Abstract
Gallibacterium spp., particularly G. anatis, have received much attention as poultry pathogens in recent years. We report here the presence and antimicrobial resistance profile of 69 Gallibacterium isolates obtained from 2,204 diagnostic submissions of broiler and layer chickens in 2019–2021. Gallibacterium-positive chickens had lesions primarily in the respiratory tract, reproductive tract, and related serosal surfaces. Gallibacterium spp. were initially identified based on their typical cultural characteristics on blood agar. The isolates were confirmed by a genus-specific PCR spanning 16S-23S rRNA and MALDI-TOF mass spectrometry. Phylogenetic analysis based on 16S rRNA gene sequence revealed distinct clades. Of the 69 isolates, 68 clustered with the reference strains of G. anatis and 1 with Gallibacterium genomospecies 1 and 2. Antimicrobial susceptibility testing of 58 of the 69 isolates by a MIC method showed variable responses to antimicrobials. The isolates were all susceptible to enrofloxacin, ceftiofur, florfenicol, and gentamicin. There was a high level of susceptibility to trimethoprim–sulfamethoxazole (98.0%), streptomycin (98.0%), amoxicillin (84.0%), sulfadimethoxine (71.0%), and neomycin (71.0%). All of the isolates were resistant to tylosin. There was resistance to penicillin (98.0%), erythromycin (95.0%), clindamycin (94.0%), novobiocin (90.0%), tetracycline (88.0%), oxytetracycline (76.0%), and sulfathiazole (53.0%). A high rate of intermediate susceptibility was observed for spectinomycin (67.0%) and sulfathiazole (40.0%). Our findings indicate a potential role of G. anatis as an important poultry pathogen and cause of subsequent disease, alone or in combination with other pathogens. Continuous monitoring and an antimicrobial susceptibility assay are recommended for effective treatment and disease control.
The genus Gallibacterium, consisting of gram-negative bacteria within the family Pasteurellaceae, has been reported historically under various names, including Pasteurella salpingitis, Actinobacillus salpingitis, Pasteurella anatis, and Pasteurella haemolytica.14,28,33 The bacteria are currently classified within the genus Gallibacterium as G. anatis biovar anatis, G. anatis biovar haemolytica, G. salpingitis, G. trehalosifermentans, G. melopsittaci, and Gallibacterium genomospecies 1–3.6,14 Among these species and biovars, G. anatis biovar haemolytica is known to be associated with salpingitis, peritonitis, and septicemia in chickens.18,19
Gallibacterium infection in chickens, particularly G. anatis, has been reported worldwide within various poultry production and management systems.10,26 There are also reports suggesting its ability to infect calves 34 and humans. 5 Experimental infections leading to peritonitis, folliculitis, increased mortality, and lowered egg production have been reported in layers9,32; experimentally infected cockerels have been reported to have epididymitis and decreased semen quality. 31 In addition, the emergence and spread of multidrug-resistant strains of Gallibacterium are increasingly matters of concern. Lower susceptibility of G. anatis to various drugs, including clindamycin, penicillin, novobiocin, sulfonamides, tylosin, and tetracycline, has been reported in poultry4,7,24,28 and non-poultry isolates. 34
Reports suggest that G. anatis plays a significant role in disease progression, alone or in combination with other pathogens.20,23,30,33 Nevertheless, the incidence of this bacterium seems underreported in poultry because of the close resemblance of clinical manifestations with those associated with other common pathogens, such as Pasteurella multocida and Escherichia coli.24,29 We report here Gallibacterium field strains in broiler and layer chicken flocks with a clinical history of bacterial infections and lesions, primarily in the respiratory tract, reproductive tract, and related serosal surfaces. We further provide information on the phylogenetic relatedness and antimicrobial resistance profiles of Gallibactreium strains isolated in our study.
Materials and methods
Avian diagnostic specimens for bacterial isolation
From August 2019 to July 2021, 2,204 clinical cases of broiler and layer chickens were submitted to the Animal Diagnostic Laboratory (ADL) at the Pennsylvania State University (University Park, PA, USA) for autopsy, as well as histologic and microbiologic investigation. These avian cases originated mostly from Pennsylvania (PA) and Ohio (OH) poultry, with some derived from Indiana (IN) and Maryland (MD). Case flocks had a clinical history of bacterial infection in the respiratory tract, reproductive tract, and other related serosal surfaces. Tissue and swab specimens were collected from each of the clinical cases and processed for the isolation and identification of suspicious pathogens, including common avian bacteria, viruses, and other organisms, such as protozoa and fungi, as per the routine diagnostic procedures in our laboratory (Table 1; Suppl. Table 1).
