Abstract
Surveillance for SARS-CoV-2 in American mink (Neovison vison) is a global priority because outbreaks on mink farms have potential consequences for animal and public health. Surveillance programs often focus on screening natural mortalities; however, significant knowledge gaps remain regarding sampling and testing approaches. Using 76 mink from 3 naturally infected farms in British Columbia, Canada, we compared the performance of 2 reverse-transcription real-time PCR (RT-rtPCR) targets (the envelope [E] and RNA-dependent RNA polymerase [RdRp] genes) as well as serology. We also compared RT-rtPCR and sequencing results from nasopharyngeal, oropharyngeal, skin, and rectal swabs, as well as nasopharyngeal samples collected using swabs and interdental brushes. We found that infected mink were generally RT-rtPCR–positive on all samples; however, Ct values differed significantly among sample types (nasopharyngeal < oropharyngeal < skin < rectal). There was no difference in the results of nasopharyngeal samples collected using swabs or interdental brushes. For most mink (89.4%), qualitative (i.e., positive vs. negative) serology and RT-rtPCR results were concordant. However, mink were positive on RT-rtPCR and negative on serology and vice versa, and there was no significant correlation between Ct values on RT-rtPCR and percent inhibition on serology. Both the E and RdRp targets were detectable in all sample types, albeit with a small difference in Ct values. Although SARS-CoV-2 RNA can be detected in multiple sample types, passive surveillance programs in mink should focus on multiple target RT-rtPCR testing of nasopharyngeal samples in combination with serology.
Although severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2; Coronaviridae) is primarily a human pathogen, mink are also highly susceptible because of homology between the angiotensin-converting enzyme 2 (ACE2) receptor in mink and humans. 13 Indeed, there have been multiple spillover events in which SARS-CoV-2 has been transmitted from farm workers to farmed mink with subsequent mink-to-mink transmission. 26 As of January 2023, outbreaks in farmed mink have been reported in Canada, Denmark, France, Greece, Italy, Latvia, Lithuania, the Netherlands, Poland, Spain, Sweden, and the United States. 34
The clinical presentation and epidemiology of these outbreaks have been somewhat variable. In some, infection of mink on affected farms has occurred in the absence of visible disease2,24; in other cases, there has been increased mortality as well as clinical signs including reduced feed intake, nasal discharge, respiratory distress, and diarrhea.4,23,33 Postmortem examinations of naturally and experimentally infected mink have indicated that SARS-CoV-2 primarily affects the respiratory tract causing rhinitis and interstitial pneumonia.1,23,28 Immunohistochemistry has demonstrated the presence of SARS-CoV-2 in the nasal mucosa as well as the lungs, which are also tissues in which the highest levels of SARS-CoV-2 RNA and live virus have been found.1,28,29
In addition to the animal health implications, SARS-CoV-2 outbreaks in farmed mink are problematic from a public health standpoint because SARS-CoV-2 can be transmitted from mink back to people.8,19,21,25 As the virus adapts to the mink host, mutations could accumulate at a faster rate than in human hosts, 25 giving rise to novel variants with the potential to impact transmissibility or immunity in humans.3,9,15 For this reason, surveillance for SARS-CoV-2 in farmed mink has been identified as a global priority.10,18
Surveillance of SARS-CoV-2 in mink is centered around reverse-transcription real-time PCR (RT-rtPCR) testing of samples obtained from live and dead mink 10 ; however, there is considerable uncertainty around the optimal sampling protocol. Oropharyngeal swabs are recommended most commonly, 10 although studies have suggested that SARS-CoV-2 is detected more easily in the nasal versus the oral cavity of infected mink.1,29 In choosing between nasal and oropharyngeal swabs, feasibility must also be considered. Mink have very small nares and nasal cavities, and it may be challenging for animal health professionals to obtain swabs that are appropriate for collecting samples from the deep nasal cavity or nasopharynx, particularly during global supply chain shortages. Interdental or gum brushes have been suggested for the collection of host DNA for genetic testing 30 and could provide an alternative to medical-grade swabs for the collection of SARS-CoV-2 samples from the nasal cavity of mink.
