a significant difference (p = 0.031;
Summary results from the 137 animals tested for Treponema sp. in our study. Results are given for the 2 rtPCR assays (FRO and TAS) in relation to the presence of BDD lesions.
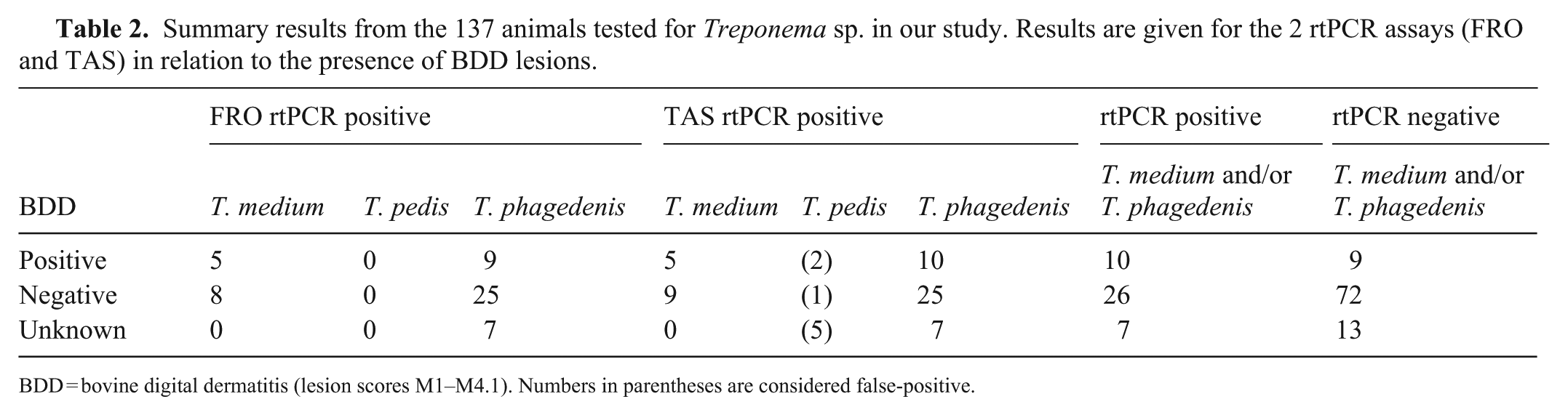
BDD = bovine digital dermatitis (lesion scores M1–M4.1). Numbers in parentheses are considered false-positive.
Discussion
We tried to optimize primers and probes for published rtPCR BDD-associated Treponema sp. targets, including the most promising published assay. 17 The latter rtPCR was used as a gold standard, given that, in our opinion, much effort had been invested during the assay development with respect to target genes, primers, and probes. 17 This assay was thoroughly evaluated with published data on specificity and sensitivity. 17 According to our routine protocols, we therefore only slightly improved and adapted this rtPCR by designing a significantly shorter probe for T. pedis, including an extraction control, and by using an optimized master mix.
As well, we tried to combine the “best of both worlds” into one by combining targets and optimizing primers and probes from 2 existing assays4,38 into the new TAS rtPCR assay. Compared with the FRO rtPCR assay, the TAS rtPCR assay has the advantage of also detecting human T. phagedenis and does not use the 16S rRNA gene as the target region. Genetic and phenotypic differences between cattle and human isolates of T. phagedenis have been reported.28,31 It is unclear if 2 distinct and host-adapted lineages of T. phagedenis exist, or whether human variants cannot be found in cattle. The use of the 16S rRNA gene in PCR might not be optimal, given that these genes are often highly conserved between related species, and the fact that many Treponema sp. might be undiscovered increases the risk for nonspecific amplification. We have observed such nonspecific amplification quite often with a nested PCR, 15 in which, after sequencing the 16S rRNA gene–based PCR products, we found uncultured Treponema sp. PT3 instead of T. phagedenis and Treponema sp. PT13 instead of T. pedis. 21
However, we found that not only the 16S rRNA, but also other genes might be prone to nonspecific amplification because of conserved primer/probe sequences. The TAS T. pedis rtPCR assay had such “false” positive amplification as a result of a probably yet uncultivated and unknown Treponemaceae species. This is clearly one of the drawbacks of our newly designed rtPCR. In addition, the performance, efficiency, and LOD of the TAS assay were generally lower than the FRO rtPCR assay. We could not show the assumed advantage of detecting (human) variants of T. phagedenis. Indeed, 2 animals were positive for T. phagedenis only in the TAS rtPCR assay, thereby increasing sensitivity. However, 1 animal was positive only in the FRO rtPCR assay; all 3 reactions were weakly positive at the LOD. In contrast to the original study, 17 we were able to detect T. medium in our sample set by the FRO rtPCR assay, further advocating the usefulness of this improved rtPCR assay.
Only clinical data are available on the prevalence of BDD in Swiss cattle. In our study, 16.2% of animals had lesions typical of BDD. This is somewhat lower than the 20.7% observed in a comprehensive study in 2021, which included more than 7,500 animals. 24 By sampling bovine feet from slaughter animals at the abattoir, we focused on beef cattle, with an average age of 28 mo. Lower prevalence of BDD in beef cows than in dairy cows has been reported,7,20 and therefore the general prevalence based on visual observation might be underestimated. The overall prevalence of Treponema sp. based on rtPCR in our study was 31.4%, about double the prevalence of lesions typical for BDD. A higher load of Treponema has been observed in active lesions,12,34 a situation not seen in our study. However, this might be because of the limited sample size we analyzed.
From the set of hindlimbs in which BDD scores were available, a significantly higher portion of affected animals was positive for Treponema sp. compared with those in which lesions were absent. This is a further indication that the detected Treponema sp. are involved in BDD. It is unclear whether healthy carrier animals later develop the disease. In contrast to ovine foot rot, in which the finding of virulent Dichelobacter nodosus is an early indicator of disease, 39 a large knowledge gap still exists in the pathogenesis of BDD. Given the multifactorial character of BDD, it might be more difficult to identify such an early indicator for this disease. Further studies monitoring the prevalence of Treponema sp. and disease status over time would help provide a clearer picture. Improvements in molecular detection and its implementation in different laboratories will substantially contribute to that aim.
Supplemental Material
sj-pdf-1-vdi-10.1177_10406387261424230 – Supplemental material for Detection of Treponema species associated with bovine digital dermatitis in cattle using novel real-time PCR assays
Supplemental material, sj-pdf-1-vdi-10.1177_10406387261424230 for Detection of Treponema species associated with bovine digital dermatitis in cattle using novel real-time PCR assays by Laura Tassera, Isabelle Brodard, Jörg Jores, and Peter Kuhnert in Journal of Veterinary Diagnostic Investigation
Footnotes
Acknowledgements
We thank all participating slaughterhouses and their employees for their help in acquiring cattle samples.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This study was supported by the research fund of the Institute of Veterinary Bacteriology, University of Bern.
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