Summary of the metadata associated with the 69 Gallibacterium isolates from layer and broiler chickens.
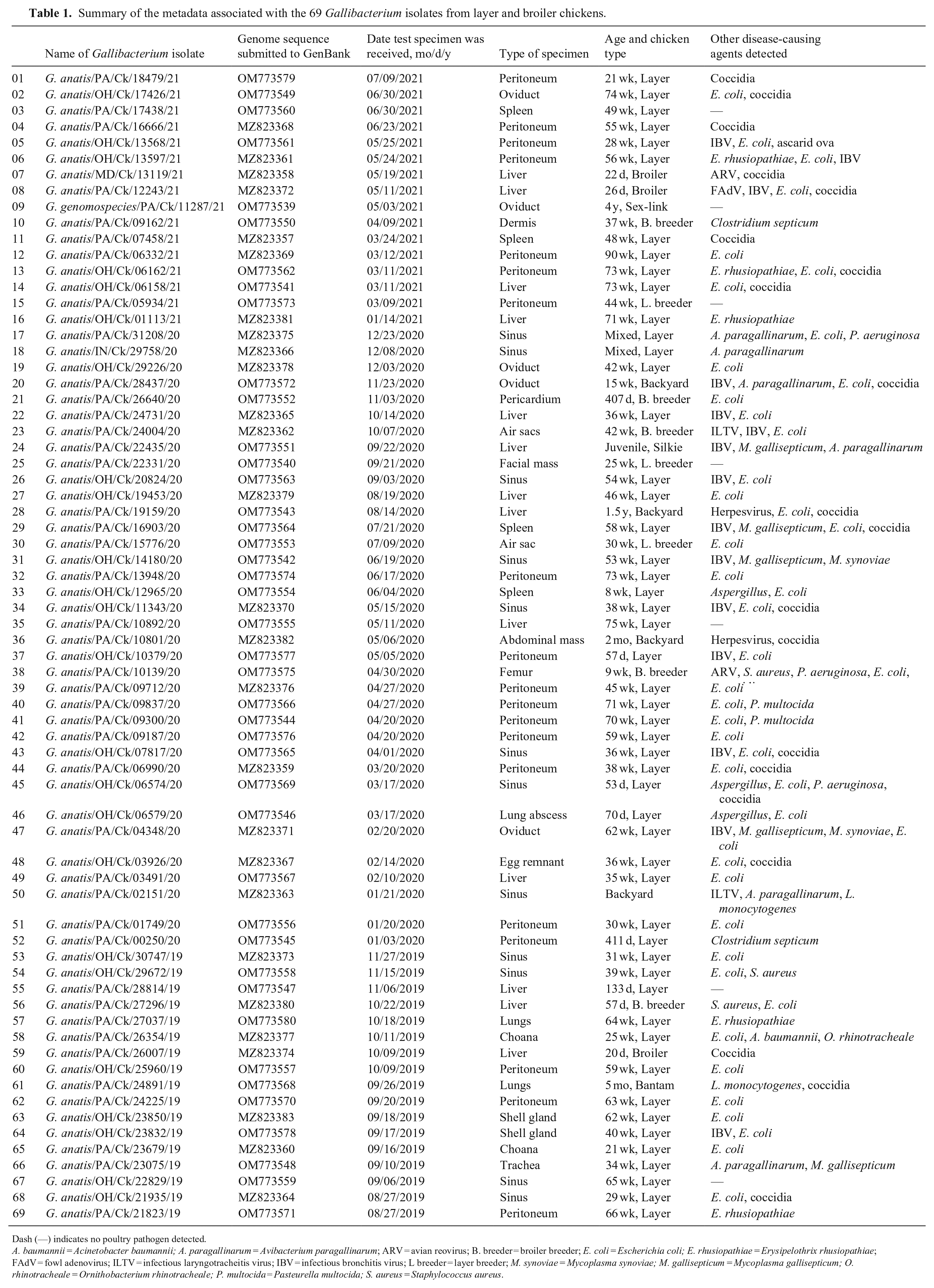
Dash (—) indicates no poultry pathogen detected.
