Abstract
A One Health approach has been key to monitoring the COVID-19 pandemic, as human and veterinary medical professionals jointly met the demands for an extraordinary testing effort for SARS-CoV-2. Veterinary diagnostic laboratories continue to monitor SARS-CoV-2 infection in animals, furthering the understanding of zoonotic transmission dynamics between humans and animals. A RT-PCR assay is a primary animal screening tool established within validation and verification guidelines provided by the American Association of Veterinary Laboratory Diagnosticians (AAVLD), World Organisation for Animal Health (WOAH), and the U.S. Food and Drug Administration (FDA). However, differences in sample matrices, RNA extraction methods, instrument platforms, gene targets, and cutoff values may affect test outcomes. Therefore, targeted validation for a new sample matrix used in any PCR assay is critical. We evaluated a COVID-19 assay for the detection of SARS-CoV-2 in feline and canine lung homogenates and oral swab samples. We used the commercial Applied Biosystems MagMAX Viral/Pathogen II (MVP II) nucleic acid isolation kit and TaqPath COVID-19 Combo kit, which are validated for a variety of human samples, including nasopharyngeal and oropharyngeal swab samples. Our masked test showed a high detection rate and no false-positive or false-negative results, supporting sample extension to include feline oral swab samples. Our study is a prime example of One Health, illustrating how a COVID-19 assay designed for human testing can be adapted and used to detect SARS-CoV-2 in oral swab samples from cats and likely dogs, but not lung homogenates.
The severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) pandemic is a prime example of human and veterinary medicine coming together in the concept of One Health, given that SARS-CoV-2 originated in animals. SARS-CoV-2 emerged rapidly, infecting >700 million people worldwide, and the resulting COVID-19 disease has been linked to >6 million deaths. 3 The rapid spread and subsequent requests for large numbers of laboratory tests throughout the world to confirm the presence of the virus or anti-viral antibodies challenged human laboratories to meet the demand for the volume of testing. Use of molecular testing led to the identification of at least 33 animal species naturally infected with SARS-CoV-2 10 ; however, there is still more to learn about species susceptibility and the zoonotic transmission dynamics between humans and animals, with few examples known. Veterinary diagnostic laboratories (VDLs) can test a variety of animal species and sample types to expand the knowledge gap on virulent strain evolution in animal species, potential reverse zoonoses, and inter- and intra-species transmission. VDLs are well equipped to respond to outbreaks with the capacity and infrastructure for testing, including for SARS-CoV-2, through high-throughput PCR testing and, in some cases, next-generation sequencing (NGS).
SARS-CoV-2 reverse-transcription PCR (RT-PCR) assays target various parts of the viral genome including envelope (E), spike (S), nucleocapsid (N), and open reading frame (ORF) genes. 23 The variation in viral genome target sites, along with different extraction methods, instrument platforms, and cutoff values coupled with the emergence of new variants, may affect test outcome and can lead to noncomparable results.8,11 Although inter-laboratory comparison tests show that multiple laboratory methods can successfully detect SARS-CoV-2, there are still variations in results between methods.5,6 In the United States, many VDLs are accredited by the American Association of Veterinary Laboratory Diagnosticians (AAVLD), which requires laboratories to validate and verify assays to provide quality assurance that results are consistent across laboratories, to minimize contamination risks, and to verify method or sample matrix extensions before use. 1 The U.S. Food and Drug Administration (FDA) provides standards and guidelines for test validation studies along with official certifications.1,21,22 Validation procedures include establishing the following performance parameters: accuracy (for quantitative assays), false-positive/negative rate (for qualitative assays), sensitivity (i.e., limit of detection [LOD]), repeatability, reproducibility, specificity, extraction efficiency, and ruggedness/robustness.17–19 Evaluation of assay performance for a new species biomatrix (i.e., a matrix extension study) is necessary because matrix constituents can significantly interfere with PCR-based detection. 15 New matrices may be of different consistencies and/or contain various enzymes or microbial communities, which may significantly interfere with the extraction of RNA or its subsequent analysis.
We evaluated the performance of an assay, utilizing the commercial Applied Biosystems (ABI) MagMAX Viral/Pathogen II (MVP II; cat. A48383; Thermo Fisher) nucleic acid isolation kit and TaqPath COVID-19 Combo kit (cat. A47814; Thermo Fisher), collectively the COVID-19 assay, for RT-PCR detection of RNA extracted from canine and feline lung homogenates and oral swabs. The TaqPath COVID-19 kit was originally designed by the manufacturer for qualitative detection of SARS-CoV-2 RNA extracted from specific human (not animal) matrices, such as nasopharyngeal swab, nasopharyngeal aspirate, or bronchoalveolar lavage specimens. 2 The COVID-19 assay was chosen because of the validation performed by the manufacturer under the Emergency Use Authorization for human testing, and the North Dakota State University–VDL, (NDSU-VDL; Fargo, ND, USA) molecular testing staff had a streamlined workflow for human testing with the assay. Our objective, using a One Health approach, was to evaluate whether a testing protocol for the detection of SARS-CoV-2 in humans could be adapted and used to detect the virus in samples from animals.
