Abstract
Trichomonosis is a venereal disease of cattle caused by the protozoan Tritrichomonas foetus. T. foetus infection in cattle herds can be economically costly for cattle producers; therefore, testing is important for detection of the agent. Given that bulls are considered to be subclinical carriers of T. foetus, it is important to detect T. foetus infection prior to movement and/or breeding season. We have described previously the development of an updated set of PCR primers and probes that offer increased sensitivity of T. foetus detection in preputial washings collected in PBS by utilizing reverse-transcription real-time PCR (RT-rtPCR) that targets the 5.8S ribosomal RNA of the T. foetus organism. Here, we report improvements in the updated RT-rtPCR reagents as well as the evaluation of testing of pooled preputial washings. We found that up to 5 preputial washings can be pooled, similar to routine testing practices (InPouch culture), without reducing the sensitivity of detection of T. foetus.
Trichomonosis, a sexually transmitted disease of cattle, is caused by infection with Tritrichomonas foetus, a flagellated protozoan parasite. T. foetus has a worldwide distribution and is endemic in areas where cows are bred by natural service, rather than by artificial insemination.12,15 T. foetus infection can cause significant economic losses to cattle producers because of the reduced reproductive performance of an infected herd, ranging from decreased calving rates and extended calving seasons, to increased costs as a result of culling and replacement of infected cattle, expenses associated with feeding and maintaining nonproductive cattle, and diagnostic testing for infection control within the herd.3,7,13,14 T. foetus–infected cows typically display clinical signs of disease, including vaginitis, abortion, infertility, and other reproductive issues.12,13,15,16 Most clinical signs clear within several weeks to months with reproductive rest, although permanent infertility is possible in some cows.4,12,13,15,16 Although several factors, including age, breed, and individual natural immunity, appear to play a role in T. foetus infection, 13 infection control is generally focused on the T. foetus status of bulls, given that bulls typically display no clinical signs of infection and are thus considered subclinical carriers of the agent. Unlike the European Union, which utilizes artificial insemination for infection control, the United States has established control programs that focus on testing and culling of positive bulls. 10
The traditional method for detection of T. foetus is culture of preputial washings, which is collected via scraping the preputial mucosa and inoculating into a culture medium (InPouch; Biomed Diagnostics); samples are then incubated at 37°C for up to 7 d and examined daily by light microscopy for the presence of motile organisms.12,16 Although culture of T. foetus is a relatively easy process, it requires several days to obtain a result, is costly given the requirement for the widely used culture medium pouches, and is prone to bacterial contamination that can inhibit growth of the organism. 1 In addition, culture inherently suffers from low sensitivity, given the requirement for motile organisms, and low specificity, because the pathogenic T. foetus cannot be visually distinguished from several other nonpathogenic, morphologically similar, trichomonad species that are also found in cattle.12,13,16 As a general practice, with culture, animals should have ≥3 negative tests, 1 wk apart, prior to classification as T. foetus–negative. 13
With the development of PCR, improvements have been made in testing methods to use either in conjunction with, or in place of, culture testing.4,11,15 We reported previously the development of an improved workflow and reverse-transcription real-time PCR (RT-rtPCR) reagents that target the T. foetus 5.8S ribosomal RNA from individual direct preputial washings collected in PBS, thus improving T. foetus detection in bovine preputial washings and eliminating the need for incubation in culture medium prior to testing. 8 However, we only evaluated the improved workflow using individual samples. We report here an update to the T. foetus workflow, which utilizes a commercial extraction kit (MagMAX CORE nucleic acid purification kit; Applied Biosystems), as well as an improved master mix chemistry (TaqMan fast virus 1-step master mix; Applied Biosystems) to enhance T. foetus detection. We used this updated workflow subsequently to evaluate testing of pooled preputial washings that had been collected in PBS.
