Abstract
Foot-and-mouth disease (FMD) is a highly contagious disease of cloven-hoofed animals. FMD poses an economic threat to the livestock industry in the United States. Due to the potential use of vaccines composed of partially purified structural proteins of the FMD virus (FMDV), it is important to test samples from infected and vaccinated animals with a competitive ELISA that detects antibodies against nonstructural proteins (NSPs) of FMDV. Our study extends the diagnostic validation of the Prionics ELISA (Thermo Fisher) and the VMRD ELISA. We used diverse serum sample sets from bovine, porcine, and other cloven-hoofed animals to evaluate the analytical specificity and sensitivity, diagnostic specificity and sensitivity, and differentiation of infected from vaccinated animals (DIVA) per validation guidelines outlined by the World Organisation for Animal Health (WOAH). The 2 tests were analytically 100% accurate. The VMRD test was diagnostically more sensitive than Prionics, but Prionics was diagnostically more specific than the VMRD test. Both tests could tell if animals were infected or vaccinated. Considering these data, both VMRD and Prionics ELISAs can be used for serodetection of FMDV antibodies at the Foreign Animal Disease Diagnostic Laboratory and within the National Animal Health Laboratory Network laboratories.
Keywords
Foot-and-mouth disease (FMD), a highly contagious viral disease of cloven-hoofed animals, causes severe economic losses because of trade limitations and recovery efforts in affected areas.14,20 The causative agent, FMD virus (FMDV; Picornaviridae, Aphthovirus, Aphthovirus vesiculae), is a positive sense, single-stranded RNA virus comprised of 8,400 nucleotides encoding 4 structural and 8 nonstructural viral proteins. 5 The 4 structural viral proteins, VP1–4, assemble to form the virus capsid and serve as the main antigenic components in all licensed FMDV vaccines. The nonstructural proteins (NSPs), L, 2A–C, and 3A–D, are required for viral replication and interactions with host cell factors, but are depleted in highly purified FMDV vaccines.5,7
Considering that conventional FMDV vaccine production utilizes NSP depletion and formulation with adjuvant, differentiation of infected and vaccinated animals (DIVA) is possible via NSP-specific antibody assays.8,22 Per the WOAH terrestrial manual, 20 highly NSP-depleted, inactivated FMDV vaccines and well-validated DIVA-capable NSP assays are crucial for control programs to re-achieve FMDV-free status in FMD-affected regions. B-cell epitopes identified in FMDV NSP 2B, 2C, 3A, 3B, and 3D have been utilized to develop DIVA-capable tests.13,17,18,21 Of those assays, a 3B core repeat epitope (QKPLK) and a monoclonal antibody (mAb) specific for this epitope were used in a competitive ELISA. 17 Another competitive ELISA utilized the 3D epitope (MRKTKLAPTVAHGVF) and the sequence-specific mAb. 21 An indirect ELISA using the same 3B core repeat epitope (QKPLK) was developed and evaluated in various regions and species.9,13,15 Among the FMDV NSPs, 3ABC is considered the most reliable antigen for DIVA testing. Two commercially available competitive ELISAs using the 3ABC antigen and specific mAbs reported higher diagnostic sensitivity and specificity than other NSP assays had reported in previous validation studies using limited sera and species in Europe.3,6
However, diagnostic evaluation studies of competitive or indirect ELISAs have demonstrated regional and species-related variation in performance. This variation can greatly complicate disease response and control1–4,6,8,9,13,15,17,18,22; therefore, method comparison studies of different ELISAs are pivotal for disease outbreak preparedness and response. The United States has FMDV vaccination capabilities that will be utilized in the event of an outbreak, making DIVA-capable assays critical in a post-outbreak environment in which both animal vaccination and disease status need to be known. The method comparison that we present here evaluates the performance characteristics of 2 FMDV DIVA-capable antibody ELISAs, the Prionics FMDV NS antibody ELISA kit (v.1.4_e, Thermo Fisher; hereafter Prionics) and another FMDV antibody ELISA kit (v.170323, VMRD; hereafter VMRD), using diverse sera, including positive sera from cattle and pigs in endemic areas and negative sera from cattle and pigs in the United States. Samples from caprine species were used when available.
Materials and methods
Production of VIAA
The virus infection–associated antigen (VIAA) test used as the reference assay to select the samples for an FMDV-positive panel of field samples was performed per the Foreign Animal Disease Diagnostic Laboratory (FADDL) standard operating procedure (SOP: RVS-0070) and published method. 1 Briefly, recombinant FMDV 3D antigen (His-Tagged 3D fusion protein) was expressed in Escherichia coli strain BL21 and purified by affinity chromatography (HisTrap FF column; GE HealthCare).
E. coli strain BL21 (in 15% glycerol) carrying recombinant plasmid vectors pET3D (cloned FMDV 3D polymerase gene) and pCG1 (cloned yeast ubiquitin proteinase gene) was cultured on a LB agar plate containing kanamycin (25 µg/mL) and chloramphenicol (20 µg/mL) at 37 ± 2°C overnight. A single colony was picked from the LB plate and inoculated into 100 mL of sterile NZCYM broth supplemented with 0.4% glucose, 25 µg/mL kanamycin, and 20 µg/mL chloramphenicol and incubated in a 30 ± 2°C shaker at 220 rpm for 16–20 h. This culture served as the starter culture.
Once the starter culture reached 1.2–1.5 OD, 1.25 mL of IPTG (isopropyl β-d-1-thiogalactopyranoside, 100 mM stock; Sigma) was added to each flask (250 mL, final IPTG concentration 0.5 mM) and incubated in a 25 ± 2°C shaker (220 rpm) for 4–5 h to induce overexpression of the recombinant VIAA antigen. After IPTG induction, the entire culture (4 × 250 mL) was transferred to 4 × 500 mL sterile polypropylene centrifuge bottles; the bacterial cells were pelleted by centrifugation at 8,000 ± 1,000 × g for 15 ± 2 min at 4°C.