Materials and methods
Heat-inactivated viral stocks
The heat-inactivated Washington isolate (VR-1986HK, strain 2019-nCoV/USA-WA1/2020) was purchased from ATCC and used during in-house experiments at the NDSU-VDL. Our masked test utilized both a heat-inactivated Washington isolate (NR-52286, SARS-related coronavirus 2 isolate USA-WA1/2020) and Omicron isolate (NR-56495, SARS-related coronavirus 2 isolate hCoV-19/USA/GA-EHC-2811C/2021) obtained from BEI Resources.
Sample acquisition
Feline and canine lungs were collected from animals submitted to the NDSU-VDL for postmortem examination. Animals had signs of respiratory infection that were mainly bacterial but were not suspected of SARS-CoV-2 infection. One set of bleach-soaked, rinsed, and autoclaved scissors, forceps, and a homogenizer probe was used per animal to avoid cross-contamination. Samples of feline or canine lung (5–15-mm cubes) were placed in 50-mL conical tubes with 40 mL of 1 × sterile PBS (pH 7.4), resulting in ~10 g per 40 mL of total volume. Tissues were homogenized (200 hand-held homogenizer probe; VWR) until larger pieces of tissue were no longer visible. After homogenization, all tubes were mixed by vortexing and held at −20°C. A pool of feline lung homogenates in PBS was made by combining equal amounts (8 mL) from each of the 5 animals and repeated for canine lung homogenates.
Feline and canine oral swab samples were collected antemortem by veterinary professionals through diagnostic submissions or by veterinary pathologists from autopsy cases. Oral swabs were chosen because they are common samples for SARS-CoV-2 testing in animals, relatively easy to collect, and have been shown to be as effective as nasal swabs in human testing.9,14,20 Sterile, rayon-tipped, plastic-shaft swabs (cat. 10808-146; VWR) were used. Up to 4 swabs were collected for some individuals. None of the animals was suspected of harboring SARS-CoV-2; all were tested using the COVID-19 RT-PCR assay prior to pooling. Samples were all negative based on no detection of SARS-CoV-2 ORF1ab, S, and N genes. Feline and canine oral swabs were placed in 1–6 mL of PBS. Samples were mixed by vortexing for at least 30 s and then frozen at −20°C. Equal amounts of samples were pooled from 5 cats and 6 dogs. Prior to spiking with heat-inactivated virus, samples were thawed at ~4°C and vortexed for at least 30 s.
RNA extraction and SARS-CoV-2 RT-PCR
RNA extraction was completed on a magnetic particle processor with 96 deep-well head (KingFisher Flex; Thermo Scientific) using the MVP II kit. The complete protocol with modifications can be found on protocols.io, an open-access repository platform. 16 Briefly, plates were set up using the manufacturers’ guidelines, and 400 µL of sample was added to each well, with one modification from the human protocol. The binding solution, magnetic beads, proteinase K, and MS2 phage control were mixed and added to wells together instead of individually, which reduced preparation time and usage of pipette tips that were in limited supply during the height of the SARS-CoV-2 pandemic. The MVP_2Wash_400_Flex program was selected on the KingFisher Flex. Once the program was complete, the elution plate was removed, covered, and stored briefly on a magnetic-ring stand (cat. AM10050; Thermo Fisher) at ~4°C until added to the SARS-CoV-2 RT-PCR reagents.
The TaqPath COVID-19 Combo kit was used for RT-PCR. The master mix was prepared according to the manufacturer’s guidelines in a designated “clean” reagent preparation room. In a separate “template” room, 5 µL of RNA was added to 20 µL of master mix. The TaqPath COVID-19 control was used per the manufacturer’s directions. Plates were run on an ABI 7500 Fast or ABI 7500 DX thermocycler according to the manufacturer’s cycling conditions, and results were interpreted using the COVID-19 Interpretive Software and manual visualization. For initial testing at the NDSU-VDL, a sample was considered positive or detected if there was an ORF1ab Ct value and sigmoidal curve because N and S genes were not detected reliably in spiked samples. This did deviate from the human protocol that required reportable Ct values for ≥2 genes. Some samples, if of human origin, would have been considered inconclusive. Additionally, the TaqPath COVID-19 Combo kit had been tested against a range of pathogens by the manufacturer in-silico for cross-reactivity that included human coronavirus strains, SARS-coronavirus, MERS-coronavirus, various parainfluenza viruses, and influenza A, B, and C viruses. 2 We determined that the chance of cross-reactivity amplification was very low or unlikely to occur.
Assay specificity
The NDSU-VDL also tested for cross-reactivity against canine and feline viruses and parasites in addition to other coronaviruses routinely tested at the NDSU-VDL using confirmed control material. These included: bovine coronavirus, canine distemper virus, canine herpesvirus, canine parvovirus, feline panleukopenia virus, canine respiratory coronavirus, feline calicivirus, feline coronavirus, feline herpesvirus, influenza A virus, Neospora caninum, porcine deltacoronavirus, and Toxoplasma gondii. Most controls were propagated from ATCC strains. The bovine coronavirus control was a modified-live virus vaccine diluted in PBS, and the canine respiratory coronavirus control was an aliquot of supernatant from homogenized tissue in Hank balance salt solution.