The utility of rectal swabs or feces is unclear. Although some studies have found that infected mink are less likely to shed virus and/or shed a lower quantity of virus in the gastrointestinal versus respiratory tract,1,23,33 other studies have shown that the ACE2 receptor is present in the mink small intestine, 20 and there have been SARS-CoV-2–infected mink farms in which diarrhea is the only clinical sign. 4 Interestingly, SARS-CoV-2 is shed in the feces of 30–60% of infected humans and can be detected in feces for a much longer time compared to respiratory samples, 17 with average fecal shedding lasting 27.9 days in one study. 35 Likewise, sample type may impact the quality of sequence data generated by whole-genome sequencing (WGS). 22 WGS has proven to be instrumental in studying the biology and epidemiology of SARS-CoV-2 and it is critical that alternative sampling methods yield high-quality genomic data.
There is a similar paucity of data regarding the presence of virus on pelts. To date, the World Organisation for Animal Health recommends against the selling of raw pelts because of biosecurity risks. 24 However, there are very few reports of SARS-CoV-2 on the fur or pelts from infected mink.1,4 The degree to which pelts from infected animals are contaminated is important because this is the purpose for which mink are produced, and the inability to sell pelts as a result of bio-security risks has adverse economic impacts on infected farms.
Serology is often used in partnership with RT-rtPCR testing given that seropositivity can indicate that animals have been exposed to SARS-CoV-2 even if they are RT-rtPCR–negative.2,24 The correlation between RT-rtPCR and serology results appears to vary based on the timeline of an outbreak, with RT-rtPCR–positive animals being more common than seropositive animals during the prodromal period and period of disease, whereas the reverse is true during periods of decline and convalescence. 4 However, little is known about the correlation between RT-rtPCR and serology results within an individual animal.
Finally, it has been suggested that the emergence of new mutations in SARS-CoV-2 could impact the sensitivity of different RT-rtPCR assays and that a multiplex RT-rtPCR approach targeting different parts of the genome should be used. 10 However, there is little evidence regarding the comparative performance of different RT-rtPCR assays in mink.
Using mink from 3 naturally infected farms in British Columbia (BC), Canada, our objectives were: 1) to compare the performance of RT-rtPCR assays targeting the envelope (E) and RNA-dependent RNA polymerase (RdRp) genes of SARS-CoV-2 in mink; 2) to compare RT-rtPCR results for nasal samples obtained using nasopharyngeal swabs and interdental bushes; 3) to determine whether SARS-CoV-2 can be detected in the pelt or subcutaneous tissue of infected mink; 4) to compare RT-rtPCR results for nasal, oropharyngeal, rectal swabs, and skin samples; 5) to compare WGS results obtained from nasal, oropharyngeal, rectal swabs, and skin samples; and 6) to compare serology and RT-rtPCR results.
Materials and methods
SARS-CoV-2 outbreaks and passive surveillance on BC mink farms
Between December 2020 and April 2022, there were SARS-CoV-2 outbreaks in 3 mink farms in BC. Outbreaks on farms A and B were short-lived, and the producers reported a significant increase in mortality rate above what would be considered “normal” for those farms (Suppl. Table 2). The outbreak on farm C was detected on passive surveillance (see below), and the producer reported no clinical signs or increased mortality rate associated with SARS-CoV-2 detection in their herd. The AW.1 strain of SARS-CoV-2 was detected on farms A and B; the B.1 and B.1.618 strains were detected on farm C. Mink on all farms had black coat color.
In response to the initial outbreak on farm A, the BC Ministry of Agriculture and Food (AFF) initiated a SARS-CoV-2 passive surveillance program for both infected and non-infected farms. That program required all mink farms in BC to submit all natural mortalities (regardless of putative cause) to the Animal Health Centre (Abbotsford, BC, Canada), which is the AFF veterinary diagnostic laboratory, for SARS-CoV-2 testing by RT-rtPCR on nasopharyngeal swabs. The cause of death was not investigated for any of the mink included in the AFF SARS-CoV-2 passive surveillance program, and AFF did not collect data on morbidity or mortality rates for any of the farms under surveillance, including the 3 infected farms. The purpose of the surveillance program was to detect newly infected farms and to monitor for ongoing SARS-CoV-2 transmission on farms on which outbreaks occurred.