The bacterial cell pellet was re-suspended in lysis/binding buffer (10 mM NaPO4, 500 mM NaCl, 50 mM imidazole, 0.5 mM EDTA, NP-40 0.1% v/v, 2.8 µg/mL pepstatin, 2 µg/mL leupeptin, 10 mM β-ME in ultrapure water) at a ratio of 5 mL of buffer per gram of cell paste. The cell suspension was adjusted to a volume of 35 mL with lysis/binding buffer, then lysozyme powder (2–5 mg) and 350 µL of 100 mM phenylmethanesulfonyl fluoride (PMSF) solution (Sigma 93482) were added and sonicated 6–8 times, 15–20 s each (sonicator; Misonix) with an interval of 1 min between pulses. DNase I powder (2–3 mg; Sigma) was added to the sonicated crude cell extracts and incubated while shaking on a flat tilting platform at room temperature for 25–30 min. The DNase-treated cell extract was centrifuged at 30,000 ± 1,000 × g in a fixed angle 45Ti rotor for 20 ± 5 min at 8°C. The supernatant was collected and passed through a 45-µm syringe filter using a 20-mL sterile syringe. The clear supernatant (cell-free extract) was used as the starting material for the recombinant VIAA antigen purification.
Purification of recombinant VIAA antigen from cell-free extracts
A column (HisTrap FF Crude column; GE HealthCare) was washed with 50 mL of Milli-Q ultrapure deionized water and equilibrated with 50 mL of binding buffer (50 mM imidazole, 10 mM β-ME, 0.1% (v/v) NP-40, 500 mM NaCl, 20 mM NaPO4, pH 8). The clear supernatant (i.e., the cell-free extract) was passed through the equilibrated column at a flow rate of 1 mL/min. The column was washed with 50 mL of binding buffer to remove any unbound protein. The recombinant VIAA 3D antigen was eluted from the column using binding buffers containing 250 and 300 mM of imidazole sequentially and characterized in sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE; 12% acrylamide NuPAGE mini gels, Thermo Fisher) and SDS buffer (NuPAGE MOPS), followed by protein staining (PageBlue protein staining solution; Thermo Fisher). Elution fractions containing the recombinant VIAA 3D protein in SDS-PAGE were pooled and dialyzed in a Tris buffer (50 mM Tris-HCl, 100 mM NaCl, 0.1% v/v NP-40, 10 mM β-ME, 10% v/v glycerol in ultrapure water [Milli-Q]) using a dialysis cassette (Side-A-Lyzer; Thermo Fisher). The dialyzed VIAA 3D antigen was further purified by an ammonium sulfate precipitation method using a 40% saturation protocol. 19
Evaluation of analytical specificity and sensitivity
Analytical specificity (ASp) of the investigational VMRD ELISA and the comparator Prionics ELISA was evaluated with sera collected from animals (bovine, equine, porcine) infected or immunized with 1 of the 24 non-FMD clinical and/or genetic look-alike viruses obtained from the National Veterinary Services Laboratories (NVSL; Ames, IA, USA). Analytical sensitivity (ASe), also known as the limit of detection (LOD), was evaluated with sera from 17 animals experimentally infected with 1 of the 7 serotypes of FMDV. Fourteen serum samples, including 2 sera from each FMDV serotype, were 2-fold serial diluted (1:2–1:1024) and tested in duplicate wells on 3 consecutive days. Three sets of porcine sera, consecutively collected from previous FMDV challenge studies using serotypes A and C viruses, were also tested at neat to represent a time point panel for ASe evaluation.
Evaluation of diagnostic specificity and sensitivity
Diagnostic specificity (DSp) was evaluated using sera collected from 2,067 animals (1,041 bovine, 1,010 porcine, and 16 other cloven-hoofed ungulates) from slaughterhouses in the United States, a FMDV-free country. Diagnostic sensitivity (DSe) was evaluated using serum collected from 571 (463 bovine, 77 porcine, and 31 other ungulates) experimentally infected animals or animals from endemic regions. These samples were tested with a VIAA reference test for FMDV to be eligible for use as the panel for DSp. 1
Evaluation of DIVA capability
DIVA capability of the 2 cELISAs was evaluated using serum samples collected from 73 vaccinated animals. Those included 58 bovine and 15 porcine sera collected at 21 d post-vaccination (pre-challenge).
Investigational cELISA protocols
The VMRD ELISA kit was used according to the manufacturer’s instructions. The cutoff for positive samples was ≥40% inhibition. The Prionics ELISA kit was used according to the manufacturer’s instructions with a ≥50% inhibition cutoff for positive samples. We used the overnight serum incubation protocol instead of the 2-h incubation for the Prionics ELISA for better sensitivity.
Data analysis
The DSp values of the VMRD and Prionics cELISAs were calculated as the percentage of U.S.-origin sera that were negative on the reference VIAA assay. DSe was the percentage of the VIAA-defined, FMDV antibody–positive sera with a positive result in the assay being evaluated. Receiver operating characteristic (ROC) curve and scatter plot analysis were performed using Excel (v.2403; Microsoft) and R (v.4.4.1, https://www.r-project.org/) to evaluate the cutoff for positive and negative detection by the 2 cELISAs through comparison with the VIAA-positive and -negative serum reference panels described above.10–12 A binomial test 16 was used to determine if there were significant (p ≤ 0.05) differences in sensitivity and specificity between the comparator cELISA when testing various sets of serum samples. All statistical analyses were performed using R.