Assay sensitivity
A 6-point, 10-fold dilution series was run for each matrix at the NDSU-VDL to evaluate sensitivity. Each matrix was spiked with the heat-inactivated Washington isolate (from ATCC) to equate to 100,000 copies per reaction (c/rxn). Then, 10-fold serial dilutions, using the appropriate stock matrix as the diluent, were prepared, equating to 10,000, 1,000, 100, 10, and 1 c/rxn (~2,000, 200, 20, 2, and 0.2 c/µL, respectively). Each dilution was run in triplicate.
Sample stability
The stability of SARS-CoV-2 in different samples (PBS, canine swabs, canine lung homogenate, feline swabs, feline lung homogenate), temperatures (4°C, –20°C, –80°C), and over time (4 wk) was investigated by spiking a pooled stock of each matrix (PBS, canine oral swabs in PBS, canine lung homogenate, feline oral swabs in PBS, feline lung homogenate with the heat-inactivated Washington isolate [VR-1986HK, strain 2019-nCoV/USA-WA1/2020]). Spiking resulted in ~10,000 c/rxn. A sample matrix baseline Ct was set at week 0 for each matrix. Single-use aliquots were prepared for each sample matrix and each temperature to avoid freeze–thaw artifacts. RT-PCR was run on 2 extractions from single-use aliquots of spiked samples stored at 4°C, –20°C, or −80°C. This was repeated once per week for 4 wk.
Masked test
To evaluate assay performance in an unbiased manner, we conducted a masked test (MT) to detect SARS-CoV-2 in feline oral swabs and feline lung homogenate in addition to PBS. Masked (i.e., anonymized) samples were prepared in the Veterinary Laboratory Investigation and Response Network (Vet-LIRN) laboratory by spiking heat-inactivated Washington and Omicron variants in feline oral swab and lung matrices, PBS, and tubes supplied by NDSU-VDL. Canine samples were not included in the MT because of limited resources. The testing laboratory (NDSU-VDL) was not aware of the identity of the samples, which included: concentration per each spike level, number of spike levels (five: 0, 100, 250, 500, and 1,000 copies/400 µL), and number of replicates per spike level. This eliminated the