T. foetus–positive and –negative preputial washings in PBS were collected over a 3-y period from specimens submitted to the Texas A&M Veterinary Medical Diagnostic Laboratory (College Station, TX, USA) for RT-rtPCR testing. Individual positive sample cycle threshold (Ct) values were 17.7–33.6. Samples had been stored at −20°C prior to use in our study. To evaluate the updated workflow, 20 individual T. foetus–positive samples were extracted, in triplicate, and tested using both the previously described workflow 8 and the new workflow, which utilized a commercial extraction kit (MagMAX CORE nucleic acid purification kit) and a newer master mix chemistry (TaqMan fast virus 1-step master mix). Total nucleic acid was extracted following the manufacturer’s instructions. Briefly, 100 µL of preputial washing in PBS was combined with 10 µL of proteinase K reagent and 720 µL of lysis binding solution (350 µL of CORE lysis solution, 350 µL of CORE binding solution, 20 µL of CORE magnetic beads, 2 µL of VetMAX Xeno internal positive control [Xeno RNA, Applied Biosystems; 10,000 copies/µL]) in a 96-well deep-well plate. Nucleic acid extraction was performed using an automated magnetic particle processor (KingFisher Flex; Thermo Scientific). The following plates were placed in the KingFisher Flex: sample plate, wash solution 1 plate (500 µL of wash solution 1/well), wash solution 2 plate (500 µL of wash solution 2/well), and elution buffer plate (90 µL of elution buffer/well).
Following nucleic acid extraction, samples were evaluated via RT-rtPCR, which utilized an improved master mix chemistry (TaqMan fast virus 1-step master mix), per the manufacturer’s instructions. T. foetus–Xeno primer/probe mix, which contains the same published T. foetus primers/probe targeting the T. foetus 5.8S ribosomal RNA, 8 as well as primers/probe targeting the VetMAX Xeno RNA, were combined with a master mix that contained all required RT-rtPCR components. Each reaction contained 6.25 µL of TaqMan fast virus 1-step master mix, 1 µL of 25 × T. foetus–Xeno primer/probe mix, and 12.75 µL of nuclease-free water. Each RT-rtPCR reaction contained 5 µL of extracted nucleic acid combined with 20 µL of complete master mix for a total reaction volume of 25 µL. All extracted samples contained Xeno RNA, at 20,000 cp/rxn, to monitor extraction and PCR efficiency. The RT-rtPCR was performed (7500 Fast thermocycler; Applied Biosystems), in standard mode, with the following cycle parameters: 50°C for 5 min (1 cycle), 95°C for 20 s (1 cycle), and 40 cycles at 95°C for 15 s and 60°C for 60 s.
For the evaluation of sample pooling, we pooled samples in groups of 5: 200 µL of each sample was combined for a total volume of 1 mL. For positive pools, 1 T. foetus–positive preputial washing was combined with 4 T. foetus–negative preputial washings (200 µL each); for negative pools, 5 negative preputial washings (200 µL each) were combined. All pools were vortexed thoroughly prior to further processing. We assessed 30 T. foetus–positive pools and 30 T. foetus–negative pools, in triplicate, with the improved workflow. We compared results with our published workflow 8 for both individual and pooled samples. Pooled samples with Ct values ≤38.0 were considered positive for T. foetus; a Ct ≤38.0 cutoff for pooled samples accounted for the ~2.5 Ct increase observed when samples were pooled at a 1:5 dilution and allowed for the detection of a weak-positive sample in a pool. Per laboratory protocol, any positive pools with a Ct value ≤38.0 would require follow-up testing of all samples within the pool to determine which sample(s) was T. foetus positive. Any pools with a Ct value >38.0 were classified as negative.
Positive T. foetus Ct values for the samples tested with our published workflow 8 were 24.8–37.9; exogenous internal positive control (XIPC) Ct values were 30.3–34.9 (x̄ Ct = 31.5). When tested with the updated workflow, the positive T. foetus Cts were 23.2–39.0; Xeno Cts were 29.2–32.8 (x̄ Ct = 30.6). Two of the positive samples were undetected utilizing our updated workflow; previous workflow Ct values were 35.7 and 36.5. The increase in Ct value is likely a result of sample degradation from long-term frozen storage at −20°C and/or freezing and thawing of the sample. Overall, samples tested with the new workflow demonstrated ~1.83 Ct mean difference in detection from our previously reported T. foetus workflow (Fig. 1A).