Sample collection
As part of the AFF surveillance program, 249 mink from the 3 infected farms were sampled. We collected an expanded sample set from each of 76 of the 249 mink, including nasopharyngeal swab, nasopharyngeal brush, oropharyngeal swab, rectal swab, subcutaneous swab, and serum. Nasopharyngeal samples were collected using both a flocked swab (Becton Dickinson) and an interdental brush (Proxabrush Go-Betweens Tight; Sunstar). Sterile polyester-tipped applicators (Puritan) were used for oropharyngeal, rectal, and skin swabs. For animals that had been pelted prior to submission, the subcutaneous swab was collected by swabbing an ~10-cm2 area of the carcass in the region of the dorsocranial thorax. For animals that were submitted with the pelt on, the mink was skinned using a sterile scalpel, and the analogous area of the subcutaneous tissue adherent to the pelt was swabbed. All swabs and brushes were placed in viral transport medium (VTM; Puritan) and stored at −20°C prior to submission to the BC Centre for Disease Control Public Health Laboratory (BCCDC-PHL, Vancouver, BC, Canada). Whole blood samples were collected using Nobuto filter strips (Fisher Scientific) by saturating the length of the strip with cardiac blood. Nobuto strips were allowed to air dry then placed into individual envelopes and stored at 4°C until shipped to the National Microbiology Laboratory (NML; Winnipeg, Manitoba, Canada) for testing.
RNA extraction, RT-rtPCR, and sequencing
The BCCDC-PHL developed a multiplex RT-rtPCR assay targeting the E 7 and RdRp 12 genes of SARS-CoV-2, as described previously (Suppl. Table 1). Briefly, total nucleic acids were extracted from 200 μL of inoculated VTM (MagMax Express 96 nucleic acid extractor, Applied Biosystems; MagMax viral/pathogen nucleic acid isolation kit, Thermo Fisher) according to the manufacturers’ recommendations. The RT-rtPCR assay was performed (TaqMan fast virus master mix, Applied Biosystems 7500 FAST real-time RT-RTPCR system; Thermo Fisher) as described previously. 12 Mink with nasal swabs with Ct values < 36.0 were considered positive, those with Ct values of 36.0–40.0 were considered suspect, and those with Ct values > 40.0 were considered negative.
RT-rtPCR–positive samples were sequenced across the whole genome using a 1,200-bp tiled amplicon approach. 11 Complementary DNA was synthesized (Superscript IV; Invitrogen), and amplicons were generated with 2 primers pools. Sequencing libraries were generated using an abbreviated version of the DNA preparation protocol (Illumina) and sequenced (MiSeq or NextSeq platform; Illumina). 14 Sequence data were analyzed (modified Nextflow bioinformatics pipeline, https://github.com/BCCDC-PHL/ncov2019-artic-nf) to assess quality and generate consensus sequence.
Serology
At the NML, to obtain the 1:9 serum dilution required for testing, saturated Nobuto strips were cut into 4–5 pieces and placed into a 2-mL Sarstedt tube containing 360 µL of PBS (pH 7.4) containing 0.05% polysorbate 20 and eluted overnight at 4°C. Following elution, samples were mixed by vortexing and tested (cPass SARS-CoV-2 neutralization antibody detection kit; GenScript) according to the manufacturer’s protocol. Briefly, 60 µL of sample was added to 60 µL of HRP-conjugated receptor-binding domain solution and incubated at 37°C for 30 min. A 100-µL aliquot of the mixture was transferred to the ELISA microwell test plate and incubated at 37°C for 15 min. Microwells were washed 4 times with 260 µL of wash buffer, then 100 µL of 3,3’,5,5’-tetramethylbenzidine (TMB) substrate was added to each well. Following a 20-min incubation in the dark at room temperature, 50 µL of stop solution was added to each well. Absorbance was read immediately at 450 nm. Each assay plate included positive and negative controls that met the required quality control parameters. Percentage inhibition was calculated for each sample using the following: %inhibition = (1 – OD sample/OD negative control) × 100%. Samples with > 30% inhibition were considered positive for SARS-CoV-2 neutralizing antibodies. Given the limited volume of eluate obtained from the Nobuto strips, each sample was tested only once.
Statistical analysis
Unless otherwise stated, results obtained from the RT-rtPCR assay for the E gene were used for statistical analysis because this is the current standard for animal testing in Canada. However, analyses were repeated using the results obtained from the RdRp RT-rtPCR assay as a sensitivity analysis (i.e., to determine if results varied by assay).