opportunity for unconscious bias. Matrices were stored at −20°C until use. Biologic and PBS samples were thawed at 4°C and vortexed for at least 30 s, then 400 µL was aliquoted into a 1.7-mL tube. Heat-inactivated viral RNA stocks were serially diluted in corresponding sample matrix and spiked into tubes by adding 50 µL of diluted RNA. A total of 180 samples (i.e., 60 for each sample type and PBS) were made. A third was tested on day 1, a third on day 2, and a third was archived by MT organizers for possible follow-up. All samples were kept on ice during spiking. Day-1 and day-2 test samples were shipped overnight to NDSU-VDL on dry ice and stored at −80°C until extraction. A single analyst performed the extraction and RT-PCR. Day-1 extraction and RT-PCR were performed on the seventh day after the sample preparation by organizers, and day-2 samples were analyzed on the twelfth day after the sample preparation. SARS-CoV-2 was considered detected if there was an ORF1ab Ct value and sigmoidal curve, not detected if there was no ORF1ab Ct value, and inconclusive if the ORF1ab curve was not clearly sigmoidal.
Results
Assay specificity
Assay specificity was verified in-house (at NDSU-VDL) using non–SARS-CoV-2 coronaviruses and pathogens that can often be found in canine and feline samples (Table 1). There was no indication of amplification of the ORF1ab, N, or S genes, and the MS2 internal control was detected, confirming that the extractions and RT-PCR were successful.
Cross-reactivity of TaqPath COVID-19 Combo kit to other pathogens.
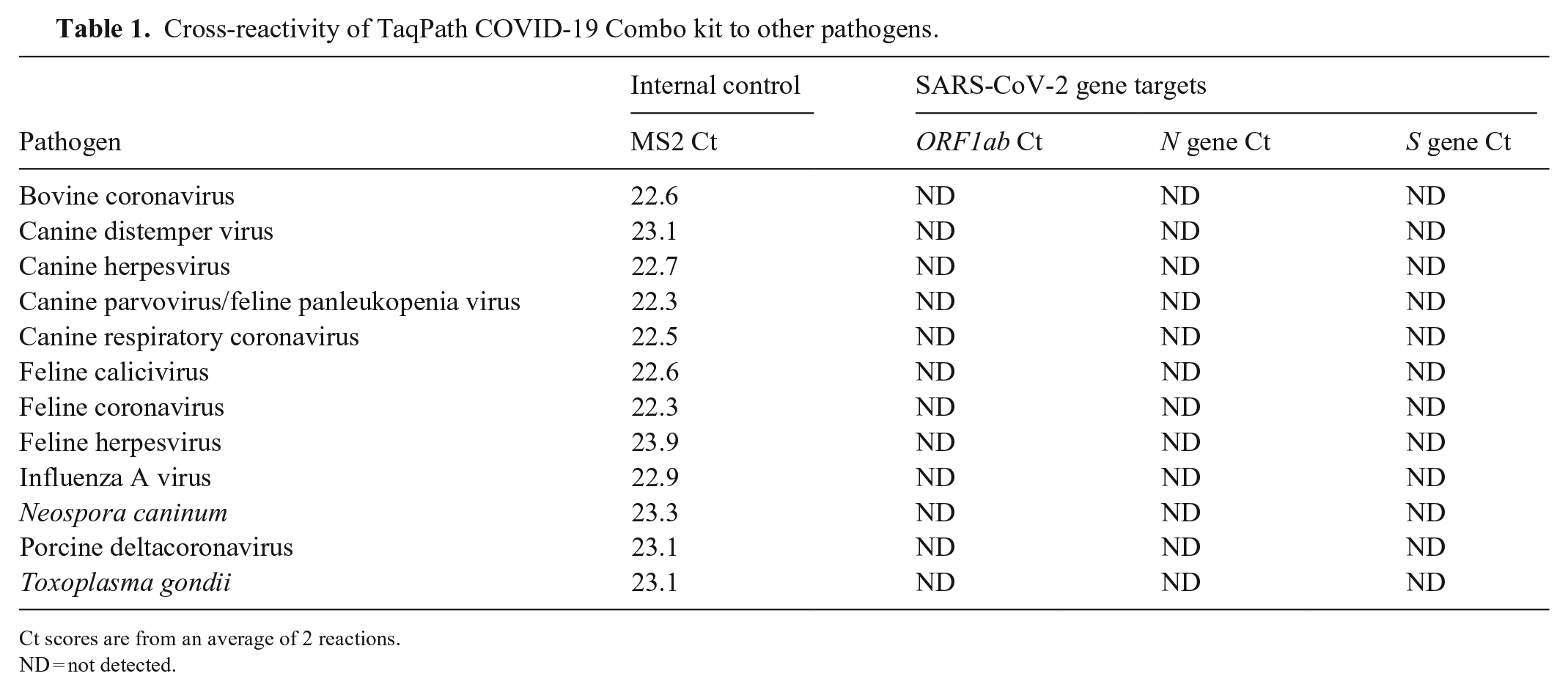
Ct scores are from an average of 2 reactions.
ND = not detected.
The ORF1ab gene was detected in all 3 replicates for each matrix at the 10 c/rxn level (Fig. 1), but in only 2 of 3 replicates at 1 c/rxn in canine lung homogenate, feline oral swabs, and PBS. Additionally, there was no detection of the N or S genes at 1 c/rxn for any of the 5 sample matrices.
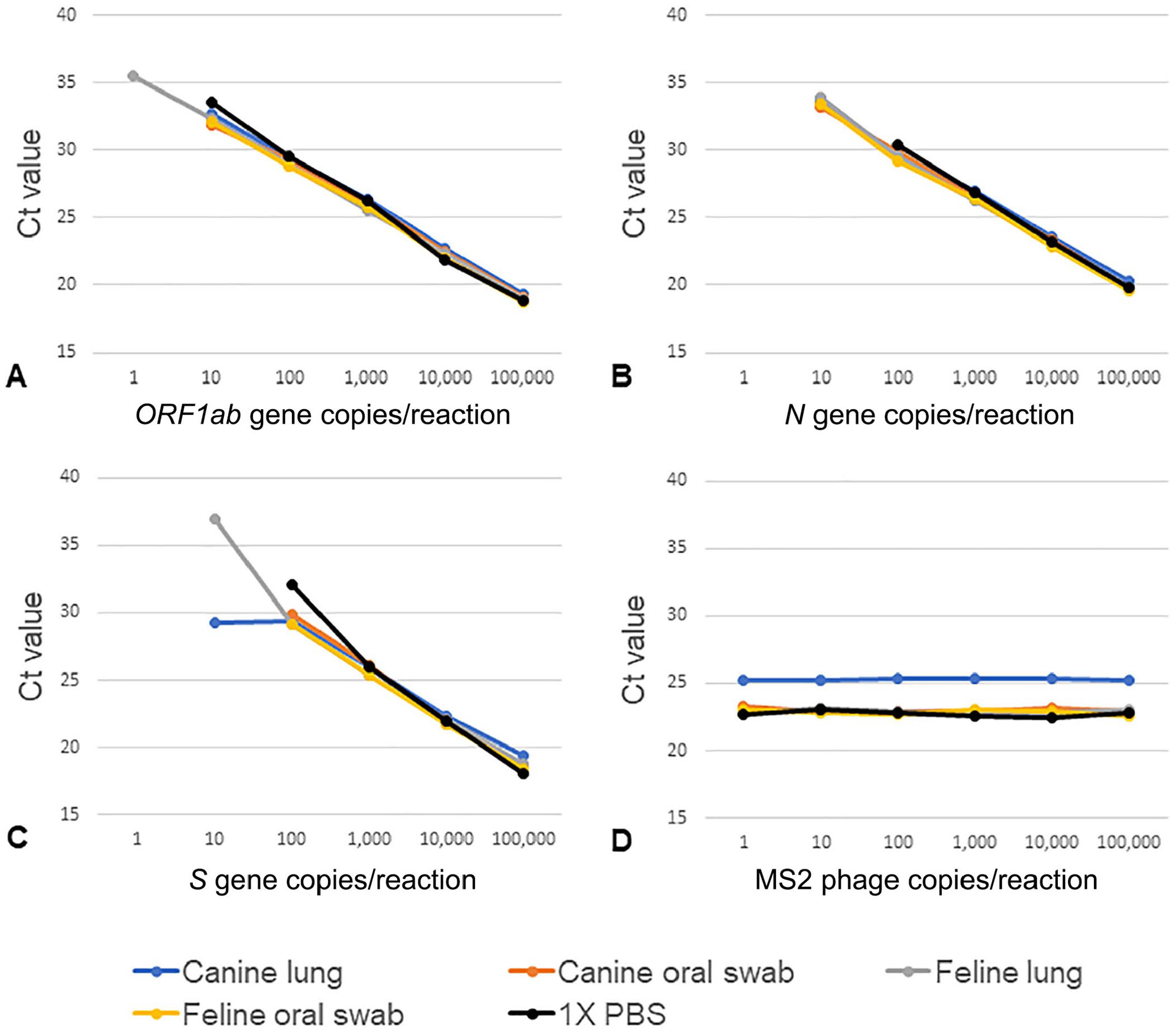
SARS-CoV-2 gene detection in sample matrices. All points are the mean of 3 replicate reactions.