A. baumannii = Acinetobacter baumannii; A. paragallinarum = Avibacterium paragallinarum; ARV = avian reovirus; B. breeder = broiler breeder; E. coli = Escherichia coli; E. rhusiopathiae = Erysipelothrix rhusiopathiae; FAdV = fowl adenovirus; ILTV = infectious laryngotracheitis virus; IBV = infectious bronchitis virus; L breeder = layer breeder; M. synoviae = Mycoplasma synoviae; M. gallisepticum = Mycoplasma gallisepticum; O. rhinotracheale = Ornithobacterium rhinotracheale; P. multocida = Pasteurella multocida; S. aureus = Staphylococcus aureus.
For the isolation and identification of bacteria, particularly Gallibacterium spp., tissue specimens or swabs were processed aseptically for aerobic cultures on blood agar plates, McConkey agar plates, and Columbia nalidixic acid agar plates (Remel; Thermo Scientific). The inoculated plates were incubated at 37°C for 48 h and observed daily to monitor bacterial growth. The observed individual colonies were processed for subcultures, and a pure culture was confirmed at the genus and/or species level by matrix-assisted laser-desorption time-of-flight mass spectrometry (MALDI-TOF MS).
MALDI-TOF mass spectrometry
Mass spectrometry (MS) was performed through a MALDI-TOF Biotyper (Bruker Scientific) per the manufacturer’s instructions, using flexControl software v.3.4 for the automatic acquisition of mass spectra in the positive linear mode. The generated mass spectrum was compared with the reference database of mass spectra available in the manufacturer-provided software with default settings that returned the top 10 identification matches, with a score ranging from 0.0 to 3.0. A MALDI-TOF score of 1.7–2.0 was used to assign a genus; a score >2.0 provided species-level identification. The pure cultures that were assigned to the genus level as “Gallibacterium species” or to the species level as “Gallibacterium anatis” were stored in trypticase soy broth containing 15% glycerol (Hardy) at −80°C.
Phenotypic characteristics and hemolytic pattern
Each archived Gallibacterium isolate was inoculated onto a blood agar plate and incubated for 24–36 h at 37°C under aerobic conditions to yield a fresh culture and determine the hemolytic pattern. Gray, smooth, shiny, convex-shaped, semi-transparent, circular, β-hemolytic colonies raised above the agar surface were considered typical of hemolytic species of Gallibacterium, particularly G. anatis biovar haemolytica.
Genome extraction and PCR
A single colony of pure culture was added to lysogeny broth (Luria broth; Amresco) and incubated at 37°C in a shaking incubator for 24 h or overnight. Broth (300 µL) was collected from each tube for genomic DNA (gDNA) extraction according to the manufacturer’s instructions (E.Z.N.A. blood DNA mini kit; OMEGA Bio-tek). The gDNA was first confirmed by a Gallibacterium-specific PCR assay using a pair of primers that spanned the 16S rRNA (1133fgal, 5′-TATTCTTTGTTACCARCGG-3′) and 23S rRNA (114r, 5′-GGTTTCCCCATTCGG-3′) genes. 11 This was followed by the amplification of a partial 16S rRNA gene (1,063 bp) of the bacterium for sequencing and subsequent phylogenomic analysis using a standard pair of primers (41F, 5′-GCTCAGATTGAACGCTGGCG-3′; 1066R, 5′-TGTAAAGTGTTGTGCTCGAC-3′). 35 The primer pair included the regions (V1, V2, V3, and V6) of the 16S rRNA sequence, which is considered significant in delineating the reads to the lowest taxonomic nodes of genus and species.12,13
For each of the primer pairs, 50 µL of the reaction mixture was formulated with 20 µmol of each primer, 1.25 U of Taq polymerase (OneTaq DNA polymerase kit; New England BioLabs), 10 mmol of each of the dNTPs (Promega), and 8–15 ng of gDNA. Amplification was performed in a thermocycler (GenAmp PCR System 9700; Applied Biosystems). The initial denaturation was carried out once at 95°C for 4 min, followed by 35 cycles of denaturation at 95°C for 30 s, an annealing step at 55°C for 1 min, and an extension step at 72°C for 2 min. The final extension was performed for 10 min at 72°C. The amplified PCR product was gel electrophoresed (1.0%), visualized, and the band size was compared with the 100-bp ladder (exACT gene 100 bp DNA ladder; Fisher Scientific) under a UV trans-illuminator and recorded.