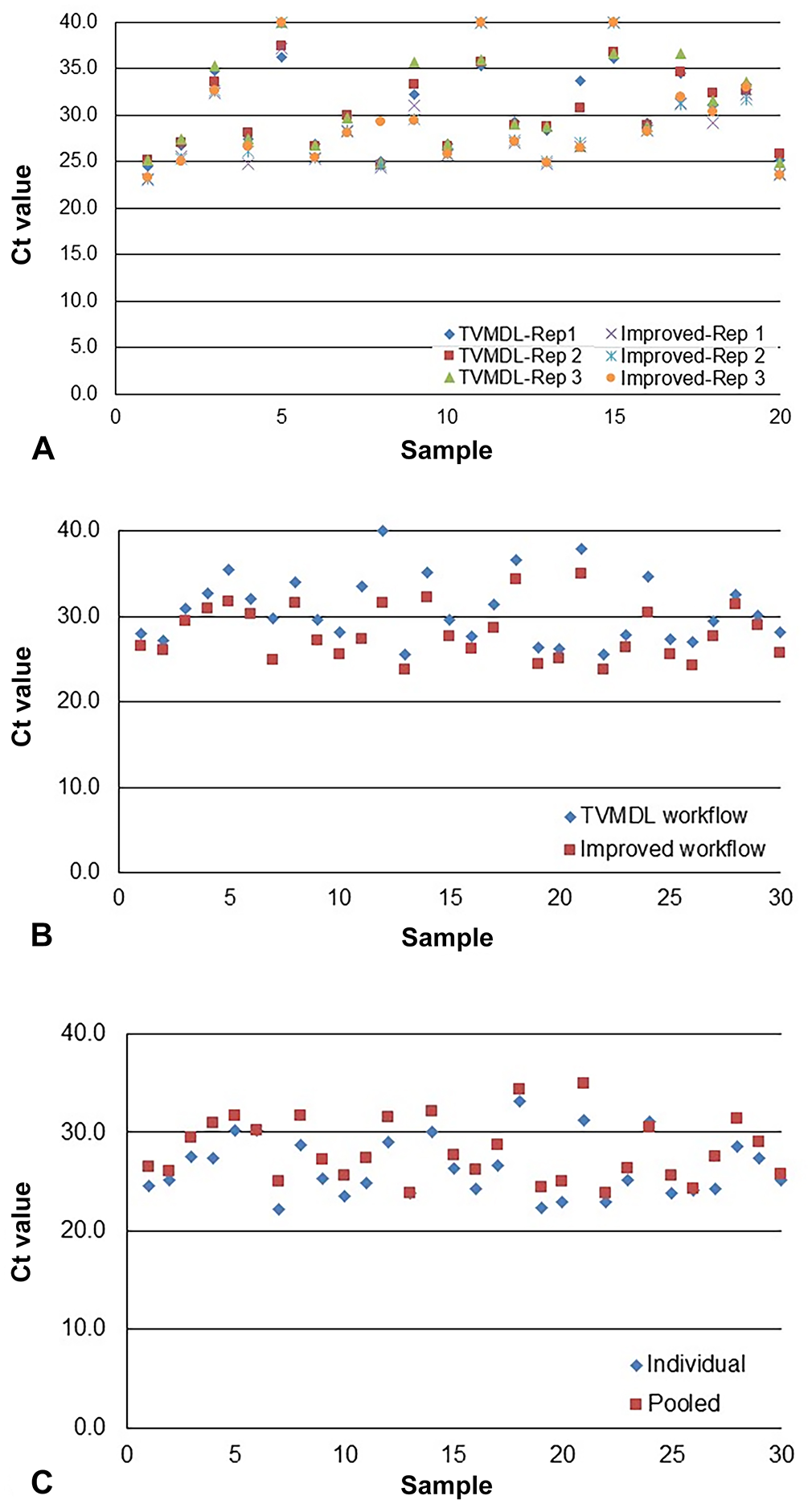
Pooled T. foetus–positive Ct values were lower when tested with our new workflow (Cts of 23.8–35.0) compared to our published workflow 8 (Cts of 25.5–40.0); the x̄ Xeno Ct was 30.5 and the average XIPC Ct was 31.4, respectively (Fig. 1B). Compared to the individually tested T. foetus–positive samples, sample pools determined to be positive were 100% in agreement with individual T. foetus–positive status (Fig. 1C). In general, pooling positive samples with negative samples at a 1:5 ratio resulted in ~2.5 Ct value higher shift than individual sample Ct values, as expected for this dilution ratio. All pooled negative samples were T. foetus–negative; x̄ Xeno Ct = 31.7.