Thermo Fisher established the LOD for the RT-PCR assay at 10 genomic copy equivalents (GCE) in human nasopharyngeal swabs (NP) and bronchoalveolar lavage (BAL). This equated to ORF1ab Ct values of ~30–31. 2 The LOD at ~10 c/rxn in each sample matrix was confirmed based on the ORF1ab gene from the dilution series and 12 RT-PCR replicates from the lowest dilution with consistent Ct values, 10 c/rxn (2 c/µL of eluate). The average Ct values over 12 replicates of 10 c/rxn in canine lung, canine swabs, feline lung, feline swabs, and PBS were 33.1, 32.6, 32.0, 31.9, and 33.1, respectively. Of note, the internal MS2 control had consistent Ct values in canine oral, feline oral, and feline lung samples across all dilutions; however, there was an overall increase of ~2 Cts, from 23 to 25, in canine lung matrix indicating potential for a matrix effect (Fig. 1; Table 2). Interestingly, the dilution series ORF1ab Ct values for canine lung samples were similar to the other sample matrices. The ORF1ab gene target was more consistently detected at lower levels of RNA, whereas N and S gene targets tended to shift or were no longer detected. When focusing on detection of the ORF1ab gene, the COVID-19 assay had high specificity and sensitivity for detection of SARS-CoV-2 in animal sample matrices.
Ct value for internal control, MS2, during SARS-CoV-2 detection across matrices.
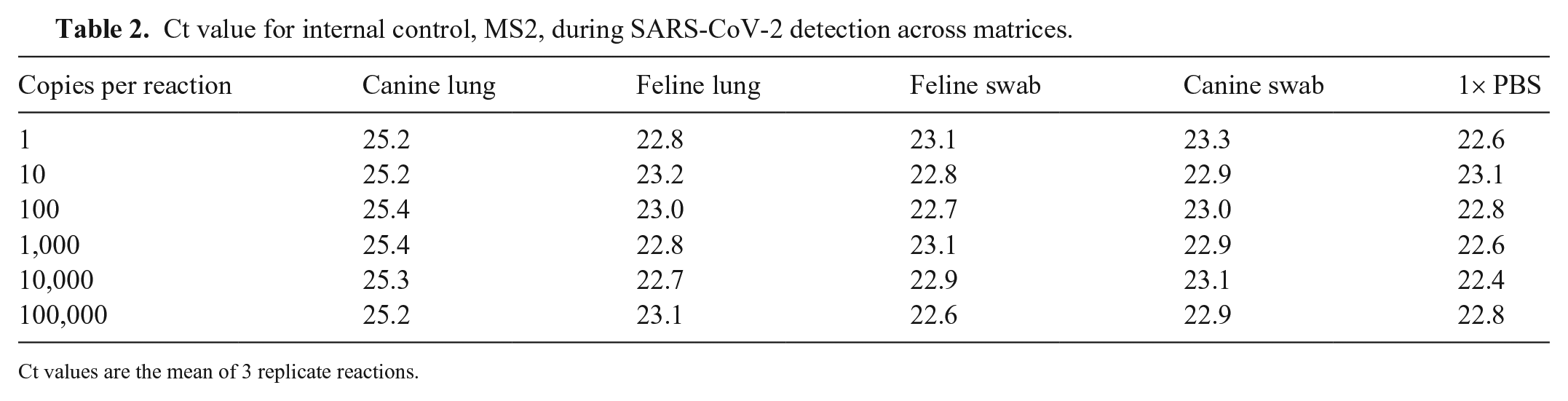
Ct values are the mean of 3 replicate reactions.
Sample stability
The Ct values changed <0.5 Cts from the baseline, after 4 wk in storage for all sample matrices at all temperatures, except canine lung at 4°C (Table 3). Canine lung samples at 4°C had a decrease of 2.3 Cts after 4 wk in storage. The lower Ct value in the fourth week was consistent between the 2 replicates, 22.4 and 22.2. Similarly, Ct values were also lower after 3 wk, 22.3 and 21.8. This could be the result of degradation of inhibitors, inconsistencies in spiking, or uneven distribution in separate aliquots.
SARS-CoV-2 stability at 4 wk in different storage temperatures based on ORF1ab gene marker.