Results
Analytical specificity of VMRD and Prionics assays
All 24 sera collected from animals (bovine, equine. porcine) infected or immunized with each of 24 non-FMD viruses were negative on both VMRD and Prionics assays (Fig. 1). Our results indicate that both tests are specific for FMDV and do not cross-react with major bovine and porcine viruses endemic in the United States.
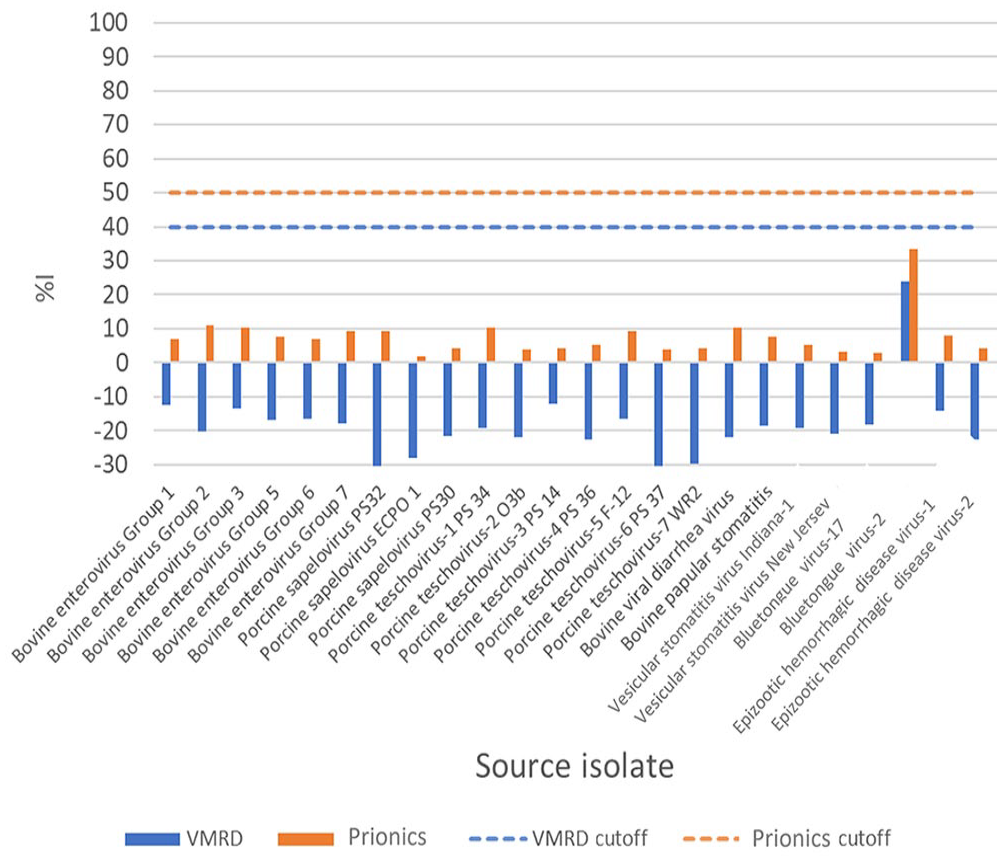
Evaluation of the analytical specificity of VMRD and Prionics FMDV antibody ELISAs using antisera to 24 bovine and porcine viruses endemic in the United States. The antisera were tested at a 1:2 dilution per kit instructions. %I = percent inhibition.
Analytical sensitivity of VMRD and Prionics assays
ASe was evaluated based on the LOD, when tested with sera from 17 animals experimentally infected with 1 of the 7 serotypes of FMDV. The mean LOD titers for a positive result to 14 FMDV-specific sera on the VMRD assay were higher than the Prionics assay in 12 of 14 sera specific to 7 FMDV serotypes (Fig. 2). This indicates that the VMRD assay has improved ASe and can detect specific antibodies at lower concentrations.