A repeated-measures ANOVA using the package rstatix was used to compare Ct values among samples obtained via nasal swab, nasal brush, oral swab, rectal swab, and skin swab. Subsequently, multiple pairwise paired t-tests were used to compare Ct values between each sample type using the Bonferroni multiple testing correction method to adjust the p-values. Paired t-tests were also used to compare Ct results obtained from the E and RdRp gene assays for each sample type. Finally, the Pearson correlation coefficient was used to compare Ct value and %inhibition on serology.
The suitability of each sample type for WGS analyses was determined by examining genome completeness. A Friedman test followed by the Nemenyi post-hoc pairwise test was conducted to identify differences in sample quality, as measured by coverage across the full viral genome among the 5 sample types. Sphericity and normality assumptions were assessed with Levene and Shapiro tests, respectively. P values ≤ 0.05 were considered statistically significant.
All analyses and visualization were performed in R v.3.5.2 (https://www.r-project.org/).
Results
Among the 76 mink included in our study, 18 were from farm A, 3 were from farm B, and 55 were from farm C (Suppl. Table 2). A total of 32 of 76 (42%) mink were RT-rtPCR–positive, and 3 of 76 (4%) were suspect-positive. Among the 32 positive mink, SARS-CoV-2 RNA was detected in all sample types from 28 (88%) animals and in nasal and oral samples from 4 (12%) animals (Suppl. Excel spreadsheet). For the 3 suspect mink, SARS-CoV-2 RNA was detected only in nasal samples from 2 animals and in nasal and oral samples from 1 animal.
Among the 28 mink in which SARS-CoV-2 RNA was detected in all samples, average Ct values (with range) for each sample type were as follows (Fig. 1): 18.4 (15.8–22.3) for nasal swab, 18.1 (13.8–22.4) for nasal brush, 21.8 (17.4–26.8) for oral swab, 30.9 (23.6–37.1) for rectal swab, and 27.4 (19.9–37.5) for skin swab. Repeated measures ANOVA revealed that Ct values were significantly different among the different sample types (p < 0.001). In the post-hoc analysis with a Bonferroni adjustment, all pairwise differences between sample types were significantly different (p < 0.001) except for the comparison between nasal swabs and nasal brushes (p = 1.0).
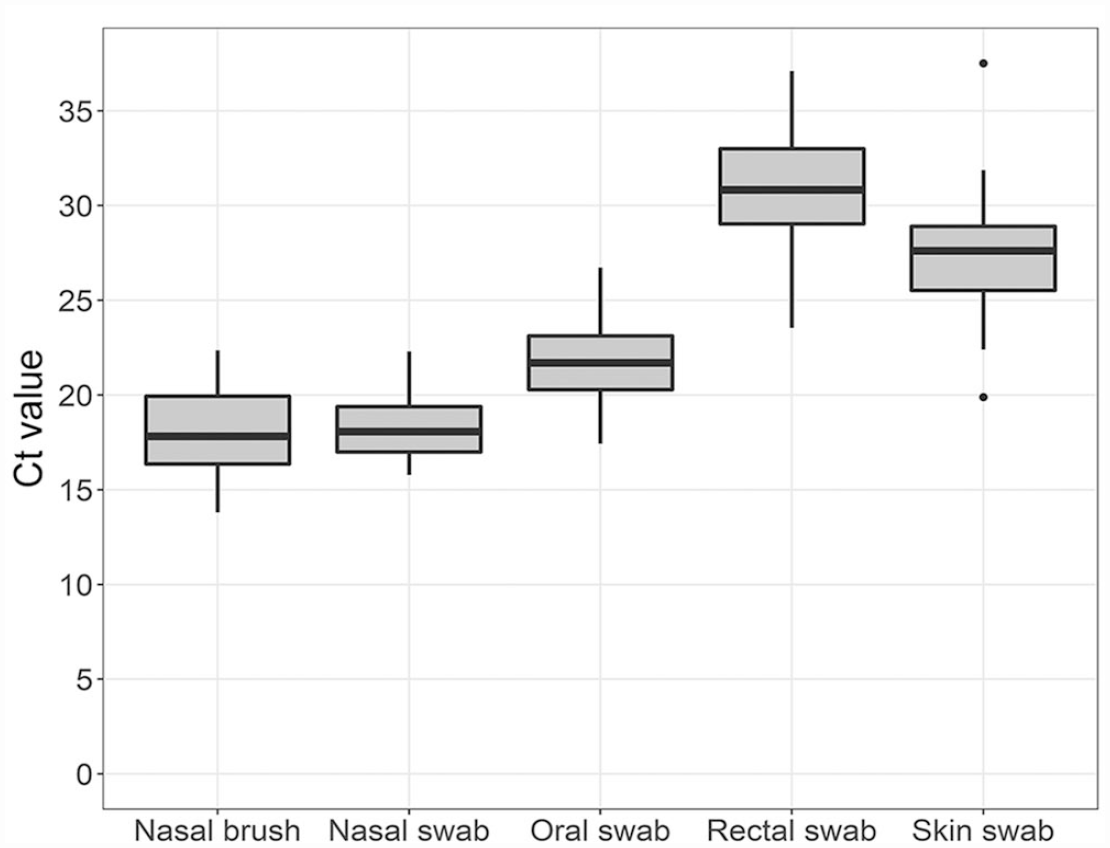
Comparison of SARS-CoV-2 E gene RT-rtPCR Ct values for different sample types obtained from mink. Midlines represent the median value, with outer bounds of boxes delineating the interquartile range (IQR) of Ct values for a sample type. Error bars show the most extreme point within 1.5*IQR; outliers beyond this range are signified by dots.