Pooling has been reported in the literature as a cost-effective method for detecting T. foetus, particularly if prevalence levels are expected to be low, with comparable detection rates between individually tested samples and pooled samples.2,5,6,9 Pooling of 5 individually cultured samples followed by extraction and PCR provided 100% sensitivity and specificity compared to individual samples. 9 A comparison of T. foetus sample pooling in groups of 5 or 10 samples, in either culture medium (InPouch) or PBS, found no significant difference in detection by pool size or sample type. 5 A follow-up study found no significant differences in detection rate when samples were pooled in various ratios, up to 25 samples in a pool; results demonstrated 100% specificity and 98.8% sensitivity with microscopic culture. 6
Interestingly, in one study, all samples tested individually were negative in some positive pools. 6 Given that these samples were collected from experimentally infected known-positive bulls, it is hypothesized that inhibitors may have been present in one or more of the samples in the pool that prevented detection of the positive sample; known internal controls were not utilized in the study, which would have aided in monitoring sample inhibition. 6 The authors postulated that the potential presence of inhibitors combined with the known variability of infection level, as well as variations in sample collection and handling that can affect pathogen detection, are representative of situations common in the field. 6 As such, they suggested that repeat testing should be a common practice prior to classification of T. foetus–positive or –negative status.5,6,13,15
To our knowledge, all previous reports of testing of sample pools focused on detection of T. foetus DNA. Our PCR assay reported previously, 8 as well as the updated reagents reported here, target detection of T. foetus RNA to further enhance pathogen detection. We used T. foetus–positive preputial washings that had been submitted routinely to our laboratory. Once identified as positive, the samples were stored at −20°C. Although samples remained positive following frozen storage, we did note increased Ct values (~2–3 Cts) for some prior to use in our study. These increased values could be the result of sample degradation caused by freezing and thawing of the sample and/or long-term storage at −20°C. To provide accurate and reliable results, samples should be shipped to the laboratory, stored cold at 4°C, and PCR tested as soon as feasible, to avoid prolonged storage and possible sample degradation.
Detection rates for T. foetus can potentially decline when testing sample pools, particularly for weak-positive or high Ct samples (i.e., low numbers of target), and should be taken into consideration when pooling samples for testing. To offset this potential loss in sensitivity, the Ct cutoff for sample pools should be increased (i.e., Ct = 38.0) to provide a greater chance of detecting weak-positive samples. Furthermore, the increased Ct value cutoff should also compensate for inhibition that might be present in the pool, which could prevent detection of a single positive sample in the pool. Of note, our updated workflow utilizes a larger sample volume input for extraction (100 µL vs. 50 µL). This increased sample volume input, combined with the enhanced sensitivity of the RT-rtPCR reagents, provides improved detection of T. foetus in sample pools. When combined with inclusion of the exogenously added Xeno RNA, and a higher Ct cutoff that accounts for dilution affects, our updated and improved workflow provides improved detection of T. foetus in both individual and pooled specimens.
Footnotes
Correction (Feb 2023):
Article updated to correct the title to “Detection of Tritrichomonas foetus by RT-rtPCR in pooled bovine preputial washings” and in the content accordingly.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