Sanger sequencing and phylogeny analysis
The 16S rRNA PCR products (1,063 bp) of 69 isolates were purified according to the manufacturer’s instructions (Gel/PCR DNA fragment extraction kit; IBI Scientific) and sequenced at the Genomics Core Facility, Pennsylvania State University. BLAST analysis of each sequence was performed (https://blast.ncbi.nlm.nih.gov/Blast.cgi) and compared with the 16S rRNA gene of the reference G. anatis strain (F149T). Ambiguous nucleotides were removed from each end of the sequence, and only the quality sequence (1,011 bp) was used for submission to the NCBI database and subsequent analysis. To determine the phylogeny and nucleotide divergence, the study sequences and 16S rRNA sequences of reference or representative strains and species of Gallibacterium were retrieved from GenBank (https://www.ncbi.nlm.nih.gov) and aligned with BioEdit v.7.2.5 21 using the default settings in the ClustalW program. We assessed the phylogenetic relationship and genetic divergence of our isolates with reference strains using the maximum-likelihood method and Tamura–Nei model with 2,000 bootstrap replicates in MEGAX software. 27
Antimicrobial susceptibility test
The broth microdilution method was used to evaluate the susceptibility of the 69 isolates using a dehydrated panel specific for poultry named AVIAN1F (Sensititre, TREK Diagnostic Systems; Table 2). Briefly, a 10-µL loop of culture with McFarland turbidity of 0.5 in demineralized water was added to 11 mL of Mueller–Hinton broth, and 50 µL of this broth was then added to each well of a 96-well plate carrying different antimicrobials at various concentrations. The inoculated plates were incubated at 37°C under aerobic conditions for 18–24 h. Before reading the plate, bacterial growth was first validated in the form of typical bead formation in each plate’s control wells (without antimicrobials). Plates with a lack of growth were excluded from further analysis. The results were read and interpreted with a software database (Sensititre SWIN; Thermo Scientific), which is based on the CLSI guidelines for bacteria isolated from animals (VET 01-A4, VET 06, VET 01S).15–17
Antimicrobial profiles of the 58 Gallibacterium isolates tested.
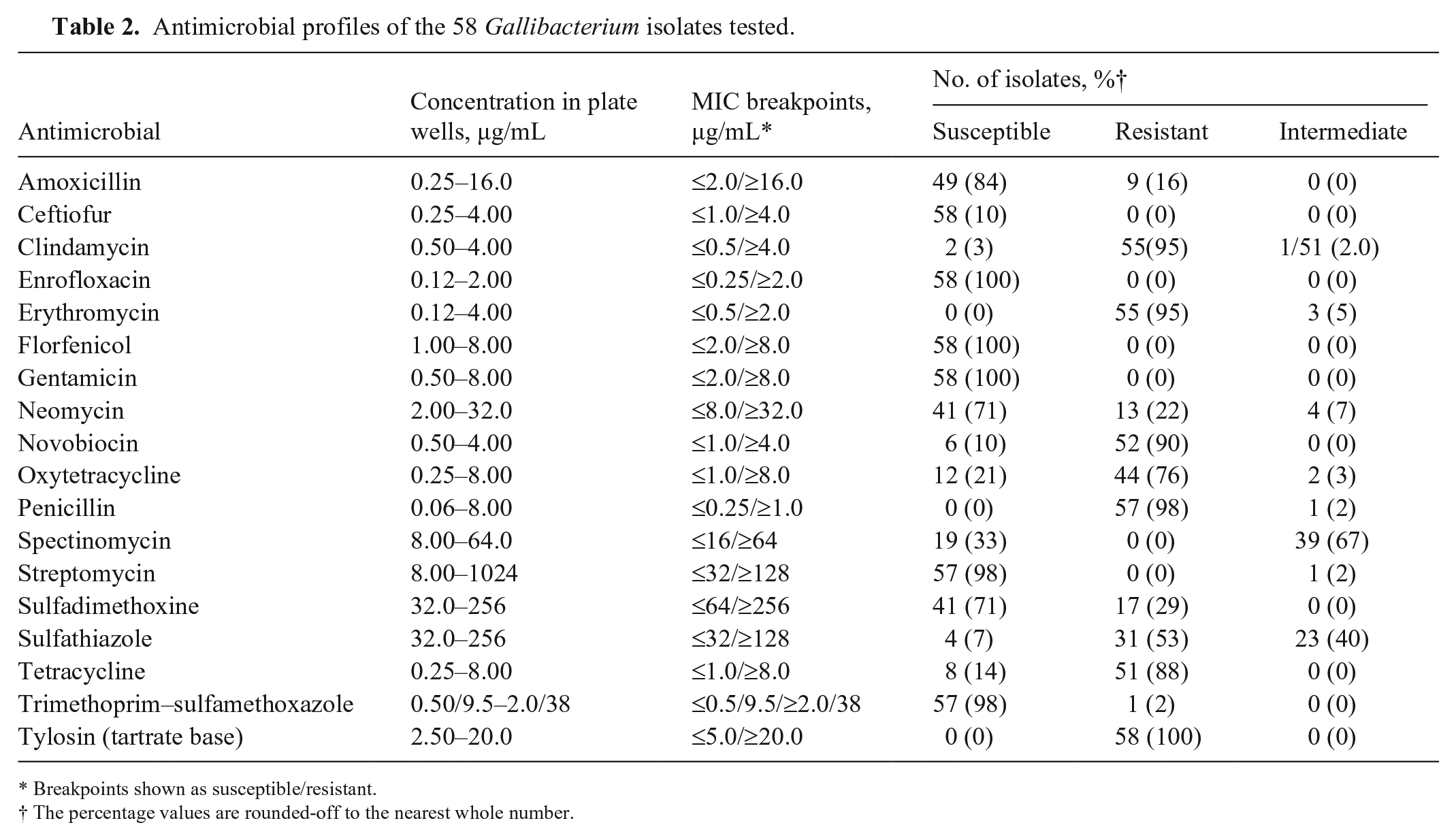
Breakpoints shown as susceptible/resistant.
The percentage values are rounded-off to the nearest whole number.
Results