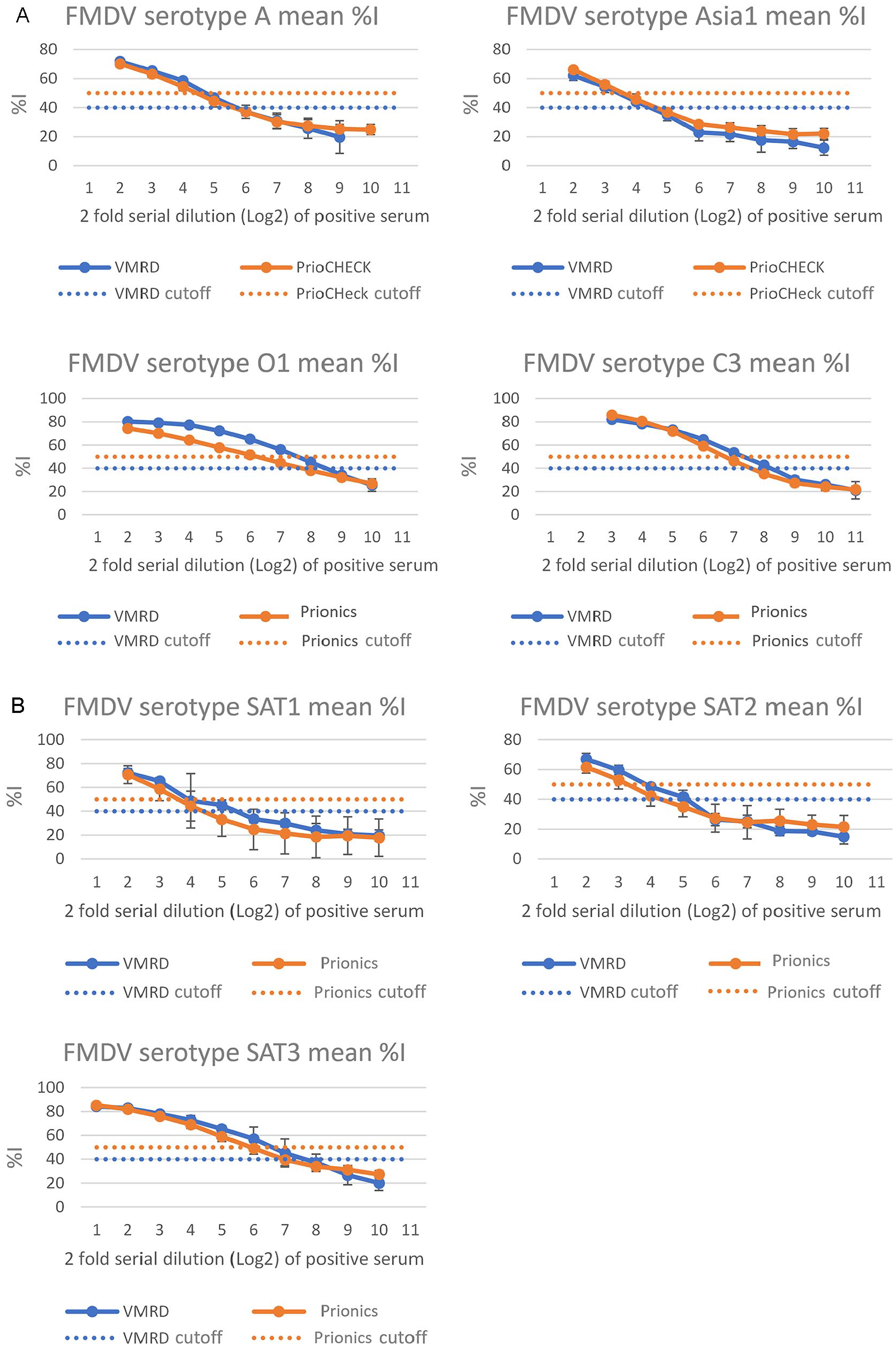
Evaluation of the analytical sensitivity of VMRD and Prionics FMDV antibody ELISAs using sera from animals infected with 1 of 7 FMDV serotypes. FMDV positive sera used for the analytical sensitivity analysis were generated using
Three sets of porcine sera, consecutively collected from FMDV challenge studies, were also tested to represent a time point panel for ASe evaluation (Fig. 3). Positive detection of serotype A sera in the VMRD and Prionics assays were observed as early as 5–6 dpi. Positive detection of serotype O1 was observed as early as 7 dpi by the VMRD kit; however, the Prionics ELISA did not detect antibodies until 8 dpi.
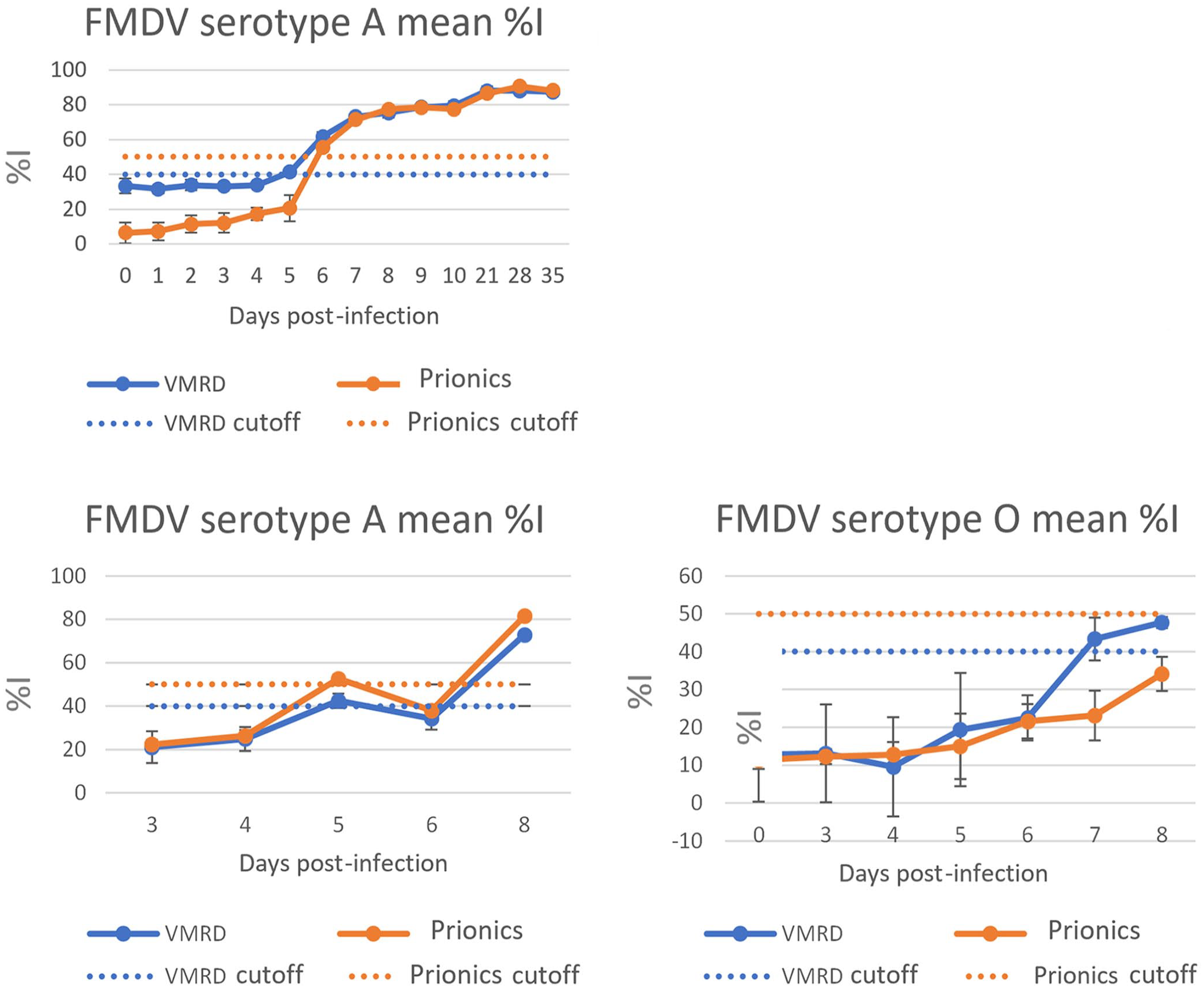
Evaluation of early detection of FMDV antibodies by the VMRD and Prionics FMDV ELISAs using sera sequentially collected from FMDV challenge animals. FMDV positive sera were collected daily from 3 animals infected (single challenge) with O1/Manisa (n = 1) or A/A24 (n = 2). The sera were tested in duplicate wells on 3 consecutive days. %I = percent inhibition.