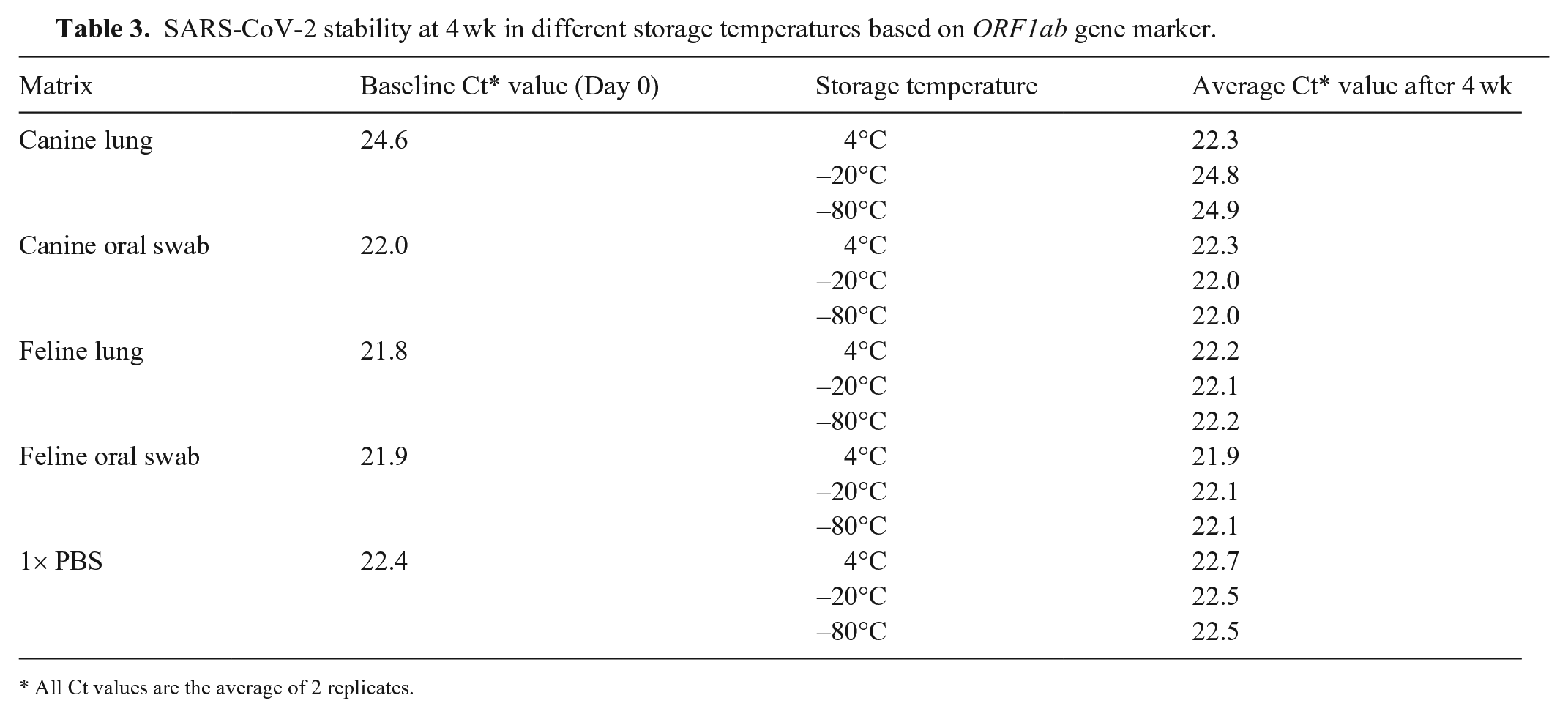

All Ct values are the average of 2 replicates.
Assay reliability for detection of SARS-CoV-2 in feline oral swabs
The rate of detection (ROD) was as expected for all PBS and feline oral swab matrix samples in the MT. Samples with WA-1 variant spiked into PBS or oral samples had a ROD of 0% for all blank samples and 100% for spike levels of 1,000, 500, 250, and 100 c/400 µL. No false-positives or false-negatives were detected. Similarly, samples spiked with the Omicron variant had a ROD of 0% for all blank samples and 100% for spike levels at 1,000, 500, and 250 c/400 µL, and 83% for PBS and 66% for oral swab matrix at 100 c/400 µL (Table 4). The ROD was not as expected for lung samples. The ROD was 50% for WA-1 variant spiked into lung at both 250 and 1,000 c/400 µL. No other spike levels were detected and, thus, resulted in false-negatives. No false-positives were detected. For Omicron variant samples, detection decreased as concentration decreased (100% for spike levels of 1,000 and 500, with 66% detection at 250 and 17% for 100 c/400 µL). No false-positives were detected (Table 4). Importantly, results matched the LOD as determined in-house by NDSU-VDL using non-masked samples.
Rate of detection (ROD) in feline samples and 1× PBS by matrix in masked test.