We obtained 69 Gallibacterium isolates from 2,204 clinical submissions, representing 1 isolate per flock. Of the 69 isolates, 45 were from PA, 22 from OH, 1 from IN, and 1 from MD poultry. Seven poultry flocks were infected with Gallibacterium exclusively; the remaining 62 flocks had mixed infections of Gallibacterium with other bacteria, viruses, protozoa, and fungi. Among these microorganisms, E. coli was the most prevalent, with coinfections detected in 47 of the 69 (68.0%) Gallibacterium-positive cases. Coccidian oocysts were the second most prevalent coinfection, detected in 19 of the 69 cases (27.5%; Table 1; Suppl. Table 1).
Tracing back the poultry production types of the 69 Gallibacterium-positive cases indicated that 51 were commercial layer chickens, 7 were backyard layer chickens, 5 were broiler breeders, 3 were layer breeders, and 3 were commercial broiler chickens (Table 1; Suppl. Table 1). The Gallibacterium-positive cases do not seem to be correlated with poultry production types and housing conditions based on the case distributions of 58 commercial poultry flocks, 4 antimicrobial-free commercial flocks, and 7 small backyard layer flocks. Among the 69 Gallibacterium-positive poultry flocks, 29 were >1-y-old, 25 were 6-mo to 1-y-old, and 14 were ≤ 6-mo-old. In the case specimens, 36 isolates were obtained from serosal surfaces or visceral organs, 20 from the respiratory tract, 9 from the reproductive tract, 2 from the skin or fascial mass, 1 from the heart, and 1 from the femur tissues.
As evidenced by MALDI-TOF MS analysis, the peptide mass fingerprints of 49 isolates corresponded to G. anatis (score > 2.0); 20 isolates matched at the level of genus Gallibacterium (score 1.7–2.0). All of the Gallibacterium isolates had colony characteristics suggestive of hemolytic species of Gallibacterium. Furthermore, a 2-band pattern typical of Gallibacterium spp. was observed when the gDNA of each isolate was amplified using the 1133fgal and 114r primer pairs. Using the primer pair 41F and 1066R, the sequencing PCR amplified a partial 16S rRNA gene (1,063 bp). The gene sequencing data of the 69 isolated field isolates were submitted to the NCBI database (GenBank MZ823357‒MZ823383, OM773539‒OM773580). When these isolate sequences were compared with the representative species of Gallibacterium, including the reference G. anatis strain F149T (NR_036870), the sequences of the field isolates corresponded to positions 42–1,053 of the 16S rRNA gene. The nucleotide identity among field isolates was 94.1–100%. Following phylogenetic analysis, 68 isolates were clustered with the reference strains of G. anatis. Given that these 68 isolates were all hemolytic, they were identified as G. anatis biovar haemolytica. In contrast, 1 field isolate (Gallibacterium genomospecies/PA/Ck/11287/21) was found to be clustered with the reference strain for Gallibacterium genomospecies 2 (CCM 5976; Fig. 1).