Diagnostic specificity of VMRD and Prionics assays
Serum samples collected from 1,041 bovine, 1,010 porcine, and 16 other ungulates in the United States were tested using the VMRD and Prionics assays. Initial DSp estimation for the VMRD and Prionics assays based on single-well per sample runs were 97.5% and 99.6%, respectively. The VMRD test had 51 non-negative results (49 bovine and 2 porcine sera), and the Prionics test had 7 non-negative results. All non-negative samples were retested in duplicate wells to check the validity of the initial results. The final DSp results, including re-test results, were 98.2% for the VMRD assay and 99.7% for the Prionics assay (Table 1).
Summary of diagnostic specificity evaluation of the VMRD and Prionics ELISAs using sera from 1,041 bovine, 1,010 U.S. porcine, and 16 other ungulates for initial testing.
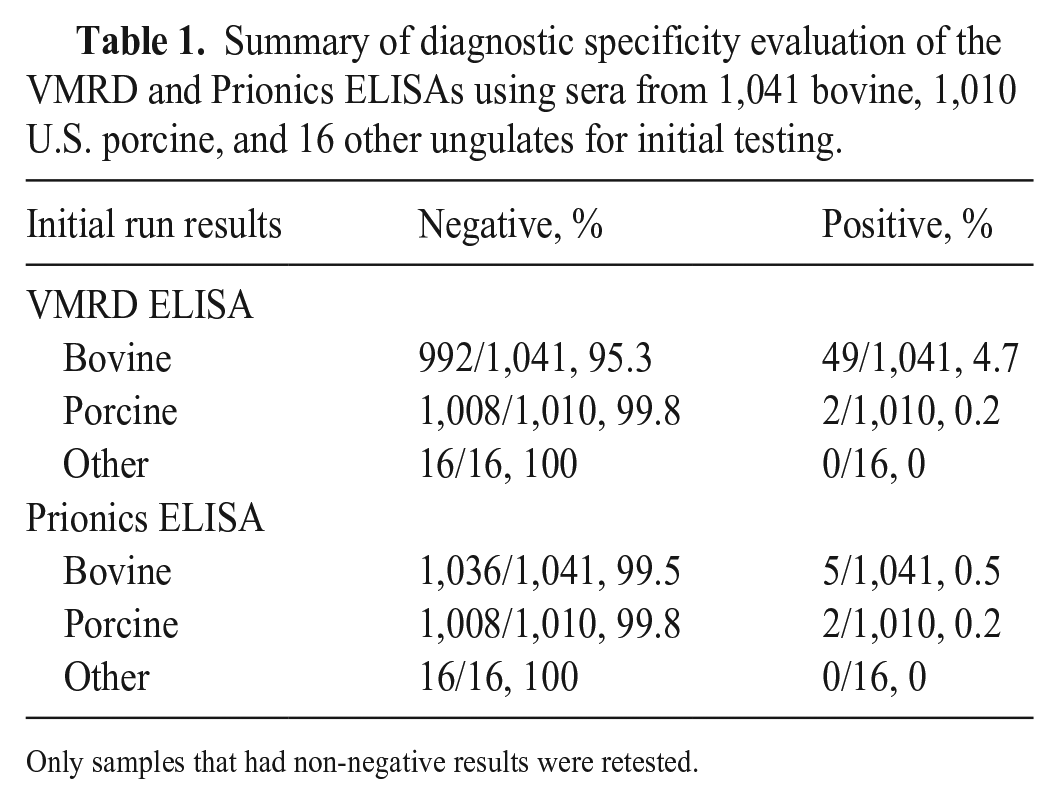
Only samples that had non-negative results were retested.
DSe was evaluated using sera from experimentally infected animals and animals from FMDV endemic areas. These samples were tested with the VIAA reference test before being used as the panel for DSe; 571 of the 671 samples were positive on the VIAA test. The initial evaluation using the 571 DSe VIAA test positives, including bovine, porcine, and other species, resulted in an initial DSe of 98.9% for VMRD and 98.1% for Prionics. The negative samples were retested in duplicate wells to check the validity of the initial results. Several negative test results in single-well runs were reversed in rerun using duplicate wells. Against the 571 sera tested, the overall DSe estimate based on the initial run and rerun results was 99.8% for VMRD and 98.9% for Prionics (Table 2).
Summary of diagnostic sensitivity evaluation of the VMRD and Prionics ELISAs using sera from 463 bovine, 77 porcine, and 31 other ungulates for the initial testing.