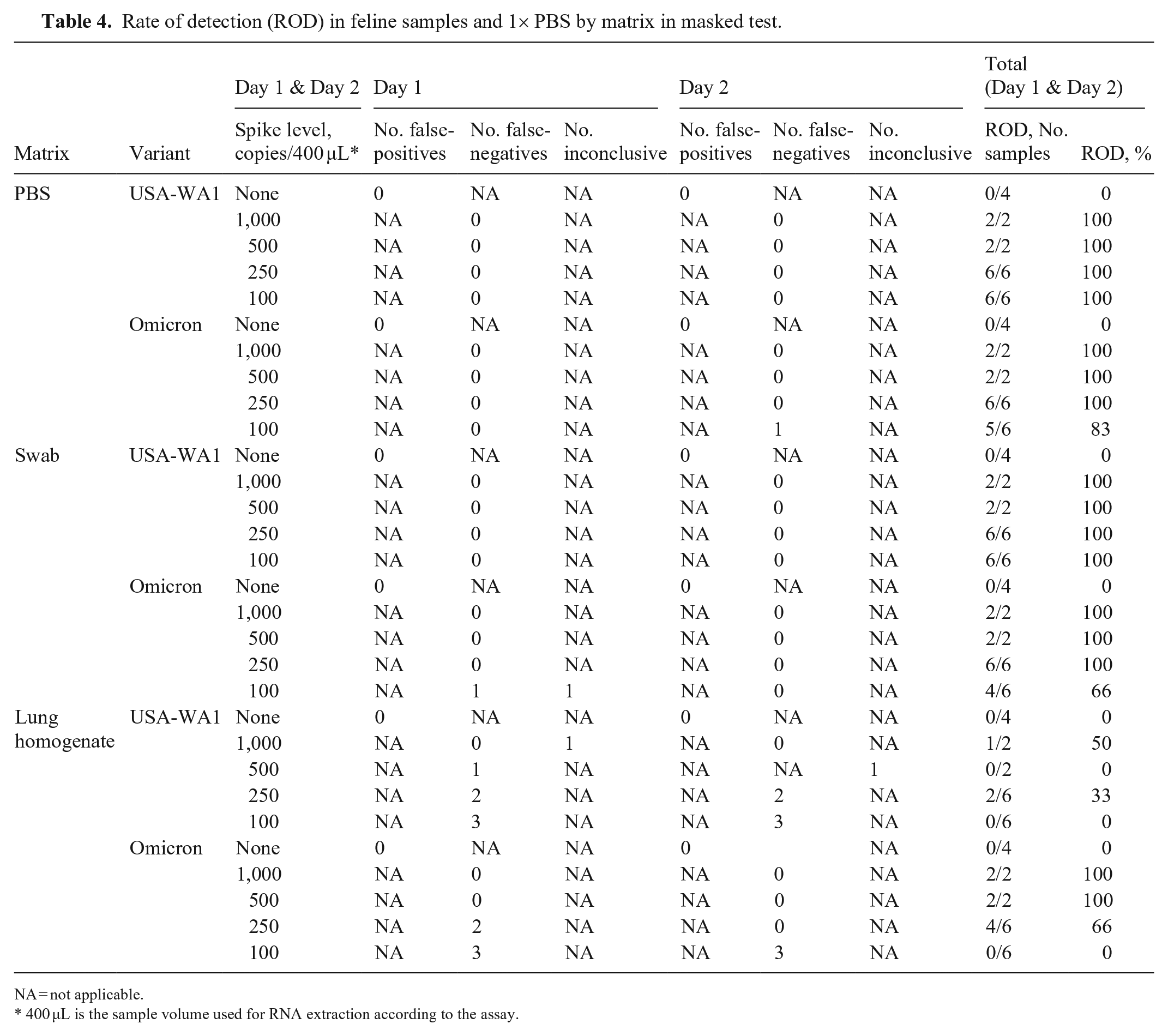
NA = not applicable.
400 µL is the sample volume used for RNA extraction according to the assay.
Discussion
In-house evaluation performed at the NDSU-VDL confirmed high specificity and sensitivity of the TaqPath COVID-19 Combo kit assay for SARS-CoV-2 in canine and feline oral and lung matrix samples in PBS. NDSU-VDL tested this kit against 13 pathogens including common coronaviruses that would likely be found in animal samples and found no amplification of any SARS-CoV-2 gene targets. Furthermore, the LOD was found to be similar for SARS-CoV-2 in animal sample matrices compared to human sample matrices, with a LOD of 10 c/rxn. Importantly, the ORF1ab gene target was more consistently detected at lower levels of RNA, whereas N and S gene Cts tended to increase or be undetected. This is consistent with other studies that have shown the lack of S-gene detection in samples that had significantly lower median Ct values of the ORF1ab and N-gene targets. 12
Therefore, we recommend focusing on the ORF1ab gene for detection along with manually checking amplification curves on the thermocycler software and Interpretive Software, and only using the N and S genes as supplemental for confirmation. This recommendation deviates from human guidelines that require ≥2 genes to be detected using the COVID-19 Interpretive Software. 2 However, the software functions used to analyze SARS-CoV-2 run files from 7500 Fast instruments is proprietary and not known. RT-PCR assay results (i.e., Cts) are usually interpreted on the software that runs a thermocycler and run template (e.g., ABI 7500 Software v.2.3; Thermo Fisher), but this is not the case with the TaqPath COVID-19 Combo kit. The run file must be imported into the Interpretive Software, which cannot be run on the same computer that operates the thermocycler software. We were not able to set a threshold on the run software that would result in the same Cts from the Interpretive Software, which is concerning. However, we see the utility of using the Interpretive Software, especially during the height of testing because it can quickly give a consistent answer for a large number of samples as detected, not detected, or inconclusive without additional human interpretation.