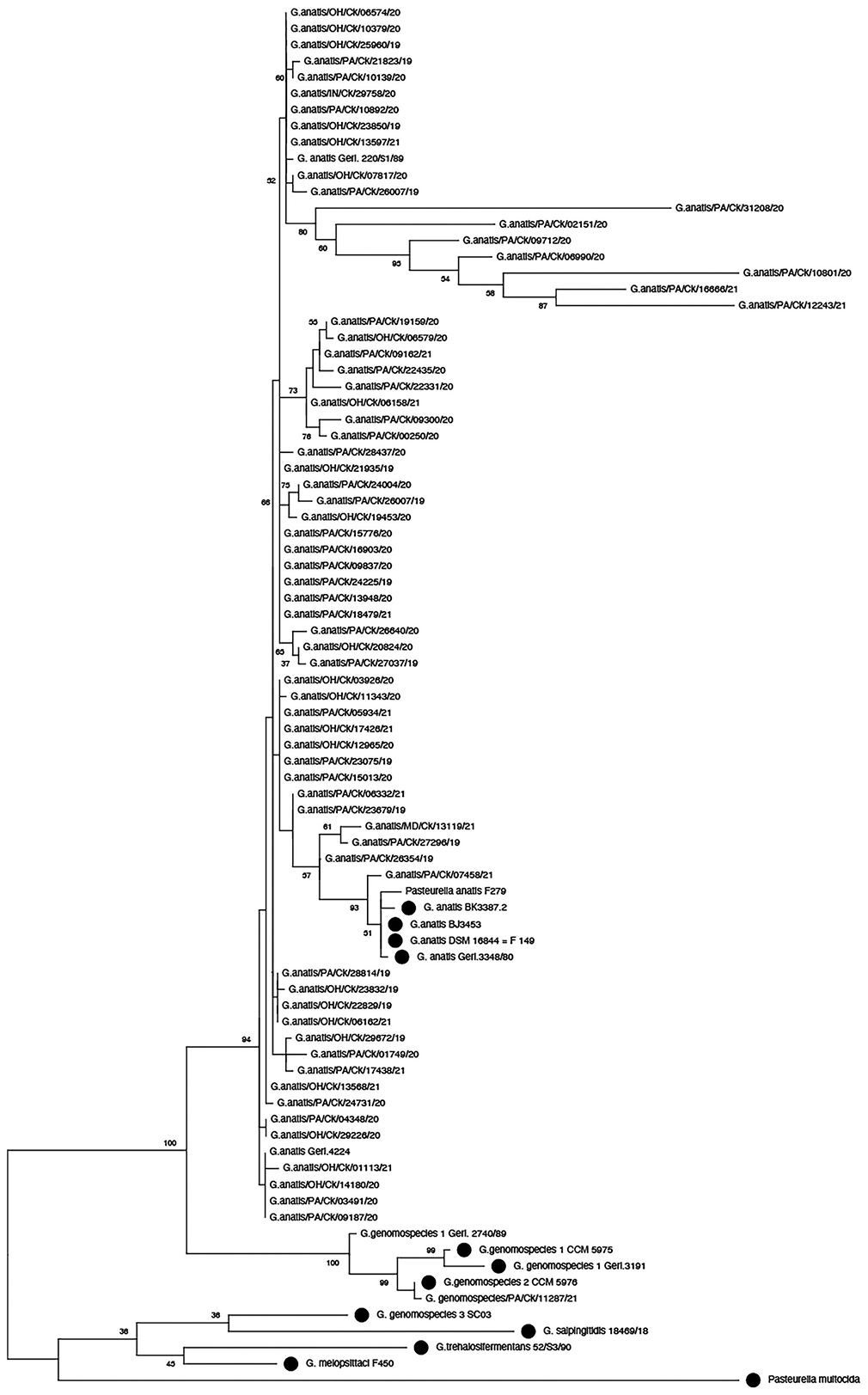
Consensus phylogenetic tree based on alignment of 16S rRNA sequences of study isolates of Gallibacterium and reference strains from GenBank (dots). The tree was constructed using the maximum-likelihood method with 2,000 bootstrap replicates.