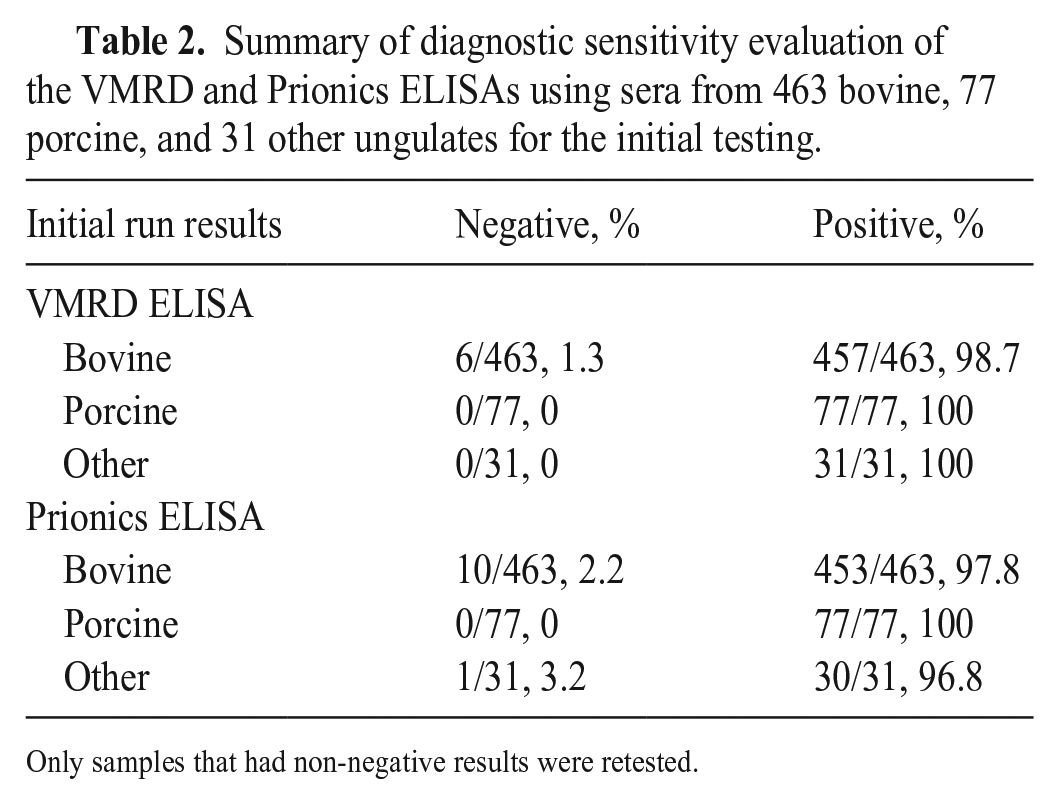
Only samples that had non-negative results were retested.
Evaluation of cutoff of the 2 ELISAs for positive or negative detection
Receiver operating characteristics (ROC) curve analysis using the percent inhibition (%I) data from the VMRD and Prionics assays was used to evaluate the cutoff values for each test kit. The optimal cutoff value to balance DSe and DSp for the VMRD test with bovine serum samples was 44–48%I (Fig. 4). The optimal cutoff for the VMRD ELISA with porcine sera was 23–45%I, consistent with the 40%I kit cutoff value (Fig. 4). The Prionics kit cutoff value (50%I) used at FADDL and in the package insert was optimal for surveillance and foreign animal disease investigation (FADI) based on sample sets