For all sample types except rectal swabs, the mean Ct value was slightly but significantly less using the RdRp gene assay compared to the E gene assay. The mean (95% CI) difference in Ct values between the 2 assays for each sample was: 0.41 (0.32–0.52) for nasal swab, 0.45 (0.37–0.54) for nasal brush, 0.38 (0.28–0.49) for oral swab, 0.43 (–0.07 to 0.93) for rectal swab, and 0.43 (0.29–0.57) for skin swab. However, the relative Ct values of the different sample types did not change regardless of which assay was used for the dependent variable.
Whole-genome sequences were obtained from the 35 SARS-CoV-2–positive and –suspect mink and there was a statistically significant difference in genome completeness of SARS-CoV-2 depending on sample type (p < 0.001). Median (IQR) genome completeness for nasal brushes, nasal swabs, oral swabs, rectal swabs, and skin swabs were 0.994 (0.978–0.994), 0.994 (0.992–0.994), 0.992 (0.974–0.994), 0.000 (0.000–0.453), and 0.891 (0.128–0.989), respectively. A statistically significant improvement in genome completeness existed between nasopharyngeal (nasal brush, nasal swab, oral swab) and skin or rectal swabs (Fig. 2). However, no significant differences were detected among the nasopharyngeal samples or among the skin and rectal swabs.
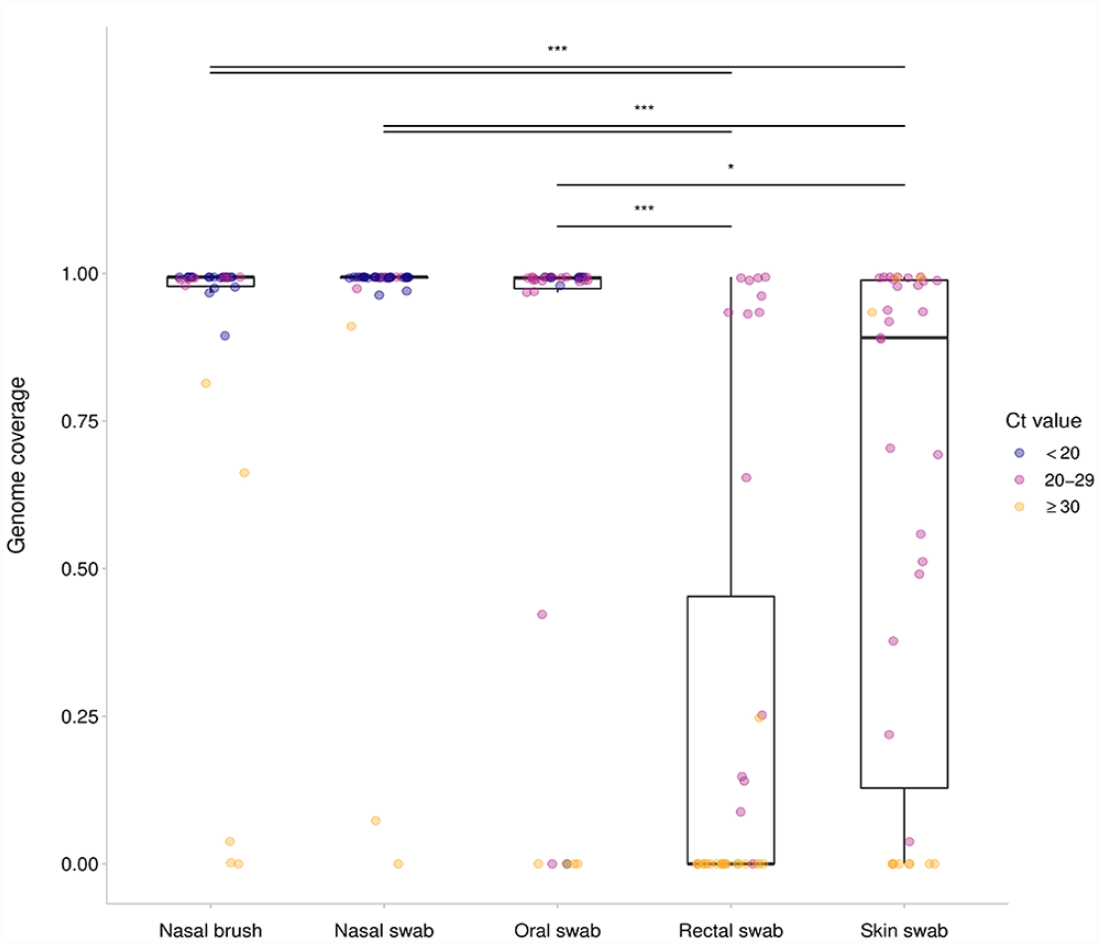
Genome completeness of SARS-CoV-2 collected using nasal brush, nasal swab, oral swab, rectal swab, and skin swab from 35 mink across 3 farms. Midlines represent the median value, with outer bounds of boxes delineating the interquartile range (IQR) of genome completeness (min. depth per site = 10 reads) for a sample type. Error bars show the most extreme point within 1.5*IQR; data beyond this range are outliers. The color of a point shows whether that sample had a Ct value of < 20 (blue), 21–29 (pink), or ≥ 30 (orange) for the RdRp gene.
Forty of 76 (53%) mink were negative on both RT-rtPCR and serology, and 29 (38%) mink were RT-rtPCR–positive or –suspect and positive on serology. Six mink were positive on RT-rtPCR and negative on serology, and one mink was negative on RT-rtPCR and positive on serology. There was no significant correlation (p = 0.95) between Ct values obtained by nasal swab and %inhibition on serology.