The 69 Gallibacterium isolates were processed to assess their antimicrobial profiles. A total of 57 isolates of G. anatis and 1 isolate of Gallibacterium genomospecies were tested successfully and allocated to susceptible, intermediate, or resistant using the interpretation criteria of the on-board software (see Table 2 for the breakpoints). The remaining 11 G. anatis isolates did not grow in the control wells of a 96-well test plate (AVIAN1F). All 57 isolates of G. anatis and the 1 isolate of Gallibacterium genomospecies were uniformly susceptible to enrofloxacin (100%), ceftiofur (100%), florfenicol (100%), and gentamicin (100%); most were susceptible to trimethoprim–sulfamethoxazole (98.0%), streptomycin (98.0%), amoxicillin (84.0%), sulfadimethoxine (71.0%), and neomycin (71.0%). Antimicrobial resistance in the isolates was frequent for tylosin (100%), penicillin (98.0%), erythromycin (95.0%), clindamycin (95.0%), novobiocin (90.0%), tetracycline (88.0%), oxytetracycline (76.0%), and sulfathiazole (53.0%). However, the Gallibacterium genomospecies isolate was resistant to clindamycin, novobiocin, oxytetracycline, penicillin, tetracycline, and tylosin. Intermediate resistance was observed exclusively with erythromycin.
Discussion
Colony characteristics and hemolysis patterns in our study were analogous for both G. anatis biovar haemolytica and Gallibacterium genomospecies. Gallibacterium-specific PCR and MALDI-TOF MS complemented each other and confirmed the field isolates as Gallibacterium. The Gallibacterium-specific PCR assay has been reported and validated on 122 strains with 100% sensitivity and specificity. 11 On the other hand, MALDI-TOF MS utilizes peptide mass fingerprints of the study isolate and the reference isolates in the spectral database to identify isolates. Although this technique has immense potential in routine testing given the smaller sample size requirements, the ability to process several samples simultaneously, and the shorter turnaround time, MALDI-TOF MS also has certain limitations. For instance, the 4 Gallibacterium species and the 3 Gallibacterium genomospecies were correctly identified by MALDI-TOF MS, but under optimal conditions. 2 However, the reproducibility of the assay may vary as a result of changes in the protein profile of the bacteria under various growth (type of media, oxygen, and incubation temperature and time) and storage conditions.2,3 Furthermore, identifying each bacterial species within a genus requires an independent reference spectral database enriched with the field isolates, 1 which may not be available in a front-line veterinary diagnostic laboratory. Excluding 1 isolate identified as Gallibacterium genomospecies, for which the reference spectral database was unavailable, this may explain why 19 of 68 isolates of Gallibacterium were not identified as G. anatis through MALDI-TOF MS. Collectively, members of Pasteurellaceae, particularly Gallibacterium spp., are phenotypically heterogeneous. 14 Accurate species-based identification of MALDI-TOF MS is limited by the need for a reference spectral database of each species and precise optimal conditions. Therefore, in a scenario in which there is a lack of a specific species-based spectral database and/or when conditions are not well optimized for an individual bacterial species, other genotypic methods or genetic markers (e.g., 16S rRNA gene) may be the methods of choice for the simultaneous identification and differentiation of bacteria.
We amplified the partial 16S rRNA gene corresponding to the hypervariable region (V1‒V6) for phylogenetic and divergence analysis. We chose this region because it provides the maximum resolution for assigning reads to lower taxonomic nodes. 12 While determining the potential utility of hypervariable regions (V1‒V9) of the 16S rRNA gene in disease diagnosis and other relevant systemic investigations, a study 13 on 110 different bacterial species, including select agents and environmental microflora, highlighted the importance of V1, V2, V3, and V6 regions when assigning reads to lower taxonomic nodes. The Gallibacterium isolates in our study were not identical but had minor genetic divergence. This divergence may explain why the isolates did not align in the same clade and showed minor variations in the clustering pattern within their respective clades.
Except for 1 isolate, the remaining 68 field isolates in our study were determined to be G. anatis biovar haemolytica. The Gallibacterium genomospecies isolate (G. genomospecies/PA/Ck/11287/21) was obtained from the oviduct of a 4-y-old backyard chicken with a history of salpingoperitonitis and septicemia. Isolates of Gallibacterium genomospecies have been obtained from chickens and pigeons with disease-linked lesions. 14 Further characterization is required before our isolate can be confidently assigned to Gallibacterium genomospecies 1, 2, or 3.