tested in our study. The optimal DSe/DSp range for bovine and porcine sera was 49–51%I (Fig. 5) and 37–58%I (Fig. 5), respectively.
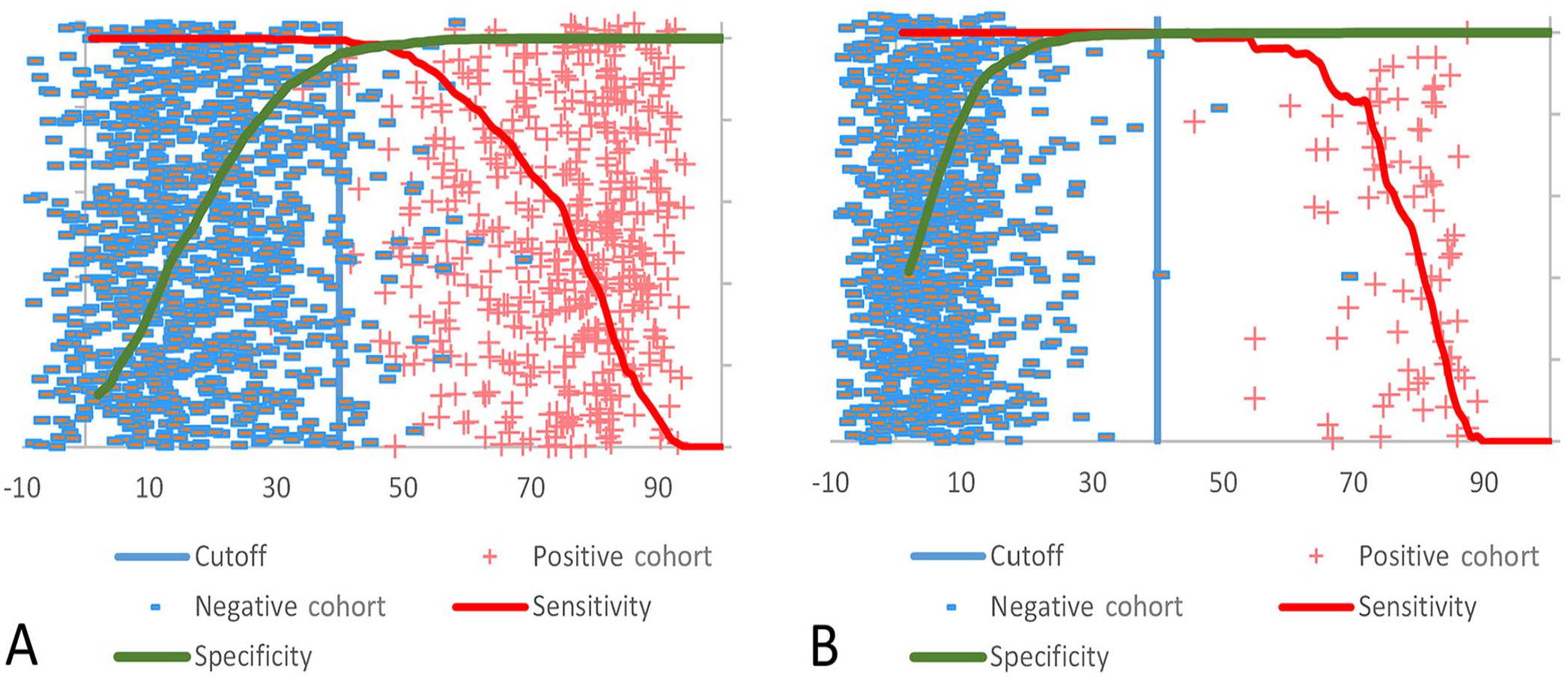
Receiver operating characteristics (ROC) curve analysis to evaluate the cutoff for the VMRD ELISA using
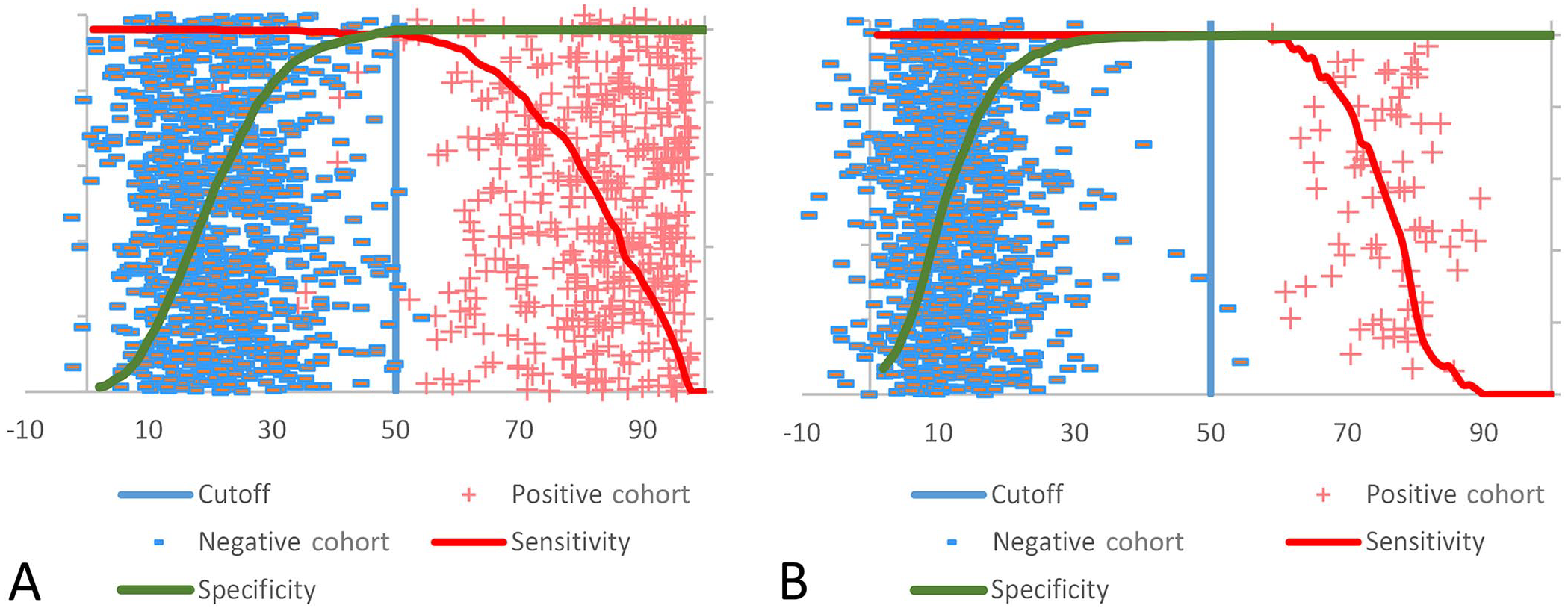
ROC curve analysis to evaluate the cutoff for Prionics ELISA using
Evaluation of DIVA capability
DIVA capability of the VMRD and Prionics ELISAs was evaluated using serum samples collected from 73 vaccinated animals (21 dpi, pre-challenge samples). Serum samples from 58 cattle and 15 pigs were tested using the VMRD and Prionics assays. Both the VMRD and Prionics assays had negative results against the 73 samples, demonstrating DIVA capability (Fig. 6).