Discussion
Comparing several sample types and diagnostic assays for the detection of SARS-CoV-2 in naturally infected mink, we found that samples obtained from the nasal cavity produced significantly lower Ct values than oropharyngeal samples. This is consistent with previous findings that the virus has particular tropism for the nasal mucosa. 20 Contrary to current recommendations,10,26 sampling of the nasal cavity may be preferable to sampling the oral cavity of dead mink. This may, however, not be feasible for live mink given the small size of mink nares and the fact that the semi-aquatic nature of mink allows them to tightly close the nares. With regard to the method of sampling the nasal cavity, we found that there was no significant difference between using swabs and interdental brushes. Interdental brushes were easier to procure during the COVID-19 pandemic when sampling consumables were in short supply and provided an effective, accessible, and cost-effective alternative to clinical-grade swabs.
Rectal swabs consistently yielded higher Ct values (lower viral loads) compared to samples from the oral and nasal cavities. This is consistent with previous findings, which have suggested that viral replication occurs mainly in the upper respiratory tract with minimal involvement of the gastrointestinal tract, and that viral RNA detected in fecal matter may be the consequence of the ingestion of respiratory tract exudate rather than infection of the intestinal mucosa.1,20,23,33 Studies have suggested that people may shed virus in the gastrointestinal tract even after clearing infections from the respiratory tract,17,35 but we did not detect a similar phenomenon in this group of mink, even during the protracted outbreak on farm C. However, it is important to note that our study was cross-sectional (i.e., each mink was sampled only once); therefore, duration of infection in any of the included mink could not be determined (i.e., even mink submitted 11 wk after the detection of the outbreak on farm C could have been acutely infected). Serial oral or nasal and rectal sampling of infected live mink may help to clarify SARS-CoV-2 shedding dynamics.