The MT tested the LOD with spiking levels of 1,000, 500, 250, and 100 RNA copies per 400 µL, or 100, 50, 25, or 10 copies per RT-PCR reaction. MT samples in PBS had a ROD of 100% at all spike levels and only 1 feline oral swab matrix sample at 100 copies/400 µL was called negative. This demonstrates excellent sensitivity of the assay for feline oral matrix samples. Results from the MT determined that lung homogenate matrix is not suitable for use with this assay. Together these data emphasize the importance of MTs to better evaluate performance characteristics and reveal possible limitations of the assay during assay development and validation.
Throughout testing at the NDSU-VDL, there were difficulties with consistent detection of SARS-CoV-2 in canine lung homogenate matrix. Even sequencing of samples from canine lung proved to be challenging, given that coverage across the genome and depth of coverage were low, resulting in erroneous clade classification compared to other sample types. MS2 phage control in feline lung homogenate was initially comparable to PBS and oral swab samples; however, the Cts increased when tested during the MT. During the MT sample preparation, several issues with feline lung sample matrix arose, which might explain the shift. First, particles (1–5 mm3) of lung tissue were visible in the lung homogenate sample supplied to the Vet-LIRN laboratory. Large pieces of lung tissue could impact viral RNA extraction, given that extraction may be efficient from the surface of those particles when handling naturally infected tissues. Second, when preparing MT samples, additional host cell lysis likely occurred at the Vet-LIRN laboratory as a result of additional freeze–thaw cycles causing additional and unexpected matrix effect and/or interference. Feline lung homogenate matrix at Vet-LIRN were exposed to an additional 2–3 freeze–thaw cycles: +4°C for 48 h, then exposure to room temperature for ~1 h during aliquoting.
A study looking at the matrix effect of saliva without RNA extraction prior to RT-PCR showed that there was a positive shift of the Ct values because of potential inhibitors in the matrix. 15 Although RNA extraction was performed on our samples before RT-PCR, the results from our study indicate that there is a matrix effect from lung tissue, which may limit detection of SARS-CoV-2. We encourage standardization of the duration, intensity (i.e., speed) of the homogenization, or type of homogenization probe used. Other studies have shown that the stability of SARS-CoV-2 within matrices dramatically decreases within 24 h under warm conditions, including room temperature. 13 Therefore, stability tests at room temperature at 1, 2, 4, 6, and 12 h would be advised for future validation of lung samples. This would provide more information on the effects of higher temperature, given that exposure to different temperatures may occur during sample collection and preparation.
The lung-matrix effects observed for the MS2 phage and SARS-CoV-2 also have implications for detection of other pathogens in various types of tissues. There is no universal standard processing method for fresh tissues across VDLs or human laboratories. This variation can also be compounded by different extraction methods and kits as well as master mix reagents.4,7 Internal validation data at the NDSU-VDL have shown that the same sample extract used in RT-PCR reactions with the same primers and probes, but different master mix reagents and/or polymerases, can result in different Ct values. We were able to narrow differences in MS2 phage Ct values to matrix effects because all of the samples were extracted and RT-PCRs were run with the same reagents. However, extraction and PCR reagents should also be carefully considered during validation. Matrix effects may not be as apparent in other situations, especially if testing different extraction methods and PCR reagents in the validation process.
We showed that the human testing workflow with the COVID-19 assay, including the TaqPath COVID-19 Combo kit, MagMAX Viral/Pathogen II, and 7500 Fast instruments, can be used to detect SARS-CoV-2 in oral swab samples from cats and likely dogs. Strict adherence to the manufacturer’s reporting instructions for humans is not appropriate for animal samples. We suggest that individual laboratories follow and establish their own parameters for determination of detection status based on Ct values and evaluations of amplification curves from run files and the Interpretive Software. Parameters could include setting specific Ct value cutoff limits, specific gene target or number of targets detected, and shape of amplification curves. The best practice is to rerun PCR, extraction + PCR, or collect a new sample. We also found that combining the binding solution, magnetic beads, proteinase K, and MS2 phage control, rather than adding individually to wells, saves time and pipette tips. Oral swab samples in PBS do not need to be fresh and can be stored cold or frozen for a month, and possibly longer. Our work emphasizes the importance of a One Health approach. Not only is SARS-CoV-2 a zoonotic pathogen that can be transmitted between species, but testing protocols developed for human samples can be adapted to reliably detect SARS-CoV-2 in animal samples.
Footnotes
Acknowledgements
We thank NDSU-VDL and West Fargo Animal Hospital staff for assistance.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
Our study was supported by the Food and Drug Administration (FDA) of the U.S. Department of Health and Human Services (HHS) as part of a financial assistance award 1U18FD007519-01 totaling $59,940 with 100% funded by FDA/HHS. The contents are those of the authors and do not necessarily represent the official views of, nor an endorsement by, FDA/HHS or the U.S. Government. Reference to any commercial materials, equipment, or process does not in any way constitute approval, endorsement, or recommendation by the FDA.