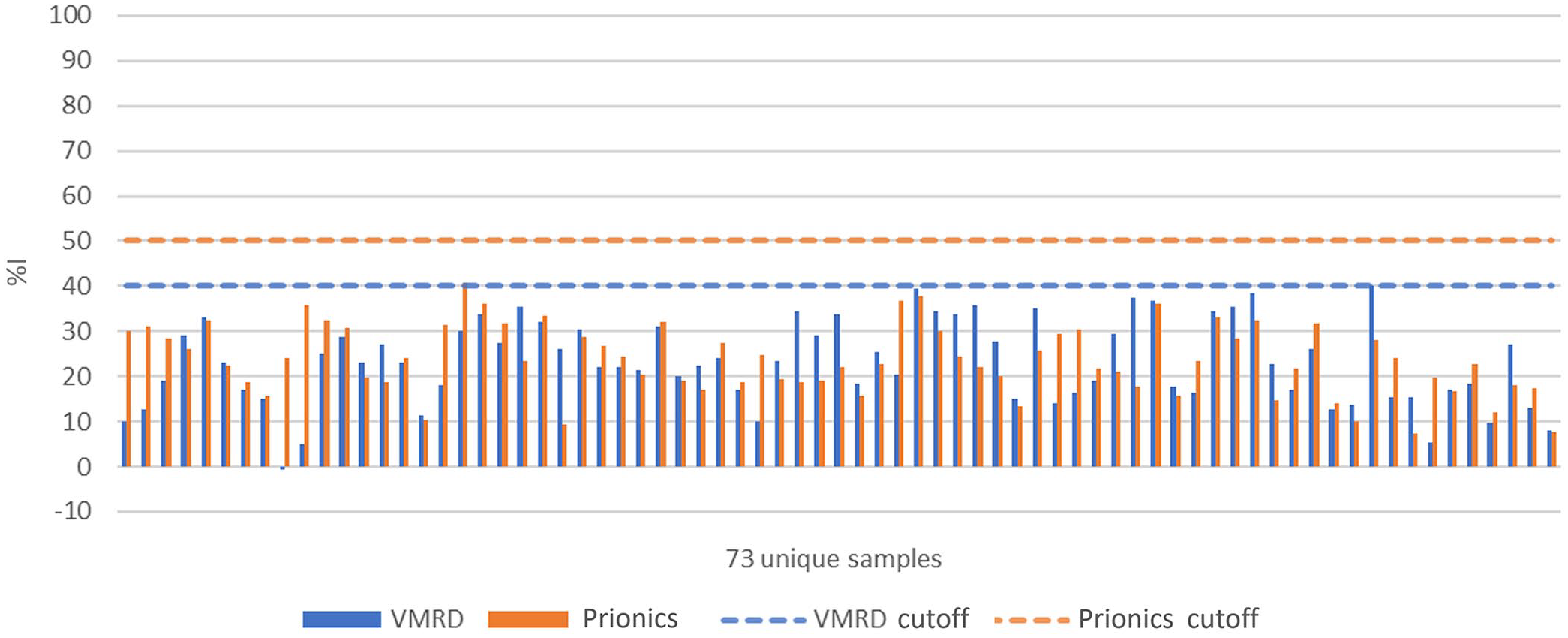
Evaluation of DIVA (differentiation between infected and vaccinated animals) capability of 2 ELISAs using sera from animals (58 cattle and 15 swine) immunized with inactivated and NSP-purified FMDV vaccines. %I = percent inhibition.
Discussion
Both the Prionics and VMRD ELISAs detect antibodies directed against the nonstructural 3ABC protein of FMDV with DIVA capability. The current cELISA employed by FADDL is the Prionics 3ABC kit, which must be sourced internationally and requires a 16–18-h overnight incubation of serum samples. A previous preliminary study indicated that the VMRD ELISA was slightly more sensitive and specific than the PrioCHECK. 6
Both the VMRD and Prionics FMDV antibody ELISAs had high analytical and diagnostic performance characteristics, which aligns with previous reports. 6 The results of the ASp indicated that both the VMRD and Prionics tests are specific for FMDV and do not cross-react with sera collected from animals infected or immunized with bovine, equine, and porcine viruses selected based on clinical look-alike or genetic closeness. The outcome of the ASe analysis indicated that both assays have a highly sensitive LOD, but the VMRD assay has higher sensitivity and can detect antibodies specific for FMDV at lower concentrations and earlier than the Prionics ELISA. The data from our DSp evaluation suggests that the Prionics assay is slightly more specific than the VMRD assay using the kit insert cutoff values and bovine sera, which differs from a previous report. 6 Our DSe evaluation results suggest that the VMRD assay is slightly more sensitive than the Prionics assay. Both assays have DIVA capability based on the evaluation using limited sera from cattle and swine vaccinated with NSP-depleted inactivated FMDVs. Our results indicate high analytical and diagnostic performance characteristics; therefore, both the VMRD and Prionics kits are acceptable for FMDV antibody detection at reference laboratories and in state diagnostic laboratories.
In ROC curve analysis, the kit cutoff of 40%I maximizes DSe (100%) with minimal loss of DSp (96%) for surveillance and FADI uses in non-endemic countries like the United States. The 40%I cutoff is preferable for FADDL and NAHLN laboratories because it minimizes the chances of a false-negative result. Additional testing on any non-negative results could be performed at a reference laboratory. In our study, the porcine serum panel from field and experimentally infected animals had higher resolution than the bovine panel and previous studies. 6 We therefore recommend 40%I as the universal cutoff for both bovine and porcine species to maximize DSe and increase the convenience for surveillance and FADI uses.
It is valuable to have 2 well-validated, high-performance assays for FMDV detection and surveillance programs for several reasons. In a potential FMDV outbreak situation, surge production and expedited delivery of assays to laboratories is crucial for rapid response and disease control. Absence of select agent materials in the final products can reduce any safety testing requirement before delivery to and use in the laboratories. Various assays may have different strengths and weaknesses in diagnostic performance and operational procedures. The USDA Center for Veterinary Biologics–licensed VMRD assay has <3-h run time and delivery to laboratories without major lead time compared to the unlicensed Prionics 3ABC kit, which contains select agent materials and requires overnight operation procedures as currently in use at FADDL.
Footnotes
Acknowledgements
We thank Dr. Jamie Barnabei and Ms. Amber Hemple for providing the samples for DIVA capability testing of the 2 assays that we evaluated. We also thank Dr. Amaresh Das for technical advice on VIAA production, Ms. Karen Moran for training on the VIAA test, and Ms. Bailey Harach and Ms. Danielle Marcone for editorial assistance on this manuscript.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
Our study was supported in part by the Inter-Agency Agreement fund between the U.S. Department of Homeland Security Science and USDA-APHIS-FADDL (project IAA 70RSAT18KPM000139).
Disclaimer
The findings and conclusions in this publication are those of the authors and should not be construed to represent any official USDA or U.S. Government determination or policy.