Interestingly, SARS-CoV-2 was detected consistently in the subcutaneous tissues of infected mink at Ct values intermediate between those associated with oral and rectal samples. As opposed to SARS-CoV-2 detection in mink fur, which is likely a result of grooming activity or contamination of the cage environment,1,4 consistent detection of SARS-CoV-2 in the subcutis of infected animals suggests viremia. Interestingly, although SARS-CoV-2 viremia is relatively common in humans, 16 it has been less well described in mink. Regardless of the source of the virus, contamination of any part of the pelt may limit options for processing and sale of pelts and thus present additional negative economic consequences for infected farms.
With regard to the various assays under consideration, although there was a high degree of agreement between qualitative (i.e., positive vs. negative) serology and RT-rtPCR results, some animals were positive on RT-rtPCR and negative on serology, and vice versa. Additionally, RT-rtPCR Ct values (a proxy for viral load) and serology %inhibition (a proxy for immunologic reaction) were not well correlated. This is likely a result of the fact that the mink included in our study were in different stages of infection or immunologic reactivity. Indeed, studies have shown that mink can begin shedding virus as soon as 1–2 d after infection, although antibodies may not be detectable until 14–18 d after infection.1,29 Similarly, viral shedding may cease quickly whereas antibody titers remain for long periods of time. 1 At a farm level, a greater prevalence of RT-rtPCR positivity has been found compared to seropositivity early in an outbreak, although the reverse is true during the later stages. 4 The correlation between serologic and RT-rtPCR results may be further obscured by the fact that mink can become re-infected over the course of an outbreak, resulting in a new period of shedding and a stronger immunologic reaction. 27 The possibility of re-infection is greater where outbreaks are prolonged, such as that on farm C. Overall, we believe that combining RT-rtPCR and serology may be helpful to maximize the detection of mink that have been exposed to or infected with SARS-CoV-2. This is particularly important given that outbreaks may not be associated with any clinical signs or increased mortality in infected mink.2,24
We found that there was a small but significant difference between RT-rtPCR assays targeting the SARS-CoV-2 E versus RdRp gene. Both assays were clearly capable of detecting the 3 different strains of virus circulating on the 3 infected farms and in different sample types. The small difference in average Ct value was not sufficient to impact diagnostic yield during these outbreak investigations. That being said, in the case of novel or evolving pathogens, it is optimal to utilize assays with 2 or more RT-rtPCR targets across the genomes to account for variations in sensitivity as well as mutations that may cause primer–probe mismatch or loss of detection. These detection failures have been observed throughout the pandemic, as was the case with the “S-gene target failure” with several SARS-CoV-2 lineages5,6 and decreased sensitivity for N-gene targets with the Omicron variants. 31
Finally, we found that sample type was associated with WGS success. Samples collected with nasal brush, nasal swab, or oral swab generated the most complete genomes. This likely reflects the relative viral load in each sample type, which can be approximated by the Ct value. 32 In our laboratory, samples with Ct < 33 were more likely to generate a complete genome. This suggests that respiratory secretions are optimal where the goal is SARS-CoV-2 detection and sequencing in an individual animal; however, manure and environmental samples may be efficient and effective for population monitoring, particularly if advanced genomic techniques can be used to obtain sequences and track viral evolution.
Supplemental Material
sj-pdf-1-vdi-10.1177_10406387231183685 – Supplemental material for A comparison of sampling and testing approaches for the surveillance of SARS-CoV-2 in farmed American mink
Supplemental material, sj-pdf-1-vdi-10.1177_10406387231183685 for A comparison of sampling and testing approaches for the surveillance of SARS-CoV-2 in farmed American mink by Chelsea G. Himsworth, Jessica M. Caleta, Michelle Coombe, Glenna McGregor, Antonia Dibernardo, Robbin Lindsay, Inna Sekirov and Natalie Prystajecky in Journal of Veterinary Diagnostic Investigation
Footnotes
Acknowledgements
We thank Christine Millar, Ciara O’Higgins, and Kirsten Tweedhope of the AHC for their assistance with sample collection. We also thank the laboratory staff at BCCDC-PHL for their assistance in carrying out RT-rtPCR and WGS.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This study was funded by Genome British Columbia (COV-200; Vancouver, BC, Canada).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