Although most of the Gallibacterium-positive flocks had a similar clinical history, the specimens used for bacterial isolation were not uniform in most cases. Indeed, typical Gallibacterium cultures were obtained from a wide range of bird tissues and swab specimens, indicating its potential to invade and grow in different organs in the body. Among the species of Gallibacterium, G. anatis biovar haemolytica has the broadest range of tissue tropism and is associated with respiratory, gastrointestinal, reproductive, and septicemic infections.10,18,19,29 This range agrees with the metadata from our study, wherein the bacterium was isolated from a wide range of tissues of clinically infected birds with a history of infection in the respiratory tract, reproductive tract, and related serosal surfaces. It is worth noting that studies have reported differing results regarding genotypic or phenotypic characteristics of the isolates recovered from different tissues in a bird. For instance, the presence of niche-independent single clones of the bacterium within a bird and flock using amplified fragment length polymorphism (AFLP)-based typing has been demonstrated. 29 In contrast, 2 closely related but niche-dependent clonal lineages, recognized using AFLP, colonized the trachea and cloaca of the same bird in a layer flock. 8 In another study, variations in the antimicrobial resistance profiles were observed within isolates originating from different organs of the same bird and even within some paired isolates from the same organ. 22
Most of the Gallibacterium isolates in our study were obtained along with other pathogens, such as Avibacterium paragallinarum, E. coli, Erysipelothrix rhusiopathiae, and viruses from birds that had a history of peritonitis, salpingitis, and serositis. This suggests that G. anatis may not necessarily be the major cause of the disease but has the potential to contribute to pronounced clinical signs in the form of coinfections. In addition to host-related factors, such as age, immune status, and stress, the experimental infection of G. anatis along with infectious bronchitis virus (Avian coronavirus), 23 A. paragallinarum, 30 or Mycoplasma gallisepticum 20 was reported to aggravate disease severity and clinical outcomes. We found that some of the Gallibacterium isolates obtained along with E. coli came from egg-laying hens with a history of reproductive disorders and reduced egg production. Although peritonitis in laying hens is attributed to invasion and infection with E. coli in birds, 25 evidence suggests a potential role of G. anatis in causing disease and further aggravating the condition in cases in which E. coli and G. anatis are isolated from the same clinical specimen. 29
We used the Sensititre SWIN software-based MIC breakpoints for Enterobacteriaceae to assess the antimicrobial activity of a range of antimicrobials against Gallibacterium spp. because agreed MIC breakpoints are not available for all of the antimicrobial substances against poultry-originated bacterial pathogens, particularly Gallibacterium spp. This may be why previous studies used different MIC breakpoints.4,7,18,24 Also, except for a few antimicrobials, there exists an inconsistency in the concentration of antimicrobials employed to assess in vitro susceptibility of G. anatis in the studies mentioned above, resulting in various MIC values and interpretation. Future investigations should be conducted to establish validated MIC breakpoints.
The lack of bacterial growth or bead formation in 11 of the Gallibacterium isolates in the control wells (without antimicrobials) indicated an inability to grow in the Mueller–Hinton broth. A similar problem of an inability to grow in Mueller–Hinton broth has been reported for some G. anatis isolates.7,22 The remaining 58 isolates were resistant to many of the antimicrobials that are used commonly in poultry production. Resistance to many antimicrobials has been reported worldwide, including in the United States,24,28 Mexico and Denmark, 7 Germany, 18 Egypt, 19 and Iran. 4 Furthermore, given that the susceptibility or resistance of Gallibacterium isolates could vary with time across various locations, there is a need to monitor these patterns before applying or administering suitable drugs to treat birds according to conditions.
Supplemental Material
sj-pdf-1-vdi-10.1177_10406387221133782 – Supplemental material for Identification, 16S rRNA–based characterization, and antimicrobial profile of Gallibacterium isolates from broiler and layer chickens
Supplemental material, sj-pdf-1-vdi-10.1177_10406387221133782 for Identification, 16S rRNA–based characterization, and antimicrobial profile of Gallibacterium isolates from broiler and layer chickens by Muhammad Z. Shabbir, Subhashinie Kariyawasam, Traci A. Pierre, Patricia A. Dunn, Eva A. Wallner-Pendleton and Huaguang Lu in Journal of Veterinary Diagnostic Investigation
Footnotes
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
Our work was supported by the U.S. Egg Industry Center at Iowa State University (grant subcontract 497-43-74A), Pennsylvania Poultry Industry Egg/Broiler Research CheckOff Program (grant 2018), Pennsylvania Soybean Board (grant PSB R2016-01), and Pennsylvania Center for Poultry and Livestock Excellence (grant CPLE21-12).
Supplemental material
Supplemental material for this article is available online.